Abstract
Antibiotic-resistance genes (ARGs) are currently discussed as emerging environmental contaminants. Hospital and municipal sewage are important sources of ARGs for the receiving freshwater bodies. We investigated the spatial distribution of different ARGs (sul1, sul2, tet(B), tet(M), tet(W) and qnrA) in freshwater lake sediments in the vicinity of a point source of treated wastewater. ARG contamination of Vidy Bay, Lake Geneva, Switzerland was quantified using real-time PCR and compared with total mercury (THg), a frequently particle-bound inorganic contaminant with known natural background levels. Two-dimensional mapping of the investigated contaminants in lake sediments with geostatistical tools revealed total and relative abundance of ARGs in close proximity of the sewage discharge point were up to 200-fold above levels measured at a remote reference site (center of the lake) and decreased exponentially with distance. Similar trends were observed in the spatial distribution of different ARGs, whereas distributions of ARGs and THg were only moderately correlated, indicating differences in the transport and fate of these pollutants or additional sources of ARG contamination. The spatial pattern of ARG contamination and supporting data suggest that deposition of particle-associated wastewater bacteria rather than co-selection by, for example, heavy metals was the main cause of sediment ARG contamination.
Similar content being viewed by others
Introduction
Since their introduction into medical practice in the 1930s, antibiotics have saved millions of lives, but the increasing numbers of bacterial pathogens that are becoming resistant to antibiotics is a growing cause of concern. Research initially focused on clinical settings as the main site of the spread and evolution of antibiotic resistance. Over the course of the past two decades, however, researchers have increasingly broadened their focus to include the environment as a source of resistance genes and as a site of antibiotic-resistance evolution (Kümmerer, 2004). The existence of a natural environmental resistance gene pool, which includes all known clinical resistance mechanisms, has been discovered (D'Costa et al., 2006; Aminov, 2009). Consequently, bacterial pathogens are assumed to acquire resistance traits from the environment via horizontal gene transfer (Aminov, 2009; Martinez, 2009a). Moreover, various studies have demonstrated that wastewater and animal waste contain large numbers of resistant bacteria, which can pass through the wastewater treatment plant (WWTP) and reach the receiving water bodies (Teuber, 2001; Kümmerer, 2004; Jury et al., 2011; Rizzo et al., 2013; Zurfluh et al., 2013). There is growing concern that continuous discharge of these contaminants may, at least locally, lead to an increase in the natural resistance background levels and thus enhance the likelihood of antibiotic-resistance genes (ARGs) being transferred back to human and animal commensals or even pathogens (Kim and Aga, 2007; Martinez, 2009b; Cantas et al., 2013). In the long term, this ‘environmental loop’ could contribute to the spread of resistance in pathogens and undermine the effectiveness of current and future antibiotics.
Lake Geneva is the largest freshwater body in Western Europe and receives treated wastewaters from the surrounding cities. The largest WWTP is located in Lausanne and discharges treated wastewater and occasionally bypass water into Vidy Bay. The bay’s sediments have been reported to be heavily polluted by fecal-indicator bacteria, heavy metals, nitrogen and phosphorus (Poté et al., 2008). Micropollutants, including antibiotics (Bonvin et al., 2011), as well as multiresistant bacteria and ARGs pass through the WWTP and are subsequently discharged in considerable amounts into Vidy Bay (Czekalski et al., 2012). Low levels of multiresistant bacteria and ARGs were found even 3.2 km from the WWTP discharge pipe, where lake water is pumped for drinking water production (Czekalski et al., 2012). However, it remained unclear whether this represented contamination or the (unknown) natural resistance background level of Lake Geneva.
Molecular tools, such as PCR and quantitative real-time PCR, have been developed for many ARGs (Heuer and Smalla, 2007; Walsh et al., 2011). A limited number of studies have applied molecular methods to aquatic environments. Primarily, rivers impacted by different contamination sources have been considered (Pei et al., 2006; Pruden et al., 2006; Graham et al., 2010). In these systems, the horizontal transport of contaminants is directional and constrained by the river channel and therefore relatively well defined. This is also true for most of the sampling points in the study of LaPara et al. (2011), which followed the input of ARGs with wastewater into the St Louis river and transport into Lake Superior. In freshwater lakes, complex transport processes prevail, dominated, for example, by temporally variable wind regimes, currents and stratification. Despite the high relevance of lakes as sources of drinking water, food and as a place for recreational activities, the dynamics of antibiotic-resistance contamination in lakes that are receiving waters for municipal wastewater has not been thoroughly investigated. It is currently unknown, for example, at which spatial scales a local source of antibiotic-resistant bacteria affects water and sediment in a receiving lake.
The present study aims to characterize the spatial impact of Lausanne’s WWTP discharge on the concentration of several plasmid-mediated ARGs in the lake sediment. The tested ARGs confer resistance to different classes of broad-spectrum antibiotics (sulfonamides: sul1, sul2, tetracyclines: tet(M), tet(B), tet(W)) and fluoroquinolones: qnrA), which differ in their origin (for example, natural versus synthetic), time of introduction, resistance mechanisms and clinical relevance. Sulfonamides were the first synthetic antibacterials. Tetracyclines, isolated from Streptomyces, followed in 1948. Fluoroquinolones were introduced during the 1970s and are, of the three substance classes, currently the most frequently used in clinics. ARGs were compared with an already well-studied WWTP-released contaminant, total mercury (THg), for which background levels in Lake Geneva are known from previous studies (Arboille et al., 1989; Poté et al., 2008). Mapping of the two-dimensional spatial pattern of the investigated ARGs and THg was carried out using a geographic information system (ArcGIS). Further data on the microbial community structure in wastewater and sediment samples and sediment properties were considered in order to interpret the observed spatial trends of ARGs in Vidy bay sediments.
Materials and methods
Study site description and sampling campaigns
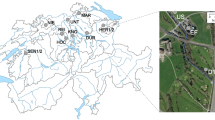
For a detailed description of the Vidy Bay study site, please refer to Czekalski et al. (2012). Lausanne (population 214 000) is the biggest city in the Lake Geneva area. The city has no pharmaceutical industries but several health-care centers. The largest is the Centre Hospitalier Universitaire Vaudois, an important site of antibiotic consumption, which discharges ∼410 m3 day−1 of raw wastewater into the municipal sewage system (Blanc, 2010). The conventional WWTP of Lausanne treats the bulk of the wastewater it receives with a chemical phosphate removal process and an aerobic biological treatment, followed by clarification. Sludge is recycled or incinerated. Only part of the water receives a more elaborate biological denitrification treatment (Supplementary Figure S1). Disinfection is not performed. Treated wastewater and occasionally untreated sewer overflow (bypass) is discharged at an average rate of 86.631 m3 day−1 (1–3 m3 s−1 up to 5-6 m3 s−1 at low and peak flow, respectively) (Vioget et al., 2011). The effluent pipe discharges the treated water 750 m offshore at a depth of 35 m into the Vidy Bay of Lake Geneva (Figure 1, sampling point STEP (‘station d’épuration’)). The rivers Chamberonne and Flon (sampling points CHAM and FLON) form part of the city’s combined sewer overflow system and also discharge into the bay (1-3 m3 s−1, Goldscheider et al., 2007). The main form of agricultural land use in their catchment areas is vineyards. Intensive animal farming is not prevalent. About 3.2 km south-west from STEP, lake water is pumped for drinking water production (sampling point DWP).
Map of Vidy Bay, indicating its location in Lake Geneva, 22 sites from which sediments were sampled, and the two river mouths of the Flon and Chamberonne (CHAM). The WWTP discharge pipe is indicated by a dashed line starting at Lausanne’s WWTP to the point of discharge, the STEP site. Bathymetric data provided by Anh-Dao Le thi and Walter Wildi, Institut F.A. Forel. Maps used with permission of swisstopo (Art. 30 GeoIV): PK50©2007, 2005©swisstopo.
Short sediment cores were taken at 22 sites during two sampling campaigns in August 2011 (Figure 1). Sediment cores were retrieved as described previously (Czekalski et al., 2012). Sediment cores were transported to the laboratory within 8 h and stored at 4 °C for a maximum of 48 h until processing. Three subsamples of 0.5 g (wet weight) from the sediment surface layer (0–3 cm) of each core were transferred to sterile 2-ml screw cap tubes. After shock-freezing in liquid nitrogen, the tubes were stored at −80 °C until DNA extraction. For a subset of sites, additional subsamples of surface sediments were taken, immediately preserved by freezing at −20 °C and freeze-dried to analyze THg, grain size and organic matter content.
Details on additional water samples taken in 2009 and 2010 from Lausanne’s sewage system and lake water in Vidy Bay that were analyzed by plate counts of resistant bacteria and qPCR assays are given in the Supplementary Information.
DNA extraction
DNA from the three subsamples from each sampling site was extracted, quantified and quality-controlled as described in Czekalski et al. (2012). Briefly, cells were lysed using bead-beating and freeze–thaw cycles, and DNA was purified using a phenol–chloroform extraction protocol. Extraction blanks omitting sediment were prepared to check for contamination sources. DNA quality was checked via agaose gel electrophoresis and ultraviolet-absorbance (Nanodrop, Thermo Fisher Scientific Inc., NanoDrop products, Wilmington, DE, USA) and quantified using the Quant-iT PicoGreen DNA quantification kit (Invitrogen, Basel, Switzerland). Extraction blanks yielded negative results. DNA pellets were resuspended in 50–100 μl of Tris–EDTA buffer and stored at −20 °C until further analysis.
qPCR assays performed with sediment, lake and wastewater DNA extracts
Six different ARGs (sul1, sul2, tet(B), tet(M), tet(W) and qnrA) and bacterial 16S rRNA gene fragments were quantified using the published protocols, primers and probes (Supplementary Table S1). Standard curves were prepared from serial 10-fold dilutions of plasmid DNA containing the respective target gene in a range of 3 × 106 to 30 gene copies. The preparation of control plasmids and standard curves for sul1 and sul2 was as described previously (Heuer and Smalla, 2007; Heuer et al., 2008). For tet(B), tet(M), tet(W) and qnrA, standard curves were prepared from cloned PCR products as described for the 16S rRNA gene in Czekalski et al. (2012). qPCR assays for sul, tet and 16S rRNA genes were performed in 96-well plates using TaqMan Environmental Master Mix 2.0 (Life Technologies Corp., Applied Biosystems, Carlsbad, CA, USA), which is optimized for samples with confirmed or expected presence of PCR inhibitors. qnrA was amplified using QuantiTect SYBR Green PCR chemistry (Qiagen, Hombrechtikon, Switzerland). In brief: each 25-μl reaction mixture contained 1 × of either Environmental or SYBR green Master Mix, primers, probe and MgCl2 according to the published methods (Supplementary Table S1) and 5 μl of template DNA or standard. All qPCR assays were performed in technical triplicates on each extract, standard and negative controls (no template control: salmon sperm DNA, 10 ng μl−1 in order to control for specificity; PCR blanks: nuclease-free water; and extraction blanks: see above). qPCR reactions were performed on a 7500 Fast Real-Time PCR System (Applied Biosystems) and analyzed using default settings.
All negative controls resulted either in no amplification or a higher threshold cycle (Ct) than the most diluted quantification standard. A sample was considered to be below limit of detection (LOD) for a target gene if ⩾2 out of 3 technical replicates were negative or if sample Ct values were ⩾Ct of negative controls. Samples above LOD were considered to be below the limit of quantification when the s.d. of Ct values of methodological triplicates was >0.5 and their Ct value was higher than the Ct of the most diluted standard whose standard deviation of Ct values was ⩽0.5.
For each PCR assay, the qPCR efficiency was calculated from the slope of the standard curve (E=10(−1/slope); Supplementary Table S2). For a subset of seven samples and for one target gene (sul2), the effect of inhibitors in the sample matrix on amplification in qPCR assays was evaluated directly. Each of the samples was spiked with 105 copies of quantification standard DNA and amplified together with the same set of non-spiked samples and control DNA. The presence of inhibitors was evaluated according to Pei et al. (2006) by calculating the suppression factor S. S varied between 1.1 and 1.4, indicating that inhibition was negligible (Supplementary Table S3).
Automated ribosomal intergenic spacer analysis (ARISA)-PCR and sequencing
In all, 1:10 dilutions of DNA extracts were subjected to ARISA using general bacterial primers (Supplementary Table S1) and PCR conditions according to Yannarell et al. (2003) with slight modifications (Bürgmann et al., 2011). A total of 10 ng μl−1 of Escherichia coli DNA served as a positive control, 10 ng μl−1 of salmon sperm DNA and nuclease-free water served as no template control and PCR blank, respectively.
Samples were run on an ABI 3130xl capillary sequencer as described previously (Bürgmann et al., 2011). Data analysis was performed using Gene Mapper Software version 4.0 (Applied Biosystems) considering only peaks with sizes between 350 and 1250 bp and a minimum peak height of 125 fluorescence units. ARISA data were subjected to the automatic and interactive binning scripts (Ramette, 2009) for R (version 2.14.2, (2012)) using a window size of 2.5 bp.
Hg analysis
THg concentrations were determined by cold vapor atomic absorption spectrophotometry after dry mineralization and pre-concentration of Hg by means of amalgamation on a gold trap (Száková et al., 2004), using an automatic mercury analyzer (Model AMA 254, Altec, Prague, Czech Republic) under the following conditions: typical sample mass 50–100 mg, drying time 45 s, decomposition time 150 s and waiting time (necessary for quantitative trapping of released mercury on the gold amalgamator) 45 s. All analyses were run in duplicates. The relative error was usually±5% and always under±10% (Roos-Barraclough et al., 2002). The limit of detection is 0.01 ng, and the working range is between 0.05 to 600 ng (Yang and Pan, 2007). Concentrations obtained for repeated analyses of certified reference materials never exceeded the published range of concentration (0.091±0.008 μg g−1 for MESS-3).
Sediment characterization and nutrient content
The sediment grain size was measured using a Coulter LS-100 laser diffractometer (Beckman Coulter, Fullerton, CA, USA), following 5-min ultrasonic dispersal in de-ionized water according to the method of Loizeau et al. (1994).
The sediment total organic matter content was estimated from loss on ignition at 550 °C for 1 h in a laboratory oven (LE14/11, Nabertherm, Lilienthal, Germany).
Data analysis, visualization and statistics
Total copy numbers of all target genes were normalized to sediment sample wet weight (g-1 wwt). This is referred to as ARG concentration from here on. Additionally, ARG copy numbers were normalized to 16S rRNA gene copy numbers and reported as percentages, as an indicator of the relative abundance of resistance genes within the bacterial population. The number copies of the 16S rRNA genes per cell varies among bacterial species and may thus vary between samples, but keeping this caveat in mind we refer to this measure as ‘ARG abundance’ from here on. Interpolation maps were created with ArcGIS (ESRIArcMap 10, Redlands, CA, USA) with the Geostatistical Analyst extension, using the Inverse Distance Weighted interpolation algorithm with power functions from 1 to 10 and a smoothing factor of 0.2.
In order to evaluate similarity of the distribution patterns between resistance genes and between resistance genes and Hg concentrations, as well as between ARGs and distance from the contamination source, correlation and regression analysis were performed in MS Excel 2010 (Microsoft, Redmond, WA, USA), using the Data Analysis Tool.
Community similarity analysis was performed based on Hellinger-transformed ARISA peak area data in R version 2.14.2 (R Core Team, 2012), using the BiodiversityR and vegan packages (Oksanen et al., 2012; Kindt and Coe, 2005). Principle coordinates analysis was performed using the Bray distance metric. ARG abundance and environmental variables were passively fitted to community principle coordinates analysis data, and significance of fit was analyzed by means of permutation analysis (n=1000).
Results and discussion
Spatial trends in ARG abundance
A central goal of this study was to characterize the spatial impact of Lausanne’s WWTP discharge pipe on the contamination of Vidy Bay sediments with different ARGs. sul genes were detected at all and tet genes at several of the sampled locations, whereas qnrA was always below detection. ARG abundance generally decreased from sul1 over sul2, tet(w), tet(M) to tet(B) (Figure 2). The prevalence of ARGs in Vidy bay sediments thus reflects their abundance in Lausanne’s wastewater: qnrA was detected in low amounts and only in hospital sewage, whereas sul and tet were more abundant in both WWTP influent and effluent samples, with sul1 abundance always being the highest (Supplementary Figure S5). This abundance pattern may be related to the longer history of clinical application of sulfonamides and tetracyclines. sul and tet genes have previously been found to be quite abundant in sewage and receiving waters (Pruden et al., 2006) and in livestock waste sludge (Zhang et al., 2013). Plasmid-mediated quinolone resistance encoded by qnrA was first reported in 1998 (Martinez-Martinez et al., 1998), which may explain its low abundance in Lausanne’s wastewater system. Other quinolone-resistance mechanisms were likewise infrequent, as shown by low colony counts on Ofloxacin- and Norfloxacin-amended agar plates inoculated with samples from Lausanne’s sewage and wastewater treatment system (Supplementary Figures S6 and S7).
Log–log plot of ARG abundance (ARG copy numbers normalized to bacterial 16S rRNA gene copy numbers) versus distance from the STEP site. STEP was arbitrarily given a distance of ½ the distance to the next closest site to allow it to be included in the analysis. Lines with R2 values are for fitted functions of type y =a × xb.
For all the detected ARGs, highest gene concentrations and abundances occurred at sites in close proximity to the WWTP discharge (Figure 2), especially at sites STEP, EG2 and NC4 (Supplementary Figures S2 and S3). At site EG2, a maximum of 2.2 × 109 (sul1) and of 1.5 × 106 (tet(B)) ARG copies g−1 wwt were determined (Supplementary Figure S2). Abundance of sul1 at this site reached 12% (Supplementary Figure S3). ARG concentration and abundance of both sul and tet genes were also high at sites NC1, NC2, NC3, NC5 and EG3 (Supplementary Figures S2 and S3), which are all located within 350 m of the STEP coordinate (Figure 1). Only sul1 was also detected at comparatively high concentration and abundance at more remote sites (EG4, NC7, NC9 and NC14).
The relationship of ARGs with distance from the WWTP discharge pipe was not linear (Pearson’s r∼−0.17 for ARG concentration and r ∼−0.36 for ARG abundance, Supplementary Tables S4 and S5), but ARG abundance followed an exponential decay function (Figure 2). The decreasing trend appeared to be intact across the entire range of distances sampled. ARG concentration trends were similar as 16S rRNA gene copy numbers did not show a clear distance-related trend (Supplementary Figure S4). However, for each ARG, the R2 was higher for abundance data compared with concentration data. A possible explanation would be sample-to-sample variations in PCR inhibition in spite of our best efforts to minimize such effects. We concluded that ARG abundance is slightly more robust for the purpose of our analysis. The R2 for ARG abundance varied between 0.48 and 0.68, indicating that considerable amounts of variation are not explained by distance (Figure 2).
We therefore used spatial mapping to investigate whether deposition patterns deviate from a simple distance relationship, indicating directional transport (Figure 3 and Supplementary Figure S5). The maps clearly show the accumulation of ARGs around the WWTP discharge pipe. Designating a sul1 abundance >1% as ‘contaminated’, Figure 3a indicates an area of at least ∼0.3 km2 has been impacted. All sampling points within this area are hereafter referred to as ARG-contaminated sediments.
ARG maps indicated a strong decrease in ARG abundance and concentration towards the deep lake (EG2 to EG9) and towards the south east (sampling locations EG14 and EG15). Ct values obtained for EG6, EG9, EG15 and the center of the lake (reference point SHL2) were either below the LOQ (sul genes) or even below LOD (tet genes). In contrast, our data indicated a less dramatic decrease in ARG levels along the transect following the shore line in south-western direction (NC5–NC13, Figures 2a–e and Figures 3a–e).
Finally, apart from site NC14 (close to the Chamberonne river—CHAM), samples taken close to shore (NC15, EG1) showed low levels of ARGs.
These observations indicate a directionality of the pollutant transport in the water column of Vidy Bay. Hydrodynamic transport in lakes is often complex. Currents are influenced by changing wind regimes, stratification and bathymetry and vary in direction and speed with depth (Wüest and Lorke, 2003; Righetti et al., 2011). For Vidy Bay, it has recently been shown that near-shore gyres can occur (which enhance the retention time of water and pollutants in the bay) and that neither of two dominant wind regimes produce a consistent current pattern (Razmi et al., 2013). However, westward shore-parallel currents can disrupt gyre formation and move water from near the STEP site to the water column above the DWP, with a dilution factor of about 100 (Razmi et al., 2013). In previous studies, tracer released from the WWTP during holomixis reached surface waters at the DWP site (NC13) within 48 h, whereas no tracers were detected when thermal stratification precluded upwelling (Wildi and Rossi, 1997; Goldscheider et al., 2007). In the case of particle-bound contaminants, the contamination record in sediments is expected to integrate over such temporally variable transport processes.
Contamination effects on the microbial community
Analysis of bacterial community similarity (Figure 4a) showed a clearly distinct community in sites close to the discharge pipe. This confirms previous clone library-based observations from Vidy Bay (Haller et al., 2011). The communities at the most strongly ARG-contaminated sites STEP and EG2 were highly similar to some untreated wastewater communities. Fitting of ARG abundances and environmental variables to the sediment community composition showed that these community changes correlated with indicators of contamination, such as ARG abundance, THg and total organic matter (Figure 4b). In contrast, particle composition parameters appeared either unrelated to the distinction between contaminated and uncontaminated sites (silt) or were not significantly correlated (clay, P>0.05). Vectors fitted for sul2 and tet(B) indicated a slightly different distribution within the microbial community compared with the other ARGs. Overall, community analysis indicated that changes in ARG abundance were related to changes in community composition, which can be explained by deposition of bacteria originating in wastewater. However, an impact of confounding factors, such as heavy metal pollution, cannot be ruled out by this analysis.
(a) Principle coordinates analysis of ARISA community profiles from sediment and water. Both biological and technical replicates are included. Dispersion ellipses are shown for sediment sampling sites near the WWTP discharge pipe and for untreated hospital and municipal wastewater (Sewage) (b) Principle coordinates analysis of ARISA community profiles from selected sediments with passively fitted environmental variables and ARG abundance. For this analysis, technical replicates were averaged, omitting peaks only present in one replicate. Samples were selected to match ARGs and sediment data availability. Symbols indicate samples grouped according to sample type and location. n.s., not significant (P>0.05).
Comparison of ARGs and Hg contamination
The spatial trends in ARG distribution were compared with THg levels (Figures 2 and 3). Hg is a pollutant that is known to be released from Lausanne’s WWTP (Loizeau et al., 2004). Background concentrations in Lake Geneva sediments are ∼0.2 mg kg−1. Although the chemistry and transport behavior of Hg is complex, particulate transport is dominant in most systems (Glass et al., 1990; Wang et al., 2004). Assuming that particle-associated bacteria or bacterial aggregates released from the WWTP are in turn primarily responsible for ARG deposition in the sediment, we expected a similar pollution pattern for THg and ARGs. However, ARG dynamics may be affected by bacterial mortality, for example, through predation, ultraviolet radiation or starvation. This could lead to a more restricted distribution compared with the heavy metal. On the other hand, released resistant bacteria could proliferate or spread ARGs to the indigenous community via horizontal gene transfer. If the latter mechanisms dominate, a diffuse and more widespread pattern of ARG concentrations compared with THg would be expected. Differences as well as similarities in the spatial patterns of both contaminants may thus be informative.
Compared with previous studies (8 mg kg−1 sediment (Poté et al., 2008)), a relatively low maximum THg concentration of 1.3 mg kg-1 sediment was measured during our survey (EG2, compare Figure 2f), indicating reduced recent deposition. We found elevated THg levels to be the highest around the STEP site but above background levels up to 600 m away in direction of the deep lake (EG4), 535 m away in the south-western direction (NC6) and 877 m in the south-eastern direction (EG14, Figure 2f).
The similarity of the vectors of THg and ARG abundances fitted to community data in Figure 4b indicated a close link, which could either be deposition from a common source or because the heavy metal is a constraining factor of community composition. Nevertheless, there are notable differences in the spatial distribution, as shown by relatively low R2 of linear regressions of THg and ARG concentration (R2=0.32–0.41, P<0.05, Supplementary Table S6) and abundance (R2=0.21–0.63, P<0.01, except for sul2 with P=0.05, Supplementary Table S7). The steeper slope of ARG abundance with distance compared with THg (Figure 2) indicates restrictions on ARG dissemination compared with THg. However, the south-western transect deviated from this general observation. Here, ARG levels decreased more slowly, whereas sediments were classified as unimpacted in terms of THg (Figure 3f).
Finally, low levels of THg were determined at near-shore sites (0.04 and 0.05 mg kg−1 for EG1 and NC14, respectively), most likely related to the comparatively large grain sizes and low organic matter content (Supplementary Table S8) of sediments deposited in this area by the Chamberonne and Flon rivers. In contrast, site NC14 near the Chamberonne river mouth showed elevated ARG levels (Figures 3a, b and f). Previous studies monitoring pollutant transport in the water column (Wildi and Rossi, 1997; Goldscheider et al., 2007; Haldimann, 2009; Bonvin et al., 2011) document that at least under certain circumstances site NC13 might be affected by contaminants released from the WWTP, including ARGs (Czekalski et al., 2012). However, the Chamberonne river is known as a discontinuous source of fecal bacteria (Haldimann, 2009; Haller et al., 2009), whereas its THg load is too low to explain elevated concentrations measured in sediments of Vidy Bay (Howa and Vernet, 1988). Our data show it to be a likely source of resistant bacteria and ARG contamination (compare Supplementary Figures S5, S6f and S7f). Low ARG levels detected at locations close to the shore further east (EG1 and NC15) would support the hypothesis that the Chamberonne plume is partly responsible for the extension of the ARG contamination in the south-western direction. This is also supported by current measurements in Vidy Bay in 2010, which revealed 45% of the currents moving west-southwest (Razmi et al., 2013).
Direct and indirect causes of ARG contamination
Independently of transport, sediment properties might also influence the natural abundance of ARGs and the further fate of deposited allochthonous ARGs. Sediments containing a high fraction of clay minerals can bind DNA after cell death and protect it from degradation by DNAse (Aardema et al., 1983). In the sampled sediments, the proportion of clay was quite low (0.01–3.92%, Supplementary Table S8). ARGs have previously been detected in extracellular DNA in livestock waste sludge but were one to two orders of magnitude less frequent than the intracellular fraction, and both fractions showed similar dynamics (Zhang et al., 2013). Therefore we assume that the detected ARGs reflect the ARGs in the viable bacterial biomass in our samples.
It has been demonstrated that heavy metals can cross- or co-select for ARGs (Berg et al., 2010) as the same mechanisms often underlie both resistances, or resistance genes are located on the same mobile genetic element (Baker-Austin et al., 2006; Knapp et al., 2011). As sediments in Vidy Bay are contaminated not only with Hg but also with other heavy metals in excess of background levels (Poté et al., 2008), it could be argued that the observed elevated ARG levels are caused by heavy metal pollution rather than bacterial deposition. In a study carried out along a section of the Almendares River, which is affected by different urban and industrial contamination sources, good correlations were demonstrated for sediment ARGs (tet and β-lactamases) and copper levels (Graham et al., 2010). Other heavy metals (Pb, Co, Zn) also partly correlated with ARGs, but neither Hg nor sul genes were considered in this study.
However, for the present study we conclude that the available evidence favors the hypothesis that the observed ARG levels are mainly a result of direct deposition of resistant bacteria rather than being due to co-selection by heavy metals: (a) The correlations of ARG levels with THg were relatively poor. The observed correlation can be explained by the parallel release of both types of contaminants from the WWTP. (b) The impact of the WWTP discharge on the deposition of fecal indicator bacteria and contrasting bacterial community composition has been shown previously (Haller et al., 2011). (c) Our analysis of community similarity using ARISA (Figure 4) confirmed a distinct community composition of ARG-contaminated sediments in the vicinity of the discharge pipe that, furthermore, exhibits a high similarity to sewage communities (Figure 4a), supporting the notion that bacteria from these environments are deposited in the sediments. (d) A more rapid decrease of ARG abundance with distance compared with THg (Figure 2) is best explained by different dynamics during transport or deposition. Finally, (e) we found highly similar patterns of ARG abundance in wastewater/sewage and sediment (Supplementary Figure S4). Nevertheless, the role of the heavy metal contamination in Vidy Bay for selection or persistence of ARGs warrants further study.
Implications
We present detailed results on the spatial distribution of different ARGs in sediments of a freshwater lake that can be convincingly linked to the inflow of municipal wastewater. Our results demonstrate that the combination of qPCR and geostatistical tools is a suitable approach to quantitatively detect and assess the dissemination of ARGs from local sources. The release of wastewater into Vidy Bay has resulted in a high ARG accumulation in sediments in the vicinity of the contamination source, but a diffuse zone of elevated ARG concentrations extends further west towards the location of the drinking water pumping station, indicating that transport takes place over considerable distances. Uptake of ARGs via drinking water is considered a risk for human health as horizontal transfer to pathogens may occur. Treatment and quality control of waste and drinking water should be reconsidered with this in mind. In Switzerland and many other European countries, the majority of WWTPs operate without a final disinfection treatment, and many rely on basic biological treatment. As modernization of many of these facilities (including Lausanne’s WWTP) is imminent or already under way, it will be important to consider modern procedures for disinfection and contaminant removal that may be able to remove ARGs from waste and drinking water (Dodd, 2012). Effective ARGs and pollutant removal may help to break the cycle of ARG exchange between the human and the aquatic environment. The available technologies should be studied with regards to their effectiveness in reducing both the load of resistant bacteria and potentially transferable DNA. It should be kept in mind, however, that chemical disinfection methods may themselves cause an undesirable selection for antibiotic resistance. Even wastewater receiving elaborate treatment and disinfection has been shown to be a source of ARG contamination (LaPara et al., 2011).
References
Aardema BW, Lorenz MG, Krumbein WE . (1983). Protection of sediment-adsorbed transforming DNA against enzymatic inactivation. Appl Environ Microbiol 46: 417–420.
Aminov RI . (2009). The role of antibiotics and antibiotic resistance in nature. Environ Microbiol 11: 2970–2988.
Arboille D, Howa H, Span D, Vernet J-P . (1989). Etude générale de la pollution par les métaux et répartition des nutriments dans les sédiments du Léman. Rapp. Comm. Int. prot. eaux Léman contre pollut. Campagne 1988: 139–172.
Baker-Austin C, Wright MS, Stepanauskas R, McArthur JV . (2006). Co-selection of antibiotic and metal resistance. Trends Microbiol 14: 176–182.
Berg J, Thorsen MK, Holm PE, Jensen J, Nybroe O, Brandt KK . (2010). Cu exposure under field conditions coselects for antibiotic resistance as determined by a novel cultivation-independent bacterial community tolerance assay. Environ Sci Technol 44: 8724–8728.
Blanc L . (2010) Characterisation of hospital wastewater: the case of the CHUV. MSc thesis, Cranfield University: Cranfield, UK and EPFL: Lausanne, Switzerland.
Bonvin F, Rutler R, Chèvre N, Halder J, Kohn T . (2011). Spatial and temporal presence of a wastewater-derived micropollutant plume in Lake Geneva. Environ Sci Technol 45: 4702–4709.
Bürgmann H, Jenni S, Vazquez F, Udert KM . (2011). Regime shift and microbial dynamics in a sequencing batch reactor for nitrification and anammox treatment of urine. Appl Environ Microbiol 77: 5897–5907.
Cantas L, Shah SQA, Cavaco LM, Manaia C, Walsh F, Popowska M et al (2013). A brief multi-disciplinary review on antimicrobial resistance in medicine and its linkage to the global environmental microbiota. Front Microbiol 4: 96.
Czekalski N, Berthold T, Caucci S, Egli A, Buergmann H . (2012). Increased levels of multiresistant bacteria and resistance genes after wastewater treatment and their dissemination into Lake Geneva, Switzerland. Front Microbiol 3: 106.
D'Costa VM, McGrann KM, Hughes DW, Wright GD . (2006). Sampling the antibiotic resistome. Science 311: 374–377.
Dodd MC . (2012). Potential impacts of disinfection processes on elimination and deactivation of antibiotic resistance genes during water and wastewater treatment. J Environ Monit 14: 1754–1771.
Glass GE, Sorensen JA, Schmidt KW, Rapp GR . (1990). New source identification of mercury contamination in the Great Lakes. Environ Sci Technol 24: 1059–1069.
Goldscheider N, Haller L, Pote J, Wildi W, Zopfi J . (2007). Characterizing water circulation and contaminant transport in Lake Geneva using bacteriophage tracer experiments and limnological methods. Environ Sci Technol 41: 5252–5258.
Graham DW, Olivares-Rieumont S, Knapp CW, Lima L, Werner D, Bowen E . (2010). Antibiotic resistance gene abundances associated with waste discharges to the Almendares River near Havana, Cuba. Environ Sci Technol 45: 418–424.
Haldimann S . (2009) Modelling the Spreading of Urban Water Discharges in the Coastal Zone of Lausanne. EPFL: Lausanne, Switzerland.
Haller L, Poté J, Loizeau JL, Wildi W . (2009). Distribution and survival of faecal indicator bacteria in the sediments of the Bay of Vidy, Lake Geneva, Switzerland. Ecol Indicators 9: 540–547.
Haller L, Tonolla M, Zopfi J, Peduzzi R, Wildi W, Poté J . (2011). Composition of bacterial and archaeal communities in freshwater sediments with different contamination levels (Lake Geneva, Switzerland). Water Res 45: 1213–1228.
Heuer H, Smalla K . (2007). Manure and sulfadiazine synergistically increased bacterial antibiotic resistance in soil over at least two months. Environ Microbiol 9: 657–666.
Heuer H, Focks A, Lamshöft M, Smalla K, Matthies M, Spiteller M . (2008). Fate of sulfadiazine administered to pigs and its quantitative effect on the dynamics of bacterial resistance genes in manure and manured soil. Soil Biol Biochem 40: 1892–1900.
Howa H, Vernet H-P . (1988). Metaux lourds dans les sediments des affluents vaudois de Léman. Institut F. A. Forel, Université de Genève: Versoix, Switzerland.
Jury KL, Khan SJ, Vancov T, Stuetz RM, Ashbolt NJ . (2011). Are sewage treatment plants promoting antibiotic resistance? Crit Rev Environ Sci Technol 41: 243–270.
Kim S, Aga DS . (2007). Potential ecological and human health impacts of antibiotics and antibiotic-resistant bacteria from wastewater treatment plants. J Toxicol Environ Health Part B Crit Rev 10: 559–573.
Kindt R, Coe R . (2005). Tree diversity analysis. A manual and software for common statistical methods for ecological and biodiversity studies: World Agroforestry Centre (ICRAF), Nairobi.
Knapp CW, McCluskey SM, Singh BK, Campbell CD, Hudson G, Graham DW . (2011). Antibiotic resistance gene abundances correlate with metal and geochemical conditions in archived Scottish soils. PLoS One 6: e27300.
Kümmerer K . (2004). Resistance in the environment. J Antimicrob Chemother 54: 311–320.
LaPara TM, Burch TR, McNamara PJ, Tan DT, Yan M, Eichmiller JJ . (2011). Tertiary-treated municipal wastewater is a significant point source of antibiotic resistance genes into Duluth-Superior Harbor. Environ Sci Technol 45: 9543–9549.
Loizeau JL, Pardos M, Monna F, Peytremann C, Haller L, Dominik J . (2004). The impact of a sewage treatment plant's effluent on sediment quality in a small bay in Lake Geneva (Switzerland-France). Part 2: Temporal evolution of heavy metals. Lakes Reserv Res Manage 9: 53–63.
Loizeau JL, Arbouille D, Santiago S, Vernet JP . (1994). Evaluation of a wide range laser diffraction grain size analyser for use with sediments. Sedimentology 41: 353–361.
Martinez-Martinez L, Pascual A, Jacoby G . (1998). Quinolone resistance from a transferable plasmid. Lancet 351: 797–799.
Martinez JL . (2009a). The role of natural environments in the evolution of resistance traits in pathogenic bacteria. Proc R Soc Lond Ser B Biol Sci 276: 2521–2530.
Martinez JL . (2009b). Environmental pollution by antibiotics and by antibiotic resistance determinants. Environ Pollut 157: 2893–2902.
Oksanen J, Blanchet FG, Kindt R, Legendre P, O'Hara RB et al (2012). vegan: Community Ecology Package. R package version 2.0-4.
Pei R, Kim S-C, Carlson KH, Pruden A . (2006). Effect of river landscape on the sediment concentrations of antibiotics and corresponding antibiotic resistance genes (ARG). Water Res 40: 2427–2435.
Poté J, Haller L, Loizeau JL, Garcia Bravo A, Sastre V, Wildi W . (2008). Effects of a sewage treatment plant outlet pipe extension on the distribution of contaminants in the sediments of the Bay of Vidy, Lake Geneva, Switzerland. Bioresour Technol 99: 7122–7131.
Pruden A, Pei R, Storteboom H, Carlson KH . (2006). Antibiotic resistance genes as emerging contaminants: studies in Northern Colorado. Environ Sci Technol 40: 7445–7450.
R Core Team. (2012) R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing: Vienna, Austria.
Ramette A . (2009). Quantitative community fingerprinting methods for estimating the abundance of operational taxonomic units in natural microbial communities. Appl Environ Microbiol 75: 2495–2505.
Razmi AM, Barry DA, Lemmin U, Bonvin F, Kohn T, Bakhtyar R . (2013). Direct effects of dominant winds on residence and travel times in the wide and open lacustrine embayment: Vidy Bay (Lake Geneva, Switzerland). Aquat Sci 1–13.
Righetti M, Toffolon M, Lucarelli C, Serafini M . (2011). Sediments as tracers for transport and deposition processes in peri-alpine lakes: a case study. J Hydrol 411: 1–11.
Rizzo L, Manaia C, Merlin C, Schwartz T, Dagot C, Ploy MC et al (2013). Urban wastewater treatment plants as hotspots for antibiotic resistant bacteria and genes spread into the environment: a review. Sci Total Environ 447: 345–360.
Roos-Barraclough F, Givelet N, Martinez-Cortizas A, Goodsite ME, Biester H, Shotyk W . (2002). An analytical protocol for the determination of total mercury concentrations in solid peat samples. Sci Total Environ 292: 129–139.
Száková J, Kolihová D, Miholová D, Mader P . (2004). Single-purpose atomic absorption spectrometer AMA-254 for mercury determination and its performance in analysis of agricultural and environmental materials. Chem Papers 58: 311–315.
Teuber M . (2001). Veterinary use and antibiotic resistance. Curr Opin Microbiol 4: 493–499.
Vioget P, Jaquerod C-A, Ruegg E, Poget E . (2011). Bilans 2010 de l'épuration vaudoise. Service des eaux, sols et assainissement, Canton de Vaud: Orbe, Savigny, Switzerland.
Walsh F, Ingenfeld A, Zampicolli M, Hilber-Bodmer M, Frey JE, Duffy B . (2011). Real-time PCR methods for quantitative monitoring of streptomycin and tetracycline resistance genes in agricultural ecosystems. J Microbiol Methods 86: 150–155.
Wang Q, Kim D, Dionysiou DD, Sorial GA, Timberlake D . (2004). Sources and remediation for mercury contamination in aquatic systems—a literature review. Environ Pollut 131: 323–336.
Wildi W, Rossi P . (1997). Assainissement de la baie de Vidy: tarcage par bactériophages 19-25 février 1997 6-11 mai 1997. Ville de Lausanne, Direction de la sécurité cociale et de lénvironnement, Service d'assainissement, Station d'épuration de Vidy, Université de Genève, Institut F.A. Forel, Section des sciences de la terre, Université Neuchâtel, Laboratoire de Microbiologie: Lausanne, Switzerland.
Wüest A, Lorke A . (2003). Small-scale hydrodynamics in lakes. Annu Rev Fluid Mech 35: 373–412.
Yang H-m, Pan W-P . (2007). Transformation of mercury speciation through the SCR system in power plants. J Environ Sci 19: 181–184.
Yannarell AC, Kent AD, Lauster DH, Kratz TK, Triplett W . (2003). Temporal patterns in bacterial communities in three temperate lakes of different trophic status. Microb Ecol 46: 391–405.
Zhang Y, Snow DD, Parker D, Zhou Z, Li X . (2013). Intracellular and extracellular antimicrobial resistance genes in the sludge of livestock waste management structures. Environ Sci Technol 47: 10206–10213.
Zurfluh K, Hächler H, Nüesch-Inderbinen M, Stephan R . (2013). Characteristics of extended-spectrum β-Lactamase- and Carbapenemase-producing Enterobacteriaceae isolates from rivers and lakes in Switzerland. Appl Environ Microbiol 79: 3021–3026.
Acknowledgements
We thank: Nina Bircher, José Santos Cáceres and Francisco Vazquez for substantial support of our laboratory work; Philippe Arpagaus and Janine Halder for assistance during field campaigns; Jean-Luc Loizeau and Andrew Clarke for kindly revising the manuscript; Holger Heuer, Fiona Walsh and Laurent Poirel for providing positive control strains; Fiona Walsh for support in establishing tet gene qPCR; Anh-Dao Le thi and Walter Wildi for providing the bathymetry map of Vidy Bay; Rosi Siber for substantial help with geostatistical analysis; Luca Rossi and Nathalie Chèvre for organizing access to sampling sites and background data; Gregor Maurer and Anoys Magnet (STEP Vidy) for access to their facilities and support; and COST action TD0803 ‘Detecting evolutionary hot spots of antibiotic resistances in Europe (DARE)’ for discussions and valuable feedback. Funding was provided by the Swiss National Science Foundation Grant PFMP2-123048.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Czekalski, N., Gascón Díez, E. & Bürgmann, H. Wastewater as a point source of antibiotic-resistance genes in the sediment of a freshwater lake. ISME J 8, 1381–1390 (2014). https://doi.org/10.1038/ismej.2014.8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2014.8
Keywords
This article is cited by
-
Synthesis and Application of MoS2 Nanosheets for the Removal of Amoxicillin from Water: Response Surface Method
Arabian Journal for Science and Engineering (2023)
-
Molecular Characterization and Antibiotic Susceptibility Profile of Acinetobacter baumannii Recovered from Hospital Wastewater Effluents
Current Microbiology (2022)
-
Antibiotic residues in environment: antimicrobial resistance development, ecological risks, and bioremediation
Environmental Science and Pollution Research (2022)
-
The occurrence of antimicrobial residues and antimicrobial resistance genes in urban drinking water and sewage in Southern Brazil
Brazilian Journal of Microbiology (2022)