Abstract
Viruses are the most abundant members of the human oral microbiome, yet relatively little is known about their biodiversity in humans. To improve our understanding of the DNA viruses that inhabit the human oral cavity, we examined saliva from a cohort of eight unrelated subjects over a 60-day period. Each subject was examined at 11 time points to characterize longitudinal differences in human oral viruses. Our primary goals were to determine whether oral viruses were specific to individuals and whether viral genotypes persisted over time. We found a subset of homologous viral genotypes across all subjects and time points studied, suggesting that certain genotypes may be ubiquitous among healthy human subjects. We also found significant associations between viral genotypes and individual subjects, indicating that viruses are a highly personalized feature of the healthy human oral microbiome. Many of these oral viruses were not transient members of the oral ecosystem, as demonstrated by the persistence of certain viruses throughout the entire 60-day study period. As has previously been demonstrated for bacteria and fungi, membership in the oral viral community was significantly associated with the sex of each subject. Similar characteristics of personalized, sex-specific microflora could not be identified for oral bacterial communities based on 16S rRNA. Our findings that many viruses are stable and individual-specific members of the oral ecosystem suggest that viruses have an important role in the human oral ecosystem.
Similar content being viewed by others
Introduction
Large communities of viruses populate the surfaces of the human body; however, the membership and role of viruses in these communities remain poorly understood. Many of the viruses in these communities are bacteriophage (Breitbart et al., 2003; Minot et al., 2011; Willner et al., 2011; Pride et al., 2012a; Minot et al., 2013), likely secondary to the abundance of bacterial cells compared with eukaryote cells in these communities. Viruses likely have a key role in microbial population structures by altering the composition of bacterial communities (Duerkop et al., 2012). The interaction of bacteria and viruses is thought to maintain broad bacterial diversity in the local environment via various ecological models, such as constant-diversity dynamics, periodic selection dynamics and kill-the-winner dynamics (Parada et al., 2008; Rodriguez-Valera et al., 2009; Sandaa et al., 2009; Rodriguez-Brito et al., 2010; Allen et al., 2011). As cellular microbiota are known to be associated with various aspects of human health, there is a potential for viruses to have a role in human health and disease by altering native microbial community diversity. In addition to these various ecological models in which viruses predate upon bacteria, viruses may have a more subtle role in bacterial communities. By contributing to the dissemination of virulence genes, including those that confer antibiotic resistance, viruses and their communities can serve as reservoirs for genetic exchange to pathogenic bacteria and contribute to the persistance of resistance genes (Paulsen et al., 2003; Pride et al., 2012a).
While viral communities of the human oral cavity are just beginning to be characterized, there is considerably more known about viral communities that inhabit the human gut. Studies of human gut viromes have revealed the presence of a diverse and largely novel population of viruses (Breitbart et al., 2003; Reyes et al., 2010). In a study of four adult pairs of twins and their mothers, gut viral communites were not associated with the genetic relationship between individuals (Reyes et al., 2010). In a more recent study of gut viromes, six individuals were sampled at four time points over a relatively short 8-day period and demonstrated substantial individuality in their virome compositions (Minot et al., 2011). In another study, a healthy individual’s colonic microbiome was sampled 16 times over 2.4 years, demonstrating that much of the viral taxonomy of the gut was persistent (Minot et al., 2013).
The oral cavity is well known to harbor a high degree of bacterial diversity (Dewhirst et al., 2010; Human Microbiome Project Consortium, 2012) but also is known to be inhabited by a highly diverse community of viruses (Willner et al., 2009; Sedghizadeh et al., 2012; Pride et al., 2012a; Robles-Sikisaka et al., 2013; Zhang et al., 2013). Previous data have indicated that human saliva harbors numerous viruses, whose presence in the community over 30-day time periods is relatively distinct (Pride et al., 2012a). That study, however, did indicate the presence of a single virus, whose entire genome structure was identified at two separate time points, suggesting that viruses are capable of persisting despite the mechanisms that oral bacteria utilize to defend against their predation. One major mechanism used by oral bacteria to resist their viruses is via the use of CRISPRs (Clustered Regularly Interspaced Short Palindromic Repeats), which employ short viral sequences called spacers to interfere with viral replication in future encounters with viruses matching those spacer sequences (Barrangou et al., 2007). CRISPRs in the human oral cavity may be limited in their ability to eradicate viruses, as it has previously been demonstrated that many spacers co-exist with matching viruses over relatively long time periods in the mouth (Pride et al., 2012a) and other environments (Weinberger et al., 2012). These data suggest that bacteria may lack the capacity to completely eradicate their viruses from the oral microbiome and could contribute to a persistence of viruses in the human oral ecosystem as long as their cellular hosts also persist.
Specific host traits, environmental exposures and disease states have been shown to be associated with constituents of the human microbiome. For example, there are identifiable differences in the bacterial biota found in human subjects with inflammatory bowel disease (Craven et al., 2012). There also are detectable differences in the gut bacterial biota in response to diet changes and obesity (Ley et al., 2005, 2006; Turnbaugh et al., 2006), which indicate that the microbiota have a role in the response to host metabolic challenges. Viromes from rodent feces harbor insect and plant viruses (Phan et al., 2011), suggesting that diet has substantial effects on viral encounters. Whereas viral communities are less well studied in humans, there also is evidence that the gut virome may respond to dietary changes (Minot et al., 2011). Host genetic factors as well as environmental variables such as sex and hormonal fluctuations have also been linked to the human microbiome, as has been demonstrated by gender-specific differences in the bacterial biota of the gut (Mueller et al., 2006; Li et al., 2008) and the oral cavity (Slots et al., 1990; Umeda et al., 1998). We previously have demonstrated that oral virome membership is determined in part by host environmental viral exposures (Robles-Sikisaka et al., 2013), and we believe membership in the human oral virome is heavily influenced by both genetic and environmental factors. In this study, we intensively sampled the saliva of eight human subjects at 11 time points over a 60-day period to improve our understanding of the dynamics of human oral DNA viruses and potential host factors that might influence viral community membership.
Materials and methods
Human subjects
Subject recruitment and enrollment were approved by the University of California, San Diego, and the Western University Administrative Panels on Human Subjects in Medical Research. All eight subjects signed informed consent indicating their willingness to participate in this study. Each subject donated saliva over a 60-day period. Saliva was collected before breakfast or oral hygiene in the AM, before lunch for the Noon time point and before dinner for the PM time point. A minimum of 3 ml of unstimulated saliva was collected on days 1, 2, 4, 7, 14, 30 and 60, and was immediately frozen at −20 °C. Exclusion criteria included pre-existing medical conditions such as diabetes, organ transplantation or other such conditions in which treatment might result in significant immunosuppression. All subjects self-reported their health status. All subjects were unrelated, and none received any antibiotics either during the study period or for 3 months before the beginning of the study. Each subject was subjected to a full baseline periodontal examination consisting of measurements of probing depths, clinical attachment loss, Gingival Index, Plaque Index and gingival irritation (Loe, 1967), and was found to have healthy oral tissues and no periodontal disease (overall clinical attachment loss of <1 mm).
Preparation and sequencing of viromes
Saliva was filtered sequentially using 0.45 and 0.2 μ filters (VWR, Radnor, PA, USA) to remove cellular and other debris and was purified on a cesium chloride gradient according to previously described protocols (Pride et al., 2012a). Only the fraction with a density corresponding to most known bacteriophage (Murphy et al., 1995) was retained, further purified on Amicon YM-100 protein purification columns (Millipore Inc, Bellerica, MA, USA), treated with DNase I and subjected to lysis and DNA purification using the Qiagen UltraSens virus kit (Qiagen, Valencia, CA, USA). Resulting DNA was amplified using GenomiPhi V2 MDA amplification (GE Healthcare, Pittsburgh, PA, USA), fragmented to roughly 200–400 bp using a Bioruptor (Diagenode, Denville, NJ, USA), and utilized as input to create libraries using the Ion Plus Fragment Library Kit according to the manufacturer’s instructions. Libraries were then sequenced using 314 or 316 chips on an Ion Torrent Personal Genome Machine (PGM; Life Technologies, Grand Island, NY, USA; Rothberg et al., 2011) producing an average read length of ∼ 172 bp for each sample.
Analysis of viromes
Owing to the error rate of Semiconductor Sequencing (Rothberg et al., 2011), we trimmed each read according to modified Phred scores of 0.5 using CLC Genomics Workbench 4.65 (CLC bio USA, Cambridge, MA, USA), removed any low complexity reads (where >25% of the length were due to homopolymer tracts) and removed any reads with substantial length variation (<50 nucleotides or >300 nucleotides) or ambiguous characters before further analysis. Each virome was screened for contaminating bacterial and human nucleic acids using BLASTN analysis (E-score <10−5) against the Ribosomal Database Project 16S rRNA database (Cole et al., 2009) and the human reference database available at ftp://ftp.ncbi.nlm.nih.gov/genomes/H_sapiens/. Any reads homologous to human sequences were removed before further analysis. Length and GC content variation among contigs were assessed using Box and Whiskers plots created using Microsoft Excel 2007 (Microsoft Corp, Redman, WA, USA). Remaining reads were assembled using CLC Genomics Workbench 4.65 based on 98% identity with a minimum of 50% read overlap, which were more stringent than criteria developed to discriminate between highly related viruses (Breitbart et al., 2002). As the shortest reads were 50 nucleotides, the minimum tolerable overlap was 25 nucleotides, and the average overlap was no less than 86 nucleotides depending on the characteristics of each virome. The consensus sequence for each contig was constructed according to majority rule, and any contigs <200 nucleotides or with ambiguous characters were removed before further analysis. The contigs from each time point in each subject were then further assembled to construct larger contigs across time points using 98% identity over a minimum 50% contig overlap. Using these criteria, the minimum tolerable overlap was 100 nucleotides for each contig, and the average overlap was no less than 438 nucleotides depending on the characteristics of each subject. For each contig contributing to these assemblies, the time point and location within the assembly was recorded, and these data were utilized to determine the first and the last time point that contributed to each assembly. These data were then compiled by hour between time points to determine the mean time difference that contributed to each assembly. Comparisons of the mean percentages of assembled contigs were determined using Microsoft Excel 2007 and statistical significance was determined by two-tailed t-tests. We also utilized a separate technique for assembly by constructing global assemblies from all reads from all subjects and time points using 98% identity over a minimum of 50% read overlap. The contribution of each subject and time point to each assembly was assessed and utilized to construct heatmaps with Java Treeview (Saldanha, 2004).
The sequences of contig89 from subject no. 1 were generated with PCR amplification of overlapping fragments using Platinum PCR SuperMix (Invitrogen, Carlsbad, CA, USA) with primers specific for contig89 (Supplementary Table 4). Each resulting amplicon was sequenced in both directions using Sanger sequencing, and for those fragments larger than 1 kb, an internal primer was utilized to completely sequence the fragment. Contigs were assembled interactively based on their overlapping sequences using Sequencher (Gene Codes Corp, Ann Arbor, MI, USA). Viral contigs were analyzed using FGenesV (Softberry Inc, Mount Kisco, NY, USA) for open reading frame prediction, and individual open reading frames were analyzed using BLASTP analysis against the NCBI non-redundant database (E-score <10−5). If the best hit was to a gene with no known function, lower level hits were used for the annotation as long as they had known putative function and still met the E-score cutoff (10−5). Polymorphisms were identified based on the majority rule consensus sequence of all 11 time points combined and were determined using CLC Genomics Workbench 4.65.
Contigs were annotated using BLASTX against the NCBI NR database with an E-score cutoff value of 10−5. Specific viral homologs were determined by parsing BLASTX results for known viral genes including replication, structural, transposition, restriction/modification, hypothetical and other genes previously found in viruses for which the E-score was at least 10−5. Each individual virome contig was annotated using this technique (Figure 3); however, if the best hit for any portion of the contig was to a gene with no known function, lower level hits were used as long as they had known function and still met the E-score cutoff. The annotation data were compiled for each subject and used to determine the relative proportions of assembled contigs that contained viral homologs.
Analysis of shared homologs present in each virome was performed by creating custom BLAST databases for each virome, comparing each database with all other viromes using BLASTN analysis (E-score <10−10). Principal coordinate analysis (PCOA) was performed on homologous virome contigs with binary Sorensen distances using Qiime (Caporaso et al., 2010b). Heatmaps were generated based on shared homologs across all subjects and time points and depicted using JAVA Treeview (Saldanha, 2004). Heatmap data were normalized based on the total number of viral contigs for each virome.
Analysis of 16S rRNA
Genomic DNA was prepared from the saliva of each subject and the time point using the Qiagen QIAamp DNA MINI kit (Qiagen). Each sample was subjected to a bead beating step before nucleic acid extraction using Lysing Matrix-B (MP Bio, Santa Ana, CA, USA). We amplified the bacterial 16S rRNA V3 hypervariable region using the forward primer 341F (5′-CCTACGGGAGGCAGCAG-3′) fused with the Ion Torrent Adaptor A sequence and one of 23 unique 10-bp barcodes, and reverse primer 514R (5′-ATTACCGCGGCTGCTGG-3′) was fused with the Ion Torrent Adaptor P1 from the saliva of each subject and time point (Whiteley et al., 2012). PCR reactions were performed using Platinum PCR SuperMix (Invitrogen) with the following cycling parameters: 94 °C for 10 min, followed by 30 cycles of 94 °C for 30 s, 53 °C for 30 s, 72 °C for 30 s and a final elongation step of 72 °C for 10 min. Resulting amplicons were purified on a 2% agarose gel-stained with SYBR Safe (Invitrogen) using the MinElute PCR Purification kit (Qiagen). Amplicons were further purified with Ampure beads (Beckman-Coulter, Brea, CA, USA), and molar equivalents were determined for each sample using a Bioanalyzer 2100 HS DNA Kit (Agilent Technologies, Santa Clara, CA, USA). Samples were pooled into equimolar proportions and sequenced on 314 chips using an Ion Torrent PGM according to the manufacturer’s instructions (Life Technologies). Resulting sequence reads were removed from the analysis if they were <130 nt, had any barcode or primer errors, contained any ambiguous characters or contained any stretch of >6 homopolymers. Sequences were assigned to their respective samples based on their 10-nt barcode sequence and were analyzed further using the Qiime pipeline (Caporaso et al., 2010b). Briefly, representative operational taxonomic units from each set were chosen at a minimum sequence identity of 97% using UClust (Edgar, 2010) and were aligned using PyNast (Caporaso et al., 2010a) against the Greengenes database (DeSantis et al., 2006). Multiple alignments were then used to create phylogenies using FastTree (Li and Godzik, 2006), and taxonomy was assigned to each operational taxonomic unit using the RDP classifier (Wang et al., 2007; Price et al., 2009). PCOA was performed based on Beta Diversity using weighted Unifrac distances (Lozupone et al., 2006).
Statistical analysis
To assess whether viromes had significant overlap within or between subjects, we performed a permutation test based on resampling (10 000 iteration). We simulated the distribution of the fraction of shared virome homologs from two different time points within individual subjects that were randomly chosen across all time points. For each set, we computed the summed fraction of shared homologs using 1000 random contigs between randomly chosen individual time points within different subjects, and from these computed an empirical null distribution of our statistic of interest (the fraction of shared homologs). The simulated statistics within each subject across all time points were referred to the null distribution of intersubject comparisons, and the P-value was computed as the fraction of times the simulated statistic for each exceeded the observed statistic. An identical analysis was performed at the operational taxonomic unit level for the 16S rRNA taxonomic assignments. For analysis of sex-specific characteristics within the viromes, a randomly chosen subject and time point from the male sex was compared with a randomly chosen subject and time from the female sex to determine the null distribution of fraction of shared contigs based on opposite sexes. We then estimated the fraction of shared homologs from randomly chosen subjects and time points within each sex and compared with the empirical null distribution from simulated inter-sex values. For all comparisons of the sexes, intrasubject comparisons were excluded. We estimated the P-value based on the fraction of times the intra-sex statistic exceeded that for the observed statistic. This technique was also utilized to assess the relative contribution of individual subjects and time points to global assemblies constructed from the reads of all subjects and time points. We assessed whether any randomly selected contig had a higher proportion of intrasubject reads than intersubject reads recruited in its assembly.
Results
Isolation of viruses and screening for contamination
We recruited eight human subjects in good overall periodontal health (Table 1) and collected saliva from each at 11 time points over a 60-day period for a total of 88 samples. We collected saliva at multiple time points on some days and only at single time points on other days as follows: Day 1 (AM, Noon, PM), Day 2 (AM only), Day 4 (AM only), Day 7 (AM and Noon), Day 14 (AM only), Day 30 (AM and Noon) and Day 60 (AM only). Our goals were to characterize oral viruses over relatively short and long time periods. Saliva was collected from each subject before meals, and for all AM time points saliva was collected before any oral hygiene practices.
We isolated viruses from all 88 samples according to our previously described protocols (Pride et al., 2012a), which involved sequential filtering to remove cellular debris, CsCl density gradient ultracentrifugation and DNA extraction from intact virions. Resulting DNA was sequenced using Semiconductor Sequencing (Rothberg et al., 2011) for a total of 38 430 905 reads after processing with a mean length of 172 nucleotides. We sequenced an average of 4 803 863 reads per subject and an average of 436 715 reads per individual time point (Supplementary Table 1). All viromes were screened for potential contaminating cellular elements by BLASTN analysis (E-score <10−5) against the NCBI Reference Human Genome Database (https://www.ncbi.nlm.nih.gov/genome/guide/human/) and the Ribosomal Database Project 16S rRNA Database (http://rdp.cme.msu.edu/). No virome had >0.5% of its reads homologous to the human reference database, and all homologous sequences were removed before further analysis. Only 3/88 (3.4%) viromes tested had any identifiable homologs to 16S rRNA, and two of those three viromes only had single 16S rRNA homologs (Supplementary Table 1). These data indicate that our collection of viromes was relatively free of contaminating cellular nucleic acids.
Identification of viruses
We assembled virome reads from all subjects and time points to construct longer contigs, as this generally results in more productive searches for homologous sequences. The mean contig length across all subjects was 1031±168 nucleotides (range from 548 to 1464 nucleotides) and the median contig length was 493±25 (Supplementary Figure 1). There was similar mean GC content (46±2%; range from 43 to 52%) and the median GC content (45±2%) among the contigs in all viromes (Supplementary Figure 2). We identified homologs to each contig by BLASTX analysis against the NCBI Non-redundant database (http://www.ncbi.nlm.nih.gov/refseq/; Supplementary Figure 3). Similar to results found for viromes from different environments (Desnues et al., 2008; Minot et al., 2011; Roux et al., 2012; Wommack et al., 2012; Minot et al., 2013; Yoshida et al., 2013), the majority of the contigs had no known viral homologues. Approximately 32.7±5.9% (range from 20.3 to 48.0%) of the virome contigs were homologous to known viruses, depending on the subject and time point analyzed (Supplementary Figure 3).
The vast majority of the assembled contigs in our cohort were homologous to bacteriophage. Many of the contigs (40.7±8.0%; range from 23 to 59%) from each subject and time point had multiple homologs along their length to different viral genes (Figure 1 and Supplementary Figure 4). We found homologs with a broad array of functions, including those involved in virus structure, virulence and replication (Supplementary Figure 5). For example, all time points evaluated for Subject no. 1 contained numerous contigs homologous to viral structural components such as head, capsid, collar and tail, virulence components such as toxins and restriction/modification enzymes and replication components such as DNA polymerases and helicases (Supplementary Figure 5). The most common identifiable viral homologs identified in all subjects were hypothetical phage genes (26.2±1.4% of the contigs), DNA polymerases (6.9±0.3%), tail fibers (6.5±0.5%), helicases (6.4±0.2%) and integrases/transposases (6.2±0.3%; Figure 1). The relatively high number of beta lactamases (present in 1.9±0.2% of the contigs) suggests that many oral viruses carry beta lactamases. Only eight of the 1084 (0.7%) beta lactamase homologs identified were homologous to plasmid beta lactamases (Supplementary Figure 6, Panel A). TEM beta lactamases (Cooksey et al., 1990) are often found on plasmids, and only represent 9.8% (97 of 1084) of the beta lactamase homologs identified in our data (Supplementary Figure 6, Panel B). The presence of multiple identifiable viral homologs in many of the assembled contigs (Figure 1 and Supplementary Figure 4) provides strong support that each subject and time point studied were highly enriched for viruses.
Viruses persist in the human oral cavity over time
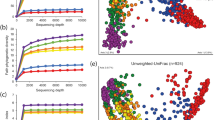
We examined the viromes from each subject and time point to determine whether specific viral genotypes persisted over time. As the analysis of viromes generally is limited by the ability to assemble reads, we first assembled reads into contigs from each subject and time point using highly stringent criteria of 98% identity over a minimum of 50% of each read (mean overlap of 86.0±2.4 nucleotides depending on the characteristics of each individual virome). We next took the contigs from each individual time point and assembled them across all time points using the same highly stringent criteria to construct larger viral contigs and determine whether any were shared within each subject over time. Utilizing this technique, we found that only 38.7±5.0% of the contigs were unique to any single time point, whereas 61.3±5.4% of the contigs were shared in two or more time points (P<0.001; Supplementary Figure 7). There was no significant difference in the percentage of contigs present in a single time point and those shared by only two time points (38.7±5.0% versus 37.4±5.0%; P=0.627), indicating that these viral contigs were just as likely to be found longitudinally as they were to be identified at any single point in time (Supplementary Figure 7). Over 4.8% of the contigs were present at over half the time points studied (⩾6 time points), indicating that a significant proportion of the viruses persisted throughout the study period. We also utilized a second assembly technique to assess whether human oral viruses were persistent members of the human oral microbiome. We created global assemblies of all reads in each subject and then measured the recruitment of reads to each contig. We found that for those contigs assembled from all time points, the relative contribution from each time point was relatively even (Figure 2, Panel b and Supplementary Figure 8), further suggesting that the same viruses were present longitudinally in each subject.
We also examined the relative time intervals in which viral contigs could be assembled from all time points in each subject to determine whether there were identifiable time intervals in which oral viruses likely persist. In all individual subjects, the majority of the viruses (33.6%±5.9%) were found to persist for 14–30 days (Figure 2, Panel a). Interestingly, 16%±4.2% of the contigs were found across the entire 60-day study period. A relative minority of the contigs persisted for 12 h or less (3.3%±1.0%). Over half of the assembled viral contigs included time points that were 7 or more days apart (69.9%±5.5% versus 30.1%±5.5%; P<0.001), highlighting the persistent nature of human oral viruses.
We examined the annotation and composition of select viral contigs to determine the relative contribution of each time point to its structure and to identify homologous sequences. Most of the structure of Subject no. 1 contig89 could be identified from four separate fragments identified on Day 4 AM. Contigs from Day 2 AM and Day 14 AM overlapped with these fragments and allowed for the full reconstruction of this virus (Figure 3, Panel a). Similar viral assemblies were assembled for all subjects (Figure 3, Panels b–h), demonstrating the persistence of viruses over the 60-day study in each subject. To identify viral homologs, we annotated the contigs based on BLASTX homology across the length of each contig. Subject no. 1 Contig89 had homologs putatively involved in virus structure, replication and virulence functions, as did contigs from all other subjects (Figure 3, Panels b–h).
Assemblies of contigs from all time points in each subject. (a) Viral contig from subject no. 1, (b) subject no. 2, (c) subject no. 3, (d) subject no. 4, (e) subject no. 5, (f) subject no. 6, (g) subject no. 7 and (h) subject no. 8. The portions of the contigs identified at each time point are represented by the colored boxes for each subject. The relative coverage of each contig is represented, along with annotated open reading frames (ORFs) above each diagram. The length of each contig is denoted at the top of each panel.
To confirm that the technique of assembling viral contigs from different time points into a single viral contig was capable of reconstructing true viruses, we verified the structure of Subject no. 1 contig89. Utilizing PCR and Sanger sequencing, we amplified and sequenced overlapping fragments of contig89 from all 11 time points. We found that the same virus was present at all time points, with few polymorphisms (19 total polymorphisms in all contigs combined for a mean of 1.73 polymorphisms per time point) over the course of the 60-day study (Figure 4). Interestingly, 53% (10 of 19) of those polymorphisms reverted to wild type (Day 1 AM) during the course of the study. Despite not identifying smaller contigs that assembled into contig89 on Day 60 (Figure 3, Panel a), we were able to verify that the full contig was present at this time point (Figure 4). We found many putative open reading frames in contig89 that had synteny with Rhodococcus phage Pepy6 and Poco6 (Summer et al., 2011), which are siphoviruses similar to others found in Lactococcus and Clostridium. These data verify that viral genotypes in the human oral cavity are highly persistent, and that some may persist for at least 60 days.
Diagram of contig89 assembled from Sanger sequences from all 11 time points in subject no. 1. Putative ORFs and their direction are indicated by the arrows at the top of the diagram. ORFs that had significant homologs (BLASTP E-score <10−5) are indicated by the text above each arrow. Those ORFs with synteny to Rhodococcus phage Pepy6 and Poco6 (Summer et al., 2011) are labeled gp054, gp065, gp067, gp068, gp069, gp071, gp072, gp105 and gp106. The location of polymorphisms (when compared with a majority rule consensus from all time points combined) are indicated by orange vertical lines.
We also used the contig89 sequences to estimate the error rate of the Semiconductor Sequencing by comparing the Sanger sequences with the semiconductor reads from each time point. We found that the mean and median error rates were 0.95% and 0.74%, respectively, consistent with those previously described (Rothberg et al., 2011).
Inter- and Intrasubject comparisons of viral genotypes
We compared viral contigs within individuals over time and among subjects over time. As revealed by heatmap analysis, we found that there were numerous viral genotypes conserved in all individuals across the entire study, suggesting that there were similar characteristics in viral communities from these healthy human subjects (Figure 5). The analysis also revealed that there was likely an even greater proportion of shared intrasubject viral genotypes than there were for intersubject genotypes, giving the heatmap the appearance of a matrix (Figure 5). We quantified the proportion of shared viral contigs and found that there was a substantial number of shared intersubject contigs but that they did not approach the high proportion of intrasubject-shared contigs over time (50.6%±6.9% shared within an individual versus 24.3%±3.3% shared between individuals; P<0.001). We also created global assemblies made from the reads of all subjects and time points, and found that the recruitment of reads to each contig also demonstrated some subject specificity (Supplementary Figure 9).
Heatmap of shared contigs across all subjects and time points. The heatmap is organized by subject and time point, where the subject number is denoted along each axis, and the individual time points are denoted by each column consecutively from Day 1 AM to Day 60 AM left to right for each subject. Each row represents an individual contig, and rows are ordered consecutively across each subject from contigs identified on Day 1 AM to Day 60 AM. The ‘matrix-like’ appearance of the heatmap is due to the high intensity of homologous sequences across all time points within each subject.
Because of the observed patterns of shared viral contigs within individuals over time (Figure 5), we tested whether homologous viral contigs were significantly subject-specific. PCOA based on beta diversity between viromes suggested that there were patterns of variations present in most viromes that were reflective of their host environment (Figure 6, Panel a). For most time points within subject numbers 1, 5, 6, 7 and 8, viromes were distinctive based on PCOA. We next utilized a permutation test to determine whether the patterns of shared homologs within each subject over time were significantly associated with their host subject. Briefly, we tested whether the fraction of shared homologs for intrasubject comparisons was greater than that for inter-subject comparisons. We performed this test by randomly sampling 1000 contigs from each subject over 10 000 iterations. While the proportions of shared homologs were only significant for subject numbers 1, 2 and 6 (P⩽0.05), the estimated fraction of intrasubject shared homologues was far greater than the fraction of intersubject shared homologues in all subjects (Table 2). Similar trends were observed when we performed a similar analysis on reads recruited to global assemblies from all subjects and time points (Supplementary Table 2). These data suggest that despite numerous shared viral genotypes among subjects, a portion of the human oral viral community is individual-specific.
Viral genotypes associated with host sex
PCOA analysis was suggestive of an association between shared oral viruses and the sex of each subject (Figure 6, Panel b). To determine whether human oral viruses were significantly associated with the sex of each subject, we performed a permutation test to determine whether the fraction of shared homologs within each sex was greater than the fraction between sexes. We treated each of the 88 viromes independently, with the exception that all intrasubject comparisons were excluded from the analysis. There was a significantly greater proportion of shared homologs within each sex (31.1±7.6 for males, 34.3±7.4 for females) when compared with the proportion between sexes (13.46±4.2 for males versus females). These data were significant for the virome contents in both males (P=0.0121) and females (P=0.0034) (Table 2 and Supplementary Table 2). Differences between the sexes in bacterial communities previously have been demonstrated for the human gut microbiome (Mueller et al., 2006; Li et al., 2008) and the subgingival crevice (Schenkein et al., 1993; Lay et al., 2005). The data presented here (Table 2) suggest that sex-based differences observed in bacteria also apply to communities of human viruses.
Characterization of bacterial communities
We found that human oral viruses in our cohort of subjects were persistent (Figure 3), largely subject-specific (Figure 6, Panel a) and were associated with the gender of each subject (Table 2). We also examined whether similar trends were present for human oral bacterial communities. Through examination of the 16S rRNA V3-hypervariable, we tested whether there was evidence of subject- or gender specificity among the bacteria in the oral microbiome. We sequenced 1 331 068 16S rRNA reads for an average of 166 384 reads per subject and 15 126 per time point (Supplementary Table 3). Much of the diversity present in the saliva of each subject was adequately sampled for each subject and time point as measured by rarefaction analysis (Supplementary Figure 10). At the phylum level, Firmicutes predominated across most subjects and time points (Supplementary Figures 11 and 12), with Bacteroidetes and Actinobacteria also accounting for a significant proportion of the bacterial community in each subject. Similar distributions of taxa abundance were produced for each subject over time, with few time points having significant variations in taxonomic relative abundances. Proteobacteria generally did not account for substantial proportions of the oral microbiome, except in Subject numbers 2 and 8 (Supplementary Figures 11 and 12).
To determine whether there were subject-specific patterns of variation present in the oral bacterial biota of each subject over time, we performed a PCOA analysis. There were no strongly observed patterns of subject specificity or gender specificity observed through 16S rRNA in our study subjects (Figure 7a and b). This was confirmed with a permutation test, which showed no significant association within or among subjects and their observed bacterial biota (P>0.05 for all subjects) (Supplementary Table 4). There also was no significant association between the bacterial biota and the sex of each subject (75.1±13.6 of the biota were shared for males, 74.9±11.6 for females, and 70.9±14.5 for males versus females; P=0.282 for males and P=0.353 for females). These data indicate that viral metagenomics provided greater resolution in identifying certain subject or subject-group-specific traits than the analysis of 16S rRNA.
Discussion
We previously demonstrated that human oral viral communities are populated by numerous viral genotypes and carry substantial gene function putatively involved in host pathogenic functions (Pride et al., 2012a). The analysis presented here has extended those findings and provided a robust data set of 88 individual viromes from eight human subjects at 11 different time points over a 60-day period. We believe that the longitudinal analysis presented here is optimal for discerning differences in viral communities between subjects or subject groups, as any individual time point may differ; however, the trends in these viral communities over time are clear (Figure 6). Our findings of a large number of viral genotypes that persist over time are contrary to that presented in our prior study (Pride et al., 2012a); however, they highlight several difficulties encountered when examining large and complex metagenome data sets. These difficulties include the following: (1) assembly related overestimation of viral genotypes, as we likely observed in our previous study (Pride et al., 2012a, 2) lack of available homologs to aid in the identification of virome constituents (Desnues et al., 2008; Minot et al., 2011; Roux et al., 2012; Wommack et al., 2012; Minot et al., 2013; Yoshida et al., 2013) and (3) the relative lack of statistical tools to discern differences in virome compositions. We did not perform detailed comparisons of viral genotypes and the bacteria present in each subject because our BLAST-based techniques for characterizing viruses generally are considered insufficient to accurately predict the hosts of each virus.
There were some shortcomings to our methodologies, including that our techniques were specific for DNA viruses, and that each virome was subjected to MDA amplification due to the small amounts of viral DNA recovered from each subject and time point. As the human oral cavity may also be home to a community of RNA viruses, it was not possible for us to decipher what proportion of human oral viruses were characterized in this study. Whereas MDA amplification is known to introduce biases into sequence data (Kim and Bae, 2011), it is unclear how MDA amplification biases could have affected these viromes. Many of the biases introduced often result when relatively low levels of starting DNA is utilized for amplification, which could have occurred in some of the viromes in this study. Our use of 11 different time points, many of which show relatively even coverage of the contigs present among multiple time points (Figure 2, Panel b), suggests that MDA amplification biases were unlikely to affect our estimates of the persistence of oral viruses in this study; however, if MDA biases were to affect our estimates of persistence, they would likely result in underestimates rather than overestimates. We specifically chose to characterize persistence of viruses rather than to focus on differences in virome composition between same day time points because heterogeneous MDA amplification could potentially explain any observed differences.
One of the primary goals of this study was to identify whether specific oral viruses were merely transient members of the oral microbiome. In our prior analyses of human viromes (Pride et al., 2012a, 2012b), we noted that many of the viral contigs were homologous to the same viruses. We now believe that those homologies reflect shortcomings in the assembly process rather than denoting the presence of distinct viral genotypes; those shortcomings likely resulted in an overestimation of the number of distinct viral genotypes observed in that study (Pride et al., 2012a, 2012b). The fact that many contigs at single time points would assemble together only after including a broader set of contigs from other time points suggested that the assembly process was a limitation in our prior analyses of human oral viromes, and strongly suggested that many of these viruses were persistent members of the oral microbiome. Whereas we used two separate methods for assembly including assembling all reads from each individual subject (Figure 5 and Table 2) and globally assembling reads from all subjects combined (Supplementary Figure 9 and Supplementary Table 2), each method produced similar results. We preferred the former method because the initial assembly of reads from a single time point might reduce the potential for chimerism when compared with the global assembly of reads from all disparate subjects and time points.
We verified that our assembly process was capable of reconstructing viruses by reproducing the 23.5-kb sequence of the assembled contig89 from each of the 11 separate time points in Subject no. 1 using traditional PCR and Sanger sequencing techniques (Figure 4). We were able to estimate that the majority of the persistent viral genotypes in all subjects were present for at least 14–30 days, and a significantly larger proportion of the viruses persisted for 7 days or longer (Figure 2, Panel a). In fact, viral genotypes were just as likely to persist over multiple time points as they were to be transiently present at a single time point (Supplementary Figure 7). We believed when we began this study that very few viral genotypes would be observed over the entire 60-day study period, and the 16%±4.2% of genotypes that persisted over this period suggests that genotypes might persist even longer.
We previously demonstrated that viral exposures as measured through CRISPR spacers were highly subject-specific (Pride et al., 2011), but little prior evidence supported that membership in the oral virome was individual-specific. Through examination of each individual at 11 time points over a 60-day period, we were able to gain a more robust picture of the virome constituents in each subject, and also were able to utilize the time points collectively to construct larger viruses (Figure 3). By measuring both relatively short (12 h or less) and relatively long time periods (days to weeks/months), we determined that many of the viruses appeared to persist over the 60-day study period (Figure 2). We did not concentrate on differences in virome content in the AM and PM time points from the same days, as greater sequencing depth than was achieved in this study would be necessary to explore those differences robustly. While there was a small proportion of the virome that were homologous between different subjects, there was a clear trend of subject specificity in all subjects (Table 2). We utilized statistical techniques (Robles-Sikisaka et al., 2013) to determine whether there were significant trends in the virome data to augment the PCOA analysis of viromes (Figure 6), as the differences visualized by PCOA only accounted for a small percentage of the variation present. Prior studies suggested that viral exposures were living environment-specific (Pride et al., 2012b; Robles-Sikisaka et al., 2013), whereas viral community membership observed in this study suggests that many viral genotypes are subject-specific. These results together suggest that the human oral viral community is formed as a result of each individual’s unique viral exposures. Other factors such as diet, age, oral health status and medical conditions might also affect oral virome composition.
Our analysis of oral viromes provides the first evidence of sex-specific differences among human viral communities but might have been predicted by gender-specific differences seen in prior studies examining bacterial biota (Kovacs et al., 2011; Gomez et al., 2012; Markle et al., 2013). Sex-specific differences previously have been observed in the bacterial biota of human hand surfaces (Staudinger et al., 2011), the forearm (Fierer et al., 2008), the gut (Mueller et al., 2006; Li et al., 2008) and the subgingival crevice (Slots et al., 1990; Schenkein et al., 1993; Umeda et al., 1998; Kumar 2013). Oral bacteria have been observed to change with periods of major hormonal fluctuations, such as puberty or pregnancy, and the hormonal changes may have a causative role in this observation of diversion of microorganisms by sex (Kumar, 2013). In this study, variance among viromes was explained by differences in sex of the host more so than variance among individuals (Table 2). Although there were only eight subjects in this study, the multitude of time points sampled allowed for comparisons among 77 different viromes (88 minus 11 viromes that would represent intrasubject comparisons), providing a robust data set for discerning differences according to gender. The P-values for both males and females were highly significant (Table 2) and were strongly suggestive of an association between host sex and virome composition.
The oral microbiome has been hypothesized to have a role in the development of periodontal disease (Huang et al., 2011; Ge et al., 2013; Wade, 2013); however, the role of viruses as members of the oral microbiome has not been elucidated. Our findings that human oral viruses are personalized features of each individual’s microbiome and that viruses are persistent members of the oral microbiome suggests that viruses have important roles in the oral ecosystem. Viruses have the potential to eradicate certain cellular hosts or to provide them with beneficial gene functions such as beta lactamases (Figure 1); thus, human oral viruses may help to shape the natural history of oral microbiome membership. The sex specificity we identified in oral viruses suggests that viral communities respond to the same potentially hormone-driven signals that govern the sex specificity also found in bacterial communities on the skin (Fierer et al., 2008; Staudinger et al., 2011), gut (Mueller et al., 2006; Li et al., 2008) and the oral cavity (Slots et al., 1990; Umeda et al., 1998). Understanding the impact of environment and genetics on human microbiota may be critical to the evaluation of disease processes such as chronic periodontitis. While human viral communities have largely been considered a biological dark matter due to a relative dearth of knowledge regarding their ecology, our data have identified a personal, persistent and gender-specific nature to human oral viruses that suggests an important interplay between host genetics and environment, which ultimately may be important for human health and disease.
References
Human Microbiome Project Consortium (2012). Structure, function and diversity of the healthy human microbiome. Nature 486: 207–214.
Allen HK, Looft T, Bayles DO, Humphrey S, Levine UY, Alt D et al (2011). Antibiotics in feed induce prophages in swine fecal microbiomes. MBio 2: e00260–11.
Barrangou R, Fremaux C, Deveau H, Richards M, Boyaval P, Moineau S et al (2007). CRISPR provides acquired resistance against viruses in prokaryotes. Science 315: 1709–1712.
Breitbart M, Salamon P, Andresen B, Mahaffy JM, Segall AM, Mead D et al (2002). Genomic analysis of uncultured marine viral communities. Proc Natl Acad Sci USA 99: 14250–14255.
Breitbart M, Hewson I, Felts B, Mahaffy JM, Nulton J, Salamon P et al (2003). Metagenomic analyses of an uncultured viral community from human feces. J Bacteriol 185: 6220–6223.
Caporaso JG, Bittinger K, Bushman FD, DeSantis TZ, Andersen GL, Knight R . (2010a). PyNAST: a flexible tool for aligning sequences to a template alignment. Bioinformatics 26: 266–267.
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK et al (2010b). QIIME allows analysis of high-throughput community sequencing data. Nat Method 7: 335–336.
Cole JR, Wang Q, Cardenas E, Fish J, Chai B, Farris RJ et al (2009). The Ribosomal Database Project: improved alignments and new tools for rRNA analysis. Nucleic Acids Res 37: D141–D145.
Cooksey R, Swenson J, Clark N, Gay E, Thornsberry C . (1990). Patterns and mechanisms of beta-lactam resistance among isolates of Escherichia coli from hospitals in the United States. Antimicrob Agents Chemother 34: 739–745.
Craven M, Egan CE, Dowd SE, McDonough SP, Dogan B, Denkers EY et al (2012). Inflammation drives dysbiosis and bacterial invasion in murine models of ileal Crohn's disease. PLoS One 7: e41594.
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K et al (2006). Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72: 5069–5072.
Desnues C, Rodriguez-Brito B, Rayhawk S, Kelley S, Tran T, Haynes M et al (2008). Biodiversity and biogeography of phages in modern stromatolites and thrombolites. Nature 452: 340–343.
Dewhirst FE, Chen T, Izard J, Paster BJ, Tanner AC, Yu WH et al (2010). The human oral microbiome. J Bacteriol 192: 5002–5017.
Duerkop BA, Clements CV, Rollins D, Rodrigues JL, Hooper LV . (2012). A composite bacteriophage alters colonization by an intestinal commensal bacterium. Proc Natl Acad Sci USA 109: 17621–17626.
Edgar RC . (2010). Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26: 2460–2461.
Fierer N, Hamady M, Lauber CL, Knight R . (2008). The influence of sex, handedness, and washing on the diversity of hand surface bacteria. Proc Natl Acad Sci USA 105: 17994–17999.
Ge X, Rodriguez R, Trinh M, Gunsolley J, Xu P . (2013). Oral microbiome of deep and shallow dental pockets in chronic periodontitis. PLoS One 8: e65520.
Gomez A, Luckey D, Yeoman CJ, Marietta EV, Berg Miller ME, Murray JA et al (2012). Loss of sex and age driven differences in the gut microbiome characterize arthritis-susceptible 0401 mice but not arthritis-resistant 0402 mice. PLoS One 7: e36095.
Huang S, Yang F, Zeng X, Chen J, Li R, Wen T et al (2011). Preliminary characterization of the oral microbiota of Chinese adults with and without gingivitis. BMC Oral Health 11: 33.
Kim KH, Bae JW . (2011). Amplification methods bias metagenomic libraries of uncultured single-stranded and double-stranded DNA viruses. Appl Environ Microbiol 77: 7663–7668.
Kovacs A, Ben-Jacob N, Tayem H, Halperin E, Iraqi FA, Gophna U . (2011). Genotype is a stronger determinant than sex of the mouse gut microbiota. Microbiol Ecol 61: 423–428.
Kumar PS . (2013). Sex and the subgingival microbiome: do female sex steroids affect periodontal bacteria? Periodontol 2000 61: 103–124.
Lay C, Rigottier-Gois L, Holmstrom K, Rajilic M, Vaughan EE, de Vos WM et al (2005). Colonic microbiota signatures across five northern European countries. Appl Environ Microbiol 71: 4153–4155.
Ley RE, Backhed F, Turnbaugh P, Lozupone CA, Knight RD, Gordon JI . (2005). Obesity alters gut microbial ecology. Proc Natl Acad Sci USA 102: 11070–11075.
Ley RE, Turnbaugh PJ, Klein S, Gordon JI . (2006). Microbial ecology: human gut microbes associated with obesity. Nature 444: 1022–1023.
Li M, Wang B, Zhang M, Rantalainen M, Wang S, Zhou H et al (2008). Symbiotic gut microbes modulate human metabolic phenotypes. Proc Natl Acad Sci USA 105: 2117–2122.
Li W, Godzik A . (2006). Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22: 1658–1659.
Loe H . (1967). The Gingival Index, the Plaque Index and the Retention Index Systems. J Periodontol 38 (Suppl): 610–616.
Lozupone C, Hamady M, Knight R . (2006). UniFrac–an online tool for comparing microbial community diversity in a phylogenetic context. BMC Bioinformatics 7: 371.
Markle JG, Frank DN, Mortin-Toth S, Robertson CE, Feazel LM, Rolle-Kampczyk U et al (2013). Sex differences in the gut microbiome drive hormone-dependent regulation of autoimmunity. Science 339: 1084–1088.
Minot S, Sinha R, Chen J, Li H, Keilbaugh SA, Wu GD et al (2011). The human gut virome: inter-individual variation and dynamic response to diet. Genome Res 21: 1616–1625.
Minot S, Bryson A, Chehoud C, Wu GD, Lewis JD, Bushman FD . (2013). Rapid evolution of the human gut virome. Proc Natl Acad Sci USA 110: 12450–12455.
Mueller S, Saunier K, Hanisch C, Norin E, Alm L, Midtvedt T et al (2006). Differences in fecal microbiota in different European study populations in relation to age, gender, and country: a cross-sectional study. Appl Environ Microb 72: 1027–1033.
Murphy FA, Fauquet CM, Bishop. DHL, Ghabrial SA, Jarvis AW, Martelli GP et al (1995) Virus Taxonomy: Sizth Report of the International Committee on Taxonomy of Viruses Vol. Supplement 10. Springer-Verlag: New York, NY, USA.
Parada V, Baudoux AC, Sintes E, Weinbauer MG, Herndl GJ . (2008). Dynamics and diversity of newly produced virioplankton in the North Sea. ISME J 2: 924–936.
Paulsen IT, Banerjei L, Myers GS, Nelson KE, Seshadri R, Read TD et al (2003). Role of mobile DNA in the evolution of vancomycin-resistant Enterococcus faecalis. Science 299: 2071–2074.
Phan TG, Kapusinszky B, Wang C, Rose RK, Lipton HL, Delwart EL . (2011). The fecal viral flora of wild rodents. PLoS Pathog 7: e1002218.
Price MN, Dehal PS, Arkin AP . (2009). FastTree: computing large minimum evolution trees with profiles instead of a distance matrix. Mol Biol Evol 26: 1641–1650.
Pride DT, Sun CL, Salzman J, Rao N, Loomer P, Armitage GC et al (2011). Analysis of streptococcal CRISPRs from human saliva reveals substantial sequence diversity within and between subjects over time. Genome Res 21: 126–136.
Pride DT, Salzman J, Haynes M, Rohwer F, Davis-Long C, White RA 3rd et al (2012a). Evidence of a robust resident bacteriophage population revealed through analysis of the human salivary virome. ISME J 6: 915–926.
Pride DT, Salzman J, Relman DA . (2012b). Comparisons of clustered regularly interspaced short palindromic repeats and viromes in human saliva reveal bacterial adaptations to salivary viruses. Environ Microbiol 14: 2564–2576.
Reyes A, Haynes M, Hanson N, Angly FE, Heath AC, Rohwer F et al (2010). Viruses in the faecal microbiota of monozygotic twins and their mothers. Nature 466: 334–338.
Robles-Sikisaka R, Ly M, Boehm T, Naidu M, Salzman J, Pride DT . (2013). Association between living environment and human oral viral ecology. ISME J 7: 1710–1724.
Rodriguez-Brito B, Li L, Wegley L, Furlan M, Angly F, Breitbart M et al (2010). Viral and microbial community dynamics in four aquatic environments. ISME J 4: 739–751.
Rodriguez-Valera F, Martin-Cuadrado AB, Rodriguez-Brito B, Pasic L, Thingstad TF, Rohwer F et al (2009). Explaining microbial population genomics through phage predation. Nat Rev Microbiol 7: 828–836.
Rothberg JM, Hinz W, Rearick TM, Schultz J, Mileski W, Davey M et al (2011). An integrated semiconductor device enabling non-optical genome sequencing. Nature 475: 348–352.
Roux S, Enault F, Robin A, Ravet V, Personnic S, Theil S et al (2012). Assessing the diversity and specificity of two freshwater viral communities through metagenomics. PLoS One 7: e33641.
Saldanha AJ . (2004). Java Treeview–extensible visualization of microarray data. Bioinformatics 20: 3246–3248.
Sandaa RA, Gomez-Consarnau L, Pinhassi J, Riemann L, Malits A, Weinbauer MG et al (2009). Viral control of bacterial biodiversity–evidence from a nutrient-enriched marine mesocosm experiment. Environ Microbiol 11: 2585–2597.
Schenkein HA, Burmeister JA, Koertge TE, Brooks CN, Best AM, Moore LV et al (1993). The influence of race and gender on periodontal microflora. J Periodontol 64: 292–296.
Sedghizadeh PP, Yooseph S, Fadrosh DW, Zeigler-Allen L, Thiagarajan M, Salek H et al (2012). Metagenomic investigation of microbes and viruses in patients with jaw osteonecrosis associated with bisphosphonate therapy. Oral Surg Oral Med Oral Pathol Oral Radiol 114: 764–770.
Slots J, Feik D, Rams TE . (1990). Age and sex relationships of superinfecting microorganisms in periodontitis patients. Oral Microbiol Immunol 5: 305–308.
Staudinger T, Pipal A, Redl B . (2011). Molecular analysis of the prevalent microbiota of human male and female forehead skin compared to forearm skin and the influence of make-up. J Appl Microbiol 110: 1381–1389.
Summer EJ, Liu M, Gill JJ, Grant M, Chan-Cortes TN, Ferguson L et al (2011). Genomic and functional analyses of Rhodococcus equi phages ReqiPepy6, ReqiPoco6, ReqiPine5, and ReqiDocB7. Appl Environ Microbiol 77: 669–683.
Turnbaugh PJ, Ley RE, Mahowald MA, Magrini V, Mardis ER, Gordon JI . (2006). An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444: 1027–1031.
Umeda M, Chen C, Bakker I, Contreras A, Morrison JL, Slots J . (1998). Risk indicators for harboring periodontal pathogens. J Periodontol 69: 1111–1118.
Wade WG . (2013). The oral microbiome in health and disease. Pharmacol Res 69: 137–143.
Wang Q, Garrity GM, Tiedje JM, Cole JR . (2007). Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73: 5261–5267.
Weinberger AD, Sun CL, Plucinski MM, Denef VJ, Thomas BC, Horvath P et al (2012). Persisting viral sequences shape microbial CRISPR-based immunity. PLoS Comput Biol 8: e1002475.
Whiteley AS, Jenkins S, Waite I, Kresoje N, Payne H, Mullan B et al (2012). Microbial 16S rRNA Ion Tag and community metagenome sequencing using the Ion Torrent (PGM) Platform. J Microbiol Methods 91: 80–88.
Willner D, Furlan M, Haynes M, Schmieder R, Angly FE, Silva J et al (2009). Metagenomic analysis of respiratory tract DNA viral communities in cystic fibrosis and non-cystic fibrosis individuals. PLoS One 4: e7370.
Willner D, Furlan M, Schmieder R, Grasis JA, Pride DT, Relman DA et al (2011). Metagenomic detection of phage-encoded platelet-binding factors in the human oral cavity. Proc Natl Acad Sci USA 108 (Suppl 1): 4547–4553.
Wommack KE, Bhavsar J, Polson SW, Chen J, Dumas M, Srinivasiah S et al (2012). VIROME: a standard operating procedure for analysis of viral metagenome sequences. Stand Genomic Sci 6: 421–433.
Yoshida M, Takaki Y, Eitoku M, Nunoura T, Takai K . (2013). Metagenomic analysis of viral communities in (hado)pelagic sediments. PLoS One 8: e57271.
Zhang Q, Rho M, Tang H, Doak TG, Ye Y . (2013). CRISPR-Cas systems target a diverse collection of invasive mobile genetic elements in human microbiomes. Genome Biol 14: R40.
Acknowledgements
This study was supported by the Robert Wood Johnson Foundation, the Burroughs Wellcome Fund and NIH 1K08AI085028 to DTP.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
All sequences including viromes and 16S rRNA are available for download in the MG-RAST database (metagenomics.anl.gov/) under the project ‘Saliva Virome Dynamics Study,’ and under consecutive individual accession numbers 4525509.3 to 4525684.3. Sequences also are available in the Short Read Archive under accession number SRP033575 . Sequences of contig89 are available in Genbank under consecutive accession numbers KF594184 to KF594194.
Author Contributions
DTP conceived and designed experiments. RR-S, SRA, ML and AGL performed the experiments. DTP, SRA and AGL analyzed the data. JS contributed statistical tools. TKB contributed reagents and performed examinations. DTP and SRA wrote the manuscript.
Supplementary Information accompanies this paper on The ISME Journal website
Rights and permissions
About this article
Cite this article
Abeles, S., Robles-Sikisaka, R., Ly, M. et al. Human oral viruses are personal, persistent and gender-consistent. ISME J 8, 1753–1767 (2014). https://doi.org/10.1038/ismej.2014.31
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2014.31
Keywords
This article is cited by
-
Toothbrush microbiomes feature a meeting ground for human oral and environmental microbiota
Microbiome (2021)
-
Characterization of oral virome and microbiome revealed distinctive microbiome disruptions in paediatric patients with hand, foot and mouth disease
npj Biofilms and Microbiomes (2021)
-
The human virome: assembly, composition and host interactions
Nature Reviews Microbiology (2021)
-
Identification of ancient viruses from metagenomic data of the Jomon people
Journal of Human Genetics (2021)
-
Integrated diversity and shared species analyses of human viromes
Archives of Virology (2021)