Abstract
Nematode predation has important roles in determining bacterial community composition and dynamics, but the extent of the effects remains largely rudimentary, particularly in natural environment settings. Here, we investigated the complex microbial–microfaunal interactions in the rhizosphere of maize grown in red soils, which were derived from four long-term fertilization regimes. Root-free rhizosphere soil samples were separated into three aggregate fractions whereby the abundance and community composition were examined for nematode and total bacterial communities. A functional group of alkaline phosphomonoesterase (ALP) producing bacteria was included to test the hypothesis that nematode grazing may significantly affect specific bacteria-mediated ecological functions, that is, organic phosphate cycling in soil. Results of correlation analysis, structural equation modeling and interaction networks combined with laboratory microcosm experiments consistently indicated that bacterivorous nematodes enhanced bacterial diversity, and the abundance of bacterivores was positively correlated with bacterial biomass, including ALP-producing bacterial abundance. Significantly, such effects were more pronounced in large macroaggregates than in microaggregates. There was a positive correlation between the most dominant bacterivores Protorhabditis and the ALP-producing keystone 'species' Mesorhizobium. Taken together, these findings implicate important roles of nematodes in stimulating bacterial dynamics in a spatially dependent manner.
Similar content being viewed by others
Introduction
Resource competition and predation are the two major driving forces underlying the dynamic changes of species composition in the biological community (Chesson and Kuang, 2008). For microorganisms inhabiting soil, the importance of resource competition has been well documented, particularly with the improved accessibility of next-generation sequencing and stable isotope techniques (Bulgarelli et al., 2013). Although the importance of predation by microfauna has long been recognized, the potential effects of predation remain poorly defined. Few studies have addressed the complex microbial–microfaunal interactions in open-field environments (Neher, 2010). For example, nematodes can stimulate microbial activity, resulting in either an increase or a decrease of microbial biomass in microcosm experiments (Trap et al., 2016). The grazing-induced influences on microbial abundance vary according to pore structure and distribution of the accessible resources such as soil organic matter and plant roots (Rønn et al., 2012).
Soils have a complex hierarchical structure including pore distribution and aggregates. Soil aggregates provide spatially heterogeneous habitats for microorganisms, which vary in nutrient availability, water potential and oxygen concentration as well as predation pressure (Ranjard, Richaume, 2001; Jiang et al., 2013). Aggregate fractions are assembled by organic matter and mineral particles (Tisdall and Oades, 1982). Macroaggregates normally contain more labile substrates predominantly derived from plant residues (Bronick and Lal, 2005), and harbor higher amounts of fungal biomass than microaggregates (MAs, Rillig and Mummey, 2006). In contrast, MAs are characterized by the highest concentration of stable organic carbon, and more importantly, they provide a protective microenvironment for microbial growth (Six et al., 2000). The relative small pore sizes make MAs inaccessible to large-sized bacterial-feeding nematodes (typically 30–90 μm in diameter; Quénéhervé and Chotte, 1996). Thus, it is crucial to understand the predator–prey interactions at the aggregate level (Ettema and Wardle, 2002).
Of particular significance is the rhizosphere soil as it functions as the critical interface for resource exchange between plants and soil. Compared with bulk soil, the rhizosphere contains a large amount of small organic compounds excreted from the roots of living plants, supporting high levels of microbial activity. Microorganisms are integral to the cycling of nutritional elements such as nitrogen and phosphorus (van der Heijden et al., 2008), and bacterial-feeding nematodes predation releases nutrients sequestered in bacterial biomass in the rhizosphere niche (Bonkowski et al., 2009). However, nematodes normally have special food preferences, and bacteria were not equally susceptible to predation by nematodes. Selective grazing by nematodes can alter the bacterial community composition (Djigal et al., 2004). This leads to an important but unexplored hypothesis that nematode grazing affects total bacterial community and specific functional groups such as phosphorus (P) cycling in different manners.
Microbial–microfaunal interactions via the microbial loop determine the rate of P cycling in the rhizosphere (Bonkowski, 2004). However, the specific effects of nematode grazing on phosphate solubilizing microbial community remain poorly understood. Phosphorus is one of the most limiting nutrients in agricultural soils (Tabatabai, 1994). Predominant enzymes involved in organic P mineralization are alkaline phosphomonoesterases (ALPs, EC 3.1.3.1) and acid phosphomonoesterases (ACPs, EC 3.1.3.2) (Nannipieri et al., 2011; Chang et al., 2015). ALPs are mostly of bacterial origin, whereas ACPs are mainly excreted by plant roots and fungi in the rhizosphere (Tabatabai, 1994; Spohn and Kuzyakov, 2013). Significantly, ALP-producing bacterial community can be quantitatively analyzed using phoD gene as a molecular marker (Sakurai et al., 2008). The phoD gene abundance is positively correlated with ALP activity as revealed by field studies with bulk soil (Tan et al., 2013; Fraser et al., 2015a, b).
Here, we investigated the reciprocal interactions between nematodes and bacteria in rhizosphere soil at the aggregate level, with an additional specific focus on ALP-producing bacteria. To this end, we performed a 13-year field experiment with red soils (Acrisol) under four fertilization regimes. Soil samples were taken from the rhizosphere of maize, and then separated into three aggregate size fractions for physiochemical and microbiological analyses. The nematode assemblages were quantitatively assessed under microscope, whereas the abundance and composition of bacterial community were examined using phospholipid fatty acid analysis and Illumina sequencing of 16S rRNA gene, respectively. Next, the abundance and composition of ALP-producing bacterial community were estimated using quantitative polymerase chain reaction (PCR) and Illumina sequencing of phoD gene. We observed significantly positive influences of nematodes on bacterial abundance and activity, which were subsequently confirmed via pot experiments under well-controlled laboratory conditions. Our findings provided insights into the microbial–microfaunal interactions in the rhizosphere at the soil aggregate level.
Materials and methods
Site description and design
The long-term fertilization experiment was conducted at the National Agro-Ecosystem Observation and Research Station in a subtropical humid monsoon climate region China (Yingtan, 28°15′N, 116°55′E) with an annual average temperature 17.6 °C and precipitation 1795 mm. The soil is an acid loamy clay-derived Quaternary red clay (Udic Ferralsols in the Chinese Soil Taxonomy and Ferric Acrisols in the FAO classification system).
Twelve concrete lysimeters, 2 m wide × 2 m long × 1.5 m deep, were used in the manure experiment since 2002. Four pig manure rates were compared in a completely randomized design with three replicates: (1) no manure (M0); (2) low manure with 150 kg N ha−1 y−1 (M1); (3) high manure with 600 kg N ha−1 y−1 (M2); and (4) high manure with 600 kg N ha−1 y−1 and lime (M3; Ca(OH)2 applied once every 3 years at 3000 kg ha−1). The pig manure contained an average total carbon of 386.5 g kg−1, total nitrogen of 36.2 g kg−1 and total phosphorus of 21.6 g kg−1 on a dry matter basis. The monoculture maize (Zea mays L.), cultivar No.11 from Denghai, was planted annually in April and harvested in July from 2002 to 2014. There were no tillage and management measures with the exception of manual weeding.
Soil sampling and aggregate fractionation
Soil sampling was conducted in late July 2014 after 13 years of fertilization. Rhizosphere soils were collected from each plot at a depth of 0−15 cm, and then were placed on ice and immediately transported to the laboratory. After shaking off the loosely adhering soil, the tightly adhering rhizosphere soil was collected with a brush, passed through a 4 mm sieve. Next, 100 g root-free soil was manually fractionated through a series of two sieves (2000 μm and 250 μm) into three aggregate sizes: large macroaggregates (>2000 μm; LMA), small macroaggregates (250–2000 μm; SMA) and MA (<250 μm; Jiang et al., 2014). Each aggregate fraction was homogenized for chemical and biological analyses. Standard methods were used to characterize soil chemical properties and phosphomonoesterase activities (Supplementary Appendix).
Characterization of total bacteria and ALP-producing bacteria
A modified method of phospholipid fatty acids (PLFAs) analysis was used to measure soil bacterial biomass, which is expressed as nanomoles of PLFA per gram of dry soil (Frostegård and Bååth, 1996). DNA was extracted from 0.5 g fresh soil using the Ultraclean Soil DNA Isolation Kit (Mo Bio Laboratories, Inc., Carlsbad, CA, USA) following the manufacturer’s instructions. Real-time quantitative PCR analysis of phoD gene was performed with primers ALPS-F730 and ALPS-R1101 (Sakurai et al., 2008). For high-throughput sequencing at the Illumina MiSeq platform, the V3–V4 region of 16S rRNA gene and phoD gene were amplified separately using primers 338 F/806 R and ALPS-F730/ALPS-R1101, respectively. Raw sequences were quality screened and trimmed, including quality trimming, de-multiplexing, taxonomic assignments, chimera detection, and screening for frame shifts. Thereafter, the 16S rDNA and phoD sequences were subjected to a similarity search against the Ribosomal Database Project database and the GenBank non-redundant nucleotide database, respectively. Finally, the sequence reads from each sample were clustered to operational taxonomic units (OTUs) at 97% similarity. Alpha diversity metrics and community relatedness were calculated at the same sequencing depth. The 16S rDNA and phoD sequences are available at the NCBI Sequence Read Archive under accession number SRP090422 and SRP044878, respectively (see details in Supplementary Appendix).
Nematode faunal composition
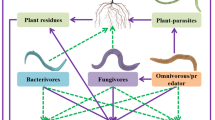
Nematodes were extracted using a modified Baermann funnel method (Barker, 1985), and visually examined with an inverted compound microscope. At least 100 nematodes were identified to the genus level for each sample. Nematodes were divided into four trophic groups: bacterivores (Ba), fungivores (Fu), plant parasites (Pp) and omnivores-predators, characterized by known feeding habitats or stoma and esophageal morphology (Yeates et al., 1993). The guilds were characterized on the colonizer-persister (c−p) scale (1−5) as previously described (Bongers and Bongers, 1998).
Microcosm experiment
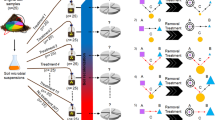
Rhizosphere soils were sterilized by acute gamma irradiation at 40 kGy doses (Buchan et al., 2012). Bacterial suspensions of fresh soils were prepared by passing through 1 μm pore-size Millipore filters (Millipore, Bedford, MA, USA) whereby nematodes and other small eukaryotes were eliminated. The dominant bacterivorous nematode, Protorhabditis spp., isolated from the experimental site was cultivated in nematode growth medium at 28 °C by feeding on Escherichia coli. Before use, nematodes were washed five times with sterile distilled water to minimize the effects of E. coli.
To set up the microcosms, 50, 150, 500 and 600 individuals were introduced into 100 g soil per pot for soils obtained from the M0, M1, M2 and M3 treatments, respectively. Nematode-free control was set up in triplicate to ensure no nematode contamination. Microcosms were incubated in the dark at 28 °C, with soil moisture being maintained at 25% (w/w). Soils were destructively sampled in 0, 3, 7, 14 and 21 days after inoculation, and then separated into three aggregate fractions for analysis of nematodes, ALP-producing bacteria, ALP and ACP activities.
Statistical analyses
The statistical procedures, including Pearson’s correlation analysis, were conducted by SPSS statistical software (SPSS Inc., Chicago, IL, USA). The aggregated boosted trees analysis was carried out to evaluate the effects of different factors on ALP-producing bacterial abundance and activity (De'Ath, 2007). The canonical analysis of principal coordinates was performed to assess the influence of different experimental factors on beta diversity (Anderson and Willis, 2003). Driving factors for nematodes and ALP-producing bacterial community composition were quantitatively evaluated using the permutational multivariate analysis of variance (ANOVA; Anderson, 2001). Structural equation modeling was used to understand how soil chemical properties altered nematodes and bacterial community in three aggregate fractions (Byrne, 2010). Interaction networks were constructed by calculating the pairwise Spearman’s rank correlations (see the details in Supplementary Appendix).
Results
Soil physiochemical properties at the aggregate level
Rhizosphere soils from the four fertilization treatments were separated into three aggregate fractions: LMA (>2 mm), SMA (0.25–2 mm) and MAs (<0.25 mm). Results of two-way ANOVA revealed significant differences in soil physiochemical properties among treatments (F(3,32)=64.59–1222.85, P<0.001) and aggregate fractions (F(2,33)=3.73–21.15, P<0.05). High levels of manure treatments (M2 and M3) contained higher proportions of LMA fractions compared with low manure treatment (M1) and the control (M0; Supplementary Figure 1a). Soil pH was significantly elevated by high manure application (Supplementary Figure 1b). The MA fraction tended to have higher nutritional substrates than the LMA and SMA fractions in terms of soil organic carbon (Supplementary Figure 1c), total nitrogen (Supplementary Figure 1d) and phosphate contents (Supplementary Figures 2a and b). The similar trend was found for soil enzymatic activities of ACP and ALP, respectively (Supplementary Figures 2c and d). Both ACP and ALP activities showed significant correlations with total phosphate (r=0.637 and r=0.953, respectively) and available phosphate (r=0.607 and r=0.958, respectively) at the level of P<0.001.
Characterizing the bacterial community in soil aggregates
Soil aggregate samples were subjected to PLFA analysis for bacterial biomass, and Illumina sequencing of 16S rRNA gene for the diversity and composition of bacterial community. The results indicated significant differences in fertilization treatments and aggregate fractions (Figure 1, P<0.05). Higher manure application resulted in higher bacterial biomass and diversity, showing a general trend of M3≈M2>M1>M0 (Figures 1a–c). The MA fraction possessed the highest bacterial biomass and diversity than the LMA fraction, with the SMA as the intermediates (Figures 1a–c).
Fertilization and aggregate fractions alter rhizosphere bacterial biomass and diversity. The biomass (a), diversity (b) and richness (c) of total bacterial community were examined together with the abundance (d), diversity (e) and richness (f) of the ALP-producing bacteria in the rhizosphere. Calculation of diversity and richness is based on OTU tables rarified to the same sequencing depth. Error bars represent standard errors of three replicates. Bars with the different letter (shown above each) are significantly different (P<0.05) by Tukey’s HSD test. ALP, alkaline phosphomonoesterase. M0, no manure; M1, low manure; M2, high manure; M3, high manure plus lime. HSD, honest significant difference; LMA, large macroaggregate; MA, microaggregate; SMA, small macroaggregate.
The bacterial communities were dominated by Chloroflexi (19.6%), Actinobacteria (19.0%), Alphaproteobacteria (11.3%), Firmicutes (10.3%), Acidobacteria (8.3%), Deltaproteobacteria (6.7%) and Gammaproteobacteria (5.1%; Figure 2a). In addition, Betaproteobacteria, Bacteroidetes, Cyanobacteria, Gemmatimonadetes, Planctomycetes and Verrucomicrobia were present at lower abundances, accounting for 14.7% of all sequences (Figure 2a). Bray-Curtis distances derived from a canonical analysis of principal coordinates were used to compare bacterial community composition between three aggregate fractions. Bacterial community composition in the MA fraction was well separated from those of the LMA and SMA fractions, mainly because of the higher abundance of Chloroflexi and Cyanobacteria but the lower abundance of Alphaproteobacteria, Gammaproteobacteria and Actinobacteria (Figure 3a). Finally, permutational multivariate ANOVA of bacterial community composition showed that 75.5% of variations could be explained by fertilization (59.3%) and aggregate fractions (14.2%; Supplementary Table 1).
Taxonomic compositions of bacterial community and bacterivores assemblages. The abundances of total bacterial (a) and ALP-producing bacterial (b) communities are based on the proportional frequencies of 16S rRNA- and phoD-like sequences. The abundance of bacterivores (c) is calculated in bacterivorous guilds. ALP, alkaline phosphomonoesterase. M0, no manure; M1, low manure; M2, high manure; M3, high manure plus lime. LMA, large macroaggregate; MA, microaggregate; SMA, small macroaggregate.
Aggregate fractions alter the bacterial community composition. The dominant OTU (relative abundance >0.1%) scores in (a) total bacterial and (b) ALP-producing bacterial community by principal coordinate analysis, which are constrained by aggregate fractions and based on Bray-Curtis distances among all the samples. The arrows point to the centroid of the constrained factor. Circle sizes correspond to the abundance of total bacterial and ALP-producing bacterial OTUs, and colors are assigned to different phyla/classes. ALP, alkaline phosphomonoesterase. M0, no manure; M1, low manure; M2, high manure; M3, high manure plus lime. LMA, large macroaggregate; MA, microaggregate; SMA, small macroaggregate.
Characterizing the ALP-producing bacterial community in soil aggregates
The abundance of ALP-producing bacteria was expressed as the phoD gene copy number per gram of dry soil as estimated by quantitative PCR analysis (Figure 1d). It followed a similar trend as total bacterial biomass, with relatively higher abundance under manure treatments (M3≈M2>M1>M0). There were significantly more ALP-producing bacteria in the MA fraction compared with the LMA and SMA fractions under M2 and M3 treatments. There was a significant positive correlation between ALP-producing bacterial abundance and ALP activity (r=0.965, P<0.001; Supplementary Figure 3). Both were most significantly influenced by soil pH, as indicted by aggregated boosted tree analysis (Supplementary Figure 4).
The diversity and composition of ALP-producing bacterial community were examined using Illumina sequencing of the phoD gene. Similar trends of the Shannon index and Chao1 richness were found for total bacteria and the specific functional group of ALP-producing bacteria: M3≈M2>M1>M0 for the effects of manure addition, and MA>SMA>LMA for variations among soil aggregates (Figure 1). The ALP-producing bacterial community was dominated by Alphaproteobacteria (45.4%), Actinobacteria (8.5%), Betaproteobacteria (7.5%) and Gammaproteobacteria (5.0%; Figure 2b). The ALP-producing bacterial communities from the three aggregate fractions were well separated by PCoA1 alone (78.8%, Figure 3b). There were significant differences among soil aggregates in regards to the abundance of ALP-producing Alphaproteobacteria (P=0.033) and Gammaproteobacteria (P=0.001; Supplementary Figure 5). The permutational multivariate ANOVA showed that the variations of ALP-producing bacterial community structure under fertilization treatments (61.5%) were much bigger than those among aggregate fractions (9.7%; Supplementary Table 1). This has been further demonstrated by hierarchical clustering analysis of dominant OTUs based on their co-occurrence (Supplementary Figure 6).
Investigating the nematode assemblages in soil aggregates
A total of 26 nematode genera were identified including nine bacterivores, five fungivores, four plant parasites and eight omnivores-predators (Supplementary Table 2). On average, bacterivores (46.6%) and plant parasites (30.5%) were the two most abundant trophic groups. The five dominant genera of nematodes were Protorhabditis, Cephalobus, Eucephalobus, Pratylenchus and Mesodorylaimus, cumulatively representing over 70% of all nematodes identified (Supplementary Table 2). The average number of total nematodes increased with increasing aggregate size, such that the LMA fraction had a significantly higher number than the SMA and MA fractions (Supplementary Table 2, P<0.05). The bacterivores, plant parasites and omnivores-predators, particularly the dominant genus within each guild (Protorhabditis, Pratylenchus and Mesodorylaimus, respectively), appeared to exhibit a general trend of LMA>SMA>MA, which was similar to that of the number of nematodes in total (Figure 2c,Supplementary Table 2). The permutational multivariate ANOVA showed that the nematode community structure was influenced by fertilization treatments (56.7%) and aggregate fractions (11.0%), which collectively explained 67.7% of the total variations (Supplementary Table 1).
Ecological interactions between nematodes and bacteria in soil aggregates
The abundance of bacterivorous nematodes were positively correlated with the two different measures of bacterial abundance, that is, total bacterial biomass (r=0.798, P<0.001) and ALP-producing bacterial abundance (r=0.783, P<0.001), and ALP activity (r=0.843, P<0.001), rather than ACP activity (r=0.318, P=0.059; Supplementary Figure 3,Supplementary Table 3). Intriguingly, the obtained data revealed positive correlations between bacterivore abundance and bacterial diversity: total bacterial diversity (r=0.655, P<0.001) and richness (r=0.430, P<0.001), as well as ALP-producing bacterial diversity (r=0.675, P<0.001) and richness (r=0.715, P<0.001; Supplementary Table 3). Furthermore, the community composition of total bacteria and ALP-producing bacteria were significantly affected by nematodes (21.0 and 22.3%), with the largest contribution from bacterivores (12.9 and 13.5%) (Supplementary Table 4).
The structural equation model was used to assess the effects of soil properties and nematodes on the bacterial community in the three aggregate fractions. Soil organic carbon produced the strongest effects on total bacterial biomass and community composition, while total bacterial diversity was primarily determined by soil pH (Figure 4). More importantly, bacterivores had positive effects on bacterial diversity (path coefficient: 0.35, P=0.017) and community composition (path coefficient: 0.48, P=0.003) in the LMA fraction (Figure 4c). For ALP-producing bacteria, soil pH was one of the most important causal factors determining ALP-producing bacterial abundance and ALP activity (Figures 4d−f). Bacterivores exerted more prominent contribution to ALP-producing bacterial abundance in the LMA fraction (path coefficient: 0.57, P<0.001) than in the SMA (path coefficient: 0.29, P=0.026) and MA (path coefficient: 0.32, P=0.022) fractions. Similar to total bacteria, the ALP-producing bacterial community composition was positively affected by bacterivores in the LMA fractions (path coefficient: 0.44, P<0.001; Figure 4f).
The effects of soil properties and nematodes on bacterial community as estimated using the structural equation model. For total bacteria, (a) microaggregates, χ2=7.969, GIF=0.977, P=0.527, AIC=37.692, RMSEA=0.001; (b) small macroaggregates, χ2=4.341, GIF=0.956, P=0.332, AIC=48.701, RMSEA=0.003; and (c) large macroaggregates, χ2=5.058, GIF=0.951, P=0.561, AIC=59.408, RMSEA=0.008. For ALP-producing bacteria, (d) microaggregates, χ2=5.949, GIF=0.957, P=0.486, AIC=57.496, RMSEA<0.001; (e) small macroaggregates, χ2=4.219, GIF=0.958, P=0.183, AIC=50.190, RMSEA=0.007; and (f) large macroaggregates, χ2=4.115, GIF=0.961, P=0.245, AIC=69.151, RMSEA<0.001. The first principal coordinates (PCoA1 explained 59.7 and 78.8% of the variations, see Figure 3) are used to represent the composition of total bacterial and ALP-bacterial community. The width of black arrows indicates the strength of significant standardized path coefficients (P<0.05). ALP, alkaline phosphomonoesterase; SOC, soil organic carbon; TN, total nitrogen.
Interaction network between nematodes and bacterial community
We sought to determine the co-occurrence patterns of nematodes and bacteria using network analysis based on strong and significant correlations. The calculated modularity index was larger than 0.4 (Table 1), indicating a typical module structure (Newman, 2006). Overall, aggregate fractions showed a remarkable effect on association networks of nematodes and bacteria, as well as ALP-producing bacteria. For total bacteria and ALP-producing bacteria, the values of average path length, average clustering coefficient (avgCC) and modularity in these empirical networks were higher than those of their respective identically sized Erdös–Réyni random networks (Table 1). Furthermore, average connectivity (avgK) and modularity was greater in the LMA than in the SMA and MA networks, whereas average path length followed the opposite trend.
The co-occurrence patterns between bacterivores and ALP-producing bacteria were further compared across three aggregate fractions. Notably, there were more positive than negative correlations in all networks, regardless of aggregate fractions (Table 1). Bacterivores were more closely (for example, more abundant nodes) correlated with ALP-producing bacteria in the LMA than in the SMA and MA fractions (Figure 5, Table 1). In particular, the dominant bacterivores Protorhabditis (degree=15) showed stronger positive correlations with ALP-producing bacteria in the LMA network (Figure 5). Topologically, the individual nodes played different roles in the networks according to two properties: the within-module degree Z and among-module degree P. The genus Mesorhizobium (class Alphaproteobacteria) was categorized as the module hub for all three networks (Figure 5, Table 2). Notably, bacterivores Protorhabditis showed strong positive correlations with module hubs OTU2517 (r=0.861, P=0.006), OTU1444 (r=0.822, P=0.009) and OTU1352 (r=0.958, P<0.001), and explained more than one-fourth of variations in the abundance of three module hubs (Supplementary Table 5).
Interaction networks between bacterivorous nematodes and ALP-producing bacterial communities. A connection stands for a strong (Spearman’s ρ>0.8) and significant (P<0.01) correlation for the MA (a), SMA (b) and LMA (c) fractions. The co-occurring networks are colored by phylum/class. For each panel, the size of each node is proportional to the number of connections (that is, degree), and the thickness of each connection between two nodes (that is, edge) is proportional to the value of Spearman’s correlation coefficients. A blue edge indicates a positive interaction between two individual nodes, while a red edge indicates a negative interaction. The numbers inside the nodes are as follows: (1) the dominant bacterivores Protorhabditis, (2) the module hub OTU2517, (3) the module hub OTU1444 and (4) the module hub OTU1352. ALP, alkaline phosphomonoesterase; LMA, large macroaggregate; MA, microaggregate; SMA, small macroaggregate.
Verifying the nematodes–bacteria interactions by soil microcosm experiments
Having found positive effects of nematode grazing on ALP-producing bacterial abundance and activity in open-field environments, we proceeded to demonstrate this in the microcosm under well-controlled laboratory conditions. We applied the natural microbial community with and without bacterivorous nematodes to pre-sterilized soils. Dynamic changes of bacterivores, ALP-producing bacteria and ALP activity were monitored over a period of 21 days. Parallel to our expectation, there were remarkable increases in nematodes, ALP-producing bacteria abundance and ALP activity over time (P<0.001), particularly under M2 and M3 treatments. Specifically, the abundance of Protorhabditis was approximately 25% higher in the LMA fraction than that in the MA fraction (Figure 6). Protorhabditis produced significant effects on ALP-producing bacterial abundance and ALP activity under M2 and M3 treatments compared to under M0 and M1 treatments (Supplementary Figure 7). After 14 days’ incubation, ALP-producing bacterial abundance and ALP activity were elevated by 23.1 and 12.3% with bacterivores addition under the M2 treatment, and increased by 30.3 and 14.1% under the M3 treatment, respectively (Figure 6). Significantly, the effects of Protorhabditis grazing were two to three times larger in the LMA fraction relative to the MA fraction under M2 and M3 treatments (Figure 6).
Microcosm experiment showing the effects of nematode grazing on ALP-producing bacterial abuncance (a) and ALP activity (b). Data are means and standard errors of three replicates with (+Nematode) and without (−Nematode) the inoculation of Protorhabditis in 14 days. Asterisks over the time indicate a significant difference (**P<0.01; *P<0.05). M0, no manure; M1, low manure; M2, high manure; M3, high manure plus lime. ALP, alkaline phosphomonoesterase; MA, microaggregate; LMA, large macroaggregate; SMA, small macroaggregate.
Discussion
Predator–prey interactions are building blocks of food webs, whose stability requires a negative feedback loop. Specifically, an increase of predator abundance causes declines in prey populations, which in turn prevents further increase of the predator population (Djigal et al., 2004). However, a great number of theoretical and empirical studies show that predators can also produce positive effects on their prey, and vice versa (Ingham et al., 1985; Brown et al., 2004; Fu et al., 2005). The underlying mechanisms include enhanced nutrient mineralization (Diehl et al., 2000), disposal to new niches for colonization (Ingham et al., 1985), as well as the emergence of novel physical and behavioral prey refuges (Cressman and Garay, 2009). The net influence between predator and prey is dependent on a combination of both positive and negative effects, and it is likely subject to temporal and spatial dynamic changes. Significantly, when the predator–prey interactions are extended from simple pairs to community levels, such as the bacterivores–bacteria relationships in soil, it remains possible that the two communities may display no ecological correlations, likely owing to the effects of species compensation. In heterogeneous soil environments, the complex bacterivores–bacteria interaction networks form the basis of the heterotrophic eukaryotic food web, ensuring energy flows through the bacterial energy channel to higher trophic levels (Bonkowski et al., 2009). It is thus important to understand the ecological relationships between the bacterial community and their predatory bacterivores in various soils.
Here, we found that bacterivores significantly enhanced bacterial abundance in the maize rhizosphere across four fertilization treatments and three aggregate fractions. This result was initially surprising, as a recent meta-analysis revealed that bacterivores caused a 16 and 17% reduction in soil microbial biomass and bacterial abundance, respectively (Trap et al., 2016). However, our findings were consistent with previous results from microcosm studies, showing that moderate grazing of microbivores on microflora could stimulate microbial growth (Fu et al., 2005). A plausible explanation is that certain bacterivores feed on senescent bacterial cells, and consequently stimulate nutrient cycling (Ingham et al., 1985). In addition, many bacteria reside at the body surfaces or in the digestive systems of nematodes (Neher, 2010). The movement of nematodes can help disperse bacteria to new niches for colonization in heterogeneous soil environments.
With regard to the effects of bacterivores on bacterial community, we observed positive correlations between bacterivores abundance and bacterial diversity in terms of both Shannon index and Chao1 richness. Moreover, bacterivores caused significant changes in bacterial community composition. These data clearly indicated that nematode predation had significant roles in driving dynamic changes of the bacterial community. Bacterivorous nematodes could potentially promote bacterial diversification by generating new ecological opportunities through the evolution of novel predator-resistant strategies or in the form of access to predator-free space (Nosil and Crespi, 2006; Meyer and Kassen, 2007). More importantly, bacterial strains were not equally susceptible to predation. They used different physical and chemical means against nematodes predation, such as bacterial cell shape, filamentation, biofilms as well as the production of pigments, polysaccharides and toxins (Jousset et al., 2009; Jousset, 2011; Bjørnlund et al., 2012). In addition, bacterivores possessed selective feeding traits, which were largely determined by physical constraints of their feeding apparatus and specific detection of chemical cues produced by taxonomically different bacteria (Bonkowski et al., 2009). Clearly, selective predation was fundamentally important for bacterivores to maximize their own fitness and elicit the influences on bacterial community dynamics.
Bulk soils are a resource-limited environment when compared with rhizosphere. Bacterivores–bacteria interactions were previously examined in bulk soils obtained from the same experimental sites (Jiang et al., 2013, 2014). The average total number of nematodes and total bacterial biomass in bulk soils were about half of those found in rhizosphere soils. Interestingly, nematodes and bacteria displayed the similar relationships in the rhizosphere and bulk soils, that is, positive correlations between bacterivores abundance and bacterial biomass as well as bacterial diversity. The data thus suggest that resource constraints may not be the major factor to determine the bacterivores–bacteria community interactions in soils.
In this work, our understanding on the ecological interactions between bacteria and bacterivorous nematodes has been extended to a specific functional group of ALP-producing bacteria. Our results revealed that bacterivores enhanced ALP-producing bacterial abundance and ALP activity, but produced no significant effects on ACP activity. This finding makes sense as soil ACPs were mostly derived from plant and fungi (Tabatabai, 1994). Selective predation likely accounted for the significant effects of nematodes on determining ALP-producing bacterial community composition. For example, bacterivores abundance was positively correlated with Gammaproteobacteria (r=0.878, P<0.001) and Betaproteobacteria (r=−0.812, P<0.001), rather than Actinobacteria (r=0.014, P=0.936). The data support the previous notion that bacterial-feeding nematodes prefer to feed on Gram-negative bacteria (for example, Pseudomonas, a typical rhizosphere colonizer) over Gram-positive bacteria, likely because their thinner cell walls are easier to be digested (Salinas et al., 2007).
We examined the interaction networks between bacteria and bacterivorous nematodes, and identified Mesorhizobium as the keystone species for ALP activity across three aggregate fractions. These keystone species served as gatekeepers in the ecological functions of the bacterial community, with important contributions to biogeochemical cycling (Lynch and Neufeld, 2015). It was different from the ammonia-oxidizing bacterial and archaeal community, which occupied two different keystone species (a module hub and a connector) in three aggregate fractions (Montoya et al., 2006; Jiang et al., 2015). This result suggested that the ALP-producing bacterial community was more susceptible to nematode predation than the ammonia oxidizers. Manure applications promoted the formation of the LMA fraction, the intra-aggregate pore spaces of which were more suitable for bacterivorous nematode survival. Higher density of bacterivores population comprised the vast and complex networks of the nematodes–bacteria associations. The stronger positive effect of bacterivores on Mesorhizobium in the LMA probably grew more predominant contribution to ALP-producing bacterial abundance and ALP activity. Thus, the LMA network could be considered as a better-organized soil food wed with more functional interrelated bacterivorous nematodes and bacteria.
In conclusion, the data presented here showed that nematode predation promotes bacterial community dynamics in red soil, and the extent of the effects varied greatly at the level of soil aggregates. Specifically, the abundance of bacterivores was positively correlated with bacterial biomass and the levels of bacterial diversity. Moreover, nematode predation produced significant influences on species compositions of the bacterial community. In regards to the specific functional groups of ALP-producing bacteria, they displayed the similar effects as the total bacterial community. There was no sufficient evidence to suggest that ALP-producing bacteria were disproportionally affected (or specifically targeted) by nematodes. Interaction network analysis revealed significant effects of nematode on the keystone species of the bacterial community. More specifically, there was a positive correlation between the most dominant nematode Protorhabditis and the ALP-producing keystone 'species' Mesorhizobium. This may explain the findings that nematode grazing stimulated ALP activity. Finally, a systematic and comprehensive understanding of nematodes–bacteria interactions has been achieved at the level of soil aggregate. In general, microaggregates contain higher levels of bacterial abundance and diversity but less amount of bacterivorous nematodes compared with large macroaggregates. Conversely, nematode predation occurred more actively in large macroaggregates than in microaggregates. Together, nematode predation has an important role in determining the composition and dynamics of bacterial community in a spatially dependent manner.
References
Anderson MJ, Willis TJ . (2003). Canonical analysis of principal coordinates: a useful method of constrained ordination for ecology. Ecology 84: 511–525.
Anderson MJ . (2001). A new method for non-parametric multivariate analysis of variance. Austral Ecol 26: 32–46.
Barker KR . (1985). Nematode extraction and bioassays. In: Barker KR (ed). An Advanced Treatise on Meloidogyne Methodology, vol 2. North Carolina State University Graphics: Raleigh, NC, USA, pp 19–35.
Bjørnlund L, Liu MQ, Rønn R, Christensen S, Ekelund F . (2012). Nematodes and protozoa affect plants differently, depending on soil nutrient status. Eur J Soil Biol 50: 28–31.
Bonkowski M . (2004). Protozoa and plant growth: the microbial loop in soil revisited. New Phytol 162: 617–631.
Bonkowski M, Villenave C, Griffiths B . (2009). Rhizosphere fauna: the functional and structural diversity of intimate interactions of soil fauna with plant roots. Plant soil 321, 213–233.
Bongers T, Bongers M . (1998). Functional diversity of nematodes. Appl Soil Ecol 10: 239–251.
Bronick CJ, Lal R . (2005). Soil structure and management: a review. Geoderma 124: 3–22.
Brown DH, Ferris H, Fu S, Plant R . (2004). Modeling direct positive feedback between predators and prey. Theor Popul Biol 65: 143–152.
Buchan D, Moeskops B, Ameloot N, de Neve S, Sleutel S . (2012). Selective sterilisation of undisturbed soil cores by gamma irradiation: effects on free-living nematodes, microbial community and nitrogen dynamics. Soil Biol Biochem 47: 10–13.
Bulgarelli D, Schlaeppi K, Spaepen S, van Themaat EVL, Schulze-Lefert P . (2013). Structure and functions of the bacterial microbiota of plants. Annu Rev Plant Biol 64: 807–838.
Byrne BM . (2010) Structural Equation Modelling with AMOS: Basic concepts, applications, and programming, 2nd edn. Routledge: New York, NY, USA.
Chang A, Schomburg I, Placzek S, Jeske L, Ulbrich M, Xiao M et al. (2015). BRENDA in 2015: exciting developments in its 25th year of existence. Nucleic Acids Res 43: 439–446.
Chesson P, Kuang JJ . (2008). The interaction between predation and competition. Nature 456: 235–238.
Cressman R, Garay J . (2009). A predator-prey refuge system: evolutionary stability in ecological systems. Theor Popul Biol 76: 248–257.
De'Ath G . (2007). Boosted trees for ecological modeling and prediction. Ecology 88: 243–251.
Diehl S, Cooper SD, Kratz KW, Nisbet RM, Roll SK, Wiseman SW et al. (2000). Effects of multiple, predator–induced behaviors on short–term producer–grazer dynamics in open systems. Am Nat 156: 293–313.
Djigal D, Brauman A, Diop T, Chotte JL, Villenave C . (2004). Influence of bacterial-feeding nematodes (Cephalobidae) on soil microbial communities during maize growth. Soil Biol Biochem 36: 323–331.
Ettema CH, Wardle DA . (2002). Spatial soil ecology. Trends Ecol Evol 17: 177–183.
Fraser TD, Lynch DH, Bent E, Entz MH, Dunfield KE . (2015a). Soil bacterial phoD gene abundance and expression in response to applied phosphorus and long-term management. Soil Biol Biochem 88: 137–147.
Fraser T, Lynch DH, Entz MH, Dunfield KE . (2015b). Linking alkaline phosphatase activity with bacterial phoD gene abundance in soil from a long-term management trial. Geoderma 257–258: 115–122.
Frostegård A, Bååth E . (1996). The use of phospholipid fatty acid analysis to estimate bacterial and fungal biomass in soil. Biol Fert Soils 22: 59–65.
Fu S, Ferris H, Brown D, Plant R . (2005). Does the positive feedback effect of nematodes on the biomass and activity of their bacteria prey vary with nematode species and population size? Soil Biol Biochem 37: 1979–1987.
Ingham RE, Trofymow J, Ingham ER, Coleman DC . (1985). Interactions of bacteria, fungi, and their nematode grazers: effects on nutrient cycling and plant growth. Ecol Monogr 55: 119–140.
Jiang Y, Jin C, Sun B . (2014). Soil aggregate stratification of nematodes and ammonia oxidizers affects nitrification in an acid soil. Environ Microbiol 16: 3083–3094.
Jiang Y, Sun B, Jin C, Wang F . (2013). Soil aggregate stratification of nematodes and microbial communities affects the metabolic quotient in an acid soil. Soil Biol Biochem 60: 1–9.
Jiang Y, Sun B, Li H, Liu M, Chen L, Zhou S . (2015). Aggregate-related changes in network patterns of nematodes and ammonia oxidizers in an acidic soil. Soil Biol Biochem 88: 101–109.
Jousset A . (2011). Ecological and evolutive implications of bacterial defences against predators. Environ Microbiol 14: 1830–1843.
Jousset A, Rochat L, Péchy-Tarr M, Keel C, Scheu S, Bonkowski M . (2009). Predators promote defence of rhizosphere bacterial populations by selective feeding on non-toxic cheaters. ISME J 3: 666–674.
Lynch MD, Neufeld JD . (2015). Ecology and exploration of the rare biosphere. Nat Rev Microbiol 13: 217–229.
Meyer JR, Kassen R . (2007). The effects of competition and predation on diversification in a model adaptive radiation. Nature 446: 432–435.
Montoya JM, Pimm SL, Solé RV . (2006). Ecological networks and their fragility. Nature 442: 259–264.
Nannipieri P, Giagnoni L, Landi L, Renella G . (2011) Role of phosphatase enzymes in soil. In: Bunemann EK et al (ed). Phosphorus in Action Soil Biology 26. Springer Verlag: Berlin/Heidelberg, Germany, pp 215–241.
Neher DA . (2010). Ecology of plant and free-living nematodes in natural and agricultural soil. Annu Rev Phytopathol 48: 371–394.
Newman MEJ . (2006). Modularity and community structure in networks. Proc Natl Acad Sci USA 103: 8577–8582.
Nosil P, Crespi BJ . (2006). Experimental evidence that predation promotes divergence in adaptive radiation. Proc Natl Acad Sci USA 103: 9090–9095.
Quénéhervé P, Chotte JL . (1996). Distribution of nematodes in vertisol aggregates under a permanent pasture in Martinique. Appl Soil Ecol 4: 193–200.
Ranjard L, Richaume AS . (2001). Quantitative and qualitative microscale distribution of bacteria in soil. Res Microbiol 152: 707–716.
Rillig MC, Mummey DL . (2006). Mycorrhizas and soil structure. New Phytol 171: 41–53.
Rønn R, Vestergård M, Ekelund F . (2012). Interactions between bacteria, protozoa and nematodes in soil. Acta Protozool 51: 223–235.
Sakurai M, Wasaki J, Tomizawa Y, Shinano T, Osaki M . (2008). Analysis of bacterial communities on alkaline phosphatase genes in soil supplied with organic matter. Soil Sci Plant Nutr 54: 62–71.
Salinas KA, Edenborn SL, Sexstone AJ, Kotcon JB . (2007). Bacterial preferences of the bacterivorous soil nematode Cephalobus brevicauda (Cephalobidae): effect of bacterial type and size. Pedobiologia 51: 55–64.
Six J, Elliott ET, Paustian K . (2000). Soil macroaggregate turnover and microaggregate formation: a mechanism for C sequestration under no-tillage agriculture. Soil Biol Biochem 32: 2099–2103.
Spohn M, Kuzyakov Y . (2013). Distribution of microbial- and root derived phosphatase activities in the rhizosphere depending on P availability and C allocation–coupling soil zymography with 14C imaging. Soil Biol Biochem 67: 106–113.
Tabatabai MA (1994). Soil enzymes. In: Weaver RW, Angle JS, Bottomley PS (eds). Methods of Soil Analysis, Part 2, Microbiological and Biochemical Properties. Soil Science Society of America: Madison, WI, USA, pp 775–833.
Tan H, Barret M, Mooij MJ, Rice O, Morrissey JP, Dobson A . (2013). Long-term phosphorus fertilisation increased the diversity of the total bacterial community and the phoD phosphorus mineraliser group in pasture soils. Biol Fert Soils 49: 661–672.
Tisdall J, Oades JM . (1982). Organic matter and water-stable aggregates in soils. J Soil Sci 33: 141–163.
Trap J, Bonkowski M, Plassard C, Villenave C, Blanchart E . (2016). Ecological importance of soil bacterivores for ecosystem functions. Plant Soil 398: 1–24.
van der Heijden MG, Bardgett RD, van Straalen NM . (2008). The unseen majority: soil microbes as drivers of plant diversity and productivity in terrestrial ecosystems. Ecol Lett 11: 296–310.
Yeates GW, Bongers T, de Goede RGM, Freckman DW, Georgieva SS . (1993). Feeding habits in soil nematode families and genera—an outline for soil ecologists. J Nematol 25: 315–331.
Acknowledgements
We are very grateful to Wenqing Fan who helped cultivate nematode in microcosm experiment, and Haiyan Qian for her assistance in field experiments. This research was funded by National Key R&D Project (2016YDFD0200309), Strategic Priority Research Program of the Chinese Academy of Sciences (XDB15030201), National Basic Research Program of China (2014CB441003), Science and Technology Service Network Initiative (KFJ-SW-STS-142), and the Youth Innovation Promotion Association of CAS (2017361).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Jiang, Y., Liu, M., Zhang, J. et al. Nematode grazing promotes bacterial community dynamics in soil at the aggregate level. ISME J 11, 2705–2717 (2017). https://doi.org/10.1038/ismej.2017.120
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2017.120
This article is cited by
-
Impact of captivity and natural habitats on gut microbiome in Epinephelus akaara across seasons
BMC Microbiology (2024)
-
Conceptualizing soil fauna effects on labile and stabilized soil organic matter
Nature Communications (2024)
-
Comparison of microbial communities in unleached and leached ionic rare earth mines
Environmental Science and Pollution Research (2024)
-
Dynamic Changes in Rhizosphere Microbial Communities of Watermelon During Continuous Monocropping with Gravel Mulch
Journal of Soil Science and Plant Nutrition (2024)
-
Introducing N2-fixing tree species into Eucalyptus plantations increases organic phosphorus transformation but decreases its accumulation within aggregates in subtropical China
Plant and Soil (2024)