Abstract
Microorganisms in biological wastewater treatment plants require adaptive strategies to deal with rapidly fluctuating environmental conditions. At the population level, the filamentous bacterium Candidatus Microthrix parvicella (Ca. M. parvicella) has been found to fine-tune its gene expression for optimized substrate assimilation. Here we investigated in situ substrate assimilation by single cells of Ca. M. parvicella using nano-scale secondary-ion mass spectrometry (nanoSIMS). NanoSIMS imaging highlighted phenotypic heterogeneity among Ca. M. parvicella cells of the same filament, whereby 13C-oleic acid and 13C-glycerol-3-phosphate assimilation occurred in ≈21–55% of cells, despite non-assimilating cells being intact and alive. In response to alternating aerobic–anoxic regimes, 13C-oleic acid assimilation occurred among subpopulations of Ca. M. parvicella cells (≈3–28% of cells). Furthermore, Ca. M. parvicella cells exhibited two temperature optima for 13C-oleic acid assimilation and associated growth rates. These results suggest that phenotypic heterogeneity among Ca. M. parvicella cells allows the population to adapt rapidly to fluctuating environmental conditions facilitating its widespread occurrence in biological wastewater treatment plants.
Similar content being viewed by others
Main
Activated sludge-based biological wastewater treatment plants (BWWTPs) rely on the substrate assimilation capabilities of microorganisms to drive metabolic transformations culminating in wastewater remediation (Daims et al., 2006). Frequent changes in the influent substrate composition and variations in environmental factors as well as alternating aerobic and anoxic phases result in BWWTPs representing highly fluctuating environments. Therefore, microbial populations in BWWTPs require adaptive strategies to deal with these continuous perturbations.
Laboratory-based studies have suggested that phenotypic heterogeneity among individual cells of isogenic populations confers adaptive advantages in fluctuating environments (De Jong et al., 2011; Levy et al., 2012). Phenotypic heterogeneity may reflect a bet-hedging strategy whereby multiple phenotypes of isogenic populations constitute a series of bets in response to rapidly changing environmental conditions (Levy et al., 2012). In particular population-level variations in the expression of genes involved in carbon assimilation allows populations to hedge their bets (De Jong et al., 2012). Single-cell approaches allow the study of within-population phenotypic heterogeneity (Grimbergen et al., 2015). Nano-scale secondary-ion mass spectrometry (nanoSIMS), which allows visualization and quantification of differences in substrate assimilation among individual microbial cells, is particularly well suited for this task (Zimmermann et al., 2015).
Candidatus Microthrix parvicella (Ca. M. parvicella) is a ubiquitous lipid-accumulating filamentous bacterium that can dominate municipal BWWTPs resulting in operational difficulties, such as sludge bulking and foaming (Rossetti et al., 2005). Based on laboratory, in situ and genomic investigations, Ca. M. parvicella appears to be metabolically versatile and can assimilate diverse carbon substrates while being adaptable to a wide range of environmental conditions, for example, oxygen concentrations and temperatures (Andreasen and Nielsen, 1998; Tandoi et al., 1998; Nielsen et al., 2002; Muller et al., 2012; McIlroy et al., 2013). A previous in situ microautoradiographic study has highlighted differences in substrate assimilation among Ca. M. parvicella filaments (Kindaichi et al., 2013). At the population-level, recent community-wide integrated omic analyses indicate that Ca. M. parvicella exhibits varying levels of expression for genes involved in substrate assimilation (primarily long-chain fatty acids; Muller et al., 2014a) but exhibits overall low levels of genetic variation (McIlroy et al., 2013; Muller et al., 2014a). Based on these observations, we hypothesized that phenotypic heterogeneity among individual Ca. M. parvicella cells might be a mechanism for the population to adapt to the rapidly changing environmental conditions encountered in BWWTPs.
Here we investigated substrate assimilation by Ca. M. parvicella cells using 13C-oleic acid, 13C-triolein, 13C-glycerol and 13C-glycerol-3-phosphate. Four independent time-series incubation experiments were performed each in duplicate (Figure 1a, details in Supplementary Methods). Single-cell substrate assimilation of Ca. M. parvicella was quantified using a combination of fluorescence in situ hybridization and nanoSIMS (Figures 1b and c) as well as bulk stable isotopic analyses using liquid chromatography coupled to tandem mass spectrometry (Supplementary Table S1). Furthermore, we verified the integrity and cellular morphology of Ca. M. parvicella cells and filaments using atomic force microscopy (Figures 1b and c).
I n situ phenotypic heterogeneity in substrate assimilation by ‘Ca. M. parvicella’. (a) Overview of the four independent isotopic incubation experiments. All experiments were conducted at 25 °C, except for the temperature-dependent experiment for which various temperature ranges were used. (b) Fluorescence in situ hybridization (FISH) with a ‘Ca. M. parvicella’-specific probe followed by atomic force microscopy (AFM) imaging to verify cellular integrity among ‘Ca. M. parvicella’ cells. (c) The same region was analyzed using nanoSIMS to obtain 13C-isotopic enrichment information. AFM and nanoSIMS images were overlayed to highlight the distribution of newly assimilated substrates among ‘Ca. M. parvicella’ cells. Regions of interest around individual ‘Ca. M. parvicella’ cells were defined manually using the corresponding FISH images and their corresponding 13C atomic percentages were subsequently calculated. (d) 13C-oleic acid assimilation at different time points under either aerobic or anoxic conditions. (e) Temperature-dependent aerobic assimilation of 13C-oleic acid by single cells of ‘Ca. M. parvicella’ after 5 h of incubation. (f) 13C-glycerol-3-phosphate assimilation under aerobic or anoxic conditions when administered as a single substrate. (g) Assimilation of 13C-oleic acid following alternating aerobic–anoxic conditions. (d–g) The dotted line indicates the 13C atomic percentage of ‘Ca. M. parvicella’ single cells from time point 0 h.
First, we investigated potential fine-scale differences in the fatty acid assimilation of 13C-triolein and 13C-oleic acid under aerobic and anoxic conditions. The 13C-oleic acid assimilation rates by Ca. M. parvicella cells were most pronounced under anoxic conditions after 1 h of incubation (Figure 1d), underlining the preference of microaerophilic conditions by Ca. M. parvicella (Rossetti et al., 2005). Thereafter, the highest rates of assimilation were attained under both aerobic and anoxic conditions after 5 h followed by a significant reduction (analysis of variance, P<0.0001) by 8 h of the experiment (Figure 1d). Importantly, 13C-oleic acid remained detectable in the supernatant fraction of the experimental samples (Supplementary Table S1) and, thus, the observed trend was not due to exhaustion of the substrate over time. In contrast to 13C-oleic acid, 13C-triolein assimilation by Ca. M. parvicella cells was minimal (Supplementary Figure S1).
The seasonal dominance of Microthrix populations during wintertime has been partially attributed to the higher bioavailability of lipid substrates when wastewater temperatures are lower (Rossetti et al., 2005; Muller et al., 2014a; Roume et al., 2015). By taking into account that 13C-oleic acid assimilation by Ca. M. parvicella was highest after 5 h with equal assimilation rates under aerobic or anoxic conditions, we performed temperature-dependent incubation experiments under aerobic conditions over a wide range of temperatures (4–35 °C), and we then compared Ca. M. parvicella 13C-oleic acid assimilation rates at the 5 h time point (Figure 1e). 13C-oleic acid assimilation was apparent at 4 °C but markedly decreased with increasing temperatures (4–20 °C). Between 25 and 30 °C, 13C-oleic acid assimilation increased significantly (analysis of variance, P<0.0001) but decreased again at 35 °C. The observed two temperature optima may be attributed to differences in the bioavailability of 13C-oleic acid (higher levels of bioavailability are expected at the lower temperatures, for example, at 4 °C; Rossetti et al., 2005) and altered activity of the acyl-CoA ligases for 13C-oleic acid assimilation (higher assimilation rates might be expected at the higher temperatures, for example, 25 °C). These wide ranges of temperature-dependent 13C-oleic acid assimilation characterized by two temperature optima emphasize Ca. M. parvicella's generalist lifestyle strategy (Muller et al., 2014a), defined as an ability to tolerate a wide range of environmental conditions.
Ca. M. parvicella encodes glycerol and glycerol-3-phosphate transporters (McIlroy et al., 2013) and can simultaneously assimilate oleic acid and glycerol (Kindaichi et al., 2013). Recent genome-scale metabolic reconstructions suggest that glycerol conversion into glycerol-3-phosphate may occur prior to its assimilation (McIlroy et al., 2013; Roume, 2013). To investigate these phenotypic traits, we carried out experiments using 13C-glycerol or 13C-glycerol-3-phosphate in combination with or without unlabeled oleic acid. Interestingly, Ca. M. parvicella cells assimilated 13C-glycerol-3-phosphate only as a single substrate measurable after 8 and 24 h of the experiment under both aerobic and anoxic conditions (Figure 1f). Although the absence of 13C-glycerol assimilation is consistent with a previous study (Tomei et al., 1999), the lack of simultaneous assimilation with oleic acid is at odds with the observations of another in situ study (Kindaichi et al., 2013), which may suggest intraspecific phenotypic differences according to geographic location. Nonetheless, the rapid assimilation of 13C-oleic acid compared with 13C-glycerol-3-phosphate underlines previous suggestions that Ca. M. parvicella engages in optimal foraging behavior (Muller et al., 2014a), which posits that, in an environment with diverse substrates, successful taxa will have a preference for the most energy-dense substrates (Frens, 2010).
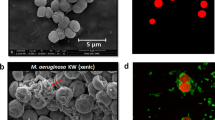
Intriguingly, nanoSIMS imaging revealed extensive phenotypic heterogeneity in substrate assimilation between individual Ca. M. parvicella cells of the same filament (Figure 2). For instance, ≈35–55% and ≈5–35% of Ca. M. parvicella cells assimilated 13C-oleic acid and 13C-glycerol-3-phosphate, respectively, whereas the remainder of cells (45–95%) did not exhibit any 13C-substrate assimilation (Supplementary Table S2). Furthermore, phenotypic heterogeneity in the 13C-oleic acid assimilation appeared to be temperature-dependent whereby relatively low phenotypic heterogeneity was observed at 4 and 30 °C, respectively (Supplementary Table S2). To date, nanoSIMS imaging of filamentous bacteria from other environments has revealed variations in substrate assimilation among cells of the same population (Popa et al., 2007; Vasquez-Cardenas et al., 2015). However, the complete absence of 13C-substrate assimilation in a substantial fraction of cells belonging to the same filament is unique to the results presented in this study. Importantly, intense fluorescence in situ hybridization signals, atomic force microscopic cell integrity results acquired prior to nanoSIMS analyses and Live-Dead staining (Boulos et al., 1999; Roume et al., 2013) did not reveal differences in terms of viability between assimilating and non-assimilating cells, suggesting that the observed intercellular phenotypic heterogeneity is an intrapopulation feature of Ca. M. parvicella (Figures 1a–c, Supplementary Figure S2).
NanoSIMS visualization of phenotypic heterogeneity with regard to substrate assimilation among “Ca. M. parvicella” filaments under aerobic or anoxic conditions. The micrographs show 13C-oleic acid assimilation after 1 h (a–d) and 8 h during the fatty acid assimilation experiment (e–h) and 13C-glycerol-3-phosphate after 24 h when administered as a single substrate during the simultaneous substrate assimilation experiment (k–n).
We further estimated Ca. M. parvicella growth rates based on cells that exhibited substrate assimilation (Foster et al., 2011) as much of newly assimilated 13C-oleic acid appeared to be utilized for cell growth rather than for triglyceride accumulation as 13C-glyceryl trioleate (Supplementary Table S2). In response to different substrates and temperature conditions tested in this study, the estimated Ca. M. parvicella growth rates ranged from 0.12 to 0.78 day−1, which are in agreement with those estimated using the total extended filament length approach (Tandoi et al., 1998; Rossetti et al., 2002).
Given the prevalence of Ca. M. parvicella phenotypic heterogeneity, we investigated Ca. M. parvicella 13C-oleic acid assimilation in response to alternating aerobic–anoxic phases, a regularly encountered fluctuation in BWWTPs in which Ca. M. parvicella can become prominent. In response to alternating anoxic phases, ≈28% of aerobically preconditioned Ca. M. parvicella cells exhibited a wider range of 13C-oleic acid assimilation rates compared with ≈3% of anoxically preconditioned Ca. M. parvicella cells which experienced alternating aerobic conditions (Figure 1g). Compared to their non-alternated controls, less 13C-oleic acid assimilation was observed among Ca. M. parvicella cells subjected to alternating conditions (Supplementary Figure S3). This was reflected in the presence of subpopulations of assimilating Ca. M. parvicella cells, which in turn suggests that an increase in phenotypic heterogeneity (Supplementary Table S2) results from fluctuating environmental conditions and reflects a possible adaptation strategy. Given the low levels of population-level genetic variation in Ca. M. parvicella (McIlroy et al., 2013; Muller et al., 2014a) as well as the expected clonality among cells of the same filament, genetic variation is unlikely to be the source for observed phenotypic heterogeneity among Ca. M. parvicella cells. However, the observed phenotypic heterogeneity among subpopulations of Ca. M. parvicella cells suggests that this population follows a bet-hedging strategy.
The adaptive function of phenotypic heterogeneity has been well described in laboratory studies, yet its significance in natural and engineered environments is poorly understood. Here we provide direct evidence for phenotypic heterogeneity among cells of Ca. M. parvicella that is independent of varied 13C-oleic acid assimilation rates in response to different temperature and alternating aerobic–anoxic regimes (Figures 1d, e and g, and Supplementary Figure S3). Given that Ca. M. parvicella intermittently blooms resulting in operational difficulties (Rossetti et al., 2005) or that it may represent a means of recovering chemical energy in the form of lipids from wastewater (Muller et al., 2014b), strategies for controlling its growth in BWWTPs are highly desirable. Our results highlight the importance of accounting for phenotypic heterogeneity in devising such schemes in the future.
References
Andreasen M, Nielsen P . (1998). In situ characterization of substrate uptake by Microthrix parvicella using microautoradiography. Water Sci Technol 37: 19–26.
Boulos L, Prevost M, Barbeau B, Coallier J, Desjardins R . (1999). LIVE/DEAD® BacLightTM: application of a new rapid staining method for direct enumeration of viable and total bacteria in drinking water. J Microbiol Methods 37: 77–86.
Daims H, Taylor MW, Wagner M . (2006). Wastewater treatment: a model system for microbial ecology. Trends Biotechnol 24: 483–489.
De Jong IG, Haccou P, Kuipers OP . (2011). Bet hedging or not? A guide to proper classification of microbial survival strategies. Bioessays 33: 215–223.
De Jong IG, Veening J, Kuipers OP . (2012). Single cell analysis of gene expression patterns during carbon starvation in Bacillus subtilis reveals large phenotypic variation. Environ Microbiol 14: 3110–3121.
Foster RA, Kuypers MMM, Vagner T, Paerl RW, Musat N, Zehr JP . (2011). Nitrogen fixation and transfer in open ocean diatom-cyanobacterial symbioses. ISME J 5: 1484–1493.
Frens KM . (2010). Effects of food type and patch location on foraging: a field test of optimal foraging predictions. PhD thesis, University of Michigan, Ann Arbor, MI, USA (http://hdl.handle.net/2027.42/69156.
Grimbergen AJ, Siebring J, Solopova A, Kuipers OP . (2015). Microbial bet-hedging: the power of being different. Curr Opin Microbiol 25: 67–72.
Kindaichi T, Nierychlo M, Kragelund C, Nielsen JL, Nielsen PH . (2013). High and stable substrate specificities of microorganisms in enhanced biological phosphorus removal plants. Environ Microbiol 15: 1821–1831.
Levy SF, Ziv N, Siegal ML . (2012). Bet hedging in yeast by heterogeneous, age-correlated expression of a stress protectant. PLoS Biol 10: e1001325.
McIlroy SJ, Kristiansen R, Albertsen M, Michael Karst S, Rossetti S, Lund Nielsen J et al. (2013). Metabolic model for the filamentous Candidatus Microthrix parvicella based on genomic and metagenomic analyses. ISME J 7: 1161–1172.
Muller EEL, Pinel N, Gillece JD, Schupp JM, Price LB, Engelthaler DM et al. (2012). Genome sequence of Candidatus Microthrix parvicella Bio17-1, a long-chain-fatty-acid-accumulating filamentous actinobacterium from a biological wastewater treatment plant. J Bacteriol 194: 6670–6671.
Muller EEL, Pinel N, Laczny CC, Hoopmann MR, Narayanasamy S, Lebrun LA et al. (2014a). Community-integrated omics links dominance of a microbial generalist to fine-tuned resource usage. Nat Commun 5: 5603.
Muller EEL, Sheik AR, Wilmes P . (2014b). Lipid-based biofuel production from wastewater. Curr Opin Biotechnol 30: 9–16.
Nielsen P, Roslev P, Dueholm T, Nielsen J . (2002). Microthrix parvicella, a specialized lipid consumer in anaerobic-aerobic activated sludge plants. Water Sci Technol 46: 73–80.
Popa R, Weber PK, Pett-Ridge J, Finzi JA, Fallon SJ, Hutcheon ID et al. (2007). Carbon and nitrogen fixation and metabolite exchange in and between individual cells of Anabaena oscillarioides. ISME J 1: 354–360.
Rossetti S, Tomei M, Levantesi C, Ramadori R, Tandoi V . (2002). Microthrix parvicella: a new approach for kinetic and physiological characterization. Water Sci Technol 46: 65–72.
Rossetti S, Tomei MC, Nielsen PH, Tandoi V . (2005). Microthrix parvicella, a filamentous bacterium causing bulking and foaming in activated sludge systems: a review of current knowledge. FEMS Microbiol Rev 29: 49–64.
Roume H, Heintz-Buschart A, Muller EEL, May P, Satagopam VP, Laczny CC et al. (2015). Comparative integrated omics: identification of key functionalities in microbial community-wide metabolic networks. Biofilms Microbiomes 1: article number 15007.
Roume H . (2013). Molecular eco-systems biology of lipid accumulating microbial communities in biological wastewater treatment plants. Doctoral thesis, University of Luxembourg, Luxembourg (http://hdl.handle.net/10993/15553.
Roume H, EL Muller E, Cordes T, Renaut J, Hiller K, Wilmes P . (2013). A biomolecular isolation framework for eco-systems biology. ISME J 7: 110–121.
Tandoi V, Rossetti S, Blackall LL, Majone M . (1998). Some physiological properties of an Italian isolate of ‘Microthrix parvicella’. Water Sci Technol 37: 1–8.
Tomei MC, Levantesi C, Rossetti S, Tandoi V . (1999). Microbiological characterisation of pure cultures and its relevance to modelling and control of bulking phenomena. Water Sci Technol 39: 21–29.
Vasquez-Cardenas D, van de Vossenberg J, Polerecky L, Malkin SY, Schauer R, Hidalgo-Martinez S et al. (2015). Microbial carbon metabolism associated with electrogenic sulphur oxidation in coastal sediments. ISME J 9: 1966–1978.
Zimmermann M, Escrig S, Hübschmann T, Kirf M, Brand A, Inglis RF et al. (2015). Phenotypic heterogeneity in metabolic traits among single cells of a rare bacterial species in its natural environment quantified with a combination of flow cell sorting and NanoSIMS. Front Microbiol 6: 243.
Acknowledgements
We thank Mr Bissen and Mr Di Pentima from the Syndicat Intercommunal à la Vocation Ecologique (SIVEC) for their permission to collect samples from the Schifflange wastewater treatment plant, Luxembourg. This work was funded by an ATTRACT program grant (ATTRACT/A09/03) and a European Union Joint Programming in Neurodegenerative Diseases grant (INTER/JPND/12/01) awarded to PW as well as postdoctoral grants Aide à la Formation Recherche (AFR) to ARS (PDR-2013-1/5748561) and EELM (PDR-2011-1/SR), all funded by the Luxembourg National Research Fund (FNR). Anne Kaysen and Kacy Greenhalgh are thanked for their help with sample transportation. We thank Dr Aidos Baumuratov and Martine Schmitz from the imaging facilities at the Luxembourg Centre for Systems Biomedicine and at the University of Luxembourg, respectively. We also thank Emmanuelle Cocco and Laurence Joly from the Luxembourg Institute of Science and Technology for their help with LC-MS/MS analyses. We are grateful to three anonymous reviewers for their very valuable comments.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Sheik, A., Muller, E., Audinot, JN. et al. In situ phenotypic heterogeneity among single cells of the filamentous bacterium Candidatus Microthrix parvicella. ISME J 10, 1274–1279 (2016). https://doi.org/10.1038/ismej.2015.181
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2015.181
This article is cited by
-
Single-cell stable isotope probing in microbial ecology
ISME Communications (2022)
-
Composting pig manure and sawdust with urease inhibitor: succession of nitrogen functional genes and bacterial community
Environmental Science and Pollution Research (2020)
-
Roles of bacteriophages, plasmids and CRISPR immunity in microbial community dynamics revealed using time-series integrated meta-omics
Nature Microbiology (2020)
-
Integration of time-series meta-omics data reveals how microbial ecosystems respond to disturbance
Nature Communications (2020)
-
Quantitative macromolecular patterns in phytoplankton communities resolved at the taxonomical level by single-cell Synchrotron FTIR-spectroscopy
BMC Plant Biology (2019)