Abstract
Microbial activities that affect global oceanographic and atmospheric processes happen throughout the water column, yet the long-term ecological dynamics of microbes have been studied largely in the euphotic zone and adjacent seasonally mixed depths. We investigated temporal patterns in the community structure of free-living bacteria, by sampling approximately monthly from 5 m, the deep chlorophyll maximum (∼15–40 m), 150, 500 and 890 m, in San Pedro Channel (maximum depth 900 m, hypoxic below ∼500 m), off the coast of Southern California. Community structure and biodiversity (inverse Simpson index) showed seasonal patterns near the surface and bottom of the water column, but not at intermediate depths. Inverse Simpson’s index was highest in the winter in surface waters and in the spring at 890 m, and varied interannually at all depths. Biodiversity appeared to be driven partially by exchange of microbes between depths and was highest when communities were changing slowly over time. Meanwhile, communities from the surface through 500 m varied interannually. After accounting for seasonality, several environmental parameters co-varied with community structure at the surface and 890 m, but not at the intermediate depths. Abundant and seasonally variable groups included, at 890 m, Nitrospina, Flavobacteria and Marine Group A. Seasonality at 890 m is likely driven by variability in sinking particles, which originate in surface waters, pass transiently through the middle water column and accumulate on the seafloor where they alter the chemical environment. Seasonal subeuphotic groups are likely those whose ecology is strongly influenced by these particles. This surface-to-bottom, decade-long, study identifies seasonality and interannual variability not only of overall community structure, but also of numerous taxonomic groups and near-species level operational taxonomic units.
Similar content being viewed by others
Introduction
Exploration of long-term diversity and temporal dynamics of marine microbial communities living near the sea surface has revealed seasonality, connectivity to ecosystem parameters and co-occurrence patterns (Fuhrman et al., 2006; Steele et al., 2011; Gilbert et al., 2012). However, relatively little is known about whether these characteristics are consistent with or influence subsurface microbial communities, especially those below the euphotic zone. The mesopelagic region is of great biogeochemical interest, as it is the site of much of the Earth’s carbon remineralization (Aristegui et al., 2009; Herndl and Reinthaler, 2013), as well as the nitrogen (Zehr and Ward, 2002) and sulfur cycles (Canfield et al., 2010), particularly in oxygen minimum zones that are found in association with coastal upwelling (Wright et al., 2012). The San Pedro Ocean Time Series (SPOT) provides an opportunity to study the entire water column, from surface to mesopelagic, in a coastal seasonal upwelling system that includes an oxygen minimum at the bottom of the water column. SPOT is located over the San Pedro Basin, in the San Pedro Channel, off the coast of Southern California. The San Pedro Channel has seasonally variable subsurface currents characterized by the surfacing of the California countercurrent during winter months (Hickey, 1992) and a seasonally variable mixed layer with maximum depth of 40 m (Chow et al., 2013). SPOT’s surface shows seasonal environmental and biological patterns, relationships between microbial communities and biotic and abiotic parameters, and a prevalence of putative interactions between microbes, all of which likely influence community structure (Fuhrman et al., 2006; Fuhrman and Steele, 2008; Steele et al., 2011; Needham et al., 2013; Chow et al., 2013, 2014). The physical and chemical environment at SPOT is regularly sampled, and previous work has identified seasonal and other scales of variability in particle flux (Collins et al., 2011), nutrient availability and biogeochemistry (Hamersley et al., 2011) and turnover of water within the San Pedro Basin (Berelson, 1991). This physiochemical and surface-biological information provides a useful background for understanding the temporal dynamics of bacteria throughout the water column.
Most surface microbial communities share common features, including, at a broad level, dominance by Alphaproteobacteria, especially the SAR11 clade, and the presence of other common groups, such as Cyanobacteria, Actinobacteria and Gammaproteobacteria (especially the SAR86 clade) (Pommier et al., 2007; Fuhrman and Hagstrom, 2008). Marine bacterial communities at the ocean’s surface vary globally (Rusch et al., 2007; Pommier et al., 2007), at mesoscales (10–100 km), across ocean fronts (Pinhassi et al., 2003; Hewson et al., 2006b), at smaller scales between river plumes, bays and estuaries and their surrounding environments (Casamayor et al., 2002; Crump et al., 2004; Fortunato et al., 2011; Yeo et al., 2013) and even down to the micrometer scale (Long and Azam, 2001); however, within a given water mass, at least up to several kilometre wide, communities are coherent (Hewson et al., 2006b).
Microbes share common community structure but vary around these common features temporally, as they do on spatial scales. At the SPOT Station, pairs of samples taken days, weeks, months or years apart share many of the same near-species level operational taxonomic units (OTUs), often at similar abundances (Needham et al., 2013; Chow et al., 2013). In fact, most pairs of samples, even those taken at opposite seasons, many years apart, have on average at least 36% similarity (Chow et al., 2013). Overlying this stability, surface assemblages the San Pedro Channel vary seasonally, with some taxa more abundant in surface waters in particular seasons. Seasonal patterns have also been observed in the surface waters of the Bermuda Atlantic Time Series (BATS) (Steinberg et al., 2001; Morris et al., 2005; Treusch et al., 2009; Giovannoni and Vergin, 2012; Vergin et al., 2013b), the Plymouth Marine Lab Western English Channel time series (Gilbert et al., 2012), as well as at other locations (Kan et al., 2007; Rich et al., 2011). The Hawaii Ocean Time Series has less variable surface conditions and less defined, but still detectable, community seasonality (Eiler et al., 2011; Giovannoni and Vergin, 2012). In the SPOT data set, at time scales of 1–4 years, samples that are taken more years apart generally have more dissimilar community structure. Meanwhile samples collected more than 4 years apart have comparable dissimilarity. Seasonal and longer-term changes in community composition were reduced in the deep chlorophyll maximum (DCM) layer compared with surface waters (Chow et al., 2013). Generally, seasonal patterns appear to be ubiquitous in surface waters, though they appear to vary in magnitude between locations. Thus, seasonality at SPOT is representative of seasonality elsewhere, making it a good environment to investigate the less explored topic of subsurface seasonality.
The microbial communities of the mesopelagic ocean (150–1000 m) are generally believed to be less seasonally variable than those found at the surface. Subeuphotic zone microbial communities vary between depths (for example, Garcia-Martinez and Rodriguez-Valera, 2000 and DeLong et al., 2006) and between water masses (Hamilton et al., 2008; Galand et al., 2009a, 2009b; Agogué et al., 2011). Microbial communities at BATS have been shown to vary seasonally from the surface down to depths of 300 m (the deepest depth studied). These patterns are driven by the winter mixing of the top 200–250 m of water, and subsequent summer stratification (Morris et al., 2005; Carlson et al., 2009; Treusch et al., 2009). The mesopelagic at Hawaii Ocean Time Series also appears to be seasonally stable (Eiler et al., 2011). Over a shorter (∼1 year) time series, microbial communities at 300 m in the Northwest Mediterranean Sea vary in abundance, activity and structure, and reportedly relate to water stratification, dissolved organic carbon and transparent exopolymeric particles (Weinbauer et al., 2013). Previous mid-water work at SPOT demonstrated repeating patterns in the abundance of nitrifying organisms including Archaeal groups and Nitrospina OTUs (Beman et al., 2010). Community structure measured over 4 years at SPOT at 500 m has been shown to be quite stable relative to spatial data sets spanning large regions of the ocean (Hewson et al., 2006a). Temporal variation occurs in the context of spatial variability, which is stronger than temporal variability in some (Fortunato et al., 2011; Yeo et al., 2013) but not in other sites (Nelson et al., 2008).
To investigate the natural history of the microbial community at SPOT, between depths, years and seasons, at multiple taxonomic levels, we ask the following questions:
-
1
What is the degree of change of the (surface to 890 m) microbial community at multiple time-scales?
-
2
At what depths and seasons do the highest and lowest levels of biodiversity occur, in terms of both richness and evenness?
-
3
Which environmental and biological parameters relate to community structure?
-
4
Which individual OTUs and taxonomic groups contribute to the seasonal and long-term variability of the microbial community?
Answering these questions will allow us a novel perspective of how microbes throughout the water column change over time and how individual groups contribute to overall shifts in community structure at these depths.
Materials and methods
Sampling
Samples were collected by Niskin bottle from five depths (5 m, DCM, 150 m, 500 m and 890 m) at the SPOT Microbial Observatory site (33° 33′N, 118° 24′W) approximately monthly from August 2000 to January 2011. Sampling at 890 m began in 2003 and samples from below 5 m were not collected in 2007.
Environmental measurements
An in situ sensor (Sea-Bird Electronics, Bellevue, WA, USA) measured temperature and salinity. Oxygen was measured using Winkler titrations from 2000 to 2006 and with an in situ oxygen electrode (Sea-bird, model 13) over the entire data set. The electrode oxygen values were linearly related to Winkler values (R2=0.93). Winkler data were always used in our analyses when available for the appropriate depths. If only electrode data were available, then oxygen values were estimated from the electrode values using a linear fit of electrode vs Winkler values.
Nitrate, nitrite and phosphate samples were stored at −20 °C and measured by standard colorimetric techniques (Parsons, 1984). Total bacterial and viral abundances were determined by SYBR green epifluorescence microscopy (Noble and Fuhrman, 1998; Patel et al., 2007). Bacterial heterotrophic productivity was measured in triplicate 10 ml seawater samples by [3H] leucine incorporation with a conversion factor of 1.5 × 10−17 cells per mol of leucine (Kirchman et al., 1985; Fuhrman et al., 2006).
We calculated values for cell turnover time (leucine), and excess phosphate concentration (P*) as follows:
Cell turnover time (days) = Bacteria concentration (cells per ml)/Productivity (cells per ml per day).
P* = [Phosphate] Concentration−16 × [Nitrate + Nitrite] Concentration (as in Deutsch et al., 2007).
Monthly and 8-day average estimates of surface chlorophyll a concentration and integrated primary productivity, 8-day average estimates of photosynthetically active radiation, colored dissolved organic matter and particulate organic carbon concentrations were downloaded as satellite data from NASA. Meteorological data including daily air temperature, precipitation (measured at Avalon airport, Santa Catalina Island, 16 km away), wave period and height (measured by a buoy in Santa Monica basin, 40 km away), upwelling and multivariate El Niño Southern Oscillation index (MEI) scores (compiled by National Oceanic and Atmospheric Administration) were downloaded and synthesized (Supplementary Information).
Bacterial community structure
For each depth, seawater (5 m—10 l, DCM—10 l, 150 m—15 l, 500 m—20 l and 890 m—20 l) was filtered sequentially through an ∼1-μm pore size, A/E filter and a 0.2-μm pore size Durapore filter (both 142 mm in diameter). Bacterial DNA was isolated as described previously (Fuhrman et al., 1988; Chow et al., 2013) from the frozen (−80 °C) 0.2 μm Durapore filter by hot SDS lysis followed by phenol-chloroform purification (Fuhrman et al., 1988) and stored in Tris-ethylenediaminetetraacetic acid solution at −80 °C. Automated Ribosomal Intergenic Spacer Analysis (ARISA) was performed by amplifying 2 ng of bacterial DNA and performing fragment analysis on 10 ng of the amplified product as initially described by Brown et al. (2005) and with modifications described by Chow et al. (2013). As in both of these articles, technical replicates were generated by running each sample in two lanes of the ARISA gel. Peaks were called in DAx (Chow et al., 2013) and ARISA fragments were normalized to relative abundances, calculated as proportions of total abundance, and dynamically binned (Ruan et al., 2006). Fragments were classified, using clone libraries, based on Greengenes (DeSantis et al., 2006), SILVA (Quast et al., 2012), and RDP (Maidak et al., 2001) taxonomies as outlined previously (Brown et al., 2005; Needham et al., 2013; Chow et al., 2013). In brief, we used existing sequenced clone libraries from SPOT and elsewhere, as well as data sets that contained both 16S ribosomal subunit and intergenic spacer (ITS) DNA sequences to map ARISA fragment sizes to taxonomic identity. Because we were investigating communities throughout the water column, we modified earlier protocols assigning identity to peaks by prioritizing clones from the depths where each ARISA fragment was most abundant on average, rather than always prioritizing surface clones (Supplementary Information). To establish maximum and minimum possible values of community similarity, we determined the mean similarity of machine replicates (maximum) and the statistical difference between 1000 arbitrarily chosen pairs of samples in which the order of the OTUs had been randomized (minimum).
Seasonality and interannual variability of environmental parameters
Each measured environmental parameter was tested for seasonal and interannual variability by way of a generalized additive mixed-effects model (GAMM) (Wood, 2004, 2006) in which data were fit with two splines, one seasonally cyclic and another that fit the overall data set (see Wood, 2006, 321–324; Ferguson et al., 2008). From this model, we report estimated degrees of freedom and P-values for both spline functions and identify the month and year of highest and lowest abundance in the data set. Seasonal and interannual parameters are defined as those whose GAMM function’s seasonal or interannual component had a P-value of <0.05, respectively. We compared this model with another model in which we attempted to fit the variable using the same cyclic seasonal spline and a long-term spline based on the MEI rather than year (Supplementary Information).
Seasonality and interannual variability of overall community structure
Seasonal and interannual patterns were investigated by determining whether samples taken in similar times of year (seasonal) or similar years (interannual) had similar microbial community structure, using both visual and statistical approaches. Visually, we compared temporal distance between pairs of samples (lag) with the Bray-Curtis similarity of those samples. For each month of possible lag, we calculated mean and 95% confidence intervals of Bray-Curtis similarities (Supplementary Information). More quantitatively, we compared the Bray-Curtis similarity of each pair of samples to whether the pair of samples was taken in similar or different times of year (seasonal) and whether the samples were taken in similar or different dates (interannual). The relationship between these temporal and community similarity was evaluated statistically using a Mantel test (Supplementary Information). Depths with seasonally or interannually variable communities were defined as those with the corresponding Mantel P-values of <0.05.
Alpha diversity
For each time point and depth, Shannon index and inverse Simpson index (ISI) were calculated using the ‘diversity’ function in the ‘vegan’ R-package (Oksanen et al., 2013). Sample richness was estimated by quantifying, at each depth, the number of OTUs with a relative abundance of more than 0.01%, 0.1% and 1% of the total community. For subsequent analysis, we used 0.1% as our richness cutoff because it was well above our ARISA detection threshold of 0.01%, giving us good confidence that the OTUs were indeed present in our data set. Pielou’s evenness was calculated as ‘H/ln(S)’, where H is the Shannon Index, S was the number of OTUs found with greater than 0.1% relative abundance. For each depth, we calculated mean values and 95% confidence intervals of each of these metrics. Analysis of variance was applied to determine whether there was statistically significant difference in richness and ISI between depths (Supplementary Information). We also determined whether richness and ISI related to season, long-term trends, the degree of change of community structure, or the similarity of community structure between depths (Supplementary Information).
Environmental parameters and community structure
Partial Mantel tests were applied to determine which environmental parameters relate to community structure at each depth. To identify which environmental variables could predict community structure beyond seasonal effects, we statistically removed the effects of seasonality and long-term variability from this environmental variability analysis (Supplementary Information).
Temporal dynamics of microbial taxa over time
Relative abundances for each taxon (at multiple taxonomic levels) were determined in each sample by summing the relative abundance of all ARISA OTUs within that group. GAMMs were used to model the abundance of taxonomic groups’ relative abundances using cyclic seasonal splines and interannually variable splines. This GAMM approach was similar in structure to that applied to environmental variables (see above). These GAMMs determined which taxonomic groups, and which of the 100 most abundant OTUs at each depth, showed seasonality and long-term variability. They also identified the months and years in which these groups were most and least abundant (Supplementary Information).
Results
Variability of environmental and biological parameters, between seasons, years and depths
Temperature and salinity varied seasonally at several depths throughout the water column, but showed greater variability in surface waters than deeper water (Figure 1, Supplementary Figure S1, Supplementary Table S1). Satellite measurements of surface chlorophyll a, primary productivity, particulate organic carbon concentration, colored dissolved organic matter concentration and sea surface height were all seasonally variable with the highest concentrations or rates in the spring. Mixed layer depth, upwelling, wave height, average wave period, dominant wave period, precipitation, daily maximum and minimum air temperatures and wind speeds all varied seasonally.
Nutrient concentrations varied with depth and season (Figure 1, Supplementary Figure S1, Supplementary Table S1). Notable among these variations was the seasonality of nitrite, despite its low abundance, at 890 m (Supplementary Figure S1, Supplementary Table S1) and the interdepth variability of oxygen which ranged from saturated (>200 μM) in surface waters to strongly hypoxic at the bottom of the water column (<10 μM) and were seasonally variable at 5 and 150 m, but not so at other depths.
Bacterial abundances were about threefold higher in the euphotic zone (5 m and DCM, ∼1 × 106 cells per ml), where they varied seasonally, than in deeper waters (150 m, 500 m, 890 m, ∼3 × 105 cells per ml) where they did not show seasonal variability. Bacterial growth rates, as measured by leucine uptake, varied seasonally at all depths except 500 m. Estimated average doubling times of bacteria were around 5 days in the euphotic zone, considerably faster than the estimated average doubling time of around 100 days in the deeper depths. At all depths, many parameters showed long-term variability over the course of the data set at most depths, showing increasing, decreasing or non-linear trends (Supplementary Table S1). There were very few cases in which using MEI rather than year to predict long-term variability, improved statistical significance of a GAMM suggesting that the El Niño Southern Oscillation had minimal effect on environmental variables in our data set, likely because there were no strong El-Niño years during the time these data were collected.
Bacterial community data overview
In all, 332 distinct OTUs were detected over the entire data set, where OTU was considered only if their peak in the electropherogram constituted at least 0.01% of the total ARISA peak area in at least 1 month. We assigned taxonomic identities to 131 OTUs. The most abundant OTUs tend to be the most easily identified via our clone libraries and sequence databases, and we assigned identity to OTUs that cumulatively comprised ∼90% of the bacterial community (5 m, 93%; DCM, 91%; 150 m, 86%; 500 m, 90%; 890 m, 85%).
Seasonal variability in microbial community structure
At all depths, samples taken 1 month apart were between 50% and 60% similar. In contrast, machine replicates (same sample) averaged 78% similar, while simulated random unrelated samples averaged 6.5% similar. As has been reported previously at the surface (Hatosy et al., 2013; Chow et al., 2013), when temporal difference between samples was plotted against Bray-Curtis dissimilarity, we observed a sinusoidal pattern with maximum Bray-Curtis similarities at multiples of 1 year, suggesting a seasonal repetition. We observed this sinusoidal pattern at 890 m but not at the intermediate depths (Figure 2). In contrast, at the DCM and 150 m, samples taken 6 months apart were not statistically dissimilar from samples taken 1 month apart (Table 1). A Mantel test comparing the Bray-Curtis dissimilarities with seasonal difference confirmed these seasonal patterns at the surface and 890 m, and suggested weaker but still statistically significant seasonality at the DCM. Meanwhile, the Mantel test did not suggest seasonality at 150 or 500 m (Table 1).
Mean Bray-Curtis similarities of all pairs of samples (y axis) separated by different intervals of time (x axis). Circles represent the mean similarity of all pairs of samples taken a given number of months apart (intermonthly; left y axis). Thus, the first circle is the mean Bray-Curtis similarity of all pairs of samples taken 15–45 days (∼1 month) apart, the second circle is the mean similarity of all pairs of samples taken 46–76 days (∼2 months), the twelfth circle (aligned with the 1-year lag tick mark) represents samples taken 12 months apart and so on. Squares represent mean similarity of samples taken a given number of years apart (interannual; right y axis). Accordingly, the first square represents the mean Bray-Curtis similarity of all pairs of samples taken less than 1 year apart, the second is all pairs of samples taken between 1 and 2 years apart and so on. Error bars, for both types of data points, represent 95% confidence intervals of the mean similarity. Points with non-overlapping error bars suggest statistically significant differences in mean similarity between samples taken different distances apart in time. For instance, samples taken 6 months apart in the surface (the sixth circular data point) are statistically less similar than samples taken 1 month apart (first circular point), while samples taken 12 months apart (twelfth circular point) are not less similar than samples taken 1 month apart.
Interannual variability in microbial community structure
In surface waters, communities sampled less than a year apart averaged 50% Bray-Curtis similarity while samples taken 1–2 years apart had only 45% similarity, a statistically significant decrease (ANOVA F=12.84, DF=11, P<10−15; Tukey corrected t-test P<10−6). Samples taken farther apart in time appeared to be less similar up through 4 years apart (Figure 2), as was found previously (Chow et al., 2013), with samples taken 4 years apart statistically significantly less similar than samples taken 2 years apart (Tukey corrected t-test P<10−4). All pairs of samples 4 or more years apart tended to be ∼37% similar on average, while oscillating seasonally. In contrast, at the DCM, 150 m and 500 m depths, communities 1 year apart were roughly as similar as communities sampled 2 and 6 years apart. Communities sampled more than 7 years apart were less similar than those taken fewer than 7 years apart at these intermediate depths (ANOVA F=6.11 DCM, 4.59 150 m, 14.37 500 m, DF=11, P<10−6; Tukey corrected t-test P<0.05). At 890 m there was a weaker but still statistically detectable decrease in similarity between samples taken 1 year apart and samples taken 4–5 years apart (ANOVA F=2.95, DF=8, P<0.01, Tukey corrected t-test P<0.05) (Figure 2). At all depths except 890 m, Mantel tests suggested that there was a statistically significant relationship between the temporal distance of samples and their Bray-Curtis distance, further indicating the presence of a long-term trend (Table 1).
Alpha diversity relates to season and certain community structure patterns
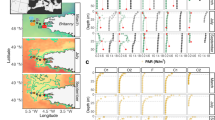
Both average richness (number of OTUs with >0.1% relative abundance) and ISI varied between depths (Richness: ANOVA F=3.81, DF=4, P<0.01; ISI: ANOVA F=37.37, DF=4, P<0.001) (Figure 3a). The 150-m depth had the highest mean richness and ISI, which was statistically significantly higher than the richness at 5 m only (Tukey corrected P<0.01) and ISI at all depths except 890 m (Tukey P<0.01) (Figure 3a). At 5 m and the DCM, ISI (but not richness) was highest in the winter, while at 890 m both richness and ISI values were highest in the spring (Figure 1, Supplementary Figure S1, Supplementary Table S1). In the DCM layer, richness was highest in winter while ISI did not vary seasonally. At 150 m and 500 m neither richness nor ISI varied seasonally.
(a) Mean biodiversity index scores by depth. X axis labels are the biodiversity metrics under evaluation. Richness is mean number of species in a given sample with greater than 1%, 0.1% and 0.01%. Inverse Simpson (ISI) and Shannon H are biodiversity indexes and Pelou J measures evenness. Y axes are the values for each metric (OTUs for richness and inverse Simpson’s index, unitless for Shannon Biodiversity and Peilou’s evenness. Symbols represent depths and are described in the legend. Bars represent 95% confidence intervals from the mean. (b) Similarity between communities at 150 and 500 m (x axis) compared with the ISI of the community at 150 m (y axis). Each point represents the relationship between ISI and similarity between depths for a single month. The solid line is a trend line and the dashed lines represent 95% confidence intervals of this trend. The R-value of this correlation is 0.49 with associated confidence intervals from 0.30 to 0.64. This R-value and its confidence interval correspond to the darkened symbol and bar in (c). This figure is provided as an example of correlations seen between depths and ISI shown in (c). Other pairs of depths, shown in (c), show similar trends. (c) R-values of correlations (y axis) between inverse Simpson index (ISI) values for each depth given as a symbol, and its similarity to other depths or to the previous month (x axis). Symbol shapes signify depth at which the biodiversity was measured. The first five parameters signify the Bray-Curtis similarity between that the depth measured (indicated by symbol) and each other depth (vs5m, vsDCM, vs150m, vs500m, vs890m). Higher R-values represent a stronger correlation between the biodiversity at the measured depth and its similarity to the depth given by the x label. For instance, the y value of the darkened diamond shape is 0.32, corresponding to slope of the relationship between biodiversity at 150 m and interdepth similarity between 150 m and 500 m depicted in (b). The ‘vsLastMonth’ column represents the relationship between ISI and the Bray-Curtis similarity between that sample and a sample taken at that same depth collected in the previous month. Bars represent 95% confidence intervals for the R-value. Confidence intervals not overlapping the x axis indicate statistically significant correlations.
We observed a positive correlation at all depths between either richness and ISI and the similarity of that sample to communities at most other depths. As an example, in months when community structure at 150 and 500 m were similar, the 150 m community had a high ISI (Figure 3b). This finding is also true of most other pairs of depths (Figure 3c, Supplementary Figure S2). We further observed that, at 150 m and 500 m, months in which community structure had changed little in the past month also had higher richness and ISI than months that had undergone larger changes in the past month (Figure 3c; Supplementary Figure S2).
Environmental parameters relate to community structure
After seasonal and long-term variability were factored out, similar surface communities generally were characterized by similar monthly mean chlorophyll a concentrations, monthly mean primary productivity, particulate organic carbon concentration, virus abundance and leucine incorporation rates (P<0.01) (Table 2). At the DCM, 150 m and 500 m, no environmental parameter related significantly to community structure with P<0.01. At 890 m, the depth of the mixed layer and the amount of precipitation on the day of sampling related to community structure (P<0.01). For these observations in which P<0.01, the false discovery rate (Q-value) was 3% in the surface and 9% at 890 m, providing us with reasonable confidence in the result, despite the multiple comparisons. Conversely, use of P<0.05 had higher false discovery rates (variable between depths but below 25% in all depths except 500 m) suggesting a proportion (though still a less than a quarter) of the weaker correlations are due to random chance. Notable statistically weaker (P<0.05) factors relating to community structure included the abundance of bacteria and the dominant (but not average) wave period at 5 m, the DCM and 150 m; surface chlorophyll a and surface primary productivity as measured by satellites to the DCM community; and the mixed layer depth to the 150 m community. The false discovery rate of 65% at 500 m suggests that the one positive relationship observed at that depth is probably due to random chance and that no measured environmental parameters explained community variability at 500 m.
Dynamics of dominant marine bacteria clades
The relative abundances of several taxonomic groups were examined both at phylum to class (Figure 4a) and at family levels (Figure 4b) at each depth (Table 3, Supplementary Figure S3). Furthermore, the subset of taxa that showed seasonal variability differed between depths.
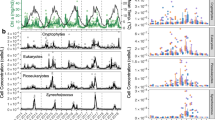
Heat map of relative abundances of bacteria from class and phylum level taxonomic groups (a) as well as family and order level groups (b). Each panel represents a different taxonomic grouping. The x axis indicates months and y axis indicates sampled depths (not to scale). Colors correspond to the summed relative abundance of all OTUs identified as falling within that taxonomic group, averaged by month. Relative abundance scores for each color are given in the scale bar at right; note that abundances are on a log scale.
Alphaproteobacteria, dominated by the SAR11 clade, were the most abundant class at every depth. The AEGEAN-169 clade, which is closely related to SAR11 (Alonso-Sáez et al., 2007), was most abundant in deeper depths where SAR11 was less abundant (Figure 4b, Table 3, Supplementary Figure S3B). Different clades of SAR11 showed different patterns with the dominant Surface 1 clade abundant regardless of season at most depths, and the less abundant Surface 2 and Surface 4 subclades seasonally variable at several depths. SAR11 Deep 1 was abundant in 150 and 500 m waters. SAR11 Surface 1 was primarily comprised of three OTUs: OTU_666.4, OTU_686.9 and OTU_670.5 (in order of abundance, and where the number indicates ARISA fragment length) (Supplementary Figures S4B and S5B) and several other less abundant OTUs. Each Surface 1 OTU seemed to exhibit different seasonal and interannual patterns from each other.
Gammaproteobacteria were abundant at all depths (Figure 4a, Table 3). They were dominated by the SAR86 group in the surface, and other OTUs such as SUP05 in deeper waters (Figure 4b, Table 3). Actinobacteria, dominated by the OCS155 clade, Cyanobacteria, which were predominantly Prochlorococcus, and Chloroplasts, from eukaryotic picophytoplankton, were all more abundant in surface waters than in deep waters (Figure 4). Both Cyanobacteria, most abundant in late fall, and Chloroplasts, abundant in late winter and early spring, were seasonal in the surface waters (Supplementary Table S2).
More abundant in deeper waters were Deltaproteobacteria, Flavobacteria and the Marine Group A (MGA) phylum (Figure 4a, Table 3), all of which were seasonally variable. Within these classes, bacteria from the NS9 (Flavobacteria), Nitrospina (Deltaproteobacteria) and Arctic 96B-7 (MGA) groups appeared to drive the seasonal variability of the broader taxonomic groups (Figure 4b, Table 3). Arctic96B-7 also showed seasonal variability at 150 and 500 m, while SAR324 appeared to drive seasonality of Deltaproteobacteria at 500 m. Investigation of genus level groups of MGA and clades Flavobacteria showed diverse seasonal and interannual patterns (Supplementary Tables S4 and S5). In addition to seasonal variability, many taxa showed long-term increases, decreases or non-linear changes in abundance.
Dynamics of abundant marine bacterial OTUs
Non-parametric regression models, using seasonal and long-term splines to fit the data, showed that for the 100 most abundant OTUs at each depth, 5 m, the DCM and 890 m had the highest fraction of seasonal OTUs, while 150 and 500 m had fewer seasonal OTUs (Table 1, Supplementary Table S5). The abundances of the five most abundant bacteria at each depth (Supplementary Figure S6) reflect the patterns seen for the 100 most abundant OTUs, with seasonal bacteria present at the surface and bottom of the water column but not at the middle water column. The five most abundant bacteria at each depth show differences in their distribution patterns across depths, with some abundant at multiple depths (for example, most abundant AEGEAN-169 OTUs), and others only abundant at one depth (for example, SAR406 with ITS length of 709.4). Chi-squared analysis confirmed that the depths did have statistically significant differences in the number of seasonal bacteria, and not in the number of bacteria that showed long-term variability. The subset of the 100 most abundant bacteria that were determined to be seasonal or interannually variable included OTUs from many abundant phylogenetic groups, including groups that taken as a whole did not vary seasonally or interannually (Supplementary Table S5).
Discussion
Seasonality is strongest at the surface and bottom of the water column
Seasonal patterns appeared to be strongest at the surface (5 m), weak in the middle water column (DCM, 150 m and 500 m), and strong again just above the sea floor (890 m) according to a number of metrics:
-
1
Community Structure: Samples collected in the same season across years were similar but samples collected from different seasons were dissimilar at 5 and 890 m (Figure 2).
-
2
Biodiversity: Richness and ISI were highest in the winter at 5 m and in the spring at 890 m (Figure 3).
-
3
Taxonomic Groups: Several broad taxonomic groups of organisms were seasonal, especially at the surface (such as SAR11 and Prochlorococcus) and at 890 m (such as Nitrospina and NS9 Flavobacteria) (Figure 4, Table 3, Supplementary Figures S2 and S3).
-
4
Individual OTUs: There were more seasonal individual OTUs at the surface, DCM and 890 m than at 150 or 500 m (Table 1, Supplementary Table S5).
Seasonality of surface communities is consistent with previous findings, both at this site (Fuhrman et al., 2006; Chow et al., 2013) and elsewhere (Morris et al., 2005; Carlson et al., 2009; Treusch et al., 2009; Gilbert et al., 2012; Vergin et al., 2013b), and is likely governed by strong seasonality in several environmental factors. Photosynthetically available radiation, day length, nutrient concentrations, seawater temperature and mixing of the water column all vary seasonally in the surface (Figure 1, Supplementary Figure S1, Supplementary Table S1) and many or all of these factors likely influence the bacterial community. However, covariance of these variables precludes identifying their individual relative contributions to seasonality. Primary productivity and concentration of chlorophyll a are also seasonally variable, and appear to relate to community structure even beyond this seasonal effect, that is, after seasonality is factored out (Table 2).
At intermediate depths, the decrease in seasonality below the photic zone is consistent with findings at Hawaii Ocean Time Series and BATS (as reviewed in Giovannoni and Vergin, 2012). Light levels, temperature and turbulence are less seasonally variable at SPOT’s deeper depths. At 500 m, temperature is stable and light intensity is insufficient for photosynthesis. Unlike BATS, the mixed layer at SPOT is almost always above 40 m, so the 150m, 500m or 890m depths likely never mix with surface water directly (even though mixed layer depth does appear to relate to community structure at 890 m; Table 2).
Seasonality of the deepest depths has not been seen previously, even in a prior analysis of a shorter subset of the SPOT 890 m data (Hatosy et al., 2013), either because the additional samples gave this study more statistical power, or because the statistical tests employed in this study are more sensitive. Particle flux, which varies seasonally in magnitude and composition (Collins et al., 2011), likely contributes to the observed seasonality at 890 m, which is at the bottom of the water column. Particles are believed to be the source of key substrates for bacteria in the deep ocean and likely transport organic nutrients from the surface to the bottom, where they may become entrained in the nepheloid layer or land on the sea floor, degrade and affect nearby overlying waters (Francois et al., 2002; Herndl and Reinthaler, 2013). Therefore, the longer residence time of particles and their associated organic matter at or near the sea floor, compared with mid-water, likely leads to stronger seasonality there. Note that our study assessed the free-living bacterial communities and not those attached to larger particles at the time of sampling. Furthermore, many particles are heavily colonized by bacteria (Simon et al., 2002) and may transport bacteria to bottom waters where they may be released (Sohrin et al., 2011). The presence of cyanobacteria (presumably photosynthetic) at 890 m reflect similar findings in other systems (Sohrin et al., 2011) and further suggests bacterial transport. Particles have been shown to sink at a rate averaging about 83 m per day (Collins et al., 2011), which suggests that it should take about 11 days to travel from the surface to the bottom of the water column. Differences in sizes and densities between particles cause different compounds or organisms to be transported from the surface to the deep at different rates. Currents in the region vary with both depth and season, (Hickey, 1991, 1992) and likely cause spatial separation between the locations where particles are generated and where they settle. Particle flux variability across space (Buesseler et al., 2009) may interact with these seasonal currents, resulting in seasonally variable particle delivery to the sea floor at SPOT. Water below the sill of the San Pedro basin at 750 m is primarily trapped (Berelson, 1991), suggesting that currents alone seem like an unlikely explanation for the greater seasonality at 890 m than at intermediate depths.
In addition to sinking particles, crustaceans and other zooplankton likely influence the subsurface communities by consuming organisms and marine aggregates at the surface and egesting rapidly sinking fecal pellets, in conjunction with diel vertical migration (Steinberg et al., 2000, 2002; Wilson and Steinberg, 2010). Other organisms that may transport nutrients or bacteria include cnidarians (Schnetzer et al., 2011) and larvaceans (Hansen, et al., 1996, Robison et al., 2005). These pelagic organisms together link the entire water column such that features of any one depth may affect other depths.
Interannual variability is apparent throughout the water column
Interannual variability of the microbial community in conjunction with observations of long-term stability may reflect seasonal trends and environmental factors vary from year to year but maintain a common ‘average’ microbial community. At all depths, long-term processes may be related to interdecadal processes such as the El Niño Southern Oscillation Index or Pacific Decadal Oscillation index. We were not able to detect a relationship between the MEI and community structure in this study (Table 2), likely because this study is only one decade long and does not encompass any major El Niño events. As time series studies continue, the longer data sets will provide information to better resolve these long time-scale trends.
Alpha-diversity patterns differ between depths
The seasonality of biodiversity at 5 m (Table 1, Supplementary Table S1) parallels trends seen in the Western English Channel (Gilbert et al., 2012) and at BATS (Vergin et al., 2013b), while the biodiversity’s seasonality at 890 m is an original observation, to our knowledge. The biodiversity maximum in the mid-water column at 150 m (Figure 3a) reflects patterns seen elsewhere in the ocean for other organisms including copepods, ostracods (Angel, 1993; Lindsay and Hunt, 2005) and fish (Badcock and Merrett, 1976). Biodiversity is often highest at ecotones, the interface between different environments, (Angel, 1993; Barton et al., 2010; Ribalet et al., 2010) and the 150 m depth is such an interface between the euphotic and disphotic zones.
The correlation, at most pairs of depths, between high similarity and high biodiversity at those depths (Figures 3b and c, Supplementary Figure S2) suggests mixing of communities between depths or immigration of microbes from one depth to another may drive increased diversity at those depths. For instance, diversity may be highest in the surface in the winter (when storms may mix water to ∼40 m) because the deeper, mixed in or upwelled, water layer brings microbes from what had been in the stratified waters below the mixed layer into the surface. Higher biodiversity at 150 and 500 m, among those samples that changed least from month to month (Figures 3b and c), suggests that particularly low disturbance allows for more niche differentiation and thus higher diversity.
Environmental variability relates to community structure
At the surface
The relationship between microbial community structure and chlorophyll a concentrations, particulate organic carbon concentrations, primary productivity, heterotrophic productivity and viral abundance at the surface (Table 2) extends Chow et al.’s (2013) observations of particular environmental variables relating to the abundance of certain OTUs by identifying variables that relate to community structure beyond the effects of seasonality. For instance, while Chow et al. (2013) observed that nitrate was related to community structure, both nitrate and community structure are seasonally variable, and in this study we were not able to differentiate nitrate effects away from seasonal effects statistically. Thus, we do not know whether nitrate itself or some other seasonal factor shapes community structure. Conversely, chlorophyll a concentrations and heterotrophic productivity (leucine incorporation) are related to the community when de-convoluted from seasonality, suggesting links between these parameters beyond season, even though these parameters also show seasonality.
Chlorophyll a concentrations likely relate to community structure because bacteria consume phytoplankton exudates (Obernosterer and Herndl, 1995; Fouilland et al., 2014), live symbiotically with phytoplankton (Caron, 2000; Aota and Nakajima, 2001) and/or respond to unmeasured variable(s) along with phytoplankton (Steele et al., 2011; Chow et al., 2014). Community structure may relate to leucine incorporation because some OTUs are more metabolically active than others. Bacterial abundance and viral abundance, but not virus-to-bacteria ratios, relate to community structure, suggesting that denser communities contain different members than less dense communities. This difference may result from greater numbers of interactions between bacteria, grazers and viruses in more dense communities. Alternatively, particular sets of environmental conditions may favor both dense microbial communities and particular microbial groups. Particulate organic carbon has been implicated as an important substrate for many marine bacteria (Azam, 1998; Simon et al., 2002) and it seems likely that the bacteria that co-occur with particles at the surface may be directly associated with (living on or eating) these particles.
At 890 m
The relationship between 890 m community structure and mixed layer depth and precipitation further suggests that conditions at the top of the water column contribute to variability at the bottom of the water column. Mixed layer depth is known to influence the range of depths at which organisms can produce biomass and the types of dominant phytoplankton and zooplankton (Sverdrup, 1953; Bissett et al., 1994; Arrigo et al., 1999; Eslinger et al., 2001) and this variation in surface productivity and plankton community structure likely in turn influences the rate of particle production, particle type and delivery to the sea floor. Collins et al. (2011) found that there was no direct correlation between rainfall and mid-water particle flux nor between rainfall and particulate carbon to nitrogen ratios, suggesting that links between rainfall and deep water community structure are by way of some process other than surface to bottom particle flux. Precipitation may trigger turbidity flows in which sediment influx from land after rainfall rapidly moves down submarine canyons, such as nearby Redondo Canyon (see Sholkovitz and Soutar, 1975). These turbidity flows likely transfer sediment to bottom waters and/or distribute it into the lower water column by entrainment (Drake and Gorsline, 1973; Drake, 1974). Correlations to rainfall on the day of sampling may relate to the observation that major rainstorms last multiple days.
Mid-water column
The absence of observed parameters strongly relating to community structure at the DCM, 150 m and 500 m depths, in combination with lack of seasonality at those depths, suggests that mid-water community structure variability is due to unmeasured influences. These influences could include unmeasured environmental factors, interactions between microorganisms, neutral processes (see Chave, 2004), or interactions between multiple environmental influences.
Variability of specific taxonomic groups drives overall community variability
We saw different seasonally variable taxonomic groups at the surface, middle and bottom of the SPOT water column (Table 3, Supplementary Tables S2–S4). The seasonal surface groups, like the surface community as a whole, likely respond to seasonality in light levels, mixing and nutrient levels. Prochlorococcus are adapted to oligotrophic stratified warm water conditions (Partensky et al., 1999) and tend to occur during the summer and fall (Figure 4), when Chlorophyll a levels are lowest (Supplementary Figure S1), as a result of this adaptation. The appearance of Prochlorococcus in deep waters may reflect transport of this organism from surface waters (see Sohrin et al., 2011). Alternatively, since no Cyanobacteria were found in our 890 m clone libraries, other bacteria with the same ITS length as cyanobacteria may inhabit the mesopelagic. The depth structure and seasonal patterning of SAR11, and their sister clade AEGEAN-169 (Supplementary Figure S4A, Supplementary Table S4) reflect patterns seen at BATS (Carlson et al., 2009, 11; Vergin et al., 2013a). The seasonality of individual OTUs within the non-seasonal SAR11 Surface 1 clade (Supplementary Tables S4, S5) suggested that this finer resolution is important in understanding the dynamics of this abundant group.
Seasonal taxa at 890 m are likely related indirectly to seasonally variable surface parameters. Flavobacteria remineralize particles and complex organic molecules (Kirchman, 2002) and are likely seasonal because the flux and content of particles or other food sources are also seasonally variable. Nitrospina are likely nitrite oxidizers (Fuessel et al., 2012) and have been implicated as drivers of nitrite oxidation in subsurface layers in much of the ocean (Mincer et al., 2007; Beman et al., 2013). At SPOT they have been shown to respond to variability of nitrite and ammonia oxidizing Archaea (Beman et al., 2010). Seasonal variability at 890 m of their substrate, nitrite, (Supplementary Figure S1, Supplementary Table S1) further supports this finding. Seasonality of clades of MGA and SAR324 at different mesopelagic depths is interesting as some strains are reported to oxidize reduced sulfur or methane (Swan et al., 2011). Because Nitrospina and SAR324 and MGA are thought to be chemoautotrophic, seasonality in these organisms’ abundances likely reflects seasonally variable biogeochemistry.
Despite these seasonal patterns, one remarkable feature of broad taxonomic groups is their stability. While communities, evaluated on an OTU level, vary significantly between months, seasons and years, communities within a given depth appear to be dominated consistently by a common set of broader taxa at similar relative abundances.
Individual OTUs drive community variability
All depths, regardless of overall seasonality, contain some OTUs that are seasonally variable and others that are not. The prevalence of non-seasonal OTUs at depths with overall community structure seasonality suggests that there are many OTUs that interact with their environment in ways not affected by this variability. Conversely, at depths in which most OTUs do not vary seasonally, there are always at least a few OTUs, making up a smaller subset of community variability, that have a seasonally variable niche. These seasonal OTUs at non-seasonal depths indicate mid-water seasonality of at least some environmental properties, such as flux of certain kinds of particles. During phytoplankton blooms, Teeling et al. (2012) observed that finer level taxonomic level groups of bacteria adapted to different ecological niches by expressing different metabolic processes. The observation that non-seasonal taxa contain seasonal OTUs (Table 3, Supplementary Tables S2–S5) supports this idea of niche differentiation within the broader taxa and is a compelling reason to focus future ecological analysis of community structure on fine, rather than coarse taxonomic groups, in order to discern mechanisms.
Advantages and considerations of using ARISA
The ARISA technique is well suited to analysis of this data set, because its reliance on the finely resolved length of the hypervariable 16S-23S ITS region allows resolution of closely related OTUs (Brown et al., 2005), often better than current next-generation 16S tag sequencing approaches (Chow et al., 2013). It is nevertheless important to consider that any PCR-based technique, including this one, has biases, such as over-representing some DNA fragments (for example, shorter ones), meaning that our relative abundance estimates may have biases toward some taxa and against others. Fortunately, any such bias is likely consistent from sample to sample, suggesting that the conclusions drawn from the seasonal and interannual dynamics, the main focus of this report, are justified. All of the statistics used to find patterns in this study are generally insensitive to such biases. It is also important to consider that this version of ARISA detects neither Archaea nor bacteria from the Planctomycetes and SAR202 phyla, all of which are members of many marine microbial communities, especially deep ones (Woebken et al., 2008; Treusch et al., 2009). It is also inevitable that some OTUs will contain members of more than one unrelated group, so the dynamics of any OTU may be the combined dynamics of any groups that share a common ITS length. Although some ARISA OTUs (∼10% of the community, on average) are currently unknown, as they do not have corresponding clones in our clone libraries, they represent a minority of our community fingerprint data. Despite being a nearly decade old, this ARISA and clone library combined approach, as shown by Brown et al. (2005) and subsequent studies, is still an effective method capable of generating high quality data suitable for this type of study.
Conclusion
Seasonal variability is most evident at 5 and 890 m while long-term variability is evident throughout the top 500 m of the water column. After seasonal and interannual variability had been factored out, several environmental parameters further explained community variability at 5 and 890 m but not at intermediate depths. These patterns suggest links between the surface environment and deep water communities, possibly driven by rapidly sinking particles and migrating plankton. This variability in community structure appears to be driven by particular OTUs and sometimes broader taxonomic groups, and likely reflects OTU-specific or group-specific responses to environmental variability. The patterns reported here present a useful initial set of expectations for similar sites, including coastal basins and oxygen minimum zones.
References
Agogué H, Lamy D, Neal PR, Sogin ML, Herndl GJ . (2011). Water mass-specificity of bacterial communities in the North Atlantic revealed by massively parallel sequencing. Mol Ecol 20: 258–274.
Alonso-Sáez L, Balagué V, Sà EL, Sánchez O, González JM, Pinhassi J et al. (2007). Seasonality in bacterial diversity in north-west Mediterranean coastal waters: assessment through clone libraries, fingerprinting and FISH. FEMS Microbiol Ecol 60: 98–112.
Angel MV . (1993). Biodiversity of the Pelagic Ocean. Conserv Biol 7: 760–772.
Aota Y, Nakajima H . (2001). Mutualistic relationships between phytoplankton and bacteria caused by carbon excretion from phytoplankton. Ecol Res 16: 289–299.
Aristegui J, Gasol JM, Duarte CM, Herndl GJ . (2009). Microbial oceanography of the dark ocean’s pelagic realm. Limnol Oceanogr 54: 1501–1529.
Arrigo KR, Robinson DH, Worthen DL, Dunbar RB, DiTullio GR, VanWoert M et al. (1999). Phytoplankton community structure and the drawdown of nutrients and CO2 in the Southern Ocean. Science 283: 365–367.
Azam F . (1998). Microbial control of oceanic carbon flux: the plot thickens. Science 280: 694–696.
Badcock J, Merrett NR . (1976). Midwater fishes in the eastern North Atlantic—I. Vertical distribution and associated biology in 30°N, 23°W, with developmental notes on certain myctophids. Prog Oceanogr 7: 3–58.
Barton AD, Dutkiewicz S, Flierl G, Bragg J, Follows MJ . (2010). Patterns of diversity in marine phytoplankton. Science 327: 1509–1511.
Beman JM, Leilei Shih J, Popp BN . (2013). Nitrite oxidation in the upper water column and oxygen minimum zone of the eastern tropical North Pacific Ocean. ISME J 7: 2192–2205.
Beman JM, Sachdeva R, Fuhrman JA . (2010). Population ecology of nitrifying Archaea and Bacteria in the Southern California Bight. Environ Microbiol 12: 1282–1292.
Berelson WM . (1991). The flushing of two deep-sea basins, Southern California Borderland. Limnol Oceanogr 36: 1150–1166.
Bissett W, Meyers M, Walsh J, Mullerkarger F . (1994). The effects of temporal variability of mixed-layer depth on primary productivity around Bermuda. J Geophys Res. 99: 7539–7553.
Brown MV, Schwalbach MS, Hewson I, Fuhrman JA . (2005). Coupling 16S-ITS rDNA clone libraries and automated ribosomal intergenic spacer analysis to show marine microbial diversity: development and application to a time series. Environ Microbiol 7: 1466–1479.
Buesseler KO, Pike S, Maiti K, Lamborg CH, Siegel DA, Trull TW . (2009). Thorium-234 as a tracer of spatial, temporal and vertical variability in particle flux in the North Pacific. Deep Sea Res Part I Oceanogr Res Papers 56: 1143–1167.
Canfield DE, Stewart FJ, Thamdrup B, Brabandere LD, Dalsgaard T, Delong EF et al. (2010). A cryptic sulfur cycle in oxygen-minimum–zone waters off the Chilean Coast. Science 330: 1375–1378.
Carlson C, Morris R, Parsons R, Treusch A, Giovannoni S, Vergin K . (2009). Seasonal dynamics of SAR11 populations in the euphotic and mesopelagic zones of the northwestern Sargasso Sea. ISME J 3: 283–295.
Caron DA . (2000). Symbiosis and mixotrophy among pelagic microorganisms. In: Microbial Ecology of the Oceans Kirchman DL, (ed.) Wiley-Liss: New York, NY, pp 495–523.
Casamayor EO, Massana R, Benlloch S, Ovreas L, Diez B, Goddard VJ et al. (2002). Changes in archaeal, bacterial and eukaryal assemblages along a salinity gradient by comparison of genetic fingerprinting methods in a multipond solar saltern. Environ Microbiol 4: 338–348.
Chave J . (2004). Neutral theory and community ecology. Ecol Lett 7: 241–253.
Chow C-ET, Kim DY, Sachdeva R, Caron DA, Fuhrman JA . (2014). Top-down controls on bacterial community structure: microbial network analysis of bacteria, T4-like viruses and protists. ISME J 8: 816–829.
Chow C-ET, Sachdeva R, Cram JA, Steele JA, Needham DM, Patel A et al. (2013). Temporal variability and coherence of euphotic zone bacterial communities over a decade in the Southern California Bight. ISME J 7: 2259–2273.
Collins LE, Berelson W, Hammond DE, Knapp A, Schwartz R, Capone D . (2011). Particle fluxes in San Pedro Basin, California: a four-year record of sedimentation and physical forcing. Deep Sea Res Part I Oceanogr Res Papers 58: 898–914.
Crump BC, Hopkinson CS, Sogin ML, Hobbie JE . (2004). Microbial biogeography along an estuarine salinity gradient: combined influences of bacterial growth and residence time. Appl Environ Microbiol 70: 1494–1505.
DeLong EF, Preston CM, Mincer T, Rich V, Hallam SJ, Frigaard N-U et al. (2006). Community genomics among stratified microbial assemblages in the Ocean’s Interior. Science 311: 496–503.
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K et al. (2006). Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72: 5069–5072.
Deutsch C, Sarmiento JL, Sigman DM, Gruber N, Dunne JP . (2007). Spatial coupling of nitrogen inputs and losses in the ocean. Nature 445: 163–167.
Drake DE . (1974). Distribution and transport of suspended particulate matter in Submarine Canyons of Southern California. In: Suspended Solids in Water Gibbs RJ, (ed.) Marine Science Springer: Santa Barbara, CA, USA, pp 133–153.
Drake DE, Gorsline DS . (1973). Distribution and transport of suspended particulate matter in Hueneme, Redondo, Newport, and La Jolla Submarine Canyons, California. Geol Soc Am Bull 84: 3949–3968.
Ducet N, Le Traon PY, Reverdin G . (2000). Global high-resolution mapping of ocean circulation from TOPEX/Poseidon and ERS-1 and -2. J Geophys Res 105: 19477–19498.
Eiler A, Hayakawa DH, Rappé MS . (2011). Non-random assembly of bacterioplankton communities in the subtropical North Pacific Ocean. Front Microbiol 2: 140.
Eslinger DL, Cooney RT, McRoy CP, Ward A, Kline TC, Simpson EP et al. (2001). Plankton dynamics: observed and modelled responses to physical conditions in Prince William Sound, Alaska. Fish Oceanogr 10: 81–96.
Ferguson CA, Carvalho L, Scott EM, Bowman AW, Kirika A . (2008). Assessing ecological responses to environmental change using statistical models. J Appl Ecol 45: 193–203.
Fortunato CS, Herfort L, Zuber P, Baptista AM, Crump BC . (2011). Spatial variability overwhelms seasonal patterns in bacterioplankton communities across a river to ocean gradient. ISME J 6: 554–563.
Fouilland E, Tolosa I, Bonnet D, Bouvier C, Bouvier T, Bouvy M et al. (2014). Bacterial carbon dependence on freshly produced phytoplankton exudates under different nutrient availability and grazing pressure conditions in coastal marine waters. FEMS Microbiol Ecol 87: 757–769.
Francois R, Honjo S, Krishfield R, Manganini S . (2002). Factors controlling the flux of organic carbon to the bathypelagic zone of the ocean. Global Biogeochem Cycles 16: 34–1–34–20.
Fu G, Baith K, McClain C . (1998) SeaDAS: The SeaWiFS data analysis system In: Proceedings of the 4th Pacific Ocean Remote Sensing Conference: Qingdao, China, pp 28–31.
Fuessel J, Lam P, Lavik G, Jensen MM, Holtappels M, Guenter M et al. (2012). Nitrite oxidation in the Namibian oxygen minimum zone. ISME J 6: 1200–1209.
Fuhrman J, Steele J . (2008). Community structure of marine bacterioplankton: patterns, networks, and relationships to function. Aquat Microb Ecol 53: 69–81.
Fuhrman JA, Comeau DE, Hagström Å, Chan AM . (1988). Extraction from Natural Planktonic microorganisms of DNA suitable for molecular biological studies. Appl Environ Microbiol 54: 1426–1429.
Fuhrman JA, Hagstrom A . (2008). Bacterial and archaeal community structure and its patterns. In Microbial Ecology of the Oceans. Wiley: Hoboken, NJ, pp 45–90.
Fuhrman JA, Hewson I, Schwalbach MS, Steele JA, Brown MV, Naeem S . (2006). Annually reoccurring bacterial communities are predictable from ocean conditions. Proc Natl Acad Sci 103: 13104–13109.
Galand PE, Casamayor EO, Kirchman DL, Potvin M, Lovejoy C . (2009a). Unique archaeal assemblages in the Arctic Ocean unveiled by massively parallel tag sequencing. ISME J 3: 860–869.
Galand PE, Lovejoy C, Hamilton AK, Ingram RG, Pedneault E, Carmack EC . (2009b). Archaeal diversity and a gene for ammonia oxidation are coupled to oceanic circulation. Environ Microbiol 11: 971–980.
Garcia-Martinez J, Rodriguez-Valera F . (2000). Microdiversity of uncultured marine prokaryotes: the SAR11 cluster and the marine Archaea of Group I. Mol Ecol 9: 935–948.
Gilbert JA, Steele JA, Caporaso JG, Steinbrück L, Reeder J, Temperton B et al. (2012). Defining seasonal marine microbial community dynamics. ISME J 6: 298–308.
Giovannoni SJ, Vergin KL . (2012). Seasonality in ocean microbial communities. Science 335: 671–676.
Hamersley MR, Turk KA, Leinweber A, Gruber N, Zehr JP, Gunderson T et al. (2011). Nitrogen fixation within the water column associated with two hypoxic basins in the Southern California Bight. Aquat Microb Ecol 63: 193–205.
Hamilton AK, Lovejoy C, Galand PE, Ingram RG . (2008). Water masses and biogeography of picoeukaryote assemblages in a cold hydrographically complex system. Limnol Oceanogr 53: 922–935.
Hansen J, Kirboe T, Alldredge A . (1996). Marine snow derived from abandoned larvacean houses: sinking rates, particle content and mechanisms of aggregate formation. Mar Ecol Prog Ser 141: 205–215.
Hatosy SM, Martiny JBH, Sachdeva R, Steele J, Fuhrman JA, Martiny AC . (2013). Beta diversity of marine bacteria depends on temporal scale. Ecology 94: 1898–1904.
Herndl GJ, Reinthaler T . (2013). Microbial control of the dark end of the biological pump. Nat Geosci 6: 718–724.
Hewson I, Steele J, Capone D, Fuhrman J . (2006a). Remarkable heterogeneity in meso- and bathypelagic bacterioplankton assemblage composition. Limnol Oceanogr 51: 1274–1283.
Hewson I, Steele JA, Capone DG, Fuhrman JA . (2006b). Temporal and spatial scales of variation in bacterioplankton assemblages of oligotrophic surface waters. Mar Ecol Prog Ser 311: 67–77.
Hickey BM . (1992). Circulation over the Santa Monica-San Pedro Basin and Shelf. Prog Oceanogr 30: 37–115.
Hickey BM . (1991). Variability in two deep coastal basins (Santa Monica and San Pedro) off southern California. J Geophys Res 96: 16689–16708.
Honaker J, King G . (2010). What to do about missing values in time-series cross-section data. Am J Pol Sci 54: 561–581.
Hooper DU, Chapin FS III, Ewel JJ, Hector A, Inchausti P, Lavorel S et al. (2005). Effects of biodiversity on ecosystem functioning: a consensus of current knowledge. Ecol Monogr 75: 3–35.
Kan J, Suzuki MT, Wang K, Evans SE, Chen F . (2007). High temporal but low spatial heterogeneity of bacterioplankton in the Chesapeake Bay. Appl Environ Microbiol 73: 6776–6789.
King G, Honaker J, Joseph A, Scheve K . (2001). Analyzing incomplete political science data: an alternative algorithm for multiple imputation. Am Pol Sci Rev 95: 49–69.
Kirchman D, K’nees E, Hodson R . (1985). Leucine incorporation and its potential as a measure of protein synthesis by bacteria in natural aquatic systems. Appl Environ Microbiol 49: 599–607.
Kirchman DL . (2002). The ecology of Cytophaga–Flavobacteria in aquatic environments. FEMS Microbiol Ecol 39: 91–100.
Ladau J, Sharpton TJ, Finucane MM, Jospin G, Kembel SW, O’Dwyer J et al. (2013). Global marine bacterial diversity peaks at high latitudes in winter. ISME J 7: 1669–1677.
Lehodey P, Alheit J, Barange M, Baumgartner T, Beaugrand G, Drinkwater K et al. (2006). Climate variability, fish, and fisheries. J Clim 19: 5009–5030.
Lindsay DJ, Hunt JC . (2005). Biodiversity in midwater cnidarians and ctenophores: submersible-based results from deep-water bays in the Japan Sea and north-western Pacific. J Mar Biol Assoc UK 85: 503–517.
Long RA, Azam F . (2001). Microscale patchiness of bacterioplankton assemblage richness in seawater. Aquat Microb Ecol 26: 103–113.
Maidak BL, Cole JR, Lilburn TG, Parker CT Jr, Saxman PR, Farris RJ et al. (2001). The RDP-II (Ribosomal Database Project). Nucleic Acids Res 29: 173–174.
Marinovic BB, Croll DA, Gong N, Benson SR, Chavez FP . (2002). Effects of the 1997-1999 El Nino and La Nina events on zooplankton abundance and euphausiid community composition within the Monterey Bay coastal upwelling system. Prog Oceanogr 54: 265–277.
Mincer TJ, Church MJ, Taylor LT, Preston C, Karl DM, DeLong EF . (2007). Quantitative distribution of presumptive archaeal and bacterial nitrifiers in Monterey Bay and the North Pacific Subtropical Gyre. Environ Microbiol 9: 1162–1175.
Morel A, Gentili B . (2009). A simple band ratio technique to quantify the colored dissolved and detrital organic material from ocean color remotely sensed data. Rem Sens Environ 113: 998–1011.
Morris RM, Vergin KL, Cho JC, Rappe MS, Carlson CA, Giovannoni SJ . (2005). Temporal and spatial response of bacterioplankton lineages to annual convective overturn at the Bermuda Atlantic Time-series Study site. Limnol Oceanogr 50: 1687–1696.
Needham DM, Chow C-ET, Cram JA, Sachdeva R, Parada A, Fuhrman JA . (2013). Short-term observations of marine bacterial and viral communities: patterns, connections and resilience. ISME J 7: 1274–1285.
Nelson JD, Boehme SE, Reimers CE, Sherrell RM, Kerkhof LJ . (2008). Temporal patterns of microbial community structure in the Mid-Atlantic Bight. FEMS Microbiol Ecol 65: 484–493.
Noble RT, Fuhrman JA . (1998). Use of SYBR Green I for rapid epifluorescence counts of marine viruses and bacteria. Aquat Microb Ecol 14: 113–118.
Obernosterer I, Herndl GJ . (1995). Phytoplankton extracellular release and bacterial growth: dependence on the inorganic N: P ratio. Mar Ecol Prog Ser 116: 247–257.
Oksanen J, Blanchet FG, Kindt R, Legendre P, Minchin PR, O’Hara RB et al. (2013). vegan: Community Ecology Package http://CRAN.R-project.org/package=vegan.
Parsons TR . (1984) A Manual of Chemical and Biological Methods for Seawater Analysis 1st edn. Pergamon Press: Oxford [Oxfordshire]: New York.
Partensky F, Hess WR, Vaulot D . (1999). Prochlorococcus, a marine photosynthetic prokaryote of global significance. Microbiol Mol Biol Rev 63: 106–127.
Patel A, Noble RT, Steele JA, Schwalbach MS, Hewson I, Fuhrman JA . (2007). Virus and prokaryote enumeration from planktonic aquatic environments by epifluorescence microscopy with SYBR Green I. Nat Protoc 2: 269–276.
Pinhassi J, Winding A, Binnerup SJ, Zweifel UL, Riemann B, Hagstrom A . (2003). Spatial variability in bacterioplankton community composition at the Skagerrak-Kattegat Front. Mar Ecol Prog Ser 255: 1–13.
Pommier T, CanbäCk B, Riemann L, BoströM KH, Simu K, Lundberg P et al. (2007). Global patterns of diversity and community structure in marine bacterioplankton. Mol Ecol 16: 867–880.
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P et al. (2012). The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41: D590–D596.
Ribalet F, Marchetti A, Hubbard KA, Brown K, Durkin CA, Morales R et al. (2010). Unveiling a phytoplankton hotspot at a narrow boundary between coastal and offshore waters. PNAS 107: 16571–16576.
Rich VI, Pham VD, Eppley J, Shi Y, DeLong EF . (2011). Time-series analyses of Monterey Bay coastal microbial picoplankton using a ‘genome proxy’ microarray. Environ Microbiol 13: 116–134.
Robison BH, Reisenbichler KR, Sherlock RE . (2005). Giant Larvacean Houses: rapid carbon transport to the deep sea floor. Science 308: 1609–1611.
Ruan Q, Steele JA, Schwalbach MS, Fuhrman JA, Sun F . (2006). A dynamic programming algorithm for binning microbial community profiles. Bioinformatics 22: 1508–1514.
Rusch DB, Halpern AL, Sutton G, Heidelberg KB, Williamson S, Yooseph S et al. (2007). The Sorcerer II global ocean sampling expedition: northwest Atlantic through eastern tropical pacific. PLoS Biol 5: e77.
Schnetzer A, Moorthi SD, Countway PD, Gast RJ, Gilg IC, Caron DA . (2011). Depth matters: Microbial eukaryote diversity and community structure in the eastern North Pacific revealed through environmental gene libraries. Deep Sea Res Part I Oceanogr Res Papers 58: 16–26.
Sholkovitz E, Soutar A . (1975). Changes in composition of bottom water of Santa-Barbara Basin—Effect. DeepSea Res 22: 13–1.
Simon M, Grossart H-P, Schweitzer B, Ploug H . (2002). Microbial ecology of organic aggregates in aquatic ecosystems. Aquat Microb Ecol 28: 175–211.
Sohrin R, Isaji M, Obara Y, Agostini S, Suzuki Y, Hiroe Y et al. (2011). Distribution of Synechococcus in the dark ocean. Aquat Microb Ecol 64: 1–14.
Steele JA, Countway PD, Xia L, Vigil PD, Beman JM, Kim DY et al. (2011). Marine bacterial, archaeal and protistan association networks reveal ecological linkages. ISME J 5: 1414–1425.
Steinberg DK, Carlson CA, Bates NR, Goldthwait SA, Madin LP, Michaels AF . (2000). Zooplankton vertical migration and the active transport of dissolved organic and inorganic carbon in the Sargasso Sea. Deep Sea Res Part I Oceanogr Res Papers 47: 137–158.
Steinberg DK, Carlson CA, Bates NR, Johnson RJ, Michaels AF, Knap AH . (2001). Overview of the US JGOFS Bermuda Atlantic Time-series Study (BATS): a decade-scale look at ocean biology and biogeochemistry. Deep Sea Res Part II Top Stud Oceanogr 48: 1405–1447.
Steinberg DK, Goldthwait SA, Hansell DA . (2002). Zooplankton vertical migration and the active transport of dissolved organic and inorganic nitrogen in the Sargasso Sea. Deep Sea Res Part I Oceanogr Res Papers 49: 1445–1461.
Sverdrup HU . (1953). On conditions for the vernal blooming of phytoplankton. J Cons 18: 287–295.
Swan BK, Martinez-Garcia M, Preston CM, Sczyrba A, Woyke T, Lamy D et al. (2011). Potential for chemolithoautotrophy among ubiquitous bacteria lineages in the Dark Ocean. Science 333: 1296–1300.
Teeling H, Fuchs BM, Becher D, Klockow C, Gardebrecht A, Bennke CM et al. (2012). Substrate-controlled succession of marine bacterioplankton populations induced by a phytoplankton bloom. Science 336: 608–611.
Treusch AH, Vergin KL, Finlay LA, Donatz MG, Burton RM, Carlson CA et al. (2009). Seasonality and vertical structure of microbial communities in an ocean gyre. ISME J 3: 1148–1163.
Vergin KL, Beszteri B, Monier A, Thrash JC, Temperton B, Treusch AH et al. (2013a). High-resolution SAR11 ecotype dynamics at the Bermuda Atlantic Time-series Study site by phylogenetic placement of pyrosequences. ISME J 7: 1322–1332.
Vergin KL, Done B, Carlson CA, Giovannoni SJ . (2013b). Spatiotemporal distributions of rare bacterioplankton populations indicate adaptive strategies in the oligotrophic ocean. Aquat Microb Ecol 71: 1–13.
Weinbauer MG, Liu J, Motegi C, Maier C, Pedrotti ML, Dai M et al. (2013). Seasonal variability of microbial respiration and bacterial and archaeal community composition in the upper twilight zone. Aquat Microb Ecol 71: 99–115.
Wilson SE, Steinberg DK . (2010). Autotrophic picoplankton in mesozooplankton guts: evidence of aggregate feeding in the mesopelagic zone and export of small phytoplankton. Mar Ecol Prog Ser 412: 11–27.
Woebken D, Lam P, Kuypers MMM, Naqvi SWA, Kartal B, Strous M et al. (2008). A microdiversity study of anammox bacteria reveals a novel Candidatus Scalindua phylotype in marine oxygen minimum zones. Environ Microbiol 10: 3106–3119.
Wood SN . (2011). Fast stable restricted maximum likelihood and marginal likelihood estimation of semiparametric generalized linear models. J R Stat Soc Ser B Stat Methodol 73: 3–36.
Wood SN . (2006) Generalized Additive Models: An Introduction with R. Chapman & Hall/CRC: Boca Raton, FL.
Wood SN . (2004). Stable and efficient multiple smoothing parameter estimation for generalized additive models. J Am Stat Assoc 99: 673–686.
Wright JJ, Konwar KM, Hallam SJ . (2012). Microbial ecology of expanding oxygen minimum zones. Nat Rev Microbiol 10: 381–394.
Wright JJ, Mewis K, Hanson NW, Konwar KM, Maas KR, Hallam SJ . (2013). Genomic properties of Marine Group A bacteria indicate a role in the marine sulfur cycle. ISME J 8 (2): 455–468.
Yeo SK, Huggett MJ, Eiler A, Rappé MS . (2013). Coastal bacterioplankton community dynamics in response to a natural disturbance. PLoS One 8: e56207.
Zehr JP, Ward BB . (2002). Nitrogen cycling in the ocean: new perspectives on processes and paradigms. Appl Environ Microbiol 68: 1015–1024.
Acknowledgements
This time series would not have been possible without the support of numerous present and past members of the SPOT team. We especially thank David Caron, Dale Kiefer, Roberta Marinelli, Troy Gunderson, Laura Gómez-Consarnau, Diane Kim, Catherine Roney, Anand Patel, Elizabeth Teel, Mahira Kakajiwala, Adriane Jones, Allie Lie, Sarah Hu, Victoria Campbell, Michael Morando, Bridget Seegers, Xiao Liu, Tu-My To, Henry Ho, Kieran Bartholow, Vartis Tsontos, and Tim Lam and the crews of the RVs Seawatch and Yellowfin. We thank William Berelson, Burt Jones, Doug Capone, John Heidelberg, Fengzhu Sun and Sergio Sañudo-Wilhelmy for advice. This work was supported by NSF grant numbers 0703159, 1136818 and from the Gordon and Betty Moore Foundation Marine Microbiology Initiative through grant GBMF3779, and the Wrigley Institute for Environmental Studies.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Cram, J., Chow, CE., Sachdeva, R. et al. Seasonal and interannual variability of the marine bacterioplankton community throughout the water column over ten years. ISME J 9, 563–580 (2015). https://doi.org/10.1038/ismej.2014.153
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2014.153
This article is cited by
-
Host-microbiota-parasite interactions in two wild sparid fish species, Diplodus annularis and Oblada melanura (Teleostei, Sparidae) over a year: a pilot study
BMC Microbiology (2023)
-
Multiyear analysis uncovers coordinated seasonality in stocks and composition of the planktonic food web in the Baltic Sea proper
Scientific Reports (2023)
-
Role of Bacterial Community Composition as a Driver of the Small-Sized Phytoplankton Community Structure in a Productive Coastal System
Microbial Ecology (2023)
-
The importance of microbiota and terrestrial inflows in controlling seston C:N:P:O:Si:Ca:Mn:Mg:Fe:K:Na:Cl:S:Cu:Zn stoichiometry of a deep coastal fjord
Biogeochemistry (2023)
-
Diversity and structure of sparids external microbiota (Teleostei) and its link with monogenean ectoparasites
Animal Microbiome (2022)