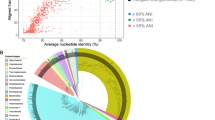
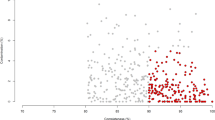
Abstract
Many microbes in complex competitive environments share genes for acquiring and utilising nutrients, questioning whether niche specialisation exists and if so, how it is maintained. We investigated the genomic signatures of niche specialisation in the rumen microbiome, a highly competitive, anaerobic environment, with limited nutrient availability determined by the biomass consumed by the host. We generated individual metagenomic libraries from 14 cows fed an ad libitum diet of grass silage and calculated functional isoform diversity for each microbial gene identified. The animal replicates were used to calculate confidence intervals to test for differences in diversity of functional isoforms between microbes that may drive niche specialisation. We identified 153 genes with significant differences in functional isoform diversity between the two most abundant bacterial genera in the rumen (Prevotella and Clostridium). We found Prevotella possesses a more diverse range of isoforms capable of degrading hemicellulose, whereas Clostridium for cellulose. Furthermore, significant differences were observed in key metabolic processes indicating that isoform diversity plays an important role in maintaining their niche specialisation. The methods presented represent a novel approach for untangling complex interactions between microorganisms in natural environments and have resulted in an expanded catalogue of gene targets central to rumen cellulosic biomass degradation.
Similar content being viewed by others
Introduction
Recent metagenomic sequencing of natural environments has revealed that microbial communities act in different trophic levels (Lauro et al., 2009), exhibit successional change (Koenig et al., 2011; Teeling et al., 2012; Huws et al., 2016) and engage in fierce competition when resources are limited (Weckx et al., 2011). This suggests that adaptation to specific niches must also occur in microbial communities, however demonstrating this in natural communities remains a challenge (Marco, 2008).
It has been proposed that radical niche diversification in prokaryotes may primarily occur through horizontal acquisition of novel genes (Jain et al., 2003; Toft and Andersson, 2010). Furthermore, the high frequency of horizontal gene transfer detected among bacteria (Gogarten et al., 1999; Creevey et al., 2004; Nelson-Sathi et al., 2014), even among ‘core’ genes (Creevey et al., 2011), suggests that horizontal gene transfer may also drive niche differentiation even within species (Coleman et al., 2006).
In contrast, microbial community profiling using 16S ribosomal RNA has revealed core communities can be defined for many environments (Turnbaugh et al., 2009; Lundberg et al., 2012; Creevey et al., 2014; Otani et al., 2014), indicating that niche specialisation of specific groups of organisms is somehow maintained even under the continued influence of horizontal gene transfer. One explanation may be that the horizontal acquisition of a single isoform of a novel gene is not enough to maintain a competitive advantage in fluctuating environmental conditions where a range of isoforms may be required to, for instance, maintain enzymatic activity (Stewart, 1976; Nevo, 2001).
The rumen microbiome of sheep and cattle is a case in point; temperature and pH fluctuate daily in response to ambient conditions and the biomass consumed (Mishra et al., 1970). As a potential source of industrially important enzymes (Hess et al., 2011; Qi et al., 2011), antimicrobial agents (Russell and Mantovani, 2002) and greenhouse gases (Qi et al., 2011), understanding how the microbiome maintains robustness to environmental fluctuations and how this may be manipulated is an important goal.
Identifying gene isoforms from metagenomic sequence data remains a difficult challenge (Scholz et al., 2012), as assembly approaches vastly oversimplify the true microbial diversity present (Temperton and Giovannoni, 2012). Recently, approaches that involve mapping metagenomic sequence reads, (which preserve the lost variability), to sequenced microbial genomes have attempted to address this by identifying single-nucleotide polymorphisms (SNPs) representing variation in the microbial population (Williams and Oleksiak, 2011; Schloissnig et al., 2013). Using methods adapted from traditional dn/ds calculations of adaptive evolution between species (Creevey and McInerney, 2003; Loughran et al., 2012; Baldwin et al., 2014) they calculated the genomic variation in the human gut microbiome based on the ratio of the rates of occurrence of SNPs that either cause a mutation in the translated amino acid sequence (non-synonymous mutations; pN) or preserve the translated amino acid sequence (synonymous mutations; pS; Schloissnig et al., 2013). The population with a higher pN/pS ratio for any function likely reflects the possession of greater numbers of distinct functional isoforms of that gene (Schloissnig et al., 2013). Furthermore, as this calculation uses the rate of synonymous substitutions as a baseline, it is robust to differences in the size of the populations compared.
We have adapted this approach to examine the rates of pN/pS for all shared functions between the two most abundant genera of bacteria in the rumen. Applied to metagenomic samples from 14 animals maintained as a single herd and provided with the same diet, we were able to estimate confidence intervals for the pN/pS ratios calculated. We then investigated whether differences in the abundance of functional isoforms existed between the two microbial populations and therefore could possibly drive niche specialisation.
Materials and methods
The animal model
The animal model on which this work is based has been described previously (Carberry et al., 2012) and was part of a larger study designed to examine the physiological control of energetic efficiency in growing beef heifers (Kelly et al., 2010). Our analysis centred on samples taken during this study from 14 Limousin x Friesian heifers following a 44-day period of ad libitum access to a high-forage diet of grass silage. Samples of rumen fluid were collected using a transesophageal sampling device (FLORA rumen scoop; Guelph, ON, Canada). Subsequently, a 20-ml aliquot was transferred using a pipette and sterilised tip into a separate labelled sterilised container, immediately frozen in liquid nitrogen, and stored at 80 °C until processing (Carberry et al., 2012).
Sampling, metagenomic library preparation and sequencing
DNA was extracted from rumen fluid samples using the repeat bead beating and column purification method of Yu and Morrison (2004). DNA was fragmented to an average size of 300 bp using a Bioruptor (Diagenode, Seraing, Belgium). The NEBNext End Repair Module was used to blunt end the fragments and purification of the reaction was performed using a QIAquick PCR purification kit (Qiagen, Hilden, Germany). The NEBNext dA-Tailing Module was used to adenylate the blunt-ended fragments and purification of the reaction was performed using a QIAquick PCR purification kit (Qiagen). Illumina standard paired end barcoded adapters were ligated onto the adenylated fragments using the Quick Ligation Kit (NEB, Ipswich, MA, USA) and purification was performed using a QIAquick PCR purification kit (Qiagen). Adapter-ligated fragments were then size selected (average insert size of 280 bp) by electrophoresis on an agarose gel, excision of a 2 mm gel slice and extraction of DNA from the agarose using the QIAquick gel extraction kit (Qiagen). PCR enrichment (12 cycles) of the library was performed using Illumina PCR Paired End Primers 1.0 and 2.0 and Phusion High-Fidelity PCR Kit (Finnzymes, Waltham, MA, USA). The library size and absence of adapter dimers were determined with a DNA1000 chip on an Agilent 2100 Bioanalyzer (Agilent, Santa Clara, CA, USA). Libraries were pooled in equimolar amounts and sequenced on an Illumina GAIIx using a standard paired end 300 cycle sequencing kit.
Assembly
The read length for this data set was 160 bp and after quality control 15 bp were trimmed from both 5′ the 3′ ends. The trimmed sequences from all 14 samples were used to create a combined metagenome assembly with SGA, version 0.9.18 (Simpson and Durbin, 2012). First, pre-processing of the reads was carried out using the preprocess module of SGA with the –phred64 flag, with settings -p1 for paired-end libraries and –permute-ambiguous to treat ambiguities as all possible combinations of characters. Second, the computationally and memory intensive indexing step was carried out using the index module of SGA with the default options. Finally, the pre-processed and indexed reads were used as input to the metagenome module of SGA to carry out the assembly with the default options, except for the -k 51 flag, for a k-mer size of 51. The resulting assembly contained all contigs ⩾200 bp in length. The raw sequence data can be accessed at the European Nucleotide Archive (http://www.ebi.ac.uk/ena) using the study ID: ERP009980.
Gene and taxonomy prediction
Gene predictions were carried out on the assembly using HMMER version 3.0 (Eddy, 2011). To do this, HMM profiles of known genes were generated for each gene family from within each group of organisms from the rumen (Creevey et al., 2014). Genes were identified from the Kegg Orthology (Kanehisa et al., 2012) database and the corresponding sequences downloaded from Uniprot (The Uniprot Consortium, 2012) using the download_profiles script of the MGKit package (Rubino and Creevey, 2014; version 0.1.13). This script automates the retrieval of all ortholog sequences available for all combinations of genes and taxa of interest. Only combinations of genes and taxa having at least two sequences in Uniprot were used (a list of taxa is available in Supplementary Table S1). Alignments were created for each gene family from a group of organisms using Clustal Omega (Sievers et al., 2011) version 1.1.0 and converted into HMMER profiles using the HMMbuild command. These profiles were used as input to HMMsearch for comparison against the contigs from the assembly, which were translated into all six reading frames using the translate_seq script in MGKit.
Converting results to annotations
Matches of contigs to the HMM profiles were converted into GFF format and combined with annotation information from KEGG using the hmmer2gff script of MGKit. They were then filtered for low-quality predictions and only matching regions in the assembly with an e-value threshold of 0.05 were retained. Furthermore, using the filter_gff script of MGKit any annotation whose length was covering <40% of the average length of the HMM profile it matched was discarded (-f option). All annotated regions that overlapped for more than 80% of their length were discarded, except the one having the lowest e-value, (-l option). Taxonomic assignments of the annotated regions were carried out using BLAST+ version 2.2.25 (Camacho et al., 2009), against a local copy of the nr protein database (May 2013; options -outfmt 6). The best match with a bit score >60 was used as simulations indicated that when using this cut-off hits were correctly assigned at the genus level 95% of the time (Supplementary File 1).
Alignment
All 14 sample reads were individually aligned to the final assembly, using Bowtie2 (Langmead and Salzberg, 2012) version 2.0.5 (options: -N 1). Possible PCR and optical duplicate reads were identified using Picard Tools (Team, 2009) version 1.58. Abundance of reads mapping to each annotation was calculated with htseq-count from the HTSeq package (Anders et al., 2015) version 0.5.4p3 (options: -s no -t gene -i ko_idx –q).
SNPs
SNPs were called from each of the alignments separately using the GATK pipeline (McKenna et al., 2010), version 2.1.8. with the UnifiedGenotyper tool (options: -dcov 50 for 4 × coverage alignment). The resulting VCF files were analysed to identify for each SNP whether it was synonymous using SNPDat version 1.0.4 (Doran and Creevey, 2013). All VCF files were then combined for analysis using the CombineVariants command of GATK (options: -genotypeMergeOptions UNIQUIFY to make all sample genotypes unique by file name). The output of SNPDat and the merged VCF file were combined using the snp_parser script of MGKit. The retained SNPs had a minimum allele frequency of 0.01, a minimum Phred quality score of 30 and supported by a minimum of four reads across all the samples.
Calculation of rates of pN/pS
The pN/pS measure (Schloissnig et al., 2013) is the ratio of two quantities, pN and pS. The former is the ratio between the number of non-synonymous amino acid changes to those expected, whereas the latter describes the same ratio for synonymous changes. The expected number of changes is calculated in a similar way as in dn/ds calculations, using a neutral model to determine the proportion of either non-synonymous or synonymous SNPs that would be expected given the reference sequence for the gene (Li, 1993). The MGKit package provides routines for these calculations on a per-isoform basis, but they can be extended to different levels of specificity, for example, using genus instead of species, or enzymes grouped into different EC levels such as 3.4.1.3 or EC 3.4.-. This involves concatenating isoforms belonging to the specific taxonomic and functional level before performing the calculation. When calculating the rates of pN/pS of individual functions within taxonomic groups, if multiple genes from the assembly were annotated with the same function and taxonomy, they were concatenated and the rates of pN/pS calculated across all the concatenated sequences.
Statistical analysis of functional divergence
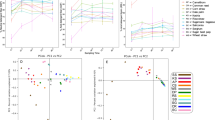
Pairwise comparisons of the distribution of pN/pS calculations for genes were carried out between taxa using a two-sample Wilcoxon rank-sum test as implemented in the Scipy package (van der Walt et al., 2011) version 0.15.1. In this analysis, pairwise statistical tests were carried out between all genes shared between Methanobrevibacter, Clostridium and Prevotella (Figure 1). All P-values were corrected for multiple testing using the Benjamini–Hochberg (Benjamini and Hochberg, 1995) method as implemented in R (R Core Team, 2014), using a significance threshold for the corrected value of 0.1.
Gene counts from the three most abundant genera in the rumen samples. (a) Venn diagram showing the overlap of genes found in at least three samples between the Prevotella, Clostridium and Metahonbrevibacter. (b) The number of genes found in at least two genera, indicating the proportion that had a significantly higher pN pS rates in one group or the other (b1) between Clostridium and Prevotella, (b2) between Clostridium and Methanobrevibacter and (b3) between Methanobrevibacter and Prevotella.
Enzyme Commission Classification
Functional diversity was also estimated for all predicted genes from the metagenomic data that had a mapping to an enzyme commission classification (EC). We concatenated all the genes from same second-level EC classification (that is, EC: 1.1.-, 1.2.- and so on), only including a predicted gene if it had been found in both Prevotella and Clostridium in at least seven samples. A single pN/pS value was calculated for each sample, representing the diversity of all the genes for any particular EC number and the distributions of the pN/pS values across samples were compared between Clostridium and Prevotella. A Wilcoxon rank-sum test as implemented in R was used to test whether the distribution of diversity estimates for any EC number was significantly different between the two groups and the P-values were corrected as previously described.
Over-representation analysis
The EC classifications that were found to have significantly different pN/pS values between Prevotella and Clostridium were used for an over-representation analysis. The significantly different EC numbers were used as foreground for the analysis, whereas the rest of the all EC numbers Prevotella and Clostridium had in common, were used as background. The files were then analysed in R, with the GOSeq package (Young et al., 2010) to perform the test and adjusting the P-values with the Benjamini–Hochberg method.
Coverage calculation
As the length of the reads was fixed at 130 bp for this data set, the coverage formula used is the following:  , where LR is the length of the reads aligned, NR is the number of reads mapping to the specific region and LA is the length of the annotation for which coverage is calculated.
, where LR is the length of the reads aligned, NR is the number of reads mapping to the specific region and LA is the length of the annotation for which coverage is calculated.
All MGKit scripts used are provided as HTML files in Supplementary Files 1–7. MGKit is available with tutorials under the GNU General Public License at https://bitbucket.org/setsuna80/mgkit/.
Results
General assembly results
The average number of reads generated per sample was 11.7 M (min 8.6 M, max 14.0 M) totalling 164.3 M reads in total. The total number of assembled contigs with a minimum size of 200 bp was 1 421 712 with an average length of 374 bp (max 62 508 bp) and an N50 of 364. The number of predicted genes passing filters from the assembly was 106 475 with 70 525 having at least 4 × coverage of aligned reads in at least one sample (30 970 in at least 3 samples, 2665 across all 14 samples). The average gene coverage per sample was 8.7 × (min 4 ×, max 1264 ×). More details are provided in Table 1.
Taxonomic and functional annotation
Following taxonomic assignments, counting only those genera with at least four observed genes and with an average coverage among all samples ⩾4 × resulted in 127 bacterial, 20 archaeal, 5 fungal and 21 protozoal genera remaining (Supplementary Table S3). Functional annotation of these genes revealed that 20.12% were involved in information storage and processing, 33.96% were involved in metabolism, 17.71% were involved in cellular processes and signalling and 28.21% were poorly characterised (Supplementary Figure S1).
This revealed that the genus with the largest number of genes in our data set was the bacterial genus Prevotella (1044 genes), followed by the archaeal genus Methanobrevibacter (533 genes) and then the bacterial genus Clostridium (409 genes). When only considering those with matches to gene families from eggNOG (Powell et al., 2012), 16.03% were from Prevotella, 8.18% from Methanobrevibacter and 6.28% from Clostridium (Figure 1a). In the following analyses we concentrated on the two most abundant bacterial genera due to their specific roles in the colonisation of plant material in the rumen (Huws et al., 2016).
Diversity estimates
pN/pS values were calculated for all genes found in a minimum of three samples and with a minimum of 4 × coverage.
The calculated pN/pS values ranged from 27.16 in gpmI (Kegg ID: K15633) from Prevotella, which is involved in carbohydrate and lipid metabolism, to many genes (59.80% of the total) with a pN/pS value of zero. When assessed by function using eggNOG the category ‘cellular processes and signalling’ had the highest average pN/pS (0.19), and ‘metabolism’ had the lowest (0.18; Figure 2). Comparison of the pN/pS values of the genes shared by the two most abundant genera of bacteria (Prevotella and Clostridium) revealed that 88 and 65 had significantly higher pN/pS values (adjusted P<0.1) in Prevotella and Clostridium, respectively (Figure 1b).
KEGG pathways
Visual comparison of the rates of pN/pS superimposed on various KEGG metabolic pathways, identified the ‘Carbon metabolism’ pathway, as showing clearly connected sub-components of the pathways where the genes involved either had a significantly higher pN/pS in Prevotella compared with Clostridium or vice versa (Figure 3). To investigate this in more detail we broke the ‘Carbon metabolism pathways’ into its constituent sub-components (called modules in KEGG). Overall pN/pS values were calculated from each of the 14 samples for each pathway module following concatenation of each of the genes in that module. The distribution of pN/pS values from the multiple samples was used to test if a significant difference existed in their functional diversity for any module between Prevotella and Clostridium. This identified the following modules as being significantly different (P<0.1 after correction) between Prevotella and Clostridium: ‘Pyruvate oxidation’, ‘Hydroxypropionate-hydroxybutyrate’, ‘Phosphate acetyltransferase-acetate kinase pathway, acetyl-CoA => acetate’ were more diverse in Prevotella; whereas ‘PRPP biosynthesis ribose 5P’, ‘Cysteine biosynthesis—serine’, ‘Formaldehyde assimilation—ribulose monophosphate pathway’, ‘Formaldehyde assimilation—serine pathway’ were found more diverse in Clostridium (Figure 4; Supplementary Table S4).
Genes marked in blue and red are significantly different between Prevotella and Clostridium and are part of the carbon metabolism pathway in KEGG. Genes in blue are significantly different between the two genera and have a higher average pN/pS in Prevotella. Genes in red have a higher average in Clostridium.
Enzyme Commission Classification
Twenty-one EC showed significantly divergent rates (P<0.05) between Prevotella and Clostridium.
The EC with significantly higher diversity in Prevotella included oxidoreductases acting on the CH-OH group of donors (EC 1.1.-), the aldehyde or oxo group of donors (EC 1.2.-), and on the CH-NH2 group of donors (EC 1.4.-); Hydrolases acting on peptide bonds (EC 3.4.-); General lyases (EC 4.99.-) and the intramolecular oxidoreductase category of isomerases (EC 5.3.-) (Supplementary Figure S2).
The EC with a significantly higher pN/pS distribution in Clostridium included: oxidoreductases acting on the CH-CH group of donors (EC 1.3.-) and on NADH or NAPH (EC 1.6.-); transferases transferring aldehyde or ketonic groups (EC 2.2.-), acyltransferases (EC 2.3.-), glycosyltransferases (EC 2.4.-) and transferring phosphorus-containing groups (EC 2.7.-); hydrolases, specifically glycosylases (EC 3.2.-), those acting on carbon–nitrogen bonds, other than peptide bonds (EC 3.5.-), and those acting on carbon–carbon bonds (EC 3.7.-); carbon–carbon lyases (EC 4.1.-); racemases and epimerases (EC 5.1.-) and cis-trans-isomerases (EC 5.2.-) from isomerases, and finally, ligases forming carbon–oxygen bonds (EC 6.1.-), carbon–sulfur bonds (EC 6.2.-) and nitrogen–metal bonds (EC 6.6.-; Supplementary Figure S3).
Supporting this, genes found to be significantly divergent between Prevotella and Clostridium were also statistically overrepresented in 14 of the EC already identified. In particular, the EC 1.1.-, 1.2.-, 1.4.-, 5.3.- and 3.4.- were significantly overrepresented in the set of genes that were more diverse in Prevotella (P<0.001) and the EC 1.3.-, 2.3.-, 2.4.-, 2.7.-, 3.2.-, 3.5.-, 4.1.- 5.1. and 6.1.- were significantly overrepresented in the set of genes that were more diverse in Clostridium (P<0.05; Supplementary Table S2).
Discussion
Taxonomic composition
The fact that there are very few fully sequenced rumen microbial genomes available have likely inflated the estimates of the number of genera we reported (Supplementary Table S3), as when the genome is not in the database, the same gene from one of the most closely related species is likely to be matched instead. However if only those genera with a minimum four genes identified are considered, then 127 bacterial and 20 archaeal genera remain, which is similar to previous 16S-based estimates of both bacteria (Creevey et al., 2014) and archaea (Janssen and Kirs, 2008).
However, the most interesting result from the taxonomic identification was from the most abundant groups. We found that Methanobrevibacter genes were more abundant than genes from Clostridium and second only to the number of genes found from Prevotella (Supplementary Table S3). These findings are contrary to estimates that would place the abundance of archaea well below that of these two groups. Previous estimates of archaeal abundance from the rumen using culture-based approaches tend to be at around 4% (Yanagita et al., 2000), with numbers from metatranscriptomics suggesting that archaeal messenger RNA represents somewhere between 0.1 (Qi et al., 2011) and 1% (Poulsen et al., 2013) of the total messenger RNA actively transcribed. We estimate from our samples that the minimum proportion of Methanobrevibacter genes in the rumen metagenome is 9.02%, following Prevotella at 17.66% and followed by Clostridium at 6.92%. The differences observed between our current and previous estimates are likely due to differences in the technologies used (Janssen and Kirs, 2008; Steinberg and Regan, 2008). For instance, biases in the primers used for ribosomal RNA amplification in archaea can under-represent certain groups and because of differences in the primers used, the comparison of abundances of archaea and bacteria is nontrivial (Tymensen and McAllister, 2012).
To our knowledge it has not previously been suggested that the Methanobrevibacter genes could make up a larger component of the rumen metagenome than Clostridium. Scaling these numbers according to average genome sizes using sequenced organisms from these groups actually increases this trend (Supplementary Table S3; Figure 5). However our estimates of the proportion of archaeal genes in the rumen metagenome from Methanobrevibacter (76.47%) and the relative proportions of genes from the major groups of bacteria are similar to those that were previously reported (Janssen and Kirs, 2008; Creevey et al., 2014), suggesting no sampling bias.
Number of genes found in each genus. For each genus the proportion of number of genes is plotted in a different colour. Each genus has two bars, the left one is the proportion of genes found and the right bar the proportion of genes scaled by the average genome size of the genus. A full colour version of this figure is available at the ISME Journal online.
Diversity estimates
We found that levels of adaptive variation (pN/pS) vary widely between microbial genera from the rumen (Figure 6) similar to that found between human gut microbes (Schloissnig et al., 2013) and that this extends to functional classes of genes (Figure 2; Supplementary Figures S2 and S3) and also for the same genes from different genera.
Variation in pN/pS calculated for all genes assigned to the genera representing the top 55% of all genes identified. Each box plot represents the distribution of pN/pS values calculated across the samples. The values for the individual samples are represented by the individual data points superimposed on the boxplots. For each genus, all genes from the same sample were concatenated and pN/pS values calculated, with the total number of (full and partially reconstructed) genes used indicated by the histogram on the right. Colours represent the group to which the genus belongs: bacteria (blue), archaea (green) and protozoa (purple).
However, the level of variation in these calculations between samples was related to the abundance of each genus in the rumen microbiome (Figure 6), and therefore the depth of sequencing achieved for each. Although the low variance in the calculations across samples of pN/pS for Prevotella (0.000033) and Clostridium (0.000363) provided enough confidence to further investigate these groups, greater depth of sequencing will be necessary for other genera. Although the pN/pS variance in the archaea Methanobrevibacter is comparable to both Prevotella and Clostridium, we focussed on comparisons between the latter two bacterial genera, which have been shown to directly compete for access to the plant fibre (Huws et al., 2016). The specific ecological niche of methanogens in the rumen is quite distinct and they do not directly compete for access to the plant fibre material. They utilise hydrogen and carbon dioxide as an energy and carbon source, respectively, and produce methane as a by-product (Carberry et al., 2014).
For Prevotella and Clostridium, we tested for significant differences in the pN/pS calculated between all shared genes, testing the hypothesis that this reflected functional importance of those genes to the organism. In this scenario a gene with significantly higher values of pN/pS in one genus compared with the other indicates higher levels of functional isoforms within its population. We hypothesise that these isoforms may allow important functions for that organism to be robust to fluctuations in the environmental conditions (or substrates) within the rumen. Functional analysis of these genes should reflect this, allowing construction of a profile of functions for each that reflects their roles in the rumen.
Metabolic adaptations in Prevotella
We found that Prevotella possess more diverse functional isoforms than Clostridium in genes involved in specific metabolic processes. For instance, pyruvate oxidation (from the carbon metabolism pathway) was significantly more diverse in Prevotella (average of 1.50 versus 0.16) as was the enzyme class 1.2.-, which contains pyruvate oxidase. Interestingly both the substrates for this pathway (pyruvate) and its product (acetyl-CoA) are central to fermentative processes and the reactions involved have been implicated in anaerobic fermentation of a diverse selection of compounds by Prevotella (Takahashi and Yamada, 2000; Purushe et al., 2010) and other species (Abdel-Hamid et al., 2001). Studies of pyruvate oxidase in E scherichia coli have suggested that it is used preferentially at low growth rates (Abdel-Hamid et al., 2001), indicating the intriguing possibility of an adaptation in Prevotella species in low nutrient conditions. Indeed, Prevotella have been identified as one of the primary colonisers of plant material consumed by the host, reaching its peak population within 1 h of incubation (Huws et al., 2016). Therefore, a strategy that would allow them to maintain a sufficient population during nutrient scarcity that enables this rapid colonisation would be advantageous.
Prevotella also possessed significantly more functional diversity in the genes belonging to the hydroxypropionate-hydroxybutyrate cycle than Clostridium (average of 0.74 versus 0.01). This is involved in the conversion of carbon compounds in fatty acids (Ellis et al., 2008; Servinsky et al., 2012), and their higher diversity suggests Prevotella has a greater repertoire of enzymes involved in this process and supports previous reports that Prevotella species are among those (including Clostridium species) that play a role in biohydrogenation of dietary polyunsaturated fatty acids to saturated fatty acids (Huws et al., 2011). This would suggest that fatty acids may be a resource for which these organisms compete and for which they have diversified niches.
Finally, when examined by their ECs, we found a significantly larger diversity of peptidases (EC 3.4) in Prevotella compared with Clostridium (average pN/pS 0.28 compared with 0.1) supporting previous reports of the importance of proteolysis as one of the main sources of metabolic energy for Prevotella (Wallace, 1996). This evidence suggests that Prevotella possess a greater diversity of functional isoforms than Clostridium that digest peptides, and may be related to the essential production of the volatile fatty acids propionate and butyrate used as nutrients by the host.
Metabolic adaptations in Clostridium
We found that Clostridium possessed a significantly higher functional diversity than Prevotella for genes involved in a range of metabolic processes including cysteine biosynthesis (average 1.64 versus 0.13) and formaldehyde assimilation/serine pathway (average 0.75 versus 0.18). Interestingly, some Clostridium species from the rumen have been characterised as having a preferential utilisation for serine, threonine, cysteine, proline and glycine in their exponential growth phase (Flythe and Kagan, 2010; Fonknechten et al., 2010). Our results suggest that this may be a specific niche specialisation by these species in the rumen.
Furthermore, the PRPP biosynthesis module that connects the non-oxidative branch of the pentose phosphate pathway to the histidine biosynthesis pathway (Nölling et al., 2001) was also significantly more diverse in Clostridium than Prevotella (P<0.01, average of 2.76 versus 0.13). This supports reports that species of Clostridium may utilise histidine as part of a strategy to overcome nutrient limitation during the stationary growth phase (Fonknechten et al., 2010).
Finally, the class of glycosylases to which the cellulosome enzymes from Clostridium species belong (EC 3.2; Ravachol et al., 2014), was more functionally divergent in Clostridium than in Prevotella. This suggests Clostridium possess a greater range of diverse functional isoforms involved in catabolism of these polysaccharides than Prevotella and may provide an advantage in competitive conditions.
Conclusion
In this study we focussed on the most abundant bacterial genera in the rumen, showing how niche specialisation is driven by resource competition. This is exemplified by the development of differences in a range of isoforms in Clostridium and Prevotella for key genes that can provide a competitive advantage over the other. For instance, although both have the potential to degrade a complex matrix like plant fibre, they show significantly different adaptations relating to the structure of the plant material they colonise. In the rumen, colonisation of plant fibre is a multi-step process and it has been noted that the abundance of Prevotella peaks before Clostridium (Huws et al., 2016). This has been hypothesised to relate to the degradation of the hemicellulose prior to cellulose in the plant. In this scenario, Prevotella would preferentially degrade hemicellulose, and when xylan compounds then become accessible, colonisation of the fibre surface is gradually taken over by Clostridium (Figure 7; Huws et al., 2016). Our results indicate that this is likely the case as Prevotella has a more diverse range of isoforms to degrade the hemicellulose matrix formed of pectins (EC 5.3.-; Oxenbøll et al., 2007) and peptides (EC 3.4.-), whereas Clostridium has a more diverse range of isoforms for degrading cellulose (EC 3.2.-). This coupled with their divergence for function in metabolic processes, indicate that isoform diversity plays an important role in maintaining their niche specialisation.
Cartoon illustration of colonisation of plant material in the rumen by Prevotella (blue) and Clostridium (red); plant fibre in green, composed by cellulose inside (bright green) and hemicellulose outside (dark green). Data suggest that the degradation process begins with Prevotella populating fibre faster than Clostridium, because its niche specialisations allowing more efficient action on hemicellulose (a) resulting in higher number of Prevotella observed in the early stages of plant fibre colonisation (b). However, as more cellulose becomes available (c) niche adaptations in Clostridium reverses this trend and increasing their numbers in the latter stages (d).
Our results demonstrate the potential power of evolutionary approaches in understanding microbial communities in vivo. Indeed, applied to less well-studied microbiomes, it should be possible to identify niche specialisation and generate novel hypotheses of the organisational structure and function of the constituents that could be tested in vitro.
References
Abdel-Hamid AM, Attwood MM, Guest JR . (2001). Pyruvate oxidase contributes to the aerobic growth efficiency of Escherichia coli. Microbiology 147: 1483–1498.
Anders S, Pyl PT, Huber W . (2015). HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31: 166–169.
Baldwin MW, Toda Y, Nakagita T, O’Connell MJ, Klasing KC, Misaka T et al. (2014). Evolution of sweet taste perception in hummingbirds by transformation of the ancestral umami receptor. Science 345: 929–933.
Benjamini Y, Hochberg Y . (1995). Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Ser B 57: 289–300.
Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K et al. (2009). BLAST+: architecture and applications. BMC Bioinformatics 10: 421.
Carberry CA, Kenny DA, Han S, McCabe MS, Waters SM . (2012). Effect of phenotypic residual feed intake and dietary forage content on the rumen microbial community of beef cattle. Appl Environ Microbiol 78: 4949–4958.
Carberry CA, Waters SM, Kenny DA, Creevey CJ . (2014). Rumen methanogenic genotypes differ in abundance according to host residual feed intake phenotype and diet type. Appl Environ Microbiol 80: 586–594.
Coleman ML, Sullivan MB, Martiny AC, Steglich C, Barry K, Delong EF et al. (2006). Genomic islands and the ecology and evolution of Prochlorococcus. Science 311: 1768–1770.
Creevey CJ, Doerks T, Fitzpatrick DA, Raes J, Bork P . (2011). Universally distributed single-copy genes indicate a constant rate of horizontal transfer. PLoS One 6: e22099.
Creevey CJ, Fitzpatrick Da, Philip GK, Kinsella RJ, O’Connell MJ, Pentony MM et al. (2004). Does a tree-like phylogeny only exist at the tips in the prokaryotes? Proc Biol Sci 271: 2551–2558.
Creevey CJ, Kelly WJ, Henderson G, Leahy SC . (2014). Determining the culturability of the rumen bacterial microbiome. Microb Biotechnol 7: 467–479.
Creevey CJ, McInerney JO . (2003). CRANN: detecting adaptive evolution in protein-coding DNA sequences. Bioinformatics 19: 1726.
Doran AG, Creevey CJ . (2013). Snpdat: easy and rapid annotation of results from de novo SNP discovery projects for model and non-model organisms. BMC Bioinformatics 14: 1.
Eddy SR . (2011). Accelerated profile HMM searches. PLoS Comput Biol 7: e1002195.
Ellis JL, Dijkstra J, Kebreab E, Bannink A, Odongo NE, McBride BW et al. (2008). Aspects of rumen microbiology central to mechanistic modelling of methane production in cattle. J Agric Sci 146: 213–233.
Flythe M, Kagan I . (2010). Antimicrobial effect of red clover (Trifolium pratense phenolic extract on the ruminal hyper ammonia-producing bacterium, Clostridium sticklandii. Curr Microbiol 61: 125–131.
Fonknechten N, Chaussonnerie S, Tricot S, Lajus A, Andreesen JR, Perchat N et al. (2010). Clostridium sticklandii, a specialist in amino acid degradation: revisiting its metabolism through its genome sequence. BMC Genomics 11: 555.
Gogarten JP, Doolittle WF, Lawrence JG . (1999). Prokaryotic evolution in light of gene transfer. Mol Biol 19: 2226–2238.
Hess M, Sczyrba A, Egan R, Kim T-W, Chokhawala H, Schroth G et al. (2011). Metagenomic discovery of biomass-degrading genes and genomes from cow rumen. Science 331: 463–467.
Huws SA, Edwards JE, Creevey CJ, Rees Stevens P, Lin W, Girdwood SE et al. (2016). Temporal dynamics of the metabolically active rumen bacteria colonizing fresh perennial ryegrass. FEMS Microbiol Ecol 92: fiv137.
Huws SA, Kim EJ, Lee MR, Scott MB, Tweed JK, Pinloche E et al. (2011). As yet uncultured bacteria phylogenetically classified as Prevotella, Lachnospiraceae incertae sedis and unclassified Bacteroidales, Clostridiales and Ruminococcaceae may play a predominant role in ruminal biohydrogenation. Environ Microbiol 13: 1500–1512.
Jain R, Rivera MC, Moore JE, Lake JA . (2003). Horizontal gene transfer accelerates genome innovation and evolution. Mol Biol Evol 20: 1598–1602.
Janssen PH, Kirs M . (2008). Structure of the archaeal community of the rumen. Appl Environ Microbiol 74: 3619–3625.
Kanehisa M, Goto S, Sato Y, Furumichi M, Tanabe M . (2012). KEGG for integration and interpretation of large-scale molecular data sets. Nucleic Acids Res 40: 109–114.
Kelly AK, McGee M, Crews DH, Fahey AG, Wylie AR, Kenny DA . (2010). Effect of divergence in residual feed intake on feeding behavior, blood metabolic variables, and body composition traits in growing beef heifers. J Anim Sci 88: 109–123.
Koenig JE, Spor A, Scalfone N, Fricker AD, Stombaugh J, Knight R et al. (2011). Succession of microbial consortia in the developing infant gut microbiome. Proc Natl Acad Sci USA 108 (Suppl): 4578–4585.
Langmead B, Salzberg SL . (2012). Fast gapped-read alignment with Bowtie 2. Nat Methods 9: 357–359.
Lauro FM, McDougald D, Thomas T, Williams TJ, Egan S, Rice S et al. (2009). The genomic basis of trophic strategy in marine bacteria. Proc Natl Acad Sci USA 106: 15527–15533.
Li WH . (1993). Unbiased estimation of the rates of synonymous and nonsynonymous substitution. J Mol Evol 36: 96–99.
Loughran NB, Hinde S, McCormick-Hill S, Leidal KG, Bloomberg S, Loughran ST et al. (2012). Functional consequence of positive selection revealed through rational mutagenesis of human myeloperoxidase. Mol Biol Evol 29: 2039–2046.
Lundberg DS, Lebeis SL, Paredes SH, Yourstone S, Gehring J, Malfatti S et al. (2012). Defining the core Arabidopsis thaliana root microbiome. Nature 488: 86–90.
Marco D . (2008). Metagenomics and the niche concept. Theory Biosci 127: 241–247.
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A et al. (2010). The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 20: 1297–1303.
Mishra M, Martz FA, Stanley RW, Johnson HD . (1970). Effect of diet and ambient temperature-humidity on ruminal pH, oxidation reduction potential, ammonia and lactic acid in lactating cows. J Anim Sci 30: 1023–1028.
Nelson-Sathi S, Sousa FL, Roettger M, Lozada-Chávez N, Thiergart T, Janssen A et al. (2014). Origins of major archaeal clades correspond to gene acquisitions from bacteria. Nature 517: 77–80.
Nevo E . (2001). Evolution of genome-phenome diversity under environmental stress. Proc Natl Acad Sci USA 98: 6233–6240.
Nölling J, Breton G, Omelchenko MV, Makarova KS, Zeng Q, Gibson R et al. (2001). Genome sequence and comparative analysis of the solvent-producing bacterium Clostridium acetobutylicum. J Bacteriol 183: 4823–4838.
Otani S, Mikaelyan A, Nobre T, Hansen LH, Koné NA, Sørensen SJ et al. (2014). Identifying the core microbial community in the gut of fungus-growing termites. Mol Ecol 23: 4631–4644.
Oxenbøll S, Scheller HV, Jensen JK, Sørensen SO, Harholt J, Geshi N . (2007). Biosynthesis of pectin. Plant Physiol 129: 283–295.
Poulsen M, Schwab C, Jensen BB, Engberg RM, Spang A, Canibe N et al. (2013). Methylotrophic methanogenic Thermoplasmata implicated in reduced methane emissions from bovine rumen. Nat Commun 4: 1428.
Powell S, Szklarczyk D, Trachana K, Roth A, Kuhn M, Muller J et al. (2012). eggNOG v3.0: orthologous groups covering 1133 organisms at 41 different taxonomic ranges. Nucleic Acids Res 40: D284–D289.
Purushe J, Fouts DE, Morrison M, White BA, Mackie RI, Coutinho PM et al. (2010). Comparative genome analysis of Prevotella ruminicola and Prevotella bryantii insights into their environmental niche. Microb Ecol 60: 721–729.
Qi M, Wang P, O’Toole N, Barboza PS, Ungerfeld E, Leigh MB et al. (2011). Snapshot of the eukaryotic gene expression in muskoxen rumen—a metatranscriptomic approach. PLoS One 6: e20521.
R Core Team. (2014). R: A Language and Environment for Statistical Computing. Available at: http://www.r-project.org.
Ravachol J, Borne R, Tardif C, de Philip P, Fierobe H-P . (2014). Characterization of all family-9 glycoside hydrolases synthesized by the cellulosome-producing bacterium Clostridium cellulolyticum. J Biol Chem 289: 7335–7348.
Rubino F, Creevey CJ . (2014). MGkit: Metagenomic Framework for the Study of Microbial Communities. Available at: https://bitbucket.org/setsuna80/mgkit.
Russell JB, Mantovani HC . (2002). The bacteriocins of ruminal bacteria and their potential as an alternative to antibiotics. J Mol Microbiol Biotechnol 4: 347–355.
Schloissnig S, Arumugam M, Sunagawa S, Mitreva M, Tap J, Zhu A . (2013). Genomic variation landscape of the human gut microbiome. Nature 493: 45–50.
Scholz MB, Lo C-C, Chain PSG . (2012). Next generation sequencing and bioinformatic bottlenecks: the current state of metagenomic data analysis. Curr Opin Biotechnol 23: 9–15.
Servinsky MD, Germane KL, Liu S, Kiel JT, Clark AM, Shankar J et al. (2012). Arabinose is metabolized via a phosphoketolase pathway in Clostridium acetobutylicum ATCC 824. J Ind Microbiol Biotechnol 39: 1859–1867.
Sievers F, Wilm A, Dineen D, Gibson TJ, Karplus K, Li W et al. (2011). Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol Syst Biol 7: 539.
Simpson JT, Durbin R . (2012). Efficient de novo assembly of large genomes using compressed data structures. Genome Res 22: 549–556.
van der Walt S, Chris Colbert S, Varoquaux G . (2011). The NumPy Array: A Structure for Efficient Numerical Computation. Computing in Science and Engineering 13: 22–30.
Steinberg LM, Regan JM . (2008). Phylogenetic comparison of the methanogenic communities from an acidic, oligotrophic fen and an anaerobic digester treating municipal wastewater sludge. Appl Environ Microbiol 74: 6663–6671.
Stewart SS . (1976). Factors affecting the cellulolytic activity of rumen contents. Appl Environ Microbiol 33: 497–502.
Takahashi N, Yamada T . (2000). Pathways for amino acid metabolism by Prevotella intermedia and Prevotella nigrescens. Oral Microbiol Immunol 15: 96–102.
Team P . (2009). Picard Tools. Available at: https://broadinstitute.github.io/picard/.
Teeling H, Fuchs BM, Becher D, Klockow C, Gardebrecht A, Bennke CM et al. (2012). Substrate-controlled succession of marine bacterioplankton populations induced by a phytoplankton bloom. Science 336: 608–611.
Temperton B, Giovannoni SJ . (2012). Metagenomics: microbial diversity through a scratched lens. Curr Opin Microbiol 15: 605–612.
The Uniprot Consortium. (2012). Reorganizing the protein space at the Universal Protein Resource (UniProt). Nucleic Acids Res 40: D71–D75.
Toft C, Andersson SGE . (2010). Evolutionary microbial genomics: insights into bacterial host adaptation. Nat Rev Genet 11: 465–475.
Turnbaugh PJ, Hamady M, Yatsunenko T, Cantarel BL, Duncan A, Ley RE et al. (2009). A core gut microbiome in obese and lean twins. Nature 457: 480–484.
Tymensen LD, McAllister TA . (2012). Community structure analysis of methanogens associated with rumen protozoa reveals bias in universal archaeal primers. Appl Environ Microbiol 78: 4051–4056.
Wallace RJ . (1996). Ruminal microbial metabolism of peptides and amino acids. J Nutr 126: 1326S–1334SS.
Weckx S, Allemeersch J, Van Der Meulen R, Vrancken G, Huys G, Vandamme P et al. (2011). Metatranscriptome analysis for insight into whole-ecosystem gene expression during spontaneous wheat and spelt sourdough fermentations. Appl Environ Microbiol 77: 618–626.
Williams LM, Oleksiak MF . (2011). Ecologically and evolutionarily important SNPs identified in natural populations. Mol Biol Evol 28: 1817–1826.
Yanagita K, Kamagata Y, Kawaharasaki M, Suzuki T, Nakamura Y, Minato H . (2000). Phylogenetic analysis of methanogens in sheep rumen ecosystem and detection of Methanomicrobium mobile by fluorescence in situ hybridization. Biosci Biotechnol Biochem 64: 1737–1742.
Young MD, Wakefield MJ, Smyth GK, Oshlack A . (2010). Gene ontology analysis for RNA-seq: accounting for selection bias. Genome Biol 11: R14.
Yu Z, Morrison M . (2004). Improved extraction of PCR-quality community DNA from digesta and fecal samples. Biotechniques 36: 808–812.
Acknowledgements
This research was conducted as part of the EU Animal-Change project, which received funding from the European Community's Seventh Framework Programme (FP7/2007-2013) under the grant agreement no. 266018. FR was supported by Teagasc and the Walsh Fellowships scheme. CJC was funded under the Science Foundation Ireland (SFI) Stokes lecturer scheme (07/SK/B1236A) and the Biotechnology and Biological Sciences Research Council (BBSRC) Institute Strategic Programme Grant, Rumen Systems Biology, (BB/E/W/10964A01).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Rubino, F., Carberry, C., M Waters, S. et al. Divergent functional isoforms drive niche specialisation for nutrient acquisition and use in rumen microbiome. ISME J 11, 932–944 (2017). https://doi.org/10.1038/ismej.2016.172
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2016.172
This article is cited by
-
A compendium of ruminant gastrointestinal phage genomes revealed a higher proportion of lytic phages than in any other environments
Microbiome (2024)
-
Colonization and development of the gut microbiome in calves
Journal of Animal Science and Biotechnology (2023)
-
Microbiome-based interventions to modulate gut ecology and the immune system
Mucosal Immunology (2022)
-
The microbiome of the buffalo digestive tract
Nature Communications (2022)
-
Genomic insights into the phylogeny and biomass-degrading enzymes of rumen ciliates
The ISME Journal (2022)