Abstract
Recent studies have provided an unprecedented view of the microbial communities colonizing captive mice; yet the host and environmental factors that shape the rodent gut microbiota in their natural habitat remain largely unexplored. Here, we present results from a 2-year 16 S ribosomal RNA gene sequencing-based survey of wild wood mice (Apodemus sylvaticus) in two nearby woodlands. Similar to other mammals, wild mice were colonized by 10 bacterial phyla and dominated by the Firmicutes, Bacteroidetes and Proteobacteria. Within the Firmicutes, the Lactobacillus genus was most abundant. Putative bacterial pathogens were widespread and often abundant members of the wild mouse gut microbiota. Among a suite of extrinsic (environmental) and intrinsic (host-related) factors examined, seasonal changes dominated in driving qualitative and quantitative differences in the gut microbiota. In both years examined, we observed a strong seasonal shift in gut microbial community structure, potentially due to the transition from an insect- to a seed-based diet. This involved decreased levels of Lactobacillus, and increased levels of Alistipes (Bacteroidetes phylum) and Helicobacter. We also detected more subtle but statistically significant associations between the gut microbiota and biogeography, sex, reproductive status and co-colonization with enteric nematodes. These results suggest that environmental factors have a major role in shaping temporal variations in microbial community structure within natural populations.
Similar content being viewed by others
Introduction
Mammals are home to trillions of microbes in their gastrointestinal tract (the gut microbiota), which impact multiple aspects of host health and disease (Sommer and Backhed, 2013). Elucidating the ecological and evolutionary processes that shape host-associated microbial communities remains a major outstanding goal (Costello et al., 2012). Laboratory rodents are a valuable tool to dissect the relative contributions of intrinsic and extrinsic factors (Carmody et al., 2015); however, it remains unclear whether these interactions can be generalized to mammals in their natural habitat. Recent studies have provided an initial view into the ecological factors linked to inter-individual variations in the gut microbiotas of wild animals. Comparative analyses suggest that diet is a major environmental factor contributing to gut microbial variation between mammalian species (Muegge et al., 2011). Diet also shapes the gut microbiota within a species, as evidenced by longitudinal analyses of the black howler monkey gut microbiota (Amato et al., 2013, 2015) and dietary perturbation experiments in wild-caught mice and fish (Bolnick et al., 2014; Wang et al., 2014). Biogeographic variation in the gut microbiota at large spatial scales has also been reported in house mice (Linnenbrink et al., 2013). Finally, host-specific factors like co-colonization with enteric parasites (Hayes et al., 2010; Keeney and Finlay, 2011) and host genetics (Benson et al., 2010; Ochman et al., 2010; McKnite et al., 2012; Goodrich et al., 2014) may also contribute to inter-individual and temporal variations in gut microbial community structure.
Yet, the relative strengths of these various factors, and their interactions, remain unclear owing to the lack of systematic analyses that monitor both intrinsic and extrinsic factors in natural populations. Such an analysis would require tractable systems wherein host factors, environmental parameters and temporal variations in the gut microbiota can be monitored in situ. Here, we report findings from such a study in well-characterized populations of wood mice (Apodemus sylvaticus) in the United Kingdom, which we monitored for 2 years. We simultaneously measured multiple environmental (season, location, population density) and host (age, sex, reproductive status, parasite infection status) parameters, and repeatedly sampled multiple individuals over time. Using these data, we examine the relative importance of environmental and intrinsic host factors in shaping gut microbial community variation between and within-individuals over time. We discovered a notable seasonal variation in gut microbial community structure, which we propose is caused by changes in host dietary intake. We also found evidence for an impact of spatial structure over a smaller scale than previously reported, reproductive status and nematode colonization. Together, our results provide an initial view of the wild wood mouse gut microbiota and support the hypothesis that environmental factors such as changes in food availability and subsequent dietary intake have a dominant role in shaping wild mammal gut microbial communities.
Materials and methods
Sample collection
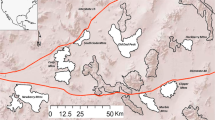
In 2010 and 2011, A. sylvaticus were trapped on six grids in two mixed woodlands (Manor and Haddon Wood; Supplementary Figure S1) on the Wirral peninsula, UK. On each grid, two live traps baited with grain and bedding material were placed every 10 meters in a 70 m × 70 m square, and trapped monthly from May to November for three consecutive nights in both years. In 2011, trapping was also performed for two consecutive nights during one additional week in each of the months August, September, October and November, though no treatments were given. Trapped animals were tagged using subcutaneous passive integrated transponder tags, so they could be individually identified upon recapture. Fecal samples were collected from all traps containing a single animal and stored in 10% buffered formalin for identification of gut parasites (Knowles et al., 2013). A sub-sample was also collected for characterization of the gut microbiota, which was frozen at −80oC within 8 h of collection. To assess the potential effect of overnight temperature on gut microbial communities, we retrieved temperature data for each sampling night from the Hawarden Chester airport weather station near our field sites between the hours of 6pm and 12pm the following day, the time from which mice could enter traps to when we collected fecal samples.
Host phenotyping
Animals were aged as either juvenile, sub-adult or adult according to pelage in the first instance, with body mass used as a secondary trait where pelage was inconclusive (juvenile <12 g, sub-adult: 12–16 g, adult >16 g). Body length, weight, sex and reproductive status were recorded. Animals were characterized as being either reproductively active (descended or protruding testes for males, pregnant or with a perforate vagina for females) or inactive. A subset of the mice was given antiparasitic treatments, including Ivermectin and Toltrazuril (2010), and Ivermectin, Fipronil, Pyrantel pamoate, or two-drug combinations (2011). We did not detect any significant impact of treatment on the gut microbiota (Table 1). Blood samples were tested for Bartonella using a nested PCR assay (Knowles et al., 2013).
16 S rRNA gene sequencing and analysis
16 S ribosomal RNA (rRNA) gene sequencing was performed on fecal samples collected from each trap to characterize the distal gut microbiota (n=481 samples, 196 555±24 236 sequences per sample; Supplementary Table S1). DNA was extracted using the PowerSoil bacterial DNA extraction kit (MoBio, Carlsbad, CA, USA), and the V4 region of the 16 S rRNA gene was PCR-amplified in triplicate using custom barcoded universal bacterial primers with the following protocol: 94 °C for 3 min, 35 cycles of 94°C for 45 s, 50 °C for 30 s, and 72 °C for 90 s, with a final extension at 72 °C for 10 min (Maurice et al., 2013). Triplicates were pooled, confirmed by gel electrophoresis, cleaned with the Ampure XP kit (Agencourt, Danvers, MA, USA), quantified using the Quant-iT Picogreen dsDNA Assay Kit (Invitrogen, Carlsbad, CA, USA), and sequenced on the Illumina HiSeq platform. 16 S rRNA gene sequences were analyzed using the QIIME software package (Caporaso et al., 2010). All sequences were used for the comparison of the relative abundance of bacterial taxa. Operational taxonomic units were assigned at 97% similarity against the Greengenes database (DeSantis et al., 2006), which we trimmed to span only the 16 S rRNA region flanked by our sequencing primers (positions 521–773). LefSe (Segata et al., 2012a) was run on sub-sampled data sets, after filtering out species-level phylotypes with <100 sequences or found in only one sample. Statistical analysis of Bray–Curtis dissimilarities calculated using the relative abundance of bacterial genera was conducted using RStudio (ver. 0.98.1091) and the adonis function in the R package ‘vegan’ (Oksanen et al., 2015). Only the first sample was included for each mouse to avoid artifacts caused by within animal comparisons. Significance values were computed using 10 000 permutations.
Parasite diagnosis
Gastrointestinal parasites (nematodes, cestodes and Eimeria protozoa) were detected using the salt flotation technique (Pritchard and Kruse, 1982). Saturated salt solution was added to formalin-preserved fecal samples, such that eggs and oocysts in each sample could be concentrated on a coverslip, and scanned for parasite detection at × 10 magnification. × 40 magnification was used for parasite identification. Coccidia (parasites belonging to the genus Eimeria) were identified using unsporulated oocyst morphology (Nowell and Higgs, 1989), and helminths using egg morphology. For each parasite species, the number of eggs or oocysts per gram of feces was calculated for each sample. When multiple samples were present for an individual within a 3-day trapping period, the arithmetic mean egg/oocyst count was taken across these days. The dominant parasites detected were nematodes (largely Heligmosomoides polygyrus) and coccidia, and thus our analyses focus on these two parasite groups.
Linear mixed models
We performed linear mixed models using the lme4 package in R v.3.0.1 (Bates et al., 2014). We controlled for repeated sampling of individual mice by including individual ID as a random intercept term. Model assumptions were checked by examining the distribution of residuals and plotting fitted values against residuals; response variables were square root or log-transformed where necessary to ensure model assumptions were met. For models of individual genera, only samples with non-zero abundance were included. In all starting models, the same set of predictors was included: external overnight temperature, grid, month, year, age, sex, nematode infection status, Eimeria infection status, drug treatment and reproductive status. Several interaction terms were included: year by month; reproductive status by sex; and parasite infection variables by treatment. Only samples for which full metadata on all the above metrics were available were included (Supplementary Table S2). All models were initially simplified by backwards-stepwise elimination of terms with P-value >0.10, beginning with interactions, and the final minimal model included only terms with P-value <0.05. Adjusted P-values (q-values) were calculated based on the 'Graphically Sharpened' false discovery rate method (Pike, 2011).
Spatial structuring of microbial communities
As wood mice are territorial and have home ranges smaller than our trapping grids (Godsall et al., 2014), fine-scale spatial variation in microhabitat and food availability could influence gut microbial ecology, both within and across our trapping grids. To test for biogeographic effects at this scale, we examined spatial autocorrelation in the gut microbiota according to mouse capture location. Spatial autocorrelations were measured using the Moran's I statistic (Moran, 1950). Only the first sample was included for each mouse to avoid artifacts caused by within animal comparisons. Genera found in ⩾10 samples (or mice) were analyzed, along with the first principal coordinates from our Bray–Curtis, unweighted UniFrac and weighted UniFrac analyses. We used a binary spatial weights matrix, with spatial neighborhoods defined as being 0–50 m apart. Data from Manor and Haddon woods were analyzed both together and separately. To control for temporal trends, we restricted our analysis to samples collected between August and November and analyzed the 2 years separately. Spatial weight matrices were row-standardized. The significance of Moran's I values was assessed with permutation tests, coded using a Markov chain Monte Carlo algorithm. For each P-value, 10 chains of length 1 000 000 were run, each starting from a random initial permutation. These settings were judged to give good chain convergence based on examination of running mean plots. We used the software packages GeoDa and PySAL (https://geodacenter.asu.edu). Batch scripts/code are available on request.
Results
The wild mouse gut harbors abundant Lactobacilli and putative enteric pathogens
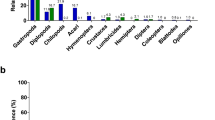
Consistent with results in captive and wild mammals (Ley et al., 2008a), wild wood mice were colonized by 10 bacterial phyla: Firmicutes (52.1±1.0% 16 S rRNA gene sequences; mean±s.d.), Bacteroidetes (37.0±0.9%), Proteobacteria (8.2±0.5%), Actinobacteria (1.1±0.2%), Tenericutes (0.9±0.1%), Deferribacteres (0.4±0.1%), Cyanobacteria (0.3±0.03%), Verrucomicrobia (0.03±0.03%), Fusobacteria (0.01±0.01%) and TM7 (0.004±0.0004%) (Figure 1a). Within the Firmicutes, the dominant bacterial order was the Lactobacillales (genus: Lactobacillus) (Figure 1b, Supplementary Figure S2). We also observed multiple α-, ɛ- and γ-proteobacterial genera that include potential bacterial pathogens: for example, Bartonella, Helicobacter, Pseudomonas, Rickettsiella and Yersinia (Figure 1b, Supplementary Figure S2, Supplementary Table S3). All nine of the mice with detectable fecal Bartonella also tested positive in time-matched blood samples, leading to a significant association between blood and fecal detection of this genus (P-value<0.05, χ2-test). Many of these genera were widespread, most notably Helicobacter (97.9% of samples), Pseudomonas (73.2%) and Yersinia (44.5%). The same was true for intestinal parasites (Supplementary Table S4), including H. polygyrus (40%) and Eimeria hungaryensis (29.3%).
Taxonomic analysis of the wild mouse gut microbiota. Pie charts represent the relative abundance of bacterial (a) phyla and (b) orders (n=481 samples). The 10 most abundant phyla and orders are shown (phyla with a mean abundance <0.001% are not included; the remaining orders are represented by the ‘other’ slice). Taxa are colored based on phylum. Sequences within the Cyanobacteria phylum could be attributed to chloroplasts (order Streptophyta), non-photosynthetic bacteria related to Cyanobacteria that are common in the mammalian gut (order YS2) (Di Rienzi et al., 2013), and algae (order Chlorophyta, family Trebouxiophyceae). We did not detect any consistent seasonal changes in the abundance or prevalence of these three groups.
Marked seasonal variation in microbial community structure
Analysis of Bray–Curtis dissimilarity among samples revealed a clear seasonal pattern differentiating samples collected in the spring/early summer (May through July) and those collected in late summer/fall (August through November) (Figure 2a, Supplementary Figure S3; P-value<0.001, PERMANOVA of Bray–Curtis distances). The observed seasonal shift in the microbiota coincides with the expected timing of an annual transition to a seed-based diet from a more insect-based diet (Watts, 1968), and may therefore be driven by a seasonal shift in food availability and diet. Consistent with this hypothesis, the mean microbial community structure for each month was significantly correlated between the 2 years (Figure 2b; R2=79%, P-value<0.01). The association between season and microbial community structure was significant in both years when considered independently, although the difference was more marked in 2010 (pseudo F-value=31.2 (2010) versus 9.9 (2011), P-value<0.001 for both years; PERMANOVA test). Statistical analysis with the LefSe software package revealed taxonomic groups ranging from the phylum to genus level that were consistently associated with season in both years (Supplementary Table S5). Lactobacillus was found at a significantly higher abundance in the spring of both years, whereas Alistipes and Helicobacter were consistently enriched in the fall (Figure 2c).
Seasonal variations in the wild mouse gut microbiota. (a) The first principle coordinate from a Bray–Curtis-based analysis of microbial community structure over time. Trend lines were generated by fitting a polynomial function to values from each year (GraphPad Prism version 6.0). Values are mean±s.e.m. (n=14–80 samples per group). Values from June and July were combined in 2011 owing to limited available samples in July (n=2). (b) Association between average monthly microbial community structures between years. Values are mean (thick black line) and 95% CI (thin grey lines) from a linear regression. (c) The relative abundance of bacterial genera in spring and fall of both years. Values are mean±s.e.m. (n=24–123 samples per group; the first sample from each mouse was included). Asterisks represent significant differences (P-value<0.05, Wilcoxon rank-sum test).
Analysis of mice captured multiple times within a year confirmed that these microbial changes occurred within individuals and were not simply due to mouse population turnover (that is, seasonal changes in the types of individual captured). We observed within individual shifts in microbiota structure in both years of the study that followed the overall population trend (Supplementary Figure S4). This was reflected by a strong positive correlation between month-to-month differences in the mean population-wide value for Bray–Curtis principal coordinates 1 and 2 (excluding repeat captured individuals) and the mean within-individual change in these metrics (PC1 R2=65%; PC2 R2=56%; both P-value<0.05, linear regression; Figures 3a and b). Analysis of 25 mice captured in both seasons confirmed that in nearly all cases there was a consistent direction of change (Figure 3c; P-value<0.0001, Wilcoxon rank-sum test).
Seasonal patterns are detectable within-individuals captured multiple times. Correlations between the mean month-to-month change in Bray–Curtis principle coordinates 1 (panel a) and 2 (panel b) within-individuals, relative to the monthly change observed at the population level (including only one pair of observations per mouse; n=2–11 paired samples per datapoint). Dots represent monthly changes seen in 2010 (black) and 2011 (white). See Supplementary Figure S4 for plots of individual animals over time. (c) We calculated the mean value of Bray–Curtis principal coordinate 1 value for each mouse in Season 1 (spring/early summer) and 2 (late summer/fall) (n=25 mice; less than equal to one sample per mouse per month included). Nearly all mice exhibited a consistent direction of change (black lines), with the exception of three animals (grey lines).
Limited spatial heterogeneity in community structure
Although microbial community structure differed significantly between the two woodlands (P-value<0.001, PERMANOVA of Bray–Curtis dissimilarities), this effect was noticeably weaker than that of season: pseudo F-value=35.3 (season) versus 4.6 (wood) when considering both years. Consistent with this weak effect, LefSe analysis only identified two nested taxa that were significantly enriched in Manor Wood: the Clostridia class and the Clostridiales order (LDA>2, P-value<0.05). We did not detect any taxa that were significantly enriched in Haddon Wood.
To quantify the spatial structure of the wild mouse gut microbiota in more detail, we evaluated spatial autocorrelation at the genus level and using community dissimilarity metrics (see Methods). In 2010, we detected significant spatial autocorrelation for the Bray–Curtis and unweighted UniFrac metrics (Figure 4; q-value<0.01). However, these patterns were weaker and only present for unweighted UniFrac in 2011, and were absent in all cases when we only considered samples from Haddon or Manor wood. Similarly, analyses of bacterial genera failed to detect significant spatial autocorrelation for 117 of the 117 tested groups during either year (q-value<0.01). Moreover, the maximum Moran's I value for this distance class was 0.138, further indicating nonexistent or weak spatial associations. Together, these analyses suggest that although the overall pattern of microbial community structure was distinct between Haddon and Manor Wood, there was no evidence for finer spatial structure within woods or between individual bacterial genera.
Spatial distribution of microbial community structure. Each circle represents the physical location of a given mouse at the time of sampling in Manor or Haddon Wood, which are subdivided into two and four trapping grids, respectively. Shading is proportional to the percentile along unweighted UniFrac principal coordinate 1 (an indication of overall microbial community membership). Between August and November in 2010 there was a slight but significant difference in community composition between Haddon and Manor woods. However, this difference was absent in August to November 2011. Within woods, no significant spatial structuring of communities was observed in either year.
Multivariate modeling reveals associations with both host and environmental factors
We next used linear mixed models (see Methods) to tease apart the relative influence of multiple environmental and host factors, and to determine their effects in isolation of confounding factors. We constructed six models for community dissimilarity metrics (principal coordinates 1 and 2 for Bray–Curtis, unweighted UniFrac and weighted UniFrac) as well as separate models for the 10 most abundant bacterial genera (Table 1). Overall, these analyses suggest that the wood mouse gut microbiota is primarily shaped by environmental factors, with significant evidence for both temporal (see ‘Year’ and ‘Month’ columns) and spatial structuring (see ‘Grid’ column). These temporal trends could not simply be explained by seasonal variation in temperature (Supplementary Figure S5), as they were unaltered by inclusion of overnight temperature as a covariate (Table 1). For all community dissimilarity metrics examined and most individual genera, our minimal models included a significant year by month interaction term, indicating seasonal differences that varied somewhat across the 2 years investigated. If these interaction terms were dissolved into their component terms, strong main effects of month were observed in nearly all models, with effects of year also common though generally weaker. Consistent with our prior analysis of spatial autocorrelation, there was a strong association between the community dissimilarity metrics and trapping grid with weaker associations at the genus level. We also detected associations between some metrics and local population density at the time of capture (Table 1).
To a lesser extent than extrinsic factors like season and year, host factors such as reproductive status and sex were associated with microbial community structure, sometimes in the form of an interaction between these two terms (Table 1). For example, the abundance of Lactobacillus was higher in reproductively active than non-active females, but did not depend on reproductive status for males (Figure 5a). We also detected associations between the gut microbiota and intestinal parasites. In particular, nematode infections were inversely associated with the abundance of the most abundant Lachnospiraceae genus and positively associated with the genus Escherichia (Figure 5b). However, no significant associations between coccidia infection or anti-parasite treatment and the gut microbiota were found, possibly due to the transient nature of the intervention (monthly treatment intervals; see Methods). Age-related differences were rare, with Alistipes the only 1 of the 10 most abundant bacterial genera associated with host age, showing an increase across the age groups from juvenile to adult (Table 1).
The gut microbiota is associated with intestinal helminth infection and reproductive state. (a) Values represent the relative abundance of Lactobacillus according to host sex and reproductive status. (b) Nematode infection is positively associated with Escherichia and negatively associated with an unclassified genus within the Lachnospiraceae family. All samples with non-zero abundance were included. Values are mean±s.e.m. (n=92–205 samples per group).
To illustrate how much variation in Bray–Curtis principal coordinates 1 and 2 was explained by environmental factors like month and year, compared with host-related factors, we calculated marginal R2 statistics from our linear mixed models (Nakagawa and Schielzeth, 2013). These are equivalent to classic R2 statistics for linear models, indicating the percentage of variation explained by a given set of predictor variables (fixed effects). For both Bray–Curtis PC1 and PC2, month (that is, seasonal differences) explained a much larger proportion of variance than year (Supplementary Table S6). Inclusion of year when month was already present in the model provided little additional explanatory power (PC1: R2GLMM(m) =43.9% with month only versus 48.7% with month and year; PC2: R2GLMM(m)=10.1% with month only versus 10.5% with month and year). Furthermore, allowing the seasonal effect to vary among years (by inclusion of a month × year interaction term) yielded limited additional explanatory power for PC1 (R2GLMM(m) =52% versus 49% variance explained), with a twofold increase in variance explained for PC2 (R2GLMM(m) =19.5% versus 10% variance explained).
Thus, seasonal differences in the gut microbiota appear to dominate the differences between years and are largely consistent across years, in agreement with our earlier analyses (Figure 2). Host-related factors (age, sex, reproductive status), enteric parasite infections and host density explained some additional variance (12% more for PC1 and 8% more for PC2 than models with only month and year terms), though their contribution was again smaller than the strong seasonal effects, particularly for PC1 (Supplementary Table S6). Individual identity explained 18% of the variation in Bray–Curtis PC1 even after including all other factors. We confirmed these trends by analyzing the entire Bray–Curtis dissimilarity matrix according to season, host sex, and wood (see Methods). Although all three factors showed a significant effect, seasonal effects explained more variation (R2=13.3%, P-value<10−4) than either host sex (R2=0.8%, P-value<0.05) or spatial structure (R2=0.8%, P-value<0.05).
Discussion
At the phylum level, the wild mouse gut microbiota is comparable to that of other mammals (including humans) with two major groups, the Firmicutes and Bacteroidetes, accounting for ~90% of the 16 S rRNA gene sequencing reads (Ley et al., 2008a; Muegge et al., 2011). We also detected high levels of the Lactobacillus genus (phylum: Firmicutes; order: Lactobacillales) constituting up to one-third of the community, similar to other omnivorous mammals, such as bears, squirrels and lemurs (Supplementary Figure S6). These results confirm that to a large degree the mammalian gut microbiota assembles in a reproducible fashion regardless of the host species (Ley et al., 2008a; Muegge et al., 2011), reflective of the restricted set of microorganisms that have adapted to life in the gastrointestinal tract (Ley et al., 2008b).
In contrast to ‘specific pathogen free’ laboratory mice, we detected widespread colonization by bacterial taxa that contain enteric pathogens, including Helicobacter and other Proteobacteria. However, given the resolution of our sequencing methods and the limited studies of wild mouse pathogens we cannot exclude the fact that these are commensal strains. Despite this important caveat, our results are consistent with previous reports, indicating that wild house mice can be reservoirs of diverse Helicobacter strains capable of infecting humans and other vertebrates (O'Rourke et al., 2001; Parker et al., 2009; Wasimuddin et al., 2012). We observed that Helicobacter abundance increased in late summer/fall, when Lactobacillus levels are low. This might suggest that Lactobacillus confers protection against infection as has been demonstrated in laboratory mice (Kabir et al., 1997; Pena et al., 2005; Medellin-Pena and Griffiths, 2009; Eaton et al., 2011). Alternatively, immune status (that is, IL-22 deficiency) has been linked to the abundance of Lactobacillus (Zenewicz et al., 2013), potentially suggesting that these seasonal changes might be in part driven by the host response to bacterial infection. Additional studies will be necessary to determine how the immune system of these mice tolerates long-term enteric pathogen colonization and to characterize the reciprocal interactions between these enteric pathogens and the commensal gut microbiota.
The wild mouse gut microbiota underwent a consistent seasonal shift in both years, with a decrease in Lactobacillus and concomitant increases in Alistipes, Helicobacter and the Lachnospiraceae family (phylum: Firmicutes). A possible explanation is that mid-summer represents a transition from a diet rich in insects to a diet primarily composed of seeds (Watts, 1968), coincident with the annual seed fall, which usually starts in late July in UK woodlands (Gurnell, 1993). Thus, we propose that seasonal patterns in dietary intake drive variations in the gut microbial community structure of wild wood mice. Differences in the timing, extent and tree species composition of seed fall, which can vary markedly between years (Gurnell, 1993), may explain the observed variation between years in the magnitude of the seasonal microbiota transition observed. Notably, a recent study of rural human subjects from South Dakota revealed differences in the gut microbiota in summer relative to winter (Davenport et al., 2014), suggesting that seasonal reconfigurations may be a conserved feature of host-associated microbial communities.
If diet is indeed the dominant factor it still remains unclear what specific components of the diet might drive the observed changes to gut microbial community structure. The elevated levels of Alistipes in the fall may be reflective of increased bile acid levels triggered by an increased consumption of fat, as seen in a recent human dietary intervention study (David et al., 2014). Members of the Lachnospiraceae family, including Eubacterium rectale and Roseburia, have been linked to the fermentation of dietary plant polysaccharides in human studies (David et al., 2014; Duncan et al., 2007), and were also enriched in the fall coinciding with the increased access to plant seeds. Similarly, the source and/or dietary trigger of Lactobacillus (often a minor member of the mammalian distal gut microbiota) also remains unclear. Lactobacillus is often found in fermented foods (Wolfe et al., 2014), raising the possibility that their elevated abundance early in the year may be driven by its cultivation in wood mouse food stores over winter.
Alternatively, seasonal changes in mouse physiology, including torpor and reduced food consumption during winter, could also have a role in the observed seasonal trends. Indeed, seasonal restructuring of the gut microbiota has recently been observed in ground squirrels under controlled laboratory conditions (Carey et al., 2013). These shifts coincided with hibernation, suggesting they are driven by a shift from dietary to host-derived substrates. We detected similar patterns in wild rodents, including a decrease in the relative abundance of Lactobacillus and an increase in Alistipes from spring/early summer to late summer/fall. Although wood mice do not hibernate, they are subject to daily torpor in conditions of low temperature and food restriction. Thus, it is possible that the seasonal microbial shifts seen here may be driven by the transition to a state of intermittent torpor.
What are the potential consequences of the observed seasonal shifts in gut microbial community structure? Recent human intervention studies have shown rapid and reproducible changes in microbial community structure and function upon consumption of an animal- versus plant-based diet (David et al., 2014). These results, considered together with the current findings from wild wood mice, make it tempting to speculate that the mammalian gut microbiota may provide a rapid way to optimize caloric intake given volatile shifts in the availability of different foods. Microbial communities that could rapidly shift their metabolic activity in response to changes in host dietary intake could have enhanced dietary flexibility, likely increasing the fitness of the host and its microbial consortia.
We also found significant but weak evidence for spatial structure, unlike the more robust associations with geographic region found in recent studies of house mice (Linnenbrink et al., 2013), wild primates (Degnan et al., 2012) and humans (Yatsunenko et al., 2012). The significant spatial structure that we did find was evident only in community-wide metrics when comparing between woods. Individual bacterial genera showed no spatial structure, and no spatial structuring was evident within woods at either the community or individual genus levels. These results emphasize that the gut microbiota of these wild mouse populations is primarily shaped by factors that are not spatially structured at the scales that we considered. These results suggest that either (i) microbial dispersal occurs efficiently over distances far greater than the host range evaluated here and/or (ii) the observed bacterial taxa are long-term and stable residents of the wild wood mouse gut microbiota. Strain-level analyses of the gut microbiota (Segata et al., 2012b; Faith et al., 2013) could help determine whether there are finer differences between woods, or among areas within each wood. Furthermore, surveying wild mice across more distant sites could provide additional insight into broader biogeographical patterns.
Our linear mixed models revealed significant associations with reproductive status and intestinal parasites. Consistent with these findings, recent studies indicate that the human gut microbiota is altered during pregnancy (Koren et al., 2012), and studies in laboratory mice have shown that infection by the nematode Trichuris muris depends on the gut microbiota (Hayes et al., 2010). The associations between intestinal nematodes and the bacterial genera Escherichia (positive) and Lachnospiraceae (negative) support recent studies in humans and animal models (Walk et al., 2010; Rausch et al., 2013), though we did not find the specific association between H. polygyrus and relative Lactobacillus abundance, as recently reported in laboratory mice (Reynolds et al., 2014). Whether the associations found result from an altered immune response of the host or from direct interactions between the intestinal parasites and the gut microbiota remains to be elucidated. Determining the causal direction and underlying mechanisms of these interactions will require more extensive longitudinal analyses of wild mice before and after helminthic infection, as well as controlled studies using captured and/or captive mice.
In conclusion, despite the common use of laboratory mice to study the environmental and host factors that shape host-associated microbial communities, we still know very little about their natural state. Our results provide an initial view of the wild-wood mouse gut microbiota, emphasizing not only commonalities between mammals, but also the importance of considering temporal variations in nutritional status, enteric pathogens, reproductive status and parasite burden in setting the stage for host-microbial interactions. Follow-up observational and interventional studies of wild mice, paired with an in-depth analysis of dietary intake, are necessary to test the hypothesis that the observed seasonal trends are due to changes in diet, and could provide a complementary and tractable approach towards better understanding the causes and consequences of inter-individual variations in the mammalian gut microbiota.
References
Amato KR, Yeoman CJ, Kent A, Righini N, Carbonero F, Estrada A et al. (2013). Habitat degradation impacts black howler monkey (Alouatta pigra) gastrointestinal microbiomes. ISME J 7: 1344–1353.
Amato KR, Leigh SR, Kent A, Mackie RI, Yeoman CJ, Stumpf RM et al. (2015). The gut microbiota appears to compensate for seasonal diet variation in the wild black howler monkey (Alouatta pigra). Microb Ecol 69: 434–443.
Bates D, Maechler M, Bolker B, Walker S . (2014). lme4: Linear mixed-effects models using Eigen and S4. R package version 0.999999-2. http://CRAN.R-project.org/package=lme4.
Benson AK, Kelly SA, Legge R, Ma F, Low SJ, Kim J et al. (2010). Individuality in gut microbiota composition is a complex polygenic trait shaped by multiple environmental and host genetic factors. Proc Natl Acad Sci USA 107: 18933–18938.
Bolnick DI, Snowberg LK, Hirsch PE, Lauber CL, Knight R, Caporaso JG et al. (2014). Individuals' diet diversity influences gut microbial diversity in two freshwater fish (threespine stickleback and Eurasian perch). Ecol Lett 17: 979–987.
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK et al. (2010). QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7: 335–336.
Carey HV, Walters WA, Knight R . (2013). Seasonal restructuring of the ground squirrel gut microbiota over the annual hibernation cycle. Am J Physiol Regul Integr Comp Physiol 304: R33–R42.
Carmody RN, Gerber GK, Luevano Jr JM, Gatti DM, Somes L, Svenson KL et al. (2015). Diet dominates host genotype in shaping the murine gut microbiota. Cell Host Microbe 17: 72–84.
Costello EK, Stagaman K, Dethlefsen L, Bohannan BJ, Relman DA . (2012). The application of ecological theory toward an understanding of the human microbiome. Science 336: 1255–1262.
Davenport ER, Mizrahi-Man O, Michelini K, Barreiro LB, Ober C, Gilad Y . (2014). Seasonal variation in human gut microbiome composition. PLoS One 9: e90731.
David LA, Maurice CF, Carmody RN, Gootenberg DB, Button JE, Wolfe BE et al. (2014). Diet rapidly and reproducibly alters the human gut microbiome. Nature 505: 559–563.
Degnan PH, Pusey AE, Lonsdorf EV, Goodall J, Wroblewski EE, Wilson ML et al. (2012). Factors associated with the diversification of the gut microbial communities within chimpanzees from Gombe National Park. Proc Natl Acad Sci USA 109: 13034–13039.
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K et al. (2006). Greengenes, a chimera-checked 16 S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72: 5069–5072.
Di Rienzi SC, Sharon I, Wrighton KC, Koren O, Hug LA, Thomas BC et al. (2013). The human gut and groundwater harbor non-photosynthetic bacteria belonging to a new candidate phylum sibling to Cyanobacteria. Elife 2: e01102.
Duncan SH, Belenguer A, Holtrop G, Johnstone AM, Flint HJ, Lobley GE . (2007). Reduced dietary intake of carbohydrates by obese subjects results in decreased concentrations of butyrate and butyrate-producing bacteria in feces. Appl Environ Microbiol 73: 1073–1078.
Eaton KA, Honkala A, Auchtung TA, Britton RA . (2011). Probiotic Lactobacillus reuteri ameliorates disease due to enterohemorrhagic Escherichia coli in germfree mice. Infect Immun 79: 185–191.
Faith JJ, Guruge JL, Charbonneau M, Subramanian S, Seedorf H, Goodman AL et al. (2013). The long-term stability of the human gut microbiota. Science 341: 1237439.
Godsall B, Coulson T, Malo AF . (2014). From physiology to space use: energy reserves and androgenization explain home-range size variation in a woodland rodent. J Anim Ecol 83: 126–135.
Goodrich JK, Waters JL, Poole AC, Sutter JL, Koren O, Blekhman R et al. (2014). Human genetics shape the gut microbiome. Cell 159: 789–799.
Gurnell J . (1993). Tree seed production and food conditions for rodents in an Oak Wood in southern England. Forestry 66: 291–315.
Hayes KS, Bancroft AJ, Goldrick M, Portsmouth C, Roberts IS, Grencis RK . (2010). Exploitation of the intestinal microflora by the parasitic nematode Trichuris muris. Science 328: 1391–1394.
Kabir AM, Aiba Y, Takagi A, Kamiya S, Miwa T, Koga Y . (1997). Prevention of Helicobacter pylori infection by lactobacilli in a gnotobiotic murine model. Gut 41: 49–55.
Keeney KM, Finlay BB . (2011). Enteric pathogen exploitation of the microbiota-generated nutrient environment of the gut. Curr Opin Microbiol 14: 92–98.
Knowles SC, Fenton A, Petchey OL, Jones TR, Barber R, Pedersen AB . (2013). Stability of within-host-parasite communities in a wild mammal system. Proc Biol Sci 280: 20130598.
Koren O, Goodrich JK, Cullender TC, Spor A, Laitinen K, Backhed HK et al. (2012). Host remodeling of the gut microbiome and metabolic changes during pregnancy. Cell 150: 470–480.
Ley RE, Hamady M, Lozupone C, Turnbaugh PJ, Ramey RR, Bircher JS et al. (2008a). Evolution of mammals and their gut microbes. Science 320: 1647–1651.
Ley RE, Lozupone CA, Hamady M, Knight R, Gordon JI . (2008b). Worlds within worlds: evolution of the vertebrate gut microbiota. Nat Rev Microbiol 6: 776–788.
Linnenbrink M, Wang J, Hardouin EA, Kunzel S, Metzler D, Baines JF . (2013). The role of biogeography in shaping diversity of the intestinal microbiota in house mice. Mol Ecol 22: 1904–1916.
Maurice CF, Haiser HJ, Turnbaugh PJ . (2013). Xenobiotics shape the physiology and gene expression of the active human gut microbiome. Cell 152: 39–50.
McKnite AM, Perez-Munoz ME, Lu L, Williams EG, Brewer S, Andreux PA et al. (2012). Murine gut microbiota is defined by host genetics and modulates variation of metabolic traits. PLoS One 7: e39191.
Medellin-Pena MJ, Griffiths MW . (2009). Effect of molecules secreted by Lactobacillus acidophilus strain La-5 on Escherichia coli O157:H7 colonization. Appl Environ Microbiol 75: 1165–1172.
Meyer F, Paarmann D, D'Souza M, Olson R, Glass EM, Kubal M et al. (2008). The metagenomics RAST server - a public resource for the automatic phylogenetic and functional analysis of metagenomes. BMC Bioinformatics 9: 386.
Moran PA . (1950). Notes on continuous stochastic phenomena. Biometrika 37: 17–23.
Muegge BD, Kuczynski J, Knights D, Clemente JC, Gonzalez A, Fontana L et al. (2011). Diet drives convergence in gut microbiome functions across mammalian phylogeny and within humans. Science 332: 970–974.
Nakagawa S, Schielzeth H . (2013). A general and simple method for obtaining R2 from generalized linear mixed-effects models. Methods Ecol Evol 4: 133–142.
Nowell F, Higgs S . (1989). Eimeria species infecting wood mice (genus Apodemus) and the transfer of two species to Mus musculus. Parasitology 98: 329–336.
O'Rourke JL, Grehan M, Lee A . (2001). Non-pylori Helicobacter species in humans. Gut 49: 601–606.
Ochman H, Worobey M, Kuo CH, Ndjango JB, Peeters M, Hahn BH et al. (2010). Evolutionary relationships of wild hominids recapitulated by gut microbial communities. PLoS Biol 8: e1000546.
Oksanen J, Blanchet FG, Kindt R, Legendre P, Minchin PR, O'Hara RB et al. (2015), vegan: Community Ecology Package. R package version 2.2-1. http://CRANR-projectorg/package=vegan.
Parker SE, Malone S, Bunte RM, Smith AL . (2009). Infectious diseases in wild mice (Mus musculus) collected on and around the University of Pennsylvania (Philadelphia) Campus. Comp Med 59: 424–430.
Pena JA, Rogers AB, Ge Z, Ng V, Li SY, Fox JG et al. (2005). Probiotic Lactobacillus spp. diminish Helicobacter hepaticus-induced inflammatory bowel disease in interleukin-10-deficient mice. Infect Immun 73: 912–920.
Pike N . (2011). Using false discovery rates for multiple comparisons in ecology and evolution. Methods Ecol Evol 2: 278–282.
Pritchard MH, Kruse GOW . (1982) The Collection and Preservation of Animal Parasites, vol. 1. University of Nebraska Press: Lincoln, NE.
Rausch S, Held J, Fischer A, Heimesaat MM, Kuhl AA, Bereswill S et al. (2013). Small intestinal nematode infection of mice is associated with increased enterobacterial loads alongside the intestinal tract. PLoS One 8: e74026.
Reynolds LA, Smith KA, Filbey KJ, Harcus Y, Hewitson JP, Redpath SA et al. (2014). Commensal-pathogen interactions in the intestinal tract: lactobacilli promote infection with, and are promoted by, helminth parasites. Gut Microbes 5: 522–532.
Segata N, Izard J, Waldron L, Gevers D, Miropolsky L, Garrett WS et al. (2012a). Metagenomic biomarker discovery and explanation. Genome Biol 12: R60.
Segata N, Waldron L, Ballarini A, Narasimhan V, Jousson O, Huttenhower C . (2012b). Metagenomic microbial community profiling using unique clade-specific marker genes. Nat Methods 9: 811–814.
Sommer F, Backhed F . (2013). The gut microbiota—masters of host development and physiology. Nat Rev Microbiol 11: 227–238.
Walk ST, Blum AM, Ewing SA, Weinstock JV, Young VB . (2010). Alteration of the murine gut microbiota during infection with the parasitic helminth Heligmosomoides polygyrus. Inflamm Bowel Dis 16: 1841–1849.
Wang J, Linnenbrink M, Kunzel S, Fernandes R, Nadeau MJ, Rosenstiel P et al. (2014). Dietary history contributes to enterotype-like clustering and functional metagenomic content in the intestinal microbiome of wild mice. Proc Natl Acad Sci USA 111: E2703–E2710.
Wasimuddin, Cizkova D, Bryja J, Albrechtova J, Hauffe HC, Pialek J . (2012). High prevalence and species diversity of Helicobacter spp. detected in wild house mice. Appl Environ Microbiol 78: 8158–8160.
Watts CHB . (1968). Foods eaten by wood mice (Apodemus sylvaticus) and bank voles (Clethrionomys glareolus) in Wytham Woods, Berkshire. J Anim Ecol 37: 25–41.
Wolfe BE, Button JE, Santarelli M, Dutton RJ . (2014). Cheese rind communities provide tractable systems for in situ and in vitro studies of microbial diversity. Cell 158: 422–433.
Yatsunenko T, Rey FE, Manary MJ, Trehan I, Dominguez-Bello MG, Contreras M et al. (2012). Human gut microbiome viewed across age and geography. Nature 486: 222–227.
Zenewicz LA, Yin X, Wang G, Elinav E, Hao L, Zhao L et al. (2013). IL-22 deficiency alters colonic microbiota to be transmissible and colitogenic. J Immunol 190: 5306–5312.
Acknowledgements
Claire Reardon and Christian Daly for sequencing support; Rachel Carmody, Lawrence David and Clint Cario for helpful discussions. This work was supported by grants from the Natural Environmental Research Council (NE/G006830/1, NE/G007349/1; ABP,AF), the National Institutes of Health (P50 GM068763; PJT), the Harvard Bauer Fellows program, the National Science Foundation (DMS-1069303; KSP), the Gordon and Betty Moore Foundation (grant #3300; KSP), the Wellcome Trust (AP fellowship from a Strategic Grant to CIIE, 095831) and a gift from the San Simeon Fund (KSP). Sequencing reads are in MG-RAST (Meyer et al., 2008) under the accession number 12064.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Maurice, C., CL Knowles, S., Ladau, J. et al. Marked seasonal variation in the wild mouse gut microbiota. ISME J 9, 2423–2434 (2015). https://doi.org/10.1038/ismej.2015.53
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2015.53
This article is cited by
-
Bacteroidetes to Firmicutes: captivity changes the gut microbiota composition and diversity in a social subterranean rodent
Animal Microbiome (2023)
-
Turnover of strain-level diversity modulates functional traits in the honeybee gut microbiome between nurses and foragers
Genome Biology (2023)
-
Effects of age, seasonality, and reproductive status on the gut microbiome of Southern White Rhinoceros (Ceratotherium simum simum) at the North Carolina zoo
Animal Microbiome (2023)
-
Discrete patterns of microbiome variability across timescales in a wild rodent population
BMC Microbiology (2023)
-
Alterations in sea urchin (Mesocentrotus nudus) microbiota and their potential contributions to host according to barren severity
npj Biofilms and Microbiomes (2023)