Abstract
Fats, oils and greases (FOG) are energy-dense wastes that can be added to anaerobic digesters to substantially increase biomethane recovery via their conversion through long-chain fatty acids (LCFAs). However, a better understanding of the ecophysiology of syntrophic LCFA-degrading microbial communities in anaerobic digesters is needed to develop operating strategies that mitigate inhibitory LCFA accumulation from FOG. In this research, DNA stable isotope probing (SIP) was coupled with metagenomic sequencing for a genome-centric comparison of oleate (C18:1)-degrading populations in two anaerobic codigesters operated with either a pulse feeding or continuous-feeding strategy. The pulse-fed codigester microcosms converted oleate into methane at over 20% higher rates than the continuous-fed codigester microcosms. Differential coverage binning was demonstrated for the first time to recover population genome bins (GBs) from DNA-SIP metagenomes. About 70% of the 13C-enriched GBs were taxonomically assigned to the Syntrophomonas genus, thus substantiating the importance of Syntrophomonas species to LCFA degradation in anaerobic digesters. Phylogenetic comparisons of 13C-enriched GBs showed that phylogenetically distinct Syntrophomonas GBs were unique to each codigester. Overall, these results suggest that syntrophic populations in anaerobic digesters can have different adaptive capacities, and that selection for divergent populations may be achieved by adjusting reactor operating conditions to maximize biomethane recovery.
Similar content being viewed by others
Introduction
Anaerobic digestion is an attractive biotechnology for recovering renewable energy as biomethane (CH4) during organic waste treatment. Worldwide interest in anaerobic digestion has led to the implementation of ~9000 full-scale facilities in Europe and over 3000 facilities in the US by 2012 (Guo et al., 2015). The conversion of organic matter into biomethane relies upon an intricate balance of multiple microbial trophic groups, including hydrolyzing and fermenting bacteria, syntrophic acetogenic bacteria and methanogenic archaea (Briones and Raskin, 2003). Despite the relatively widespread adoption of anaerobic digestion, the variety of its contributing genetic and metabolic systems still remain relatively underexplored (Vanwonterghem et al., 2016), owing to the vast phylogenetic and metabolic diversity (Sundberg et al., 2013) and general non-culturability of a majority of the microorganisms present in digesters (Vanwonterghem et al., 2014). An improved understanding of the key ecological niches within anaerobic digesters and the metabolic and physiological attributes of microbial populations occupying those niches could help to facilitate new process designs and operational strategies that maximize biomethane recovery from organic waste.
Adding fats, oils and greases (FOG) as a co-substrate during the anaerobic digestion of livestock manure or wastewater treatment solids has gained popularity to increase biomethane recovery due to the high biological methane potential of FOG (Alves et al., 2009; Luostarinen et al., 2009; Wang et al., 2013). During anaerobic degradation, FOG is rapidly hydrolyzed into glycerol and long-chain fatty acids (LCFA) (Hanaki et al., 1981). Because the degradation of LCFA in methanogenic environments relies on low hydrogen, formate and acetate levels for favorable thermodynamics, LCFA are converted into methane through an obligatory syntrophic partnership between acetogenic β-oxidizing bacteria and methanogenic archaea (Schink, 1997; Sousa et al., 2009). So far, only seven isolated bacterial species are capable of β-oxidizing LCFA (>12 carbons in length) in syntrophy with methanogens, and all belong to the families of Syntrophomonadaceae and Syntrophaceae (Sousa et al., 2009; Zhang et al., 2012). However, the generally slow growth rates of syntrophic bacteria coupled with the toxicity of LCFA has limited the number of cultured representatives obtained (Hatamoto et al., 2007).
Biomethane recovery from FOG can be limited by the reduction of microbial activity attributed to the toxicity of LCFAs at high concentrations (Hanaki et al., 1981; Angelidaki and Ahring, 1992; Rinzema et al., 1994; Davidsson et al., 2008; Luostarinen et al., 2009; Wan et al., 2011; Wang et al., 2013). Our recent work indicated that the maximum FOG loading rate to an anaerobic codigester was dependent on the LCFA-degrading population abundance and specific LCFA bioconversion kinetics (Ziels et al., 2016, 2017). The codigester LCFA feeding frequency was found to impact the composition and diversity of β-oxidizing Syntrophomonas species, with LCFA pulse feeding leading to higher bioconversion kinetics in comparison to semi-continuous LCFA feeding (Ziels et al., 2017). However, those detected differences in the syntrophic community structure between codigesters were based on time-series 16S rRNA gene amplicon sequencing, and thus the impact of feeding frequency on the active fraction of the community remained unknown. A comparison of the active LCFA-degrading populations selected with different feeding frequencies would improve our understanding of relationships between operating parameters and physiological diversity within digester syntrophic populations.
Stable isotope probing (SIP) is a cultivation-independent method that can be used to elucidate links between microbial activity (function) and identity within environmental samples (Radajewski et al., 2000; Dumont and Murrell, 2005). DNA-SIP relies on the incorporation of heavy isotopes (for example, 13C) into microbial DNA during growth on labeled substrates, and thus acts as a ‘filter’ to enrich the DNA of active populations (Chen and Murrell, 2010). DNA-SIP has been combined with shotgun metagenomic sequencing approaches to discover novel enzymes and operons within microbial communities (Saidi-Mehrabad et al., 2013; Chemerys et al., 2014; Verastegui et al., 2014), and to identify new functional traits of microbial taxa (Eyice et al., 2015). Recently-developed differential coverage binning techniques for recovering population genomes from shotgun metagenomes (Albertsen et al., 2013; Kang et al., 2015; Karst et al., 2016) could be amenable to DNA-SIP due to the expected DNA sequence composition divergence between heavy density fractions of 12C- and 13C-incubated samples (Coyotzi et al., 2016). However, there have been no reported studies coupling differential coverage binning with DNA-SIP to recover population genome bins from metagenome sequence data thus far.
The objective of this study was to apply DNA-SIP combined with shotgun metagenomic sequencing for a genome-centric comparison of the active oleate (C18:1 LCFA) degrading populations in two parallel anaerobic codigesters operated with different oleate feeding frequencies. Differential coverage binning was employed to retrieve population genome bins from the DNA-SIP 13C and 12C-metagenomes. 16S rRNA gene amplicon sequencing was also conducted on 13C and 12C heavy DNA fractions to verify key populations involved in converting oleate into biomethane within the anaerobic codigesters.
Materials and methods
Anaerobic bioreactor operation
To examine the selective impacts of feeding frequency on the microbial community, two parallel anaerobic codigesters treating manure and oleate were operated at 35 °C for over 200 days, as described previously (Ziels et al., 2017). The digesters had a 4-liter working volume and a 20-day hydraulic and solids retention time (SRT), and were fed with blended manure feedstock collected from a nearby dairy farm. Manure was fed daily at a VS loading rate of 1.3±0.2 g VS/l-d, and an aqueous solution of sodium oleate (>97%, Tokyo Chemical Industry Co.) was added to the influent feed at 10% of the daily feed volume. The daily average oleate organic loading rate (OLR) to both codigesters was identical, but their oleate feeding frequency differed. One codigester received the oleate feed semi-continuously every 6-h, and is termed the continuous-fed (CF) codigester. The other codigester received the oleate feed in a pulse every 2 days, and is termed the pulse-fed (PF) codigester. At day 205 of operation, the oleate OLR to both codigesters was reduced from 3.2 to 1.8 g COD/l-d to promote stable digestion, as acetate was accumulating in the CF codigester effluent by day 200 (Ziels et al., 2017). The resulting oleate concentration in the feed was 44 mM after day 205. From days 205–230, the pH of both codigesters averaged 7.5, and effluent VFA (acetate) was below 400 mg l−1. The codigesters were not fed between days 228-230 to reduce the level of any background substrates before commencing SIP incubations on day 230. At that time, total effluent VFA and LCFA (liquid+sorbed) levels were below 70 mg l−1.
Stable isotope probing incubations with anaerobic digester sludge
Sludge was sampled directly from the PF and CF codigesters on day 230. Microcosms were established in 35 ml glass serum bottles pre-purged with N2:CO2 (80:20), amended with 10 ml of codigester sludge (0.17 g VS total), sealed and capped with butyl rubber septa, and purged again with N2:CO2 (80:20). All microcosms were maintained at 35 °C, and stirred at 150 rpm on an orbital shaker. Microcosms were grouped into duplicate vials that were supplemented with either 12C-sodium oleate or 13C-labeled sodium oleate at an initial concentration of 8 mm. The 13C-labeled sodium oleate was universally labeled at all 18 carbons (>98% atom purity, Cambridge Isotope Laboratories, Tewksbury, MA, USA). Triplicate un-supplemented controls were incubated in parallel to measure background methane production. Gas production was measured based on the headspace pressure increase using a digital manometer (Series 490 A, Dwyer Instruments), and the headspace methane content was measured with a GC-FID (SRI 8610C) equipped with a Supelco Alumina Sulfate Plot column (50 m, 0.53 mm i.d.). Headspace pressure and composition were sampled about every 5 h using sterile syringes. Background methane production was subtracted from oleate-amended assays to measure the overall rate of substrate conversion.
After the first addition of oleate was converted at >95% efficiency into methane over 48 h (Figure 1), 12C- and 13C-labeled sodium oleate were re-supplemented at 8 mm after purging the headspace with N2:CO2 (80:20). Following another 48-h for oleate conversion into methane, a third supplement of 8 mm 12C- and 13C-labeled sodium oleate was added to the microcosms after purging the headspace with N2:CO2 (80:20). The third incubation lasted 48 h, after which biomass from all bottles was sacrificed for DNA extraction. In total, 240 μmol of either 12C- or 13C-labeled oleate was added to the microcosms over the course of 6 days (144 h).
Cumulative methane production (minus blank controls) for the microcosms fed with 12C- and 13C-labeled oleate over three repeated batch feeding periods. The number above each plot indicates the batch oleate feed round. The black dashed line shows the theoretical methane potential of the added oleate (25.5 ml CH4; based on 2.9 g COD/g oleate, 8 mm concentration and 35 °C temperature). The black solid line represents the predicted methane production based on non-linear model fitting with a modified Gompertz equation (Supplementary Table 1). Error bars represent the standard deviation of the biological replicates.
DNA extraction and density-gradient centrifugation
Anaerobic codigester sludge samples were collected for DNA extraction at the start of the SIP incubation (time zero), as well as from each 12C-oleate and 13C-oleate fed microcosm after 6 days of incubation. DNA was extracted and subjected to density-gradient centrifugation and fractionation, as described in detail in the Supplementary Information.
Quantitative PCR
To further detect differences in DNA buoyant density (BD) values between the 12C- and 13C-incubated microcosms, quantitative PCR (qPCR) was conducted on DNA from gradient fractions with BDs ranging from 1.68 to 1.75 g ml−1. The qPCR targeted 16S rRNA gene fragments of the syntrophic LCFA β-oxidizing bacterial genus of Syntrophomonas using primers and probes developed by Ziels et al. (2015). More detailed descriptions of the reaction conditions, primers and probes, and calibration standards are given in the Supplementary Information.
16S rRNA gene amplicon sequencing
DNA from ‘heavy’ gradient fractions with BDs ranging from 1.70 to 1.73 (4 fractions) were pooled at equal volumes and subjected to 16S rRNA gene amplicon sequencing for each microcosm (Figure 2). Briefly, 16S rRNA gene fragments were amplified with barcoded and indexed universal prokaryotic V4-V5 primers 515 F-Y (5′-GTGYCAGCMGCCGCGGTAA) and 926 R (5′-CCGYCAATTYMTTTRAGTTT) (Parada et al., 2016), and the products were pooled and sequenced on a 2x300 bp Illumina MiSeq run by the United States Department of Energy (DOE) Joint Genome Institute (JGI). Raw sequence data is available through the JGI Portal (http://jgi.doe.gov) under project ID number 1105527.
Total copies of Syntrophomonas 16S rRNA genes measured by qPCR for each density-gradient fraction recovered from isopycnic separation of DNA from 13C-incubated microcosms and 12C-controls for both anaerobic codigesters. The filled circles indicate gradient fractions that were pooled for subsequent 16S rRNA gene amplicon sequencing and metagenomic sequencing. Both biological replicate microcosms are shown, and each point represents an average of duplicate technical replicates.
Amplicon reads were denoised into their exact sequences using the DADA2 pipeline v.1.2.1 (Callahan et al., 2016) before clustering into 99.5% OTUs with UPARSE v.8.1 (Edgar, 2013), as described in detail in the Supporting Information.
Metagenomic sequencing, assembly and annotation
Pooled volumetric fractions of heavy DNA from density-gradient centrifugation (Figure 2), as well as codigester DNA samples from time zero, were used to generate an Illumina sequence library with an average insert size of 250 bp that was sequenced on an Illumina HiSeq-2500 with paired-end 150 bp reads at the DOE JGI. The metagenomic sequencing produced an average of 88±11 M reads for SIP DNA samples and 140±13 M reads for time zero DNA samples. Reads were trimmed and screened to remove adapters as well as filter sequences with a minimum Phred quality of 12, minimum length of 51 and no ambiguous bases using BBDuk v.36.02 (https://sourceforge.net/projects/bbmap/). Trimmed and screened paired-end reads from the codigester time zero metagenomes were individually assembled into contigs using MEGAHIT v.1.0.3 (Li et al., 2015a) with default parameters and the kmer list: 23, 43, 63, 83, 103, 123. The assembled contigs were annotated with the Integrated Microbial Genomes Metagenome (IMG/M) platform (Chen et al., 2016) by the DOE JGI, and KEGG annotation information (KO and EC numbers) were extracted. Raw sequence reads and assemblies are available through the DOE JGI portal under project IDs 1105480, 1105482, 1105484, 1105486, 1105488, 1105490, 1105494, 1105496 and 1105498.
Metagenomic binning
Quality-filtered reads from all metagenomes, including the heavy gradient fractions of the 12C- and 13C-oleate fed microcosms and from codigester time zero samples, were individually mapped to contigs from the time zero assemblies using BBMap v.36.02 (https://sourceforge.net/projects/bbmap/) in paired-end mode with default parameters. SAMtools v.1.4 (Li et al., 2009) was used to sort and index the mapping files and extract contig coverage information across samples. The mapping data and coverage information were used to bin contigs into population genome bins (GBs) separately for each codigester sample set using MetaBat v.0.26.3 (Kang et al., 2015) in ‘superspecific’ mode. CheckM v.1.0.6 was used to evaluate the level of bin completeness and contamination based on domain-level single-copy marker genes (Parks et al., 2015). Taxonomic classification of GBs was conducted by extracting a set of 107 conserved marker genes within open reading frames using mmgenome (Karst et al., 2016), running BLASTP on the extracted marker genes against the RefSeq v.52 protein database with a maximum e-value cutoff of 1e-5, and finding the least common ancestor (LCA) of the top 5 blast hits for each marker gene with MEGAN v.6.4 (Huson et al., 2007). The finest taxonomic ranking was assigned for which at least 70% of the LCA results agreed. Phylogenetic trees were generated for GBs that were over 40% complete using PhyloPhlAn v.1.3 based on 400 conserved marker genes (Segata et al., 2013), as well as with UPGMA clustering of distance estimates from in silico DNA–DNA hybridization with the Genome-to-Genome Distance Calculator v.2.1 (Auch et al., 2010).
Statistical analysis
Statistically significant differences in raw read counts of OTUs (from 16S rRNA gene amplicon libraries) and GBs (from shotgun metagenome libraries) within the 12C and 13C DNA-SIP samples were detected with the negative binomial Wald test in DESeq2 v.1.14.1 separately for each codigester data set (Love et al., 2014). Raw sequence counts of GBs were obtained from parsing the mapping files. An adjusted P-value of 0.01 and a log2 abundance increase greater than 2.0 (or >4-fold increase) were used to identify features (for example, OTUs or GBs) that significantly increased between the 12C and 13C DNA-SIP samples.
Regularized log-transformed sequence counts of 16S rRNA gene amplicon OTUs were determined with DESeq2 for use in principal component analysis. Permutational multivariate ANOVA (ADONIS) was conducted on Euclidian distance values from transformed sequence counts using the vegan package v.2.4.2 in R (Oksanen et al., 2007).
Results
Conversion of oleate into methane and enrichment of 13C DNA
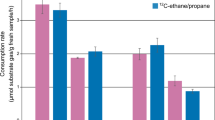
Metabolism of 12C- and 13C-labeled oleate was confirmed with 97% and 95% COD recoveries as methane in the headspace for the PF codigester and CF codigester microcosms, respectively (Figure 1). Biokinetic modeling of methane production (described in Supplementary Information) showed that the maximum methane production rate increased with each subsequent batch feeding of 12C- and 13C-labeled oleate (Supplementary Table 1). The maximum methane production rate in the PF codigester microcosms was 26%, 21% and 17% higher than the CF codigester microcosms for the first, second and third batch incubation rounds, respectively (Supplementary Table 1).
Total DNA concentrations were measured in 24 density-gradient fractions to detect BD shifts after the consumption of 240 μmol 12C- or 13C-labeled oleate after 6 days. The heavy density fractions with BDs from 1.70 to 1.725 g ml−1 contained 2.2-times and 1.6-times more DNA in the 13C-incubated samples than in the 12C-controls for the PF and CF codigester microcosms, respectively (Supplementary Figure 2). To further confirm the enrichment of 13C-labeled DNA, 16S rRNA genes of the LCFA-degrading bacterial genus Syntrophomonas were quantified in density fractions between 1.66 and 1.76 g ml−1 (Figure 2). The four heavy density-gradient fractions with BDs ranging from 1.70 to 1.725 g ml−1 contained over 12-times more Syntrophomonas 16S rRNA genes in the 13C-incubated samples than that in the 12C-controls for both codigesters (Figure 2). The four heavy gradient fractions showing high levels of 13C incorporation were therefore pooled for each microcosm for subsequent 16S rRNA gene amplicon sequencing and shotgun metagenomic sequencing (Figure 2).
16S rRNA gene amplicon sequencing of DNA-SIP samples
The microbial community profiles of pooled heavy density-gradient fractions (Figure 2) were assessed through paired-end 16S rRNA gene amplicon sequencing. After qualify-filtering, denoising, and merging paired-end amplicon reads, the number of sequences retained per sample averaged 164 000±39 000. Principal component analysis (PCA) showed that community profiles of heavy density fractions from the 12C- and 13C-incubated microcosms were distinctly clustered for both codigesters (Supplementary Figure 3). The grouping of community profiles was significant within both codigester sample sets according to the oleate isotope (ADONIS, P<0.05). Additionally, samples within each isotope group were significantly grouped according to the codigester sludge source (ADONIS, P<0.05), indicating that the 13C-enriched communities from each codigester biomass were distinct.
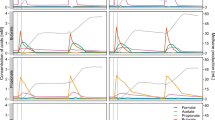
Differential abundance analysis was conducted to detect 16S rRNA gene OTUs that significantly increased in 13C-incubated heavy gradient fractions relative to 12C-controls. A total of 59 differentially abundant OTUs were identified in the PF codigester samples, 42 of which were classified to the Syntrophomonas genus (Supplementary Figure 4). Within the CF codigester samples, a total of 40 differentially abundant OTUs were identified, with 36 classified as Syntrophomonas (Supplementary Figure 5). Of the total number of differentially abundant Syntrophomonas OTUs identified, 19 were shared between the codigesters while the remaining OTUs were unique to either codigester (Supplementary Figure 6). About 48 and 35% of the total 16S rRNA gene counts of differentially abundant Syntrophomonas OTUs were attributed to unique OTUs within the 13C-samples for the CF and PF codigesters, respectively (Supplementary Figure 7). The relative fraction of Syntrophomonas 16S rRNA gene counts increased on average 2.6- and 2.9-times in the 13C-incubated libraries relative to the 12C-controls for the PF codigester and CF codigester, respectively (Figure 3). Cumulative Syntrophomonas genus 16S rRNA gene counts increased on average 5.0- and 5.5-times in the 13C-incubated samples relative to the 12C-controls for the PF codigester and CF codigester, respectively (Supplementary Figure 8). In contrast, the relative abundance of all other bacterial genera decreased in the 13C 16S rRNA gene libraries relative to 12C-controls for both codigesters, with the exception of the Ruminococcaceae NK4A214 group in the PF codigester (Figure 3). Other differentially abundant OTUs were detected belonging to the genera: Thermovirga, Aminivibrio, Candidatus Cloacamonas, Anaerofustis, Syntrophothermus and Ruminococcaceae NK4A214 group, along with unclassified members of Planctomycetes, Spirochaetae, Synergistes, Actinobacteria and Bacteriodetes (Supplementary Figures 4 and 5). No methanogenic archaeal OTUs were identified as differentially abundant in either codigester sample set. Nonetheless, the relative abundance of Methanosaeta 16S rRNA genes increased from 2.2% in the 12C-controls to 3.2% in the 13C-samples for the CF codigester libraries (Figure 3). Cumulative 16S rRNA gene counts of Methanosaeta increased by 1.7- and 2.7-times in the 13C-incubated samples relative to 12C-controls for the PF codigester and CF codigester libraries, respectively, and Methanobacterium read counts increased by 1.7- and 1.9-times, respectively (Supplementary Figure 8).
Relative fractions of the 11 most abundant genera in the heavy gradient fractions of 13C-incubated samples and 12C-controls for sample sets from both codigesters, based on 16S rRNA gene amplicon sequencing. Relative fractions were determined using DESeq2 normalized read counts for all OTUs (Love et al., 2014), and were then aggregated at the genus level. Both biological replicate microcosms are shown.
DNA-SIP metagenomic sequencing
Metagenomes from the PF and CF codigester sludges before SIP incubation (that is, time zero) were assembled to produce contigs for differential coverage binning. The PF and CF codigester time zero metagenome assemblies amounted to 536 947 079 bp and 680 484 169 bp within contigs longer than 1 kb, and had maximum contig lengths of 692 259 bp and 485 835 bp, respectively. Over 200 GBs were recovered through differential coverage binning for each codigester DNA-SIP sample set. Differential abundance analysis of GB read counts in the 13C-incubated metagenomes relative to 12C-controls using DESeq2 showed that 16 and 18 GBs were significantly enriched in the 13C-metagenomes of the PF and CF codigesters, respectively (Supplementary Table 2). Due to the low completeness of some enriched GBs (Supplementary Table 2), a threshold of 40% genome completeness was established for further phylogenetic analysis. Assigned taxonomies and genomic characteristics of differentially abundant GBs with over 40% completeness are summarized in Table 1.
Approximately 75% of the differentially abundant GBs (6 of 8) in the PF codigester samples were taxonomically assigned to Syntrophomonas, along with 71% (5 of 7) of the differentially abundant GBs in the CF codigester (Table 1). Both codigesters contained one differentially abundant GB that could only be assigned at the phylum-level to Firmicutes. In addition, one differentially abundant GB in the PF codigester was classified to the family-level of Syntrophomonadaceae, and one GB in the CF codigester was assigned to the phylum Candidatus Parcubacteria (Table 1). Multi-gene alignments based on a set of 400 conserved marker genes showed that some differentially abundant GBs from both codigesters were closely related, while each codigester also contained a set of unique 13C-enriched GBs (Figure 4). GBs that formed sister groups based on both the conserved marker gene alignments and UPGMA clustering of in silico DNA–DNA hybridization distance values included Syntrophomonas sp. PF07 and Syntrophomonas sp. CF14, as well as Syntrophomonas sp. PF15 and Syntrophomonas sp. CF22 (Figure 4; Supplementary Figure 9). Syntrophomonas sp. PF103 was consistently clustered with the reference genomes of Syntrophomonas wolfei subsp. wofei and Syntrophomonas wolfei subsp. methylbutyrica (Figure 4; Supplementary Figure 9). Firmicutes sp. CF263 was placed on a branch outside of Syntrophomonadaceae-affiliated genomes based on marker gene alignments (Figure 4), yet it clustered with the reference genome of Syntrophomonas zehnderi OL-4 based on in silico DNA–DNA hybridization (Supplementary Figure 9). Both GBs of Firmicutes sp. PF36 and Candidatus Parcubacteria sp. CF250 were placed as outgroups in both phylogenetic clustering approaches (Figure 4, Supplementary Figure S9).
Phylogenetic overview of differentially abundant genome bins identified in both anaerobic codigester DNA-SIP metagenomes. The tree was constructed based on a concatenated alignment of conserved marker genes within the metagenomic contigs of each bin using PhyloPhlAn v.1.3 (Segata et al., 2013). The color of each genome bin node represents the codigester biomass source, and the height of the outer bars represent the genome bin coverage in the heavy gradient fractions of the 13C-incubated samples and 12C-controls. The tree was illustrated using GraPhLan v.0.9.7 (Asnicar et al., 2015).
The largest GB coverage fold-change in the 13C-incubated samples was observed for the sister group of Syntrophomonas sp. PF15 & Syntrophomonas sp. CF22, which were not closely related to previously-sequenced Syntrophomonas isolates (Figure 4; Supplementary Table 2). Other GBs that had high fold-changes in the 13C-incubated samples (log2 change >3.0) included Syntrophomonas sp. PF75, Syntrophomonas sp. PF62, Syntrophomonas sp. PF103, Firmicutes sp. PF36, Syntrophomonas sp. PF111, and Syntrophomonas sp. CF196 (Figure 4; Supplementary Table 2). In comparison to the CF codigester, the PF codigester had a greater number of GBs that were enriched above a log2 change of 3.0 in the 13C-incubated samples (Figure 4).
To assess the functional potential of differentially abundant GBs in 13C-incubated samples, coverage values were extracted and summarized for KEGG ECs within the LCFA degradation pathway (KEGG Pathway Map 00071). Long-chain fatty acid CoA ligase (EC:6.2.1.3), which is responsible for activating free-fatty acids to acyl-CoA thioesters (Sousa et al., 2009), had the highest EC coverage in differentially abundant GBs within 13C-samples from both codigesters (Figure 5). The EC categories with the next highest coverage were enoyl-CoA hydratase (EC:4.2.1.17) and acetyl-CoA acetyltransferase (EC:2.3.1.9). Genes encoding for the entire fatty acid β-oxidation cycle were identified within the differentially abundant GBs for both codigesters (Figure 5). However, genes for acyl-CoA dehydrogenase (EC: 1.3.99.-) were observed in the PF codigester GBs but not in the CF codigester GBs (Figure 5). The GBs in the PF codigester also had substantially higher gene coverage for 3-hydroxyacyl-CoA dehydrogenases (EC:1.1.1.35 and EC:1.1.1.36) in 13C-incubated samples than the CF codigester GBs (Figure 5). Additionally, genes encoding for Δ2,Δ4-dienoyl-CoA reductase (EC:1.3.1.34) involved in the metabolism of unsaturated fatty acids (Ren et al., 2004) were detected in the differentially abundant GBs for both codigesters, and had a coverage that was an order of magnitude higher in the PF codigester GBs (Supplementary Table 3).
Cumulative coverage values of KEGG ECs potentially involved in LCFA degradation (KEGG map 00071), based on the coverage of all differentially abundant genome bins in the 13C-incubated DNA-SIP metagenomes. Values from duplicate biological replicates are shown for both codigesters, and the size of each marker is proportional to the log10 of the EC read coverage.
Syntrophic conversion of LCFA under methanogenic conditions involves interspecies electron transfer between Bacteria and Archaea, which commonly involves hydrogen and/or formate as electron shuttles (Stams and Plugge, 2009). Several formate dehydrogenases and hydrogenases were detected in the enriched GBs of both codigesters (Supplementary Figure 10). Formate dehydrogenases (EC:1.2.1.2, EC:1.2.1.43) were more abundant than hydrogenases (EC:1.12.1.2, EC:1.12.7.2, EC:1.12.1.3), and had similar coverage in the differentially abundant GBs from both codigesters (Supplementary Figure 10). In contrast, hydrogenases (EC:1.12.1.2, EC:1.12.1.3) had lower coverage in the CF codigester GBs relative to the PF codigester (Supplementary Figure 10).
Discussion
This research conducted a genome-centric comparison of the active populations between two anaerobic codigesters that were either pulse-fed or continuously-fed with oleate. DNA-SIP was combined with shotgun metagenomic sequencing and differential coverage binning to provide novel insights into active LCFA-degrading populations, and highlighted the potential for physiological diversity at the trophic level of syntrophy within anaerobic digesters.
As the average specific growth rate (μ) in the codigesters was 0.05 d−1 based on the 20-d SRT, higher growth rates would have been needed for cell duplication and sufficient DNA labeling during the 6-d incubation time, which could have been promoted by batch feeding oleate at initial concentrations of 8 mM in the SIP incubations. A growth rate of 0.11 d−1 would be required for cell doubling within a 6-d period, which is within the range of maximum specific growth rates (μmax) for co-cultures of syntrophic fatty acid oxidizing bacteria and methanogenic archaea of 0.10 to 0.19 d−1 (Stams et al., 2012). While a longer incubation time would have contributed to further DNA labeling, longer incubation times in DNA-SIP can be problematic because they can lead to cross-labeling of peripheral populations due to endogenous cell decay (Neufeld et al., 2006). In contrast to our study, previous studies employing DNA-SIP for syntrophic propionate- and butyrate-degrading communities in soil utilized incubation times ranging from 17 to 69 days (Lueders et al., 2004; Chauhan and Ogram, 2006; Liu et al., 2011; Gan et al., 2012; Li et al., 2015b). These longer incubation times may have been necessitated due to the low active biomass fractions in soil, which would result in low substrate conversion rates. The higher active syntrophic biomass fraction in the anaerobic codigester sludges in this study promoted more rapid oleate bioconversion in the microcosms, which permitted efficient DNA-SIP labeling within a 6- day incubation period. DNA-SIP based metagenomics could thus be potentially applied with relatively short SIP incubation times to study other biological wastewater treatment process that have relatively high active biomass fractions.
Syntrophic fatty acid conversion requires that free energy from catabolism is shared between syntrophic bacteria and methanogenic archaeal partners (Schink, 1997; Stams and Plugge, 2009). In this study, all of the 16S rRNA gene OTUs that were differentially abundant in 13C-incubated DNA-SIP samples for both codigesters belonged to the domain Bacteria, while no Archaea OTUs were differentially abundant in 13C-incubated samples. Likewise, all of the differentially abundant GBs recovered from the DNA-SIP shotgun metagenomes were assigned to bacterial taxa. However, the higher cumulative 16S rRNA gene read counts of Methanosaeta and Methanobacterium in 13C-incubated samples relative to 12C-controls indicates that they were likely methanogenic partners involved in syntrophic oleate degradation in both codigesters, but the increases in their OTU abundances were not statistically significant. The relatively high level of 12C-background substrates for methanogenesis (from the digester inoculum) likely diluted the 13C-labeled DNA of methanogenic archaea, as was also observed with protein-SIP in anaerobic digester communities (Mosbæk et al., 2016).
To the best of our knowledge, this is the first study to couple DNA-SIP with differential coverage binning to recover population GBs from environmental metagenomes. Differential abundance analysis of GB read counts was an effective technique coupled to DNA-SIP metagenomic binning to identify GBs that were significantly enriched within the heavy gradient fractions of 13C-incubated samples versus 12C-controls. The taxonomic assignment of differentially abundant GBs agreed with the results of 16S rRNA gene amplicon sequencing, as over 70% of the GBs belonged to the genus Syntrophomonas. This finding corroborates previous indications that Syntrophomonas are key syntrophic LCFA-degrading bacteria in mesophilic anaerobic digesters (Shigematsu et al., 2006; Hatamoto et al., 2007; Sousa et al., 2007, 2009; Ziels et al., 2015). However, this is the first study to determine the relative contribution of Syntrophomonas toward LCFA degradation using DNA-SIP. The finding that some phylogenetically distinct Syntrophomonas GBs and 16S rRNA gene OTUs were unique to each codigester further suggests that microbial communities involved in LCFA conversion can be physiologically diverse, and that the LCFA feeding frequency selected for unique active LCFA-degrading syntrophic taxa. Syntrophic bacteria have traditionally been considered metabolic-specialists (McInerney et al., 2009; Sieber et al., 2012), which has led to the general conclusion that they have low functional redundancy in anaerobic digesters (Werner et al., 2011; Carballa et al., 2015). A previous genome-centric analysis of anaerobic digester metagenomes found low diversity within the trophic level of syntrophic acetogenesis (Vanwonterghem et al., 2016). Using transcriptomic read mapping, Treu et al. (2016a) could only identify two active Syntrophomonas GBs in an anaerobic digester after pulsing oleate. In this study, 12 active Syntrophomonas GBs were identified using targeted DNA-SIP metagenomic analysis. DNA-SIP based metagenomic binning may thus improve our resolution of anaerobic digester trophic groups by enriching metagenome libraries with actively growing population genomes (Coyotzi et al., 2016), which may be especially useful to study low-abundance groups like syntrophic bacteria (Stams et al., 2012). The recent recovery of 236 microbial GBs from anaerobic digester metagenomes led to conclusion that Syntrophomonas were among a core essential microbial group that is present independent of digester operational conditions (Treu et al., 2016b). Our finding that the anaerobic digester feeding frequency can impact the genomic composition of Syntrophomonas thus has implications for developing operating strategies that steer the core digester community to obtain better biokinetics and process stability.
In prior research, it was observed that the frequency of LCFA feeding led to different biokinetics between the PF and CF codigesters, which were correlated with individual Syntrophomonas taxa abundances between the two systems (Ziels et al., 2017). In this study, biokinetic modeling of methane production in the DNA-SIP microcosms fed with either 13C- or 12C- oleate showed that the PF codigester sludge had higher maximum oleate conversion rates than CF codigester sludge during all three DNA-SIP batch degradation periods. This result further corroborates that biological selection from the LCFA feeding frequency altered LCFA bioconversion kinetics in the anaerobic codigesters (Ziels et al., 2017). The different oleate bioconversion kinetics observed between the two codigesters could have been attributed to differences in the functional gene repertoires of their active LCFA-degrading populations. The differentially abundant GBs in the PF codigester had more redundant KEGG ECs (for example, multiple ECs for a single reaction step) in the LCFA degradation pathway compared to the CF codigester GBs, which could indicate that pulse feeding led to a higher level of metabolic flexibility. The PF codigester GBs also had a greater distribution of formate dehydrogenase and hydrogenase encoding genes, which have been suggested to afford syntrophs with metabolic flexibility in fluctuating environmental conditions (Meyer et al., 2013; Sieber et al., 2014). The greater metabolic flexibility identified in the 13C-enriched GBs of the PF codigester in this study suggests that pulse feeding could promote a greater physiological adaptive capacity by supporting a higher level of functional redundancy within the ecological niches of anaerobic digesters, which in turn may have supported its higher oleate loading tolerance (Ziels et al., 2017). A greater tolerance to overloading and toxicity has also been observed in other pulse-fed anaerobic digesters compared to continuous-fed systems (Conklin et al., 2006; De Vrieze et al., 2013; Mulat et al., 2016). Although stochastic processes cannot be ruled out because the bioreactors were not replicated, the fact that these bioreactors were inoculated with the same seed source and Syntrophomonas was the predominant group impacted by oleate addition for over 230 days (Ziels et al., 2017) indicates that the feeding frequency likely imparted a deterministic selective pressure on the syntrophic community structure.
In summary, this study demonstrated the application of DNA-SIP combined with metagenomics and differential coverage binning to identify active populations in an engineered biological treatment process. The results indicated that the operating conditions, specifically the feeding frequency, of the methanogenic bioreactors impacted the genomic composition of active syntrophic populations. The feeding frequency also impacted the level of functional gene redundancy within the active populations, which had implications for increased physiological adaptation using a pulse feeding strategy. Thus, DNA-SIP metagenomics is a powerful tool to help understand how operating strategies may improve the performance of biological wastewater treatment processes, as was shown here for increased methane recovery from high-strength organic wastes.
References
Albertsen M, Hugenholtz P, Skarshewski A, Nielsen KL, Tyson GW, Nielsen PH . (2013). Genome sequences of rare, uncultured bacteria obtained by differential coverage binning of multiple metagenomes. Nat Biotechnol 31: 533–538.
Alves MM, Pereira MA, Sousa DZ, Cavaleiro AJ, Picavet M, Smidt H et al. (2009). Waste lipids to energy: how to optimize methane production from long-chain fatty acids (LCFA). Microb Biotechnol 2: 538–550.
Angelidaki I, Ahring BK . (1992). Effects of free long-chain fatty acids on thermophilic anaerobic digestion. Appl Microbiol Biotechnol 37: 808–812.
Asnicar F, Weingart G, Tickle TL, Huttenhower C, Segata N . (2015). Compact graphical representation of phylogenetic data and metadata with GraPhlAn. Peer J 3: e1029.
Auch AF, von Jan M, Klenk H-P, Göker M . (2010). Digital DNA-DNA hybridization for microbial species delineation by means of genome-to-genome sequence comparison. Stand Genomic Sci 2: 117–134.
Briones A, Raskin L . (2003). Diversity and dynamics of microbial communities in engineered environments and their implications for process stability. Curr Opin Biotechnol 14: 270–276.
Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJA, Holmes SP . (2016). DADA2: High-resolution sample inference from Illumina amplicon data. Nat Methods 13: 581–583.
Carballa M, Regueiro L, Lema JM . (2015). Microbial management of anaerobic digestion: exploiting the microbiome-functionality nexus. Curr Opin Biotechnol 33: 103–111.
Chauhan A, Ogram A . (2006). Fatty acid-oxidizing consortia along a nutrient gradient in the Florida Everglades. Appl Environ Microbiol 72: 2400–2406.
Chemerys A, Pelletier E, Cruaud C, Martin F, Violet F, Jouanneau Y . (2014). Characterization of novel polycyclic aromatic hydrocarbon dioxygenases from the bacterial metagenomic DNA of a contaminated soil. Appl Environ Microbiol 80: 6591–6600.
Chen I-MA, Markowitz VM, Chu K, Palaniappan K, Szeto E, Pillay M et al. (2016). IMG/M: integrated genome and metagenome comparative data analysis system. Nucleic Acids Res 45: D507–D516.
Chen Y, Murrell JC . (2010). When metagenomics meets stable-isotope probing: progress and perspectives. Trends Microbiol 18: 157–163.
Conklin A, Stensel HD, Ferguson J . (2006). Growth kinetics and competition between methanosarcina and methanosaeta in mesophilic anaerobic digestion. Water Environ Res 78: 486–496.
Coyotzi S, Pratscher J, Murrell JC, Neufeld JD . (2016). Targeted metagenomics of active microbial populations with stable-isotope probing. Curr Opin Biotechnol 41: 1–8.
Davidsson Å, Lövstedt C, la Cour Jansen J, Gruvberger C, Aspegren H . (2008). Co-digestion of grease trap sludge and sewage sludge. Waste Manag 28: 986–992.
De Vrieze J, Verstraete W, Boon N . (2013). Repeated pulse feeding induces functional stability in anaerobic digestion. Microb Biotechnol 6: 414–424.
Dumont MG, Murrell JC . (2005). Stable isotope probing — linking microbial identity to function. Nat Rev Microbiol 3: 499–504.
Edgar RC . (2013). UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods 10: 996–998.
Eyice Ö, Namura M, Chen Y, Mead A, Samavedam S, Schäfer H . (2015). SIP metagenomics identifies uncultivated Methylophilaceae as dimethylsulphide degrading bacteria in soil and lake sediment. ISME J 9: 2336–2348.
Gan Y, Qiu Q, Liu P, Rui J, Lu Y . (2012). Syntrophic oxidation of propionate in rice field soil at 15 and 30 °C under methanogenic conditions. Appl Environ Microbiol 78: 4923–4932.
Guo M, Song W, Buhain J . (2015). Bioenergy and biofuels: History, status, and perspective. Renew Sustain Energy Rev 42: 712–725.
Hanaki K, Matsuo T, Nagase M . (1981). Mechanism of inhibition caused by long-chain fatty acids in anaerobic digestion process. Biotechnol Bioeng 23: 1591–1610.
Hatamoto M, Imachi H, Ohashi A, Harada H . (2007). Identification and cultivation of anaerobic, syntrophic long-chain fatty acid-degrading microbes from mesophilic and thermophilic methanogenic sludges. Appl Environ Microbiol 73: 1332–1340.
Huson DH, Auch AF, Qi J, Schuster SC . (2007). MEGAN analysis of metagenomic data. Genome Res 17: 377–386.
Kang DD, Froula J, Egan R, Wang Z . (2015). MetaBAT, an efficient tool for accurately reconstructing single genomes from complex microbial communities. PeerJ 3: e1165.
Karst SM, Kirkegaard RH, Albertsen M . (2016). mmgenome: a toolbox for reproducible genome extraction from metagenomes. BioRxiv, doi:https://doi.org/10.1101/059121.
Li D, Liu C-M, Luo R, Sadakane K, Lam T-W . (2015a). MEGAHIT: an ultra-fast single-node solution for large and complex metagenomics assembly via succinct de Bruijn graph. Bioinformatics 31: 1674–1676.
Li H, Chang J, Liu P, Fu L, Ding D, Lu Y . (2015b). Direct interspecies electron transfer accelerates syntrophic oxidation of butyrate in paddy soil enrichments. Environ Microbiol 17: 1533–1547.
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N et al. (2009). The Sequence Alignment/Map format and SAMtools. Bioinformatics 25: 2078–2079.
Liu P, Qiu Q, Lu Y . (2011). Syntrophomonadaceae-affiliated species as active butyrate-utilizing syntrophs in paddy field soil. Appl Environ Microbiol 77: 3884–3887.
Love MI, Huber W, Anders S . (2014). Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15: 1.
Lueders T, Pommerenke B, Friedrich MW . (2004). Stable-isotope probing of microorganisms thriving at thermodynamic limits: syntrophic propionate oxidation in flooded soil. Appl Environ Microbiol 70: 5778–5786.
Luostarinen S, Luste S, Sillanpää M . (2009). Increased biogas production at wastewater treatment plants through co-digestion of sewage sludge with grease trap sludge from a meat processing plant. Bioresour Technol 100: 79–85.
McInerney MJ, Sieber JR, Gunsalus RP . (2009). Syntrophy in anaerobic global carbon cycles. Curr Opin Biotechnol 20: 623–632.
Meyer B, Kuehl JV, Deutschbauer AM, Arkin AP, Stahl DA . (2013). Flexibility of syntrophic enzyme systems in desulfovibrio species ensures their adaptation capability to environmental changes. J Bacteriol 195: 4900–4914.
Mosbæk F, Kjeldal H, Mulat DG, Albertsen M, Ward AJ, Feilberg A et al. (2016). Identification of syntrophic acetate-oxidizing bacteria in anaerobic digesters by combined protein-based stable isotope probing and metagenomics. ISME J 10: 2405–2418.
Mulat DG, Jacobi HF, Feilberg A, Adamsen APS, Richnow H-H, Nikolausz M . (2016). Changing feeding regimes to demonstrate flexible biogas production: effects on process performance, microbial community structure, and methanogenesis pathways. Appl Environ Microbiol 82: 438–449.
Neufeld JD, Dumont MG, Vohra J, Murrell JC . (2006). Methodological considerations for the use of stable isotope probing in. Microb Ecol 53: 435–442.
Oksanen J, Kindt R, Legendre P, O’Hara B, Stevens MHH, Oksanen MJ et al. (2007). The vegan package. Community Ecol Package 10: 631–637.
Parada AE, Needham DM, Fuhrman JA . (2016). Every base matters: assessing small subunit rRNA primers for marine microbiomes with mock communities, time series and global field samples. Environ Microbiol 18: 1403–1414.
Parks DH, Imelfort M, Skennerton CT, Hugenholtz P, Tyson GW . (2015). CheckM: assessing the quality of microbial genomes recovered from isolates, single cells, and metagenomes. Genome Res 25: 1043–1055.
Radajewski S, Ineson P, Parekh NR, Murrell JC . (2000). Stable-isotope probing as a tool in microbial ecology. Nature 403: 646–649.
Ren Y, Aguirre J, Ntamack AG, Chu C, Schulz H . (2004). An alternative pathway of oleate β-oxidation in Escherichia coli involving the hydrolysis of a dead end intermediate by a thioesterase. J Biol Chem 279: 11042–11050.
Rinzema A, Boone M, Knippenberg K, van, Lettinga G . (1994). Bactericidal effect of long chain fatty acids in anaerobic digestion. Water Environ Res 66: 40–49.
Saidi-Mehrabad A, He Z, Tamas I, Sharp CE, Brady AL, Rochman FF et al. (2013). Methanotrophic bacteria in oilsands tailings ponds of northern Alberta. ISME J 7: 908–921.
Schink B . (1997). Energetics of syntrophic cooperation in methanogenic degradation. Microbiol Mol Biol Rev 61: 262–280.
Segata N, Börnigen D, Morgan XC, Huttenhower C . (2013). PhyloPhlAn is a new method for improved phylogenetic and taxonomic placement of microbes. Nat Commun 4: 2304.
Shigematsu T, Tang Y, Mizuno Y, Kawaguchi H, Morimura S, Kida K . (2006). Microbial diversity of mesophilic methanogenic consortium that can degrade long-chain fatty acids in chemostat cultivation. J Biosci Bioeng 102: 535–544.
Sieber JR, Le HM, McInerney MJ . (2014). The importance of hydrogen and formate transfer for syntrophic fatty, aromatic and alicyclic metabolism. Environ Microbiol 16: 177–188.
Sieber JR, McInerney MJ, Gunsalus RP . (2012). Genomic insights into syntrophy: the paradigm for anaerobic metabolic cooperation. Annu Rev Microbiol 66: 429–452.
Sousa DZ, Pereira MA, Smidt H, Stams AJM, Alves MM . (2007). Molecular assessment of complex microbial communities degrading long chain fatty acids in methanogenic bioreactors. FEMS Microbiol Ecol 60: 252–265.
Sousa DZ, Smidt H, Alves MM, Stams AJM . (2009). Ecophysiology of syntrophic communities that degrade saturated and unsaturated long-chain fatty acids. FEMS Microbiol Ecol 68: 257–272.
Stams AJM, Plugge CM . (2009). Electron transfer in syntrophic communities of anaerobic bacteria and archaea. Nat Rev Microbiol 7: 568–577.
Stams AJM, Sousa DZ, Kleerebezem R, Plugge CM . (2012). Role of syntrophic microbial communities in high-rate methanogenic bioreactors. Water Sci Technol 66: 352.
Sundberg C, Al-Soud WA, Larsson M, Alm E, Yekta SS, Svensson BH et al. (2013). 454 pyrosequencing analyses of bacterial and archaeal richness in 21 full-scale biogas digesters. FEMS Microbiol Ecol 85: 612–626.
Treu L, Campanaro S, Kougias PG, Zhu X, Angelidaki I . (2016a). Untangling the effect of fatty acid addition at species level revealed different transcriptional responses of the biogas microbial community members. Environ Sci Technol 50: 6079–6090.
Treu L, Kougias PG, Campanaro S, Bassani I, Angelidaki I . (2016b). Deeper insight into the structure of the anaerobic digestion microbial community; the biogas microbiome database is expanded with 157 new genomes. Bioresour Technol 216: 260–266.
Vanwonterghem I, Jensen PD, Ho DP, Batstone DJ, Tyson GW . (2014). Linking microbial community structure, interactions and function in anaerobic digesters using new molecular techniques. Curr Opin Biotechnol 27: 55–64.
Vanwonterghem I, Jensen PD, Rabaey K, Tyson GW . (2016). Genome-centric resolution of microbial diversity, metabolism and interactions in anaerobic digestion. Environ Microbiol 18: 3144–3158.
Verastegui Y, Cheng J, Engel K, Kolczynski D, Mortimer S, Lavigne J et al. (2014). Multisubstrate isotope labeling and metagenomic analysis of active soil bacterial communities. mBio 5: e01157–14.
Wan C, Zhou Q, Fu G, Li Y . (2011). Semi-continuous anaerobic co-digestion of thickened waste activated sludge and fat, oil and grease. Waste Manag 31: 1752–1758.
Wang L, Aziz TN, de los Reyes FL . (2013). Determining the limits of anaerobic co-digestion of thickened waste activated sludge with grease interceptor waste. Water Res 47: 3835–3844.
Werner JJ, Knights D, Garcia ML, Scalfone NB, Smith S, Yarasheski K et al. (2011). Bacterial community structures are unique and resilient in full-scale bioenergy systems. Proc Natl Acad Sci USA 108: 4158–4163.
Zhang F, Liu X, Dong X . (2012). Thermosyntropha tengcongensis sp. nov., a thermophilic bacterium that degrades long-chain fatty acids syntrophically. Int J Syst Evol Microbiol 62: 759–763.
Ziels RM, Beck DAC, Martí M, Gough HL, Stensel HD, Svensson BH . (2015). Monitoring the dynamics of syntrophic β-oxidizing bacteria during anaerobic degradation of oleic acid by quantitative PCR. FEMS Microbiol Ecol 91: fiv028.
Ziels RM, Karlsson A, Beck DAC, Ejlertsson J, Yekta SS, Bjorn A et al. (2016). Microbial community adaptation influences long-chain fatty acid conversion during anaerobic codigestion of fats, oils, and grease with municipal sludge. Water Res 103: 372–382.
Ziels RM, Beck DAC, Stensel HD . (2017). Long-chain fatty acid feeding frequency in anaerobic codigestion impacts syntrophic community structure and biokinetics. Water Res 111: 218–229.
Acknowledgements
We thank David Stahl for his helpful input and review of this manuscript. This research was funded by the US EPA STAR [grant RD835567] (RMZ and HDS), the US DOE JGI [grant CSP 2791] (DACB), and the ERC [grant 323009] (DZS).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Ziels, R., Sousa, D., Stensel, H. et al. DNA-SIP based genome-centric metagenomics identifies key long-chain fatty acid-degrading populations in anaerobic digesters with different feeding frequencies. ISME J 12, 112–123 (2018). https://doi.org/10.1038/ismej.2017.143
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2017.143