Abstract
Understanding the evolutionary history and potential of bacterial pathogens is critical to prevent the emergence of new infectious bacterial diseases. Xanthomonas axonopodis subsp. citri (Xac) (synonym X. citri subsp. citri), which causes citrus canker, is one of the hardest-fought plant bacterial pathogens in US history. Here, we sequenced 21 Xac strains (14 XacA, 3 XacA* and 4 XacAw) with different host ranges from North America and Asia and conducted comparative genomic and evolutionary analyses. Our analyses suggest that acquisition of beneficial genes and loss of detrimental genes most likely allowed XacA to infect a broader range of hosts as compared with XacAw and XacA*. Recombination was found to have occurred frequently on the relative ancient branches, but rarely on the young branches of the clonal genealogy. The ratio of recombination/mutation ρ/θ was 0.0790±0.0005, implying that the Xac population was clonal in structure. Positive selection has affected 14% (395 out of 2822) of core genes of the citrus canker-causing Xanthomonas. The genes affected are enriched in ‘carbohydrate transport and metabolism’ and ‘DNA replication, recombination and repair’ genes (P<0.05). Many genes related to virulence, especially genes involved in the type III secretion system and effectors, are affected by positive selection, further highlighting the contribution of positive selection to the evolution of citrus canker-causing Xanthomonas. Our results suggest that both metabolism and virulence genes provide advantages to endow XacA with higher virulence and a wider host range. Our analysis advances our understanding of the genomic basis of specialization by positive selection in bacterial evolution.
Similar content being viewed by others
Introduction
Approximately 160 emerging infectious bacterial diseases have been discovered in the past 70 years (Jones et al., 2008). The recent increase in emerging bacterial diseases is partly because of agricultural practices and climate change, among other factors (Wolfe et al., 2007; Jones et al., 2008). The agroecosystem has played a critical role in the recent emergence and spread of pathogenic bacteria in plants. Prevention of emergence of new bacterial diseases is one essential task of the scientific community. Emergence of plant bacterial diseases depends largely on the life history traits and evolutionary potential of the corresponding bacteria. Among plant pathogenic bacteria, those of the genus Xanthomonas in the gamma division of Proteobacteria can cause disease in virtually all plant species (Leyns et al., 1984; Chan and Goodwin, 1999). Individual members of Xanthomonas are highly host specific (Hayward, 1993). Evolutionary analysis of X. axonopodis, which is a complex species composed of multiple pathovars (host-specific subspecies), identified two main diversification steps based on sequence analysis of selected loci including seven housekeeping genes and several virulence-associated genes (Mhedbi-Hajri et al., 2013). The first diversification led to a clustering of generalist pathogens over the past 25 000 years with no apparent connection to host or geography. The second step led to specialized pathotypes grouped according to their host range and their symptomatology over the past two centuries. Eventually, secondary contacts likely occurred between host-specialized bacteria that enabled genetic exchange of virulence-associated genes. For pathogens, acquisition of virulence genes is not the end, but instead, the beginning of the process of host specialization (Rohmer et al., 2011). During the specialization process, these pathogens experience new niches and strong directional selection from hosts, and eventually the more adapted lineages will emerge and sweep through evolution, including accumulating beneficial mutations, acquiring beneficial genes, losing antivirulence genes and so on (Lieberman et al., 2011; Rohmer et al., 2011). The process of host specialization of Xanthomonas has not been well understood. In this study, we investigated the evolutionary history and potential of X. axonopodis pv. citri (Xac) (synonym X. citri subsp. citri) by genome-wide analysis.
Xac is the causal agent of citrus canker that infects most commercial citrus cultivars and causes distinctive raised necrotic lesions on leaves, stems and fruit. Severe infections can cause defoliation, blemished fruit, premature fruit drop, twig dieback and general tree decline (Gottwald et al., 2002). Citrus canker is distributed worldwide and found in most citrus-producing countries. Importantly, it is endemic in the top three citrus-producing countries: Brazil, United States and China. Owing to its importance, citrus canker is one of the hardest-fought plant bacterial diseases in the US history. Citrus originated in Southeast Asia, that is, Northeast India, Burma and the Yunnan province of China (Scora, 1975; Gmitter and Hu, 1990). Xac is also suggested to have originated in the same area (Gottwald, 2000). Xac was introduced into the United States in at least three different events in 1910, 1986 and 1995. These introductions were followed by eradication programs, in which billions of dollars were spent (Gottwald and Irey, 2007).
Xac consists of multiple pathotypes, that is, A, A* and Aw. XacA is widespread worldwide and can infect many citrus species, hybrids between citrus species and the citrus relative trifoliate orange Poncirus trifoliate as well as some other plants in the Rutaceae family (Graham et al., 2004), whereas XacAw and XacA* have a restricted host range of Mexican lime (Citrus aurantifolia) and alemow (Citrus macrophylla) (Sun et al., 2004). However, the genetic basis of the host range and virulence difference between them has not been well determined (Jalan et al., 2013a).
In addition to Xac, X. fuscans subsp. aurantifolii (Xau) (previously named as X. axonopodis pv. aurantifolii) also causes citrus canker disease. Xau causes canker B and canker C on restricted hosts and these strains have only been found in South America (Graham et al., 2004). Given that Xau B and C have evolved to be citrus canker-causing pathogens from a separate lineage from Xac strains (Mhedbi-Hajri et al., 2013), analysis of how these Xanthomonas pathogens have evolved to specialize on citrus and associated relatives to cause citrus canker disease via convergent evolution may reveal common mechanisms underlying this disease.
The evolution of Xac has only been examined using multilocus sequence analysis or genotyping (Ngoc et al., 2009; Mhedbi-Hajri et al., 2013). To investigate the evolutionary history and potential of Xac, we sequenced 21 representative Xac strains from North America and Asia. We included strains from the XacA* and XacAw pathotypes to identify potential mechanisms underlying the host range difference within Xac. The importance and relative contribution of recombination and mutation in evolutionary history of Xac was analyzed. Furthermore, the importance of positive selection in the process of host specialization for the citrus canker-causing Xanthomonas was discussed.
Materials and methods
Strains
The strains for sequencing were chosen to represent Xac that are genetically and geographically diverse. We selected 21 strains of Xac from North America (United States) and Asia, with a time span of over two decades for genome sequencing (Supplementary Table S1). The complete genomes of strain A306 (da Silva et al., 2002) and Aw12879 (Jalan et al., 2013b) were also used in this analysis.
Sequencing, assembly and annotation
Quantity and quality of the DNA were measured using Agilent 2100 BioAnalyzer (Agilent Technologies, Inc., Santa Clara, CA, USA). Whole-genome sequencing of all strains was performed using 1/4 of a lane of paired-end 100-cycle sequencing using an Illumina genome analyzer IIx (Illumina, Hayward, CA, USA) at the Interdisciplinary Center for Biotechnology Research, University of Florida. In addition, for XacA*270 paired-end 454 pyrosequencing was performed using Roche 454 GS-FLX plus system (454 Life Sciences, Branford, CT, USA) at the Interdisciplinary Center for Biotechnology Research. A de novo BamHI optical map of the genome of XacA*270 was generated by OpGen Technologies (Madison, WI, USA). The draft genome of XacA*270 was closed using the method described in our previous study (Jalan et al., 2011).
All reference-based assemblies were performed on CLC Genomics Workbench 6.0 (CLC bio, Cambridge, MA, USA), using default parameters, a length fraction of 0.9 and similarity of 0.9 for reference assemblies to XacA306, XacAw12879 and XacA*270. Assembly of all strains was annotated using the ‘isolated genome gene calling’ pipeline from the Integrated Microbial Genomes–Expert Review (IMG/ER; Markowitz et al., 2009).
Comparative genomic analysis
The ANIm values between genomes were calculated using the NUCmer algorithm v3.1 integrated in Jspecies v1.2.1 (Richter and Rossello-Mora, 2009). The pan-genome and core genome were calculated by using OMCL and bidirectional best hit (BDBH) methods implemented in get_homologues package (Contreras-Moreira and Vinuesa, 2013) with parameters: e-value: 1e−5, identity: 60% and coverage: 75%. For pan-genome size calculation, the power law model was employed. In short, according to the power law model, the new genes N(n) found when the nth genome is added to the pan-genome can be described as N(n)=A × nα, and the pan-genome is ‘open’ if α>0, is ‘logarithmical’ if α=0 or is ‘closed’ if α<0, while A is a constant (Donati et al., 2010).The core genes were aligned using MUSCLE (Edgar, 2004) and concatenated using Gblocksb0.91 (Castresana, 2000). The best nucleotide substitution model was obtained by MODELTEST analysis (Posada and Crandall, 1998), and the subsequent maximum likelihood phylogenetic tree was constructed by using PAUP4b10 (Swofford, 1998).
To calculate the ratio of recombination rate to mutation rate (ρ/θ) and the relative contribution of recombination and mutation (r/m), a whole-genome sequence alignment of the selected strains was created using progressiveMauve (Darling et al., 2010) and the sequences of the first nine longest alignments (3.06 Mb, ∼58% of the whole genomes) for the 23 Xac strains or three longest alignments (1.77 Mb, ∼33% of each genome) for the 25 strains (23 Xac plus XauB and XauC) were concatenated using Gblocksb0.91 (Castresana, 2000), and then calculated by using ClonalFrame v1.2 (Didelot and Falush, 2007). Four independent runs of ClonalFrame were performed, each consisting of 40 000 iterations, and the first half was discarded as MCMC burn-in. The convergence and mixing properties revealed by manual comparison between the four runs suggested that the obtained parameters were confident. The presence or absence of intragenic recombination was assessed by the single breakpoint method (Pond et al., 2006) (P<0.05). Fast Unconstrained Bayesian AppRoximation (FUBAR) (Murrell et al., 2013) implemented in HYPHY2.2 package was used to evaluate the contribution of positive selection on the citrus canker-causing Xanthomonas. For each gene cluster, 400 Grid points were used, and five independent runs of FUBAR were performed, each consisting of 2 000 000 iterations. The first half was discarded as MCMC burn-in.
Effector prediction was conducted as previously described (Bart et al., 2012). In short, a data set of all known effectors from animal and plant pathogens was obtained and tblastn (Blast+ 2.2.28) was used to identify potential effectors in Xac strains with at least 45% amino acid homology and 80% coverage length.
LPS extraction and host response analysis
Lipopolysaccharide (LPS) extractions were performed as previously described with modifications (Marolda et al., 2006). Briefly, bacterial cells from 20 ml culture (OD600=1) were centrifuged and resuspended in 500 μl distilled water. LPSs were extracted with hot phenol (70 °C) three times and aqueous phases were subjected to dialysis against distilled water at 4 °C (Pur-A-Lyzer Midi 1000 Dialysis kit, cutoff=1 kDa, Sigma-Aldrich, St Louis, MO, USA). Prepared LPSs (∼50 μl per leaf) were infiltrated into plant leaves using a needleless syringe. Leave samples were collected at 0 (right after treatment), 6 and 24 h post inoculation. Time point 0 was used as the control. The expression levels of host defense-related genes were performed using a QuantiTect SYBR green RT-PCR kit (Qiagen, Valencia, CA, USA) on a 7500 fast real-time PCR system (Applied Biosystems, Foster City, CA, USA). Citrus GAPDH gene was used as an internal control. The relative fold change was calculated as previously described (Livak and Schmittgen, 2001).
Data access
The finished genomes of 14 XacA and 4 XacAw strains and draft genomes of 3 XacA* strains studied in this project have been deposited in GenBank under the project accession number PRJNA255042 (Supplementary Table S1).
Results and discussion
Genomic features of citrus canker-causing Xac strains
We sequenced 21 representative Xac strains (14 XacA strains, 3 XacA* strains and 4 XacAw strains) encompassing three countries (United States, China and Saudi Arabia), and isolated over a period of more than two decades (Supplementary Table S1). The reads from XacA and XacAw strains were reference assembled against XacA306 and XacAw12879 genome (da Silva et al., 2002; Jalan et al., 2013b), respectively. As no XacA* genome was publically available, De Novo assembly was conducted for the genome of XacA*270 (Supplementary Table S2) using previously described methodology (Jalan et al., 2011), and the two remaining XacA* strains were reference assembled based on the XacA*270 genome. The comparison of our reads and assemblies (Supplementary Table S3) with other published assemblies for Xanthomonas (Bart et al., 2012) indicated that our assemblies are of high quality. Assembly of all strains was annotated using the IMG/ER annotation pipeline and submitted to GenBank, the results for which are presented in Supplementary Table S1.
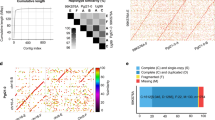
The ANIm values of the 23 Xac strains (21 strains sequenced in this study and XacA306 and XacAw12879 sequenced previously; da Silva et al., 2002; Jalan et al., 2013b) were 99.65–99.94% (Supplementary Table S4), demonstrating highly conserved genomic backgrounds among these strains, despite differences in host ranges. To reveal the mechanisms underlying differential host ranges of the three pathotypes A, A* and Aw, we calculated the pan-genome of the 23 Xac strains using the OMCL method. The core genome comprised 3912 orthologous clusters, whereas the pan-genome contained 5147 orthologous clusters. Interestingly, hierarchical clustering of the accessory orthologous clusters (that is, the clusters not harbored by all of the 23 strains) of these Xac strains produced a distribution in which the XacAw and XacA* strains were grouped together and formed a separate clade from the XacA strains (Figure 1), consistent with the phylogenetic tree constructed based on the Xac/Xau core genomes (Figure 2 and Supplementary Figure S1). Given that XacA strains have wide host ranges whereas the host range of XacAw and XacA* strains is restricted, we infer that the clade-specific genes probably contributed to the host range difference between the XacA group and XacAw and XacA* groups. Through comparative genome analysis, 89 XacA- and 121 XacAw- and XacA*-specific genes were identified (Supplementary Data Set 1).
Hierarchical clustering of 23 Xac strains based on heat map of 1235 accessory orthologous clusters. Presence and absence of the homolog for each cluster are indicated in yellow and green, respectively. A total of 100 bootstrap replicates were made, and bootstrap values of >50% were indicated at each node. The UPGMA tree was generated by using DendroUPGMA (http://genomes.urv.es/UPGMA/) with Jaccard coefficient.
Maximum likelihood (ML) tree reconstructed based upon the concatenated sequences of 2822 core genes of the 25 citrus canker-causing Xanthomonas strains. A total of 100 bootstrap replicates were made, and bootstrap values are indicated at the branch points. The parameters on the left were calculated using Modeltest 3.7 and were used by PAUP4.0b10 to construct the ML tree.
The gene avrGf1 (xopAG), which is known as the type III effector restricting the host range of XacAw strains (Rybak et al., 2009), was absent in the three XacA* strains. A previous study also reported that some XacA* strains harbor the avrGf1 gene whereas others do not (Escalon et al., 2013). Furthermore, the XacAw ΔavrGf1 mutant obtained the ability to infect more citrus species and cause canker on hosts other than limes, but the virulence was not as strong as XacA strains (Rybak et al., 2009), suggesting that other genomic feature(s) might contribute to the host range and virulence difference between XacA and XacAw. Among the type III effector genes identified in the 23 Xac genomes, only one effector, xopAF, was found to be common and specific to XacAw and XacA* (Supplementary Table S5); however, mutation of xopAF in XacAw12879 resulted in lower virulence and no change of host range (Jalan et al., 2013a), suggesting that xopAF is not related to the host range determination. Two other effectors, xopC1 and xopL, were found to differentiate XacA from Aw or A*; xopC1 was specifically harbored by the three XacA* strains whereas all the 23 strains harbored xopL, but it was frameshift mutated in XacA* strains (Supplementary Table S5). However, XopC1 and XopL do not contribute to virulence or host range determination (Dunger et al., 2012; Escalon et al., 2013). Thus, there should be other mechanisms other than the known type III effectors underlying the difference in host range and virulence between XacA and XacAw and XacA* strains. Notably, compared with the specific genes belonging to XacAw and XacA*, XacA-specific genes were significantly enriched in the COG category ‘metabolism’ (Fisher’s exact test, P=0.007; Table 1). Therefore, we reasoned that XacA-specific genes, especially the ‘metabolism’ genes, provide advantages to endow XacA with higher virulence and a wider host range (Rohmer et al., 2011).
We identified a gene region involved in LPS biosynthesis (J151_03781-J151_03787, equal to XAC3596-XAC3601, totally 7 genes, nlxA is recently determined in the intergenic region of XAC3597 and XAC3598; Yan et al., 2012) in XacA that was highly variable from that of XacAw and XacA*, but the upstream and downstream genes were conserved (Figure 3a). Three of the genes in this region, nlxA, wzt and wzm, are ‘metabolism’ genes. This variable region in XacA, XacAw and XacA* was all identified as acquired genomic content via horizontal transfer as predicted by both Alien_Hunter and IslandViewer programs (Vernikos and Parkhill, 2006; Langille and Brinkman, 2009), suggesting XacA acquired this region from different source(s) as compared with XacAw and XacA*. Some of the genes in this region, such as wxacO, rfbC, nlxA and wzt, have been demonstrated to be involved in O-antigen synthesis and transport and be associated with biofilm formation on the host and contribute to virulence in XacA (Casabuono et al., 2011; Li and Wang, 2011; Petrocelli et al., 2012; Yan et al., 2012). Among the three components, lipid A, core and O-antigen that are composed of the entire Xac LPS, the O-antigen region is probably the most important for host basal response during citrus–pathogen interaction (Casabuono et al., 2011). O-antigen is highly variable with regard to its composition, length and the branching of carbohydrate subunits, whereas the core and lipid A are conserved among different bacterial species (Nicaise et al., 2009). LPS analysis by SDS–polyacrylamide gel electrophoresis clearly demonstrated that the O-antigen region of XacA group was different from that of XacAw (Figure 3b). We hypothesized that the O-antigen differences between XacA and XacAw or A* group may induce different defence responses in different host plants. As shown in Figure 3c, LPS isolated from XacA and XacAw acted as pathogen-associated molecular pattern and induced the gene expression of pathogen-associated molecular pattern-triggered immunity markers GST1 and WRKY22 (Asai et al., 2002), salicylic acid metabolism PAL1 (Greenberg et al., 2009) and pathogen-associated molecular pattern-triggered immunity signaling kinase MKK4 (Zhao et al., 2014) in both sweet orange and Mexican lime leaves. Defence-related gene expression triggered by XacA LPS is significantly lower than that triggered by XacAw LPS in sweet orange. This is consistent with the fact that XacAw is not pathogenic on sweet orange whereas XacA causes disease on sweet orange. LPS of XacA or XacAw induced the gene expression of GST1, WRKY22, PAL1 and MKK4 in Mexican lime leaves despite both strains are pathogenic on Mexican lime. The defence-related gene expression triggered by XacA LPS is slightly higher than or at similar level as that triggered by XacAw LPS in Mexican lime. This indicates that both XacA and XacAw could suppress the pathogen-associated molecular pattern-triggered immunity induced by LPS in Mexican lime, probably using type III effectors (Jones and Dangl, 2006).
(a) Comparison of the LPS gene clusters of X. axonopodis subsp. citri str. A306 (XacA306), XacAw12879 and XacA*270. Conserved and homologous genes (>50% identity) are colored. The nlxA, wzt and wzm are ‘metabolism’ genes. (b) SDS–polyacrylamide gel electrophoresis (SDS-PAGE) analysis of LPSs extracted from XacA306 and XacAw12879. LPSs were isolated using hot phenol method, subjected to analysis on polyacrylamide gel (12% in the resolving gel and 5% in the stacking gel) and visualized by silver staining. The different regions of LPSs are shown as indicated. (c) Expression analysis of four pathogen-associated molecular pattern-triggered immunity (PTI)-related genes using quantitative reverse-transcription PCR. Relative expressions (calculated using the formula 2–ΔΔCT) were monitored from young orange leaves inoculated with XacA306 and XacAw12879 LPSs. Leaves were sampled at 0, 6 and 24 h post inoculation. Bars are means±s.e. (n=3). The experiment was repeated once with similar results. Student’s t-test was conducted. The asterisks indicate a statistically significant difference (*P<0.01 vs A strain).
In addition, several genes that have been shown to contribute to biofilm formation, including XAC1469, XAC3285, XACb0003-0004 and XACb0050, were also found to be specific to XacA strains (Laia et al., 2009; Li and Wang, 2011; Malamud et al., 2013). It remains to be determined how the specific genes other than known type III effectors for XacA, A* and Aw contribute to the host range of Xac.
Evolution of the core and accessory genomes of Xac
We hypothesize that XacA evolved to be a separate group from XacAw and XacA*. Our evidence is that the XacA strains formed a separate clade from the XacAw and XacA* strains as revealed by the phylogenetic tree reconstructed based on the core genome (Figure 2 and Supplementary Figure S1), the phyletic distribution of accessory orthologous clusters (Figure 1) of the strains analyzed and the different origin of the LPS genomic island of the two groups. This finding is consistent with the previous report based on MLVA (multilocus variable number of tandem repeat Analysis) genotyping method (Pruvost et al., 2014). The slight but significant difference of genome size (5.24±0.02 Mb (mean±s.d., same for following ones) for XacA and 5.36±0.05 Mb for Aw and A* (P=0.0003)), CDS number (4443±22 for XacA and 4591±39 for Aw and A* (P=3.7E−6)) and percentage of GC content (64.754±0.001 for XacA and 64.656±0.001 (P=7.5E−6 for Aw and A*)) of XacA strains and XacAw and XacA* also suggested the divergence of the two groups (Supplementary Figure S2). During their divergent evolutionary history, the acquisition of beneficial genes and loss of genes that restrict the host range of the pathogens (for example, avrGf1) may have provided XacA advantages to become more adapted to diverse citrus hosts than XacAw and XacA* (Rohmer et al., 2011; Merhej et al., 2013). Noteworthy, the phyletic distribution of accessory genes was globally similar to the phylogeny of core genomes, suggesting that the gain and loss of accessory genes for each strain was also under selection and might be driven by similar evolution mechanism as core genomes. In line with the positive selection of accessory genes, compared with XacAw and XacA*, the more adapted XacA strains were found to harbor much fewer ‘No COG’ genes as well as genes involved in signal transduction (COG category ‘T’) (Table 1). This phenomenon probably results from loss of useless and/or superfluous genes (for example, those involved in process such as signal transduction) for the pathogens after the pathogens have specialized on certain hosts (Merhej et al., 2013). Furthermore, our data suggest that the pan-genome of the 23 Xac strains is closed (A=155.01±35.65 and α=−1.69±0.09). The depletion curves of core genome and accumulation curves of pan-genome shows that these 23 strains comprehensively sample the pan-genome of Xac (Supplementary Figure S3). These results suggest infrequent import of new genes from outside species to Xac and there was loss of genes (for example, those involved in process such as signal transduction) in the recent evolution of Xac during host specialization (Merhej et al., 2013). One explanation for gene loss may be intensified citrus culture, allowing Xac to have a complete infection cycle on citrus without the need for an alternative host or survival in nonhost environments (Gottwald et al., 2002). The constant citrus hosts provide a relatively stable environment and nutrition source, and environmental threats and antagonists would be relatively rare.
To evaluate the frequency and importance of recombination and mutation in the evolution of Xac, the ratio of recombination rate to mutation rate (ρ/θ) and the relative contribution of recombination and mutation (r/m) were calculated on the basis of ungapped sequences of the longest 9 aligned blocks of the 23 genomes, generated by progressive MAUVE (Darling et al., 2010) (3.06 Mb, ∼58% of the whole genomes), using ClonalFrame (Didelot and Falush, 2007). The clonal genealogy calculated by ClonalFrame (Supplementary Figure S4) was consistent with the phylogenetic tree based on the core genome (Figure 2). The r/m value was 1.061±0.006, suggesting that recombination contributed as much as mutation to the observed diversity of Xac as recombination can introduce a segment of foreign DNA into the genome. However, recombination was found to have only occurred frequently on the relative ancient branches representing the ancestors of the XacA and XacAw/A* lineages, but rarely on the young branches (Supplementary Figure S4). The ratio ρ/θ is 0.0790±0.0005, implying that the Xac population was effectively clonal in structure (Fraser et al., 2007). Although other genotyping methods have also suggested a clonal population structure for Xac (Ngoc et al., 2009, Vernière et al., 2014), these results imply that the Xac pathotypes have been clonal and recombination deficient across a majority of the core genome.
Phylogenetic analysis of citrus canker-causing Xanthomonas
To uncover the mechanism underlying infection and disease development of citrus canker-causing pathogens, we conducted a comparative genomic analysis of 23 Xac strains as well as two other sequenced citrus canker-causing Xanthomonas, XauB and XauC (Moreira et al., 2010). Core orthologous gene clusters were computed for 25 total genomes using the BDBH method, and 2822 gene clusters were identified, accounting for 61.0–74.2% of the total protein coding sequences of each genome. A maximum likelihood phylogenetic tree was reconstructed based on concatenated sequences of these 2822 clusters generated by Gblocks0.91b (Castresana, 2000) using PAUP4b10 (Swofford, 1998) with the best model suggested by ModelTest 3.7 (Posada and Crandall, 1998). The high bootstrap values suggested that the topology structure of this tree was highly confident (Figure 2). The neighbor-joining tree reconstructed using MEGA6 (Tamura et al., 2013) (Supplementary Figure S1) also gave a consistent topology. The clade formed by XauB and XauC was separate from the clade of XacA, XacA* and XacAw strains, demonstrating the variability of genome content between these two groups. Based on the phylogenetic tree, XacA and variants in Florida were grouped into three separate clades that most likely correspond to three independent introductions that took place in Florida (Gottwald et al., 2002). The XacAw strains in Florida may have been introduced from India (Schubert et al., 2001). Previous studies demonstrated that XacAw strains collected from Florida were closely related to some XacA* strains isolated from India (Ngoc et al., 2009). The XacA strains from central Florida and Miami grouped together with the Brazilian strain XacA306 and formed a separate branch from other isolates from Florida and China, suggesting that the central Florida, Miami and Brazilian strains share closer ancestry than other strains. The Manatee (MN) strains of Florida are closely related to strains from China, indicating that they share closer ancestry than other strains.
Positive selection analysis of the core genome of citrus canker-causing Xanthomonas
When XauB and XauC were taken into account, the citrus canker-causing population was still effectively clonal (the ρ/θ was 0.0545±0.0006). Furthermore, when tested on gene level, only one gene encoding an aminoacyl-tRNA synthetase, a gene family known to have already undergone complex horizontal gene transfer events in its evolutionary history (O'Donoghue and Luthey-Schulten, 2003), was found to have undergone intragenic recombination (P<0.05) among the 2822 core genes. These results emphasized the rare occurrence of recombination with relative high occurrence of mutation across the core genome of the citrus canker-causing Xanthomonas.
Positive selection is an important driving force for pathogens in adaptation process (Lieberman et al., 2011; Trivedi and Wang, 2013). Knowledge of the genes affected by positive selection across citrus canker-causing species can show which genes may be involved in citrus canker development and host specificity. To evaluate the contribution of positive selection to the citrus canker-causing Xanthomonas population, FUBAR (Murrell et al., 2013) method implemented in HYPHY2.2 package (Pond et al., 2005) was employed. This uses a site-specific model of dN/dS. Out of 2822 core genes, 395 genes (14%) were identified as containing codons under positive selection (posterior probability >0.90) (Table 1). These genes were found to be enriched in the ‘carbohydrate transport and metabolism’ (Fisher’s exact test, P=0.017; Table 1), further demonstrating the specialization of the pathogens on citrus, and how citrus exerts selection on the pathogens. The ‘DNA replication, recombination and repair’ (3R) genes were also enriched in the gene set under positive selection (Fisher’s exact test, P=0.047; Table 1). The 3R genes are crucial for stable maintenance and propagation of DNA for organisms. Interestingly, an important gene involved in DNA repair, mutY (XAC2553), the defects of which can cause excess mutations but not intergenomic recombination (Huang et al., 2006), was found under positive selection. In addition to the two enriched categories, multiple genes known to contribute to virulence were detected. For example, gumC and gumB were found to be under positive selection. The gum genes, responsible for biosynthesis of xanthan gum, are crucial for Xac to form biofilms and survive on the surface of citrus tissues to initiate infection (Facincani et al., 2014). Xanthomonas have a diverse complement of T3SEs that contribute to overall pathogenicity and host range variation (Ryan et al., 2011). Among all the effectors identified in the strains analyzed as previously described (Bart et al., 2012) (Supplementary Table S5), 10 common effectors that were shared by all 25 strains were identified in the 2822 core genes. Although none of the 10 effectors was found to have been affected by intragenic recombination as suggested by single breakpoint method result, the maximum likelihood gene trees of all the 10 effectors were found to be incongruent with the maximum likelihood reference tree based on 2822 core genes revealed by Shimodaira–Hasegawa test (P<0.001; Table 2) (Shimodaira and Hasegawa, 1999). Four of them, including xopA (hpa1), xopV, xopX and hpaA, were found to be affected by positive selection. Interestingly, the four effector genes contained more codons under positive selection (xopA (2 codons), xopV (4), xopV (10) and hpaA (7)) than average (1.59 codons per gene). T3SEs play important roles in the stepwise arms race that has been taking place between pathogens and their plant hosts during their co-evolutionary history as described by the so-called ‘zig-zag’ model (Jones and Dangl, 2006), and positive selection of effector genes is an important driving force for host adaptation (Dong et al., 2014). These results suggest that these effectors underwent horizontal gene transfer or intergenic recombination during their evolutionary history, but they had been strongly affected by positive selection during the arms race after their acquirement. In addition, T3SS is the essential machinery responsible for delivering effectors from the bacterial cytosol directly into the interior of host cells (Ghosh, 2004), and its necessary role in virulence and host adaptation makes the involved genes a hot spot of selection (McCann and Guttman, 2008). This study revealed that six T3SS genes, hpaB, hrpD5, hrcQ, hpaP, hrpB7 and hpa2, have been under positive selection, hrpB7 was found to contain four codons under positive selection and the remaining ones contained one codon affected by positive selection. Positive selection was found for the T3SS and effector genes that are important for virulence and the capacity to overcome the host immune system. They could thus be good candidates for further experimental studies.
Taken together, these results indicate that the evolution of citrus canker-causing Xanthomonas is characterized by selection on new mutations that enhance virulence.
Conclusions
We sequenced and conducted genomic comparison and evolutionary analyses of 21 representative Xac strains. We found that the recent evolution of XacA is characterized by loss of genes and selection on new mutations rather than recombination. Early in its divergence from the narrow host range Xac pathotypes, XacA likely acquired beneficial genes through recombination, but the XacA genome has diversified through clonal expansion as it has colonized citrus worldwide. Mutation occurred much more frequently than recombination for Xac, and also for all the citrus canker-causing Xanthomonas, suggesting selection, rather than recombination, has been likely important in host range and virulence. Positive selection was observed for ‘carbohydrate transport and metabolism’ genes, ‘DNA replication, recombination and repair’ genes and genes involved in T3SS and T3SEs, likely providing a fitness advantage to drive adaptation of these pathogens to their citrus hosts.
Given the closed nature of the pan-genome and the clonal structure of the Xac population, the future evolutionary potential of this citrus canker pathogen may be limited. However, rare horizontal transfer events as well as selection on beneficial mutations in the core or accessory genome will likely continue to challenge management of this pathogen.
Accession codes
References
Asai T, Tena G, Plotnikova J, Willmann MR, Chiu WL, Gomez-Gomez L et al. (2002). MAP kinase signaling cascade in Arabidopsis innate immunity. Nature 415: 977–983.
Bart R, Cohn M, Kassen A, McCallum EJ, Shybut M, Petriello A et al. (2012). High-throughput genomic sequencing of cassava bacterial blight strains identifies conserved effectors to target for durable resistance. Proc Natl Acad Sci USA 109: E1972–E1979.
Casabuono A, Petrocelli S, Ottado J, Orellano EG, Couto AS . (2011). Structural analysis and involvement in plant innate immunity of Xanthomonas axonopodis pv. citri lipopolysaccharide. J Biol Chem 286: 25628–25643.
Castresana J . (2000). Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol 17: 540–552.
Chan JW, Goodwin PH . (1999). The molecular genetics of virulence of Xanthomonas campestris. Biotechnol Adv 17: 489–508.
Contreras-Moreira B, Vinuesa P . (2013). GET_HOMOLOGUES, a versatile software package for scalable and robust microbial pangenome analysis. Appl Environ Mmicrobiol 79: 7696–7701.
da Silva AR, Ferro JA, Reinach F, Farah C, Furlan L, Quaggio R et al. (2002). Comparison of the genomes of two Xanthomonas pathogens with differing host specificities. Nature 417: 459–463.
Darling AE, Mau B, Perna NT . (2010). ProgressiveMauve: multiple genome alignment with gene gain, loss and rearrangement. PLoS One 5: e11147.
Didelot X, Falush D . (2007). Inference of bacterial microevolution using multilocus sequence data. Genetics 175: 1251–1266.
Donati C, Hiller NL, Tettelin H, Muzzi A, Croucher NJ, Angiuoli SV et al. (2010). Structure and dynamics of the pan-genome of Streptococcus pneumoniae and closely related species. Genome Biol 11: R107.
Dong S, Stam R, Cano LM, Song J, Sklenar J, Yoshida K et al. (2014). Effector specialization in a lineage of the Irish potato famine pathogen. Science 343: 552–555.
Dunger G, Garofalo CG, Gottig N, Garavaglia BS, Rosa MCP, Farah CS et al. (2012). Analysis of three Xanthomonas axonopodis pv. citri effector proteins in pathogenicity and their interactions with host plant proteins. Mol Plant Pathol 13: 865–876.
Edgar RC . (2004). MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32: 1792–1797.
Escalon A, Javegny S, Verniere C, Noel LD, Vital K, Poussier S et al. (2013). Variations in type III effector repertoires, pathological phenotypes and host range of Xanthomonas citri pv. citri pathotypes. Mol Plant Pathol 14: 483–496.
Facincani AP, Moreira LM, Soares MR, Ferreira CB, Ferreira RM, Ferro MIT et al. (2014). Comparative proteomic analysis reveals that T3SS, Tfp, and xanthan gum are key factors in initial stages of Citrus sinensis infection by Xanthomonas citri subsp citri. Funct Integr Genomics 14: 205–217.
Fraser C, Hanage WP, Spratt BG . (2007). Recombination and the nature of bacterial speciation. Science 315: 476–480.
Ghosh P . (2004). Process of protein transport by the type III secretion system. Microbiol Mol Biol Rev 68: 771–795.
Gmitter FG, Hu X . (1990). The possible role of Yunnan, China, in the origin of contemporary Citrus species (Rutaceae). Econ Bot 44: 267–277.
Gottwald TR . (2000). Citrus canker. The Plant Health Instructor.10.1094/PHI-I-2000-1002-01 Updated 2005.
Gottwald TR, Graham JH, Schubert TS . (2002). Citrus canker: the pathogen and its impact. Plant Health Progressdoi:10.1094/PHP-2002-0812-01-RV.
Gottwald TR, Irey M . (2007). Post-hurricane analysis of citrus canker II: Predictive model estimation of disease spread and area potentially impacted by various eradication protocols following catastrophic weather events. Plant Health Progress 10.
Graham JH, Gottwald TR, Cubero J, Achor DS . (2004). Xanthomonas axonopodis pv. citri: factors affecting successful eradication of citrus canker. Mol Plant Pathol 5: 1–15.
Greenberg JT, Jung HW, Tschaplinski TJ, Wang L, Glazebrook J . (2009). Priming in systemic plant immunity. Science 324: 89–91.
Hayward AC . (1993). The hosts of Xanthomonas. In Swings JG, Civerolo EL, (eds) Xanthomonas. Springer: Netherlands, pp 1–17.
Huang S, Kang J, Blaser MJ . (2006). Antimutator role of the DNA glycosylase mutY gene in Helicobacter pylori. J Bacteriol 188: 6224–6234.
Jalan N, Aritua V, Kumar D, Yu F, Jones JB, Graham JH et al. (2011). Comparative genomic analysis of Xanthomonas axonopodis pv. citrumelo F1, which causes citrus bacterial spot disease, and related strains provides insights into virulence and host specificity. J Bacteriol 193: 6342–6357.
Jalan N, Kumar D, Andrade MO, Yu FH, Jones JB, Graham JH et al. (2013a). Comparative genomic and transcriptome analyses of pathotypes of Xanthomonas citri subsp. citri provide insights into mechanisms of bacterial virulence and host range. BMC Genomics 14: 551.
Jalan N, Kumar D, Yu F, Jones JB, Graham JH, Wang N . (2013b). Complete genome sequence of Xanthomonas citri subsp. citri strain AW12879, a restricted-host-range citrus canker-causing bacterium. Genome Announc 1: e00235–00213.
Jones JDG, Dangl JL . (2006). The plant immune system. Nature 444: 323–329.
Jones KE, Patel NG, Levy MA, Storeygard A, Balk D, Gittleman JL et al. (2008). Global trends in emerging infectious diseases. Nature 451: 990–993.
Laia M, Moreira L, Dezajacomo J, Brigati J, Ferreira C, Ferro M et al. (2009). New genes of Xanthomonas citri subsp. citri involved in pathogenesis and adaptation revealed by a transposon-based mutant library. BMC Microbiol 9: 12.
Langille MGI, Brinkman FSL . (2009). IslandViewer: an integrated interface for computational identification and visualization of genomic islands. Bioinformatics 25: 664–665.
Leyns F, De Cleene M, Swings J-G, De Ley J . (1984). The host range of the genus Xanthomonas. Bot Rev 50: 308–356.
Li J, Wang N . (2011). Genome-wide mutagenesis of Xanthomonas axonopodis pv. citri reveals novel genetic determinants and regulation mechanisms of biofilm formation. PLoS One 6: e21804.
Lieberman TD, Michel JB, Aingaran M, Potter-Bynoe G, Roux D, Davis MR et al. (2011). Parallel bacterial evolution within multiple patients identifies candidate pathogenicity genes. Nat Genet 43: 1275–1280.
Livak KJ, Schmittgen TD . (2001). Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 25: 402–408.
Malamud F, Homem RA, Conforte VP, Yaryura PM, Castagnaro AP, Marano MR et al. (2013). Identification and characterization of biofilm formation-defective mutants of Xanthomonas citri subsp citri. Microbiology 159: 1911–1919.
Marolda CL, Lahiry P, Vines E, Saldias S, Valvano MA . (2006). Micromethods for the characterization of lipid A-core and O-antigen lipopolysaccharide. Methods Mol Biol 347: 237–252.
Markowitz VM, Mavromatis K, Ivanova NN, Chen I-MA, Chu K, Kyrpides NC . (2009). IMG/ER: a system for microbial genome annotation expert review and curation. Bioinformatics 25: 2271–2278.
McCann HC, Guttman DS . (2008). Evolution of the type III secretion system and its effectors in plant-microbe interactions. New Phytol 177: 33–47.
Merhej V, Georgiades K, Raoult D . (2013). Postgenomic analysis of bacterial pathogens repertoire reveals genome reduction rather than virulence factors. Brief Funct Genomics 12: 291–304.
Mhedbi-Hajri N, Hajri A, Boureau T, Darrasse A, Durand K, Brin C et al. (2013). Evolutionary history of the plant pathogenic bacterium Xanthomonas axonopodis. PLoS One 8: e58474.
Moreira LM, Almeida NF Jr ., Potnis N, Digiampietri LA, Adi SS, Bortolossi JC et al. (2010). Novel insights into the genomic basis of citrus canker based on the genome sequences of two strains of Xanthomonas fuscans subsp aurantifolii. BMC Genomics 11: 238.
Murrell B, Moola S, Mabona A, Weighill T, Sheward D, Pond SLK et al. (2013). FUBAR: a fast, unconstrained bayesian approximation for inferring selection. Mol Biol Evol 30: 1196–1205.
Nicaise V, Roux M, Zipfel C . (2009). Recent advances in PAMP-triggered immunity against bacteria: pattern recognition receptors watch over and raise the alarm. Plant Physiol 150: 1638–1647.
Ngoc LBT, Verniere C, Jarne P, Brisse S, Guerin F, Boutry S et al. (2009). From local surveys to global surveillance: three high-throughput genotyping methods for epidemiological monitoring of Xanthomonas citri pv. citri pathotypes. Appl Environ Mmicrobiol 75: 1173–1184.
O'Donoghue P, Luthey-Schulten Z . (2003). Evolution of structure in aminoacyl-tRNA synthetases. Microbiol Mol Biol Rev 67: 550–573.
Petrocelli S, Laura Tondo M, Daurelio LD, Orellano EG . (2012). Modifications of Xanthomonas axonopodis pv. citri lipopolysaccharide affect the basal response and the virulence process during citrus canker. PLoS One 7: e40051.
Pond SLK, Frost SDW, Muse SV . (2005). HyPhy: hypothesis testing using phylogenies. Bioinformatics 21: 676–679.
Pond SLK, Posada D, Gravenor MB, Woelk CH, Frost SDW . (2006). Automated phylogenetic detection of recombination using a genetic algorithm. Mol Biol Evol 23: 1891–1901.
Popa O, Hazkani-Covo E, Landan G, Martin W, Dagan T . (2011). Directed networks reveal genomic barriers and DNA repair bypasses to lateral gene transfer among prokaryotes. Genome Res 21: 599–609.
Posada D, Crandall KA . (1998). MODELTEST: testing the model of DNA substitution. Bioinformatics 14: 817–818.
Pruvost O, Magne M, Boyer K, Leduc A, Tourterel C, Drevet C et al. (2014). A MLVA genotyping scheme for global surveillance of the citrus pathogen Xanthomonas citri pv. citri suggests a worldwide geographical expansion of a single genetic lineage. PLoS One 9: e98129.
Richter M, Rossello-Mora R . (2009). Shifting the genomic gold standard for the prokaryotic species definition. Proc Natl Acad Sci USA 106: 19126–19131.
Rohmer L, Hocquet D, Miller SI . (2011). Are pathogenic bacteria just looking for food? Metabolism and microbial pathogenesis. Trends Microbiol 19: 341–348.
Ryan RP, Vorhölter F-J, Potnis N, Jones JB, Van Sluys M-A, Bogdanove AJ et al. (2011). Pathogenomics of Xanthomonas: understanding bacterium–plant interactions. Nat Rev Microbiol 9: 344–355.
Rybak M, Minsavage GV, Stall RE, Jones JB . (2009). Identification of Xanthomonas citri ssp citri host specificity genes in a heterologous expression host. Mol Plant Pathol 10: 249–262.
Schubert TS, Rizvi SA, Sun XA, Gottwald TR, Graham JH, Dixon WN . (2001). Meeting the challenge of eradicating citrus canker in Florida - Again. Plant Dis 85: 340–356.
Scora RW . (1975). On the history and origin of Citrus. Bull Torrey Bot Club 102: 369–375.
Shimodaira H, Hasegawa M . (1999). Multiple comparisons of log-likelihoods with applications to phylogenetic inference. Mol Biol Evol 16: 1114–1116.
Sun XA, Stall RE, Jones JB, Cubero J, Gottwald TR, Graham JH et al. (2004). Detection and characterization of a new strain of citrus canker bacteria from key Mexican lime and Alemow in South Florida. Plant Dis 88: 1179–1188.
Swofford DL . (1998) PAUP*. Phylogenetic Analysis Using Parsimony (*and Other Methods). Version 4. Sinauer: Sunderland, MA.
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S . (2013). MEGA6: molecular evolutionary genetics analysis version 6.0. Mol Biol Evol 30: 2725–2729.
Trivedi P, Wang N . (2013). Host immune responses accelerate pathogen evolution. ISMEJ 8: 727–731.
Vernière C, Bui Thi Ngoc L, Jarne P, Ravigné V, Guérin F, Gagnevin L et al. (2014). Highly polymorphic markers reveal the establishment of an invasive lineage of the citrus bacterial pathogen Xanthomonas citri pv. citri in its area of origin. Environ Microbiol 16: 2226–2237.
Vernikos GS, Parkhill J . (2006). Interpolated variable order motifs for identification of horizontally acquired DNA: revisiting the Salmonella pathogenicity islands. Bioinformatics 22: 2196–2203.
Wolfe ND, Dunavan CP, Diamond J . (2007). Origins of major human infectious diseases. Nature 447: 279–283.
Yan Q, Hu X, Wang N . (2012). The novel virulence-related gene nlxA in the lipopolysaccharide cluster of Xanthomonas citri ssp. citri is involved in the production of lipopolysaccharide and extracellular polysaccharide, motility, biofilm formation and stress resistance. Mol Plant Pathol 13: 923–934.
Zhao C, Nie H, Shen Q, Zhang S, Lukowitz W, Tang D . (2014). EDR1 physically interacts with MKK4/MKK5 and negatively regulates a MAP kinase cascade to modulate plant innate immunity. PLoS Genet 10: e1004389.
Acknowledgements
This study was supported by Citrus Research and Development Foundation and USDA- CSREES Special Citrus Canker Grant Project 2011-73402.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Rights and permissions
About this article
Cite this article
Zhang, Y., Jalan, N., Zhou, X. et al. Positive selection is the main driving force for evolution of citrus canker-causing Xanthomonas. ISME J 9, 2128–2138 (2015). https://doi.org/10.1038/ismej.2015.15
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2015.15
This article is cited by
-
Insights into Superinfection Immunity Regulation of Xanthomonas axonopodis Filamentous Bacteriophage cf
Current Microbiology (2024)
-
Herbarium specimen sequencing allows precise dating of Xanthomonas citri pv. citri diversification history
Nature Communications (2023)
-
Origin and diversification of Xanthomonas citri subsp. citri pathotypes revealed by inclusive phylogenomic, dating, and biogeographic analyses
BMC Genomics (2019)
-
Pacbio sequencing of copper-tolerant Xanthomonas citri reveals presence of a chimeric plasmid structure and provides insights into reassortment and shuffling of transcription activator-like effectors among X. citri strains
BMC Genomics (2018)
-
Proteomics-based identification of differentially abundant proteins reveals adaptation mechanisms of Xanthomonas citri subsp. citri during Citrus sinensis infection
BMC Microbiology (2017)