Abstract
The majority of nitrogen in forest soils is found in organic matter–protein complexes. Ectomycorrhizal fungi (EMF) are thought to have a key role in decomposing and mobilizing nitrogen from such complexes. However, little is known about the mechanisms governing these processes, how they are regulated by the carbon in the host plant and the availability of more easily available forms of nitrogen sources. Here we used spectroscopic analyses and transcriptome profiling to examine how the presence or absence of glucose and/or ammonium regulates decomposition of litter material and nitrogen mobilization by the ectomycorrhizal fungus Paxillus involutus. We found that the assimilation of nitrogen and the decomposition of the litter material are triggered by the addition of glucose. Glucose addition also resulted in upregulation of the expression of genes encoding enzymes involved in oxidative degradation of polysaccharides and polyphenols, peptidases, nitrogen transporters and enzymes in pathways of the nitrogen and carbon metabolism. In contrast, the addition of ammonium to organic matter had relatively minor effects on the expression of transcripts and the decomposition of litter material, occurring only when glucose was present. On the basis of spectroscopic analyses, three major types of chemical modifications of the litter material were observed, each correlated with the expression of specific sets of genes encoding extracellular enzymes. Our data suggest that the expression of the decomposition and nitrogen assimilation processes of EMF can be tightly regulated by the host carbon supply and that the availability of inorganic nitrogen as such has limited effects on saprotrophic activities.
Similar content being viewed by others
Introduction
Most of the nitrogen (N) in forest soils exists in organic form, primarily as proteins but also as amino sugars and heterocyclic molecules (Nannipieri and Eldor, 2009). These N-containing compounds are found in a complex mixture of plant and microbial degradation products that are present in soil organic matter (SOM). The depolymerization of organic N compounds and complexes is a critical step in N cycling, because it regulates the rate of N movement from SOM into bioavailable forms that can be used by either plants or microbes (Schimel and Bennett, 2004). In boreal forests, mutualistic ectomycorrhizal fungi (EMF) are thought to have a key role in these processes. Indeed, studies of soil mesocosms show that EMF can assimilate the N present in complex organic material (plant litter, pollen grains and necromass of fungal mycelia or soil mesofauna) and are able to transfer a significant fraction of this N to the host plant (reviewed in Read et al. (2004)). Field studies using isotope analyses also provide evidence that EMF mobilize N during litter decomposition in boreal forest soils (Hobbie and Horton, 2007; Lindahl et al., 2007).
Because of their reduced set of genes encoding plant cell wall-degrading enzymes (Martin et al., 2008), the impact of EMF as saprotrophs on the turnover of carbon (C) in forest soils has been considered to be limited. However, we recently showed that the EMF fungus Paxillus involutus was able, while assimilating organic N, to significantly modify organic matter using a free-radical-based mechanism similar to that of saprophytic, brown-rot fungi (Rineau et al., 2012). Unlike saprophytic fungi, P. involutus did not show any expression of genes encoding extracellular enzymes needed to metabolize the released C. This suggests that the degradation mechanism of this EMF fungus has evolved to assimilate organic N rather than C; and that P. involutus no longer has the ability to assimilate C as an adaptation for symbiotic growth on host plants. The C delivered by the plant partner is in the form of soluble hexoses, most likely glucose (Nehls et al., 2010). Our first hypothesis was therefore that the addition of glucose would significantly stimulate degradation of organic matter by P. involutus.
The fact that the decomposition of SOM by EMF is linked to the degradation and mineralization of organic N suggests that the degradation activity should be controlled by environmental factors known to regulate the activity of enzymes involved in the breakdown of soil organic N sources such as proteins. In most filamentous fungi, including the EMF fungus Hebeloma crustuliniforme, the expression of proteolytic activity is repressed in the presence of a readily available N source such as ammonium (Zhu et al., 1994; Marzluf, 1996). Moreover, studies in saprophytic fungi have shown that N availability may also affect the expression levels of lignocellulolytic enzymes including cellulases, peroxidases and laccases (Fenn and Kirk, 1981; Chen et al., 2003; Aro et al., 2005; Edwards et al., 2011). Hence, our second hypothesis was that the presence of ammonium (the most abundant inorganic N form in forest soils) would downregulate SOM degradation by P. involutus.
To test these two hypotheses, in this study we examined how the decomposition of litter material and the assimilation of N from organic matter by P. involutus are regulated by glucose and ammonium. The modification of organic matter was measured using a combination of spectroscopic techniques, and the transcriptional response of the fungus was assessed using DNA microarray analyses. The results showed that the decomposition of the organic matter and the assimilation of N by P. involutus were significantly stimulated by glucose addition, whereas modifying the organic matter by adding inorganic N had only a minor effect. Among the genes that were upregulated by glucose, we identified a core set that encodes 39 enzymes whose expression levels were correlated with the degradation of organic litter material. Our findings support the hypothesis that the plant photosynthate, and ammonium to a lesser extent, controls the decomposition activity of EMF.
Materials and methods
Fungal strain and culture conditions
Cultures of P. involutus (Batsch) Fr. (strain ATCC 200175, Manassas, VA, USA) (Basidiomycota, Boletales) were maintained aseptically on 1.5% agar plates containing minimum Melin–Norkrans medium (MMN) (composition: 2.5 g l−1 glucose, 500 mg l−1 KH2PO4, 200 mg l−1 NH4Cl, 150 mg l−1 MgSO4.7H2O, 25 mg l−1 NaCl, 50 mg l−1 CaCl2, 12 mg l−1 FeCl3.6H2O and 1 mg l−1 thiamine-HCl; pH 4.0). The fungus was grown in petri dishes on a layer of glass beads immersed in liquid medium (Rineau et al., 2012). After 9 days of incubation (18 °C and in the dark), the MMN medium was removed with a sterile pipette. The glass beads and the mycelium were washed with 10 ml of sterile MilliQ (MQ) water, and 10 ml of MMN medium without N were added to induce a N-deprived mycelium. After 24 h, the mycelium was again washed in MQ water, and finally the organic matter extracts (10 ml) were added.
Organic matter was extracted from maize compost material by boiling 120 g of compost in 600 ml of MilliQ water for 1 h. Particles were removed by filtering and low molecular weight metabolites were removed by ultrafiltration (cutoff 10 kDa) (Rineau et al., 2012). The degradation of this material was studied in the presence/absence of an easily assimilated C source (glucose, 2.5 g l−1, providing 1 g l−1 of elemental C), and an inorganic N source (ammonium supplied as NH4Cl, 0.2 g l−1, providing 53 mg l−1 of elemental N) using four different conditions: (i) organic matter alone (OM), (ii) organic matter and glucose (OM+G), (iii) organic matter and NH4Cl (OM+N) and (iv) organic matter, glucose and NH4Cl (OM+G+N). The extracts were sterilized after addition of glucose and NH4Cl by filtration at 0.2 μm. Each treatment was then run in triplicate (one replicate was the result of pooling medium from three-petri dishes; we therefore had nine-petri dishes per treatment); each replicate is referred to as ‘sample’ throughout. The concentrations of added glucose and NH4 were similar to the ones in the MMN medium. The cultures were incubated for 7 days at 18 °C in the dark. Samples where the fungus has been growing for 7 days are referred to as ‘inoculated’ samples, while the non-inoculated OM extract are referred to as ‘reference’.
Chemical analysis
Samples of reference and inoculated material were collected in sterile Falcon tubes and diluted eightfold (in order to contain between 0 and 100 mg l−1 of C). A fraction of the material was oxidized by combustion at high temperature (720 °C) and total organic carbon concentration (TOC) was measured using a TOC Analyzer (Shimadzu, Kyoto, Japan). Total nitrogen content was measured using the same apparatus with a TNM-1 detector. The assimilation of N by P. involutus was estimated by subtracting the total N content of the substrate at the start of the experiment and after 7 days of incubation.
Fourier Transform Infrared (FTIR) spectra were recorded using a Bruker IFS66 v/s spectrometer (Bruker Scientific Instruments, Billerica, MA, USA). Data were collected in diffuse reflectance mode. Each spectrum was the result of 1000 consecutive scans at a resolution of 4 cm−1. Synchronous fluorescence spectra were recorded using a PerkinElmer LS50B spectrophotometer (Perkin-Elmer, Waltham, MA, USA). Pyrolysis-gas chromatography/mass spectrometry was performed using a PerkinElmer TurboMass/Autosystem XL with Frontier Lab double Shot pyrolyser (Perkin-Elmer). As addition of NH4 can bind to different organic molecules during the pyrolysis step in an unpredictable way (Nierop and Van Bergen, 2002), it was impossible to calculate the proportions of guaiacol, syringol, propenyl-guaiacol and 3-methylcatechol, which are proxies of lignin oxidation, in the extracts. Instead, the ratios of oxidized guaiacol to guaiacol and of oxidized syringol to syringol were calculated. Further details are given in Rineau et al. (2012).
Microarray experiments
Fungal mycelia from each of the treatments (OM, OM+G, OM+N and OM+G+N) and the replicates were collected from the bead plates and immediately placed into a clean mortar filled with liquid N2 and homogenized using a pestle. Total RNA was isolated using the RNeasy Plant Mini Kit (Qiagen, Venlo, Netherlands), the RLC buffer and the one-column DNase treatment according to the manufacturer’s instructions. Total RNA was eluted in either 60 or 100 μl of H2O and stored at −20 °C. For quality and concentration assessment all samples were tested using a 2100 Bioanalyzer and a RNA 6000 Nano kits (Agilent, Santa Clara, CA, USA).
We used a custom-designed microarray (12-plex 135K oligonucleotide microarray, Design ID: 546871) (NimbleGen/Roche, Basel, Switzerland) with probes representing 12 214 transcripts (isotigs) obtained by 454/Roche DNA sequencing of transcriptomes expressed by P. involutus during growth on various OM extracts and reference MMN medium (Rineau et al., 2012). Sanger sequencing (Applied BioSystems, Foster City, CA, USA) of expressed sequence tags expressed in free-living mycelium, mycorrhizal roots and non-mycorrhizal roots (Le Quéré et al., 2005) were also used. Each isotig was represented with up to 10 probes (60-mers) in a tiled design. This complete design has been deposited at the NCBI Gene Expression Omnibus (GEO) (Edgar et al., 2002) (accession code GPL14950). The isotig sequences are also available at the Paxillus EST database: http://mbio-serv2.mbioekol.lu.se/Paxillus/Hybrid/ (to search the database add ‘paxillus_’ to the isotig ID) and at NCBI GenBank (accession code SRA046093). The microarray analyses were performed as single-label hybridizations. For each hybridization and each sample 10 μg of total RNA was used for cDNA synthesis using the SuperScript Double-Stranded cDNA Synthesis Kit (Invitrogen, Carlsbad, CA, USA) according to the manufacturer’s instructions. For quality assessment, the produced cDNA was analyzed on a 2100 Bioanalyzer and DNA 7500 kit (Agilent). For sample labeling the One-Color DNA Labeling Kit (Cy3) (NimbleGen/Roche) was used according to the manufacturer’s instructions. After labeling, each sample received a Sample Tracking Control (NimbleGen/Roche) and hybridizations were immediately performed in a Hybridization System 4 (NimbleGen/Roche) for at least 16 h and washed according to the manufacturer’s instructions. The slides were scanned using an Agilent High-Resolution Microarray Scanner set at 2 μm resolution. The raw images were burst and processed using the NimbleScan software v. 2.5 (NimbleGen/Roche) and the built-in Robust Multichip Average algorithm including quantile normalization, to remove the effects of systematic variation in the measured fluorescence intensities (Bolstad et al., 2003; Irizarry et al., 2003a, 2003b). To this data set we also introduced previously published data representing six biological replicates on MMN medium (Rineau et al., 2012) that are available at NCBI GEO (accessions GSM848412-GSM848414 and GSM848421-GSM848423). Normalized (log2-transformed) transcriptional values were brought into the Omics Explorer ver. 2.2 (Qlucore, Lund, Sweden) for PCA and statistical analyses. In the analysis, we removed 686 isotigs that had hybridizations signals close to the background levels (<log2=5.0) and were found only in the Sanger sequenced libraries containing host plant material (thus, 11 528 isotigs were analyzed). The data for OM, OM+G, OM+N and OM+G+N have been deposited at NCBI GEO and are accessible through GEO Series accession number GSE45303.
Bioinformatics analysis
We have previously identified 267 transcripts among the 12 214 isotigs that encode enzymes and proteins with a possible role in the degradation of lignocellulose (polysaccharide modifications, lignin degradation, iron reduction and homeostasis, or oxalate metabolism and H2O2 production) (Rineau et al., 2012). Transcripts potentially encoding peptidases (including proteases) were retrieved by searching the MEROPS database (Rawlings et al., 2012) for putative homologs using the P. involutus transcript sequences as a query. This returned a list of 312 transcript sequences of putative peptidases including 26 aspartic, 73 cysteine, 139 metallo, 63 serine, 17 threonine and one glutamic acid peptidase (Shah et al., unpublished work). Nitrogen (N) transporters were identified using annotation from the genome of the EMF Laccaria bicolor (Basidiomycota, Agaricales) (Lucic et al., 2008) (Supplementary Table S1, Supplementary Information). To identify transcripts encoding enzymes in the carbohydrate and amino-acid metabolism pathways, information of the L. bicolor genome was again searched and relevant gene models were used as queries to identify homologs in the P. involutus EST database (Supplementary Table S2). Prediction of the presence of secretory signal peptides was performed using the SignalP 3.0 algorithm (Emanuelsson et al., 2007). The probability (P) of observing a given number of transcripts within a given functional category by chance was estimated using the hypergeometric distribution (Arvas et al., 2007).
To infer the regulation of metabolic pathways from the log2-transformed normalized microarray data, we used the relative expression compared with the mineral medium (MMN) for each of the transcripts and treatments. We estimated the regulation of a pathway as the mean expression change of its transcripts, but applied our prior knowledge and hypotheses to make the estimate more robust. Each pathway consists of several Enzyme Commission (EC) numbers, which in turn may be represented by several transcripts. We can write the mean expression level for a pathway as a weighted mean over its EC numbers, with weights equal to the number of transcripts in each. However, if there is complete redundancy between transcripts that have the same EC number, all weights should be 1. Expecting some redundancy, we used the geometric mean of the two options (that is, we weighted each EC number by the square root of its number of transcripts). EC numbers that appear in multiple pathways may be less relevant in regulating the individual pathways. To avoid assigning these EC numbers undue importances, we divided their weights by the number of pathways they appear in. Other EC numbers were manually identified as likely to be more important in regulating a pathway, for example, in regulatory steps or in the first step of a degradation or transformation pathway. We gave these EC numbers a threefold higher weight. These weight corrections allow us to compromise between the more extreme choices of discarding or keeping the less-trusted EC numbers.
Correlation of gene expression with substrate modifications
Canonical correspondence analysis (CCA) was used to correlate expression patterns with modifications in the substrate. This statistical tool represents objects (isotigs) and variables (gene expression levels) in a new system of coordinates that (i) depicts as much of the variation from the original data set as possible and (ii) consists of linear combinations of explanatory variables (modifications of the substrate) (Ramette, 2007).
Transcripts with expression levels strongly correlated with each type of substrate modification (arrows in the CCA plot, Figure 6) were selected using the following procedure. The transcripts’ coordinates on the CCA plot were orthogonally projected on the substrate modification arrows, generating an orthogonal coordinate of each transcript along each arrow. This gives a good approximation of the degree of correlation between gene expression level and the intensity of substrate modification (Ramette, 2007). For each substrate modification type, the transcripts were sorted based on their orthogonal coordinate, and only the most positively correlated transcripts were retained (having an orthogonal coordinate of at least half of the maximal orthogonal coordinate measured for this axis). This selection procedure retained at most 20 transcripts per variable and was considered more biologically relevant than the more mathematical procedure selecting only the top 5% of the normally distributed values, which resulted in a list of only two transcripts and was therefore judged too conservative. The statistical calculations and representations were done using the R language (http://www.r-project.org) (Ihaka and Gentleman, 1996).
Results
Substrate modifications
The percentages of N taken up from the OM, OM+N, OM+G and OM+G+N substrates were 5.3% (removal of 5 mg l−1 of N; s.d.=2), 5.3% (7 mg l−1 of N; s.d.=2), 50.3% (49 mg l−1 of N; s.d.=3) and 54.4% (39 mg l−1 of N; s.d.=1 ), respectively. Thus, the assimilation of N was significantly enhanced by adding glucose to the OM extracts at the beginning of the incubation. We have previously shown that the added glucose was not detected in the OM+G treatment at the end of the incubation (Rineau et al., 2012). The mycelium grew in all treatments, but the growth enhancement was largest in the media containing glucose (Supplementary Figure S1).
The FTIR spectra of the OM and OM+N media before and after inoculation were almost identical (Figures 1a and b), which indicates that the chemical structures of these substrates were not significantly altered during the incubation. Note that the spectral discrepancies between these media are due to the added ammonium ions displaying a strong band at 1402 cm−1. In contrast, adding glucose (OM+G) resulted in pronounced differences between FTIR spectra collected before and after incubation (Figures 1c and d). The changes observed in the sugar modes (970–1200 cm−1) originate predominantly from glucose that is completely degraded after incubation. These changes were accompanied by an increase of relative band intensities in the region 1500–1800 cm−1 and a decrease at 1350–1450 cm−1. These regions contain carboxyl+carbonyl modes and nitrate+ammonium modes, respectively (Rineau et al., 2012). The most pronounced intensity increase was observed at 1595 and 1665 cm−1. Similar, but not identical, modifications of the FTIR spectra were obtained in the treatments combining addition of glucose and NH4+ (OM+G+N). In this case, the 1720 cm−1 band displayed the most substantial relative increase.
FTIR spectra of the samples before (red lines) and after 7 days (black lines) of incubation with P. involutus for each treatment. (a) Organic matter extract (OM); (b) Organic matter extract+NH4+ (OM+N); (c) Organic matter extract+glucose (OM+G); and (d) Organic matter extract+glucose+NH4+ (OM+G+N). Each system includes three-inoculated samples and one reference sample. All spectra have been normalized to the same total area over the wavenumber region displayed.
Synchronous fluorescence spectroscopy was used to obtain complementary information about the changes in the abundances and the chemical structures of aromatic compounds in the samples. The most prominent peak in all extracts was in the 330–360 nm region, which is associated with compounds containing two condensed aromatic rings, and a peak located at 360–400 nm, which can be attributed to more complex aromatic ring systems bearing carbonyl or carboxyl groups (Rineau et al., 2012). No prominent changes were detected in the intensity of this peak after incubation in the OM and OM+N extracts (Figure 2), while addition of glucose (OM+G) resulted in a major decrease in its size. In contrast to the findings using FTIR, these spectral changes were not detected after adding NH4+ (OM+G+N). The fluorophores affecting the synchronous fluorescence spectra are thought to constitute only a minor component in humic substances, and the signal varies extensively with the nature of the fluorophore (Miano and Senesi, 1992). Thus, the decrease in fluorescence intensity in the ammonium-containing samples might be due to small chemical changes in the structure of aromatic compounds that cannot be detected by FTIR spectroscopy.
Pyrolysis-GC/MS data showed that the ratio of oxidized guaiacol to guaiacol was significantly higher in the samples where glucose has been added (OM+G and OM+G+N) (Figure 3), showing that guaiacol, a lignin constituent, was oxidized in these samples. However, we did not detect any significant oxidation of syringol, another lignin constituent.
Oxidation of lignin-building blocks measured by Py GC/MS. The size of the peaks from molecules issued from pyrolysis of oxidized guaiacol (OxG) have been summed and divided by the size of the peaks from molecules issued from pyrolysis of non-oxidized guaiacol (G). The same calculations were made with oxidized syringol (OxS) and non-oxidized syringol (S). n=3, error bars denote s.e.. Analysis of variance (ANOVA) showed a significant effect of treatment on guaiacol oxidation (P=0.01) but no significant effect on syringol oxidation. Different letters denote significant differences (P<0.05) based on Tukey post hoc test for comparisons between means across the four treatment combinations.
Transcriptional responses
Microarray analysis showed that the transcriptome profile of the mycelium of P. involutus grown on the OM extracts was distinctly different from that grown on the MMN medium (Figure 4a). The PCA analysis showed that the glucose-containing samples (OM+G and OM+G+N) were clearly separated from those without glucose (OM and OM+N). Scatter plots of the gene expression in mycelium grown on the OM substrates versus MMN showed that a majority of genes had similar expression levels (Figure 4b). Compared with the MMN medium, 962 transcripts were significantly upregulated at least twofold on the OM+G medium, 906 on the OM+G+N medium, 122 on the OM medium and 64 on the OM+N medium (false-discovery rate, q⩽0.01). Transcripts predicted to encode enzymes involved in the degradation of lignocellulose and peptidases, and enzymes of the C and amino-acid metabolism, were significantly enriched among the twofold upregulated genes in the treatments with glucose (Figure 4c).
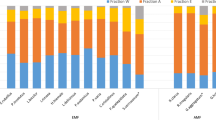
Transcriptional responses in the mycelium of P. involutus when grown in organic matter (OM) extracts, with or without glucose (G) and with or without NH4+ (N) as shown by microarray analysis. (a) Principal component analysis (PCA) was performed on the expression levels of 10 443 transcripts out of 11 528 analyzed on the DNA microarray that had a false-discovery rate of q⩽0.01. MMN, minimum Melin–Norkrans (that is, mineral nutrient) medium. n=3, except for MMN (n=6); each point in the PCA represents a replicate. (b) Differentially expressed genes in OM versus MMN. The scatter plots show the intensity of the normalized and log2-transformed hybridization signals. The diagonal line (y=x) shows genes with near identical hybridization signals. The lines at y=x+1 and y=x−1 correspond to a log2 relative expression between the OM substrates and MMN of +0.5 and −0.5, respectively. The numbers of genes upregulated more than twofold (q⩽0.01) are shown in parentheses. (c) The numbers of transcripts in various functional categories that were upregulated more than twofold (q⩽0.01) in pairwise comparisons of transcription levels after growth in media containing OM versus MMN. The total numbers of transcripts annotated to each functional category among the 11 528 probe sequences are given in parentheses. The significance of the enrichments are shown in Supplementary Table S3.
Only 17 transcripts were differentially regulated in the pairwise comparison of OM+G+N versus OM+G (q⩽0.01). The magnitude of regulation was small; only two transcripts were upregulated and six were downregulated more than twofold (Supplementary Table S4). Two of the downregulated transcripts encoded proteins with a putative role in the assimilation of N; an oligopeptidase and an OPT oligopeptide transporter protein. No transcripts were significantly down- or upregulated when ammonium was added to the OM extracts containing no glucose (that is, comparing OM+N with OM).
The transcriptional responses in the C and N metabolism were further analyzed by estimating the expression levels of specific metabolic pathways (Figure 5). Adding glucose to the OM extracts significantly stimulated the expression of the central carbohydrate pathways: glycolysis, pyruvate metabolism and TCA (citric acid) cycle as well as several linked pathways such as fructose mannose metabolism, amino sugar and nucleotide sugar metabolism and the pentose phosphate pathway. Concomitantly, various pathways in amino-acid metabolism were upregulated, including alanine, aspartate and glutamate metabolism; arginine, aspartate and proline metabolism; histidine metabolism; valine, leucine and isoleucine biosynthesis; cysteine and methionine metabolism; phenylalanine, tyrosine and tryptophan metabolism; and lysine biosynthesis. Several of the amino-acid degradation pathways, such as the lysine degradation and valine, leucine and isoleucine degradation, were upregulated when the fungus was grown in the media containing the OM substrates.
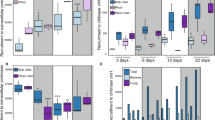
Transcriptional regulation of pathways in the carbohydrate and nitrogen metabolism of P. involutus. Each transcript was annotated with an EC number, and the change in expression level between the sample and MMN was averaged for each EC number. These relative expressions were combined with EC-dependent weights into a weighted mean and s.e.m. for each pathway (green-black-red boxes; the middle of each box shows ±s.e.m.). Red shows upregulation compared with MMN. Asterisks below the colored boxes indicate whether a majority of the EC numbers (and their transcripts) are regulated in the same direction as with MMN, without regard to the magnitude of the regulation (*P<0.01, **P<0.001).
In total, 20 out of the 129 transcripts encoding putative N transporters were upregulated more than twofold in at least one of the pairwise comparisons between media containing OM with MMN. Apart from the YaaH family and the peptide transporter (PTR) family, upregulated genes were found in all other families of N transporters including those for amino acids, oligopeptides, ammonium, urea, polyamine and allantoate/allantoin (Supplementary Table S5).
Correlating substrate modification and gene expression
In total, 308 transcripts encoding lignocellulose degrading enzymes and peptidases were significantly (q⩽0.01) upregulated when comparing the transcriptome of P. involutus grown in the OM, OM+N, OM+G and OM+N+G media. A CCA analysis showed that the expression levels of a subset of these lignocellulose and protein-modifying enzymes were positively correlated with the modification of the OM material. The first axis in the CCA plot explained a large part (75%) of the variance, and clearly separated the treatments with and without added glucose, while the second axis differentiated the treatments with and without ammonium (Figure 6). The addition of glucose was therefore the parameter that most strongly modulated the pattern of expression of genes involved in OM degradation, and inorganic N addition had only a minor effect on the regulation of those genes. The arrows representing most of the substrate modification categories (TOC, total nitrogen and modifications of the carboxyl/carbonyl, ammonium/nitrate and sugar regions in the FTIR spectra, or oxidized guaiacol/guaiacol ratio) were also strongly correlated with the first (horizontal) axis, and therefore with glucose addition. Modifications in the aliphatic region of the FTIR spectra and of the condensed/substituted aromatic molecules were positively and negatively correlated with ammonium addition (vertical axis), respectively.
CCA of expression level of the lignocellulose and protein-modifying genes and measurements of substrate modification. Each black dot represents an isotig (transcript). TOC, decrease in total organic carbon concentration; TN, decrease in total nitrogen concentration; condensed/substituted aromatic (SF), decrease in size of the highest aromatic peak (synchronous fluorescence); [COOH/C=O] [NH4/NO3] [Sugar] (FTIR), change in shape of the FTIR peaks in the corresponding regions; OxG/G(Py) and OS/S(Py), ratio of oxidized on non-oxidized guaiacol (G) and syringol (S), respectively, measured by pyrolysis-GC/MS (Py). The isotigs whose expression levels were most correlated with the condensed/substituted aromatic (SF) variable are represented by black diamonds, the aliphatic (FTIR) variable by black triangles and a group of variables (OxG/G(Py), [COOH/C=O] [NH4/NO3] [sugar] (FTIR), TOC and TN) by gray disks. Each gray square represents a sample. The correlations between gene expression levels and measured substrate modifications are presented as dashed arrows. Substrate modifications were sorted into seven categories. The percentage of total variance explained by each canonical component is indicated in parentheses.
On the basis of the CCA analysis, we identified 39 transcripts whose expression patterns were most correlated with the modifications of the substrate (Figure 7). These were divided into three groups. The first contained 20 transcripts with expression levels positively correlated with glucose addition (and therefore with the modification carboxyl/carbonyl, ammonium/nitrate and sugar regions, with TOC and total nitrogen decrease, and with oxidized guaiacol/guaiacol ratio). These transcripts consisted of eight lignin-degrading enzymes, four proteases, three endoglucanases, three Fenton-reaction related transcripts, one microbial cell wall degrading enzyme and one polysaccharide degrading enzyme (GH5: mannosidase). The second group contained seven transcripts with expression levels positively correlated with both glucose and N addition (modification of the aliphatic C–C region, labeled ‘A (FTIR)’). These transcripts consisted of three lignin-degrading enzymes, two proteases, one plant cell wall degradation enzyme and one β-glucosidase. The third group contained 12 transcripts with expression levels negatively correlated with N addition (aromatic signal in synchronous fluorescence). These transcripts consisted of six lignin-degrading enzymes, four Fenton-reaction related transcripts and two proteases.
Regulation of the lignocellulose and protein-modifying genes that were most correlated with substrate modification. The degree of correlation was calculated by orthogonal projection of the transcript coordinates on the variable arrow. The regulation pattern was calculated as a ratio between the average expression in the OM, OM+N, OM+G or OM+N+G and MMN (n=3). Relative expressions are indicated both as numbers and by a color code (darker indicates a higher relative expression). S=N-terminus with predicted secretion signal; NS, no secretion signal; NTM, sequences likely to have an incomplete N-terminus). *Indicates the ID of the family in the MEROPS peptidase database (Rawlings et al., 2012). The projection column gives a score that is proportional to the degree of correlation of the gene expression levels with the substrate modification category (cf. Figure 6).
Discussion
We recently showed that the EMF fungus P. involutus degrades polysaccharides and modifies the structures of polyphenols when acquiring N from litter material, using a trimmed brown-rot biodegradation mechanism (Rineau et al., 2012). As the fungus did not express the glycoside hydrolases (GHs) needed for decomposing the released polysaccharide fragments, we proposed that the saprophytic activity is dependent on sugars from the host plant. Assuming that glucose is the main form of plant C supply to the EMF fungus (Nehls et al., 2010), data from this study supports this hypothesis by demonstrating that glucose addition triggered the Fenton-based decomposition of organic litter extracts and N assimilation by P. involutus. The spectroscopic analyses showed that the litter material was not degraded without adding glucose. The triggering of the decomposing activity was not related to growth only because the mycelium was growing—albeit at slower rates—in cultures without glucose additions. Concomitant with the stimulation of the decomposing activity, the fungus differentially expressed a large number of transcripts encoding enzymes that are involved in the degradation process and in the mobilization of N.
The physiological mechanism providing the stimulating effect of glucose is not known. Adding glucose to the organic material significantly stimulated the expression levels of metabolic pathways involved in energy generation (glycolysis, pyruvate metabolism and the TCA cycle). Transcripts encoding the enzymes mediating the oxidative parts of the pentose phosphate pathway were also upregulated in the glucose-containing samples (Supplementary Table S6). Therefore, apart from generating energy, the added glucose also most likely stimulated the production of reducing power (NADPH) needed for biosynthetic reactions. Another possibility is that the added glucose triggered the activity of extracellular enzymes involved in the oxidation of the SOM. Such mechanisms have been proposed to explain the so-called priming effects, that is, the stimulation of SOM turnover caused by the addition of easily available organic C or N sources (Bengtson et al., 2012). It therefore remains to be determined whether the triggering effect of glucose on the decomposition activities in P. involutus is due to internal or external mechanisms, and whether any low molecular weight substrate can act as trigger.
The chemical modifications of the OM occurring in the presence of glucose were correlated with the expression of several oxidases such as multicopper oxidases (including laccases), cytochrome-p450 monooxygenases, a heme peroxidase, and an aromatic peroxigenase. Heme peroxidase is an enzyme that degrades high-redox potential substrates, such as lignin-building blocks (Liers et al., 2011), and is therefore likely involved in the guaiacol oxidation observed in the glucose-containing OM. This hypothesis was supported by the fact that its expression level was highly correlated with the modification of the substrate. Moreover, peroxidase activity has been recently linked to EMF species richness in bulk soil (Talbot et al., 2013). Aromatic peroxygenases have an ether-cleaving activity (Hofrichter and Ullrich, 2010), and therefore could break ether links within polyphenolic polymers or between polysaccharides and polyphenols (Baldrian, 2008). However, their potential contribution to the OM degradation observed here is difficult to assess, because ether bonds are also found in sugars and the high amount of glucose added to the OM+G and OM+G+N media probably masked any ether cleavage in the FTIR spectra. Glyoxal oxidase has been extensively studied in saprotrophic fungi and is associated with wood degradation by generation of H2O2 (Kersten, 1990). In the Fenton reaction, this H2O2 reacts with Fe2+ to generate highly-oxidizing free radicals. The overexpression of glyoxal oxidases in the OM-containing glucose suggests that the fungus was producing H2O2, and therefore supports the hypothesis that glucose addition and the fungal-induced Fenton reaction are correlated. The degradation of OM-containing glucose was also correlated with the expression of three GHs with a possible role in plant cell wall degradation, one belonging to the GH9 family and two to the GH61 family. The endoglucanase (GH9) was the only potential cellulase sensu stricto. The predicted protein lacks a carbohydrate-binding module and is therefore expected to be a processive endoglucanase (Zhang et al., 2010). The two GH61s are cellulose-oxidizing enzymes and are thus good candidates for a role in plant cell wall degradation (Langston et al., 2011).
The addition of glucose was correlated with a coordinated expression of gene-encoding enzymes and transporters of all the major steps of organic N assimilation by EMF. This included degradation of proteins, uptake of released amino acids and peptides, and internal transformation of amino acids (Chalot and Brun, 1998; Talbot and Treseder, 2010). Among the most upregulated genes encoding extracellular proteases were two aspartate proteases (Figure 7), which displayed sequence similarity to the aspartic endopeptidase AmProt1 of Amanita muscaria (Nehls et al., 2001). A gene encoding an endopeptidase of the fungalysin metallopeptidase (M36) family was also upregulated. Transcripts of several putative amino-acid and oligopeptide transporters were upregulated, which suggests that both amino acids and peptides were assimilated by the fungus. The most upregulated amino-acid transporter (Supplementary Table S5) was a member of the yeast amino-acid transporter (APC/YAT) family, and is homologous to the high-affinity amino-acid transporter AAT1 (Nehls et al., 1999). High expression of intracellular peptidases (a S9 prolyl oligopeptidase and a glutamate carboxypeptidase) indicates that the internalized peptides were hydrolyzed before being metabolized in the amino-acid pathways.
Glutamate/glutamine, aspartate/asparagine and alanine are thought to be the main sinks for the N assimilated by EMF (Martin and Canet, 1986; Finlay et al., 1988). Many of the genes encoding enzymes in the metabolism of these amino acids were stimulated in the OM medium with added glucose (Supplementary Table S6). One of the most upregulated genes in amino-acid metabolism encoded a NADPH-dependent glutamate synthase (EC 1.4.1.13), which together with glutamine synthetase (EC 6.3.1.2), is a key enzyme in ammonium assimilation in most EMF including in P. involutus (Morel et al., 2006). The glucose addition correlated with upregulation of glutamine-fructose-6-phosphate transaminase (EC 2.6.1.16), amidophosphoribosyl transferase (EC 2.4.2.14) and carbamoyl-phosphate synthase (EC 6.3.5.5). This suggests that the N of glutamine and glutamate were utilized for the synthesis of amino sugars and heterocyclic aromatic compounds (purine and pyrimidine). Upregulation of aminotransferases such as aspartate transaminase (EC 2.6.1.1), alanine transaminase (EC 2.6.1.2) and alanine-glyoxylate transaminase (EC 2.6.1.44) indicates an active translocation of amino groups. Analysis of the expression patterns of enzymes in the alanine, aspartate and glutamate metabolism pathways also provided evidence that the assimilated amino acids were utilized as carbon sources by P. involutus. The most upregulated transcript in the pathway encoded a glutamate dehydrogenase (EC 1.4.1.2), which catalyzes the deamination of glutamate to ammonium and oxoglutarate. Degradation of glutamate into TCA intermediates was also indicated by the upregulation of succinate-semialdehyde dehydrogenase (EC 1.2.1.16).
Readily available N sources such as ammonium have been shown to affect the expression of many fungal enzymatic activities involved in the decomposition of protein and plant-litter material (Zhu et al., 1994; Marzluf, 1996; Chen et al., 2003; Aro et al., 2005; Baldrian and Valaskova, 2008; Piscitelli et al., 2011). Ammonium has also been shown to repress the expression of amino-acid transporters and enzymes in the N assimilation pathways in EMF (Nehls et al., 1999; Javelle et al., 2003). Therefore, the limited transcriptional response in P. involutus when adding ammonium to the OM-containing glucose was not anticipated. However, we observed that the addition of ammonium- and glucose-induced specific modifications on OM, compared with addition of glucose alone, for which the aliphatic signal (observed in the FTIR spectra) decreased and the reduction of the condensed/substituted aromatic peak (in the synchronous fluorescence spectra) was repressed. These two OM modifications were accompanied by the regulation of two specific sets of genes. Aliphatic signature modification was correlated with elevated expression of an aspartic protease, a beta-glucosidase and a GH30, and repression of condensed/substituted aromatic modification with the downregulation of many lignin-degrading enzymes, including laccases.
Laccases have been used as a proxy of ligninolytic activity in the field. Recently Edwards et al. (2011) showed that elevated N deposition repressed the expression of a laccase gene, and therefore concluded that N addition slowed down lignin decomposition. In contrast, our results show that the expression levels of gene-encoding laccases are in fact not an adequate proxy of ligninolytic activity, because each fungus has many different laccases and they can perform many different functions that are not always related to lignin degradation (Baldrian, 2004). Indeed, we show that P. involutus expressed two types of laccase: one type (three genes) was unaffected by NH4 addition and involved in lignin modification (oxidation of guaiacol); the other (two genes) was downregulated by NH4 addition but its expression level was not correlated with lignin modification.
Our results also suggest that the regulation of the processes that decompose the plant cell wall may differ significantly between saprophytic and EMF. In saprophytic fungi, the expression of plant cell wall-degrading enzymes, including cellulases, hemicellulases, ligninases and pectinases, is commonly repressed in the presence of glucose (for a review, see Aro et al. (2005)). Furthermore, several studies of the brown-rot fungus Postia placenta have shown that the oxidative degradation of cellulose and wood substrates can occur without glucose being added to the medium (Martinez et al., 2009; Vanden Wymelenberg et al., 2010). We suggest that both a reduction in gene content of plant cell wall-degrading enzymes (Martin et al., 2008; Eastwood et al., 2011) and mutations affecting the transcriptional regulation of extracellular oxidases and peptidases have contributed to the symbiotic adaptations of the decomposing mechanism in EMF.
The accumulating evidence that mycorrhizal fungi may decompose SOM have led to a new model of soil C cycling in which mycorrhizal fungi influence both the inputs and losses of soil C from ecosystems (Talbot et al., 2008). Decomposition by mycorrhizal fungi implies that soil C balance is subjected to ecological factors that affect both the plant and fungal partners. Based on this concept Talbot et al. (2008) proposed three hypothesis for mechanisms by which mycorrhizal fungi act as decomposers. The first hypothesis (‘Plan B’ hypothesis is that mycorrhizal fungi metabolize significant quantities of SOM when the supplies of plant photosynthate is low. The second hypothesis (‘Coincidental Decomposer’ hypothesis is that mycorrhizal fungi decompose soil C as a consequence of mining SOM for organic nutrients. The third hypothesis (‘Priming effects’ hypothesis) is that mycorrhizal fungi decompose SOM when allocation of plant C to mycorrhizal roots is high. The findings in this study provide supports for both the ‘Priming effects’ and ‘Coincidental Decomposer’ hypotheses. Results from field studies based on isotope analysis (Hobbie and Horton, 2007; Lindahl et al., 2007), as well as enzyme assays (Talbot et al., 2013) are also in agreement with the hypothesis that the decomposing activity of EMF are largely devoted to the mobilization of N. In the present study, we examined the decomposition of a relatively nutrient-rich litter substrate extracted from maize compost. In previous experiments, we have also studied the degradation of extracts from relatively nutrient-poor forest litter material (Rineau et al., 2012). However, although the nutrient content in the forest litter material is significantly lower than in the maize compost material used here, spectroscopic analyses and transcriptome profiling showed that the mechanisms of decomposition of polysaccharides and polyphenols was similar in both extracts. We therefore predict that the stimulatory effects of glucose would be observed in both nutrient-rich and nutrient-poor litter material.
EMF takes up nutrients including N from the soil and exchange them against photosynthetically fixed carbon. The mechanisms that regulate this exchange are not known, but it has been proposed that the plant reduces C supply to the fungus if it fails to supply adequate amount of nutrients (Nehls et al., 2007). Recent studies on the arbuscular mycorrhizal symbiosis have demonstrated that the symbiosis is stabilized by reciprocal rewarding of nutrients and C resources according with the nutritional benefit provided by the other partner (Kiers et al., 2011). Moreover, experiments using arbuscular mycorrhizal mycorrhizal root organ cultures have shown that the carbon availability of the plant triggers the uptake and transport of ammonium uptake by the arbuscular mycorrhizal fungus (Fellbaum et al., 2012). The results from this study suggest that the C flux from the host plant can control the assimilation of organic N by an EMF. Studies of EMF in association with a host plant will be needed to verify this hypothesis.
Accession codes
References
Aro N, Pakula T, Penttilä M . (2005). Transcriptional regulation of plant cell wall degradation by filamentous fungi. FEMS Microbiol Rev 29: 719–739.
Arvas M, Kivioja T, Mitchell A, Saloheimo M, Ussery D, Penttila M et al (2007). Comparison of protein coding gene contents of the fungal phyla Pezizomycotina and Saccharomycotina. BMC Genomics 8: 325.
Baldrian P . (2004). Fungal laccases—occurrence and properties. FEMS Microbiol Rev 30: 215–242.
Baldrian P . (2008). Enzymes of saprotrophic basidiomycetes. In: Boddy L, Frankland JC, van West British P, (eds) Mycological Society Symposia Series Volume 28. Elsevier: Amsterdam, The Netherlands, pp 19–41.
Baldrian P, Valaskova V . (2008). Degradation of cellulose by basidiomycete fungi. FEMS Microbiol Rev 32: 501–521.
Bengtson P, Barker J, Grayston SJ . (2012). Evidence of a strong coupling between root exudation, C and N availability, and stimulated SOM decomposition caused by rhizosphere priming effects. Ecol Evol 2: 1843–1852.
Bolstad BM, Irizarry RA, Åstrand M, Speed TP . (2003). A comparison of normalization methods for high density oligonucleotide array data based on bias and variance. Bioinformatics 19: 185–193.
Chalot M, Brun A . (1998). Physiology of organic nitrogen acquisition by ectomycorrhizal fungi and ectomycorrhizas. FEMS Microbiol Rev 22: 21–44.
Chen DM, Bastias BA, Taylor AFS, Cairney JWG . (2003). Identification of laccase-like genes in basidiomycetes and transcriptional regulation by nitrogen in Piloderma byssinum. New Phytol 157: 547–554.
Eastwood DC, Floudas D, Binder M, Majcherczyk A, Schneider P, Aerts A et al (2011). The plant cell wall–decomposing machinery underlies the functional diversity of forest fungi. Science 333: 762–765.
Edgar R, Domrachev M, Lash AE . (2002). Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res 30: 207–210.
Edwards IP, Zak DR, Kellner H, Eisenlord SD, Pregitzer KS . (2011). Simulated atmospheric N deposition alters fungal community composition and suppresses ligninolytic gene expression in a northern hardwood forest. PLos One 6: 1–10.
Emanuelsson O, Brunak S, von Heijne G, Nielsen H . (2007). Locating proteins in the cell using TargetP, SignalP and related tools. Nat Protoc 2: 953–971.
Fellbaum CR, Gachomo EW, Beesetty Y, Choudhari S, Strahan GD, Pfeffer PE et al (2012). Carbon availability triggers fungal nitrogen uptake and transport in arbuscular mycorrhizal symbiosis. Proc Natl Acad Sci USA 109: 2666–2671.
Fenn P, Kirk TK . (1981). Relationship of nitrogen to the onset and suppression of ligninolytic activity and secondary metabolism in Phanerochaete chrysosporium. Arch Microbiol 130: 59–65.
Finlay RD, Odham G, Söderström B . (1988). Uptake, translocation and assimilation of nitrogen from 15N-labelled ammonium and nitrate sources by intact ectomycorrhizal systems of Fagus sylvatica infected with Paxillus involutus. New Phytol 113: 47–55.
Hobbie EA, Horton TR . (2007). Evidence that saprotrophic fungi mobilize carbon and mycorrhizal fungi mobilize nitrogen during litter decomposition. New Phytol 173: 447–449.
Hofrichter M, Ullrich R . (2010). New trends in fungal biooxidation. In: Hofrichter M, (ed) The Mycota Volume X, Industrial Applications. Springer-Verlag: Berlin, Germany, pp 425–449.
Ihaka R, Gentleman R . (1996). R: A Language for data analysis and graphics. J Comput Graph Stat 5: 299–314.
Irizarry RA, Hobbs B, Collin F, Beazer-Barclay YD, Antonellis KJ, Scherf U et al (2003a). Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4: 249–264.
Irizarry RA, Bolstad BM, Collin F, Cope LM, Hobbs B, Speed TP . (2003b). Summaries of Affymetrix GeneChip probe level data. Nucleic Acids Res 31: e15.
Javelle A, Morel M, Rodríguez-Pastrana B-R, Botton B, André B, Marini A-M et al (2003). Molecular characterization, function and regulation of ammonium transporters (Amt) and ammonium metabolizing enzymes (GS, NADP-GDH) in the ectomycorrhizal fungus Hebeloma cylindrosporum. Mol Microbiol 47: 411–430.
Kersten PJ . (1990). Glyoxal oxidase of Phanerochaete chrysosporium: its characterization and activation by lignin peroxidase. Proc Natl Acad Sci USA 87: 2936–2940.
Kiers ET, Duhamel M, Beesetty Y, Mensah JA, Franken O, Verbruggen E et al (2011). Reciprocal rewards stabilize cooperation in the mycorrhizal symbiosis. Science 333: 880–882.
Langston JA, Shaghasi T, Abbate E, Xu F, Vlasenko E, Sweeney MD . (2011). Oxidoreductive cellulose depolymerization by the enzymes cellobiose dehydrogenase and glycoside hydrolase 61. Appl Environ Microbiol 77: 7007–7015.
Le Quéré A, Wright DP, Söderström B, Tunlid A, Johansson T . (2005). Global patterns of gene regulation associated with the development of ectomycorrhiza between birch (Betula pendula Roth.) and Paxillus involutus (Batsch) Fr. Mol Plant Microbe In 18: 659–673.
Liers C, Arnstadt T, Ullrich R, Hofrichter M . (2011). Patterns of lignin degradation and oxidative enzyme secretion by different wood-and litter-colonizing basidiomycetes and ascomycetes grown on beech wood. FEMS Microbiol Ecol 78: 91–102.
Lindahl BD, Ihrmark K, Boberg J, Trumbore SE, Hogberg P, Stenlid J et al (2007). Spatial separation of litter decomposition and mycorrhizal nitrogen uptake in a boreal forest. New Phytol 173: 611–620.
Lucic E, Fourrey C, Kohler A, Martin F, Chalot M, Brun-Jacob A . (2008). A gene repertoire for nitrogen transporters in Laccaria bicolor. New Phytol 180: 343–364.
Maino TM, Senesi N . (1992). Synchronous excitation fluorescence spectroscopy applied to soil humic substances chemistry. Sci Total Environ 117/118: 41–51.
Martin F, Canet D . (1986). Biosynthesis of amino acids during 13C-glucose utilization by the ectomycorrhizal ascomycete Cenococcum geophilum monitored by 13C nuclear resonance. Physiol Veg 24: 209–218.
Martin F, Aerts A, Ahren D, Brun A, Danchin EG, Duchaussoy F et al (2008). The genome of Laccaria bicolor provides insights into mycorrhizal symbiosis. Nature 452: 88–92.
Martinez D, Challacombe J, Morgenstern I, Hibbett D, Schmoll M, Kubicek CP et al (2009). Genome, transcriptome, and secretome analysis of wood decay fungus Postia placenta supports unique mechanisms of lignocellulose conversion. Proc Natl Acad Sci USA 106: 1954–1959.
Marzluf GA . (1996). Regulation of nitrogen metabolism in mycelial fungi. In: Brambl B, Marzluf GA, (eds) The Mycota Volume III, Biochemistry and Molecular Biology. Springer- Verlag: Berlin, Germany, pp 357–368.
Morel M, Buée M, Chalot M, Brun A . (2006). NADP-dependent glutamate dehydrogenase: a dispensible function in ectomycorrhizal fungi. New Phytol 169: 179–190.
Nannipieri P, Eldor P . (2009). The chemical and functional characterization of soil N and its biotic components. Soil Biol Biochem 41: 2357–2369.
Nehls U, Kleber R, Wiese J, Hampp R . (1999). Isolation and characterization of an general amino acid permease from the ectomycorrhizal fungus Amanita muscaria. New Phytol 142: 331–341.
Nehls U, Bock A, Einig W, Hampp R . (2001). Excretion of two proteases by the ectomycorrhizal fungus Amanita muscaria. Plant Cell Environ 24: 741–747.
Nehls U, Grunze N, Willmann M, Reich M, Kuster H . (2007). Sugar for my honey: carbohydrate partitioning in ectomycorrhizal symbiosis. Phytochemistry 68: 82–91.
Nehls U, Göhringer F, Wittulsky S, Dietz S . (2010). Fungal carbohydrate support in the ectomycorrhizal symbiosis: a review. Plant Biol 12: 292–301.
Nierop KG, Van Bergen PF . (2002). Clay and ammonium catalyzed reactions of alkanols, alkanoic acids and esters under flash pyrolytic conditions. J Anal Appl Pyrol 63: 197–208.
Piscitelli A, Giardina P, Lettera V, Pezzella C, Sannia G, Faraco V . (2011). Induction and transcriptional regulation of laccases in fungi. Curr Genomics 12: 104–112.
Ramette A . (2007). Multivariate analyses in microbial ecology. FEMS Microbiol Ecol 62: 142–160.
Rawlings ND, Barrett AJ, Bateman A . (2012). MEROPS: the database of proteolytic enzymes, their substrates and inhibitors. Nucleic Acids Res 40: D343–D350.
Read DJ, Leake JR, Perez-Moreno J . (2004). Mycorrhizal fungi as drivers of ecosystem processes in heathland and boreal forest biomes. Can J Bot 82: 1243–1263.
Rineau F, Roth D, Shah F, Smits M, Johansson T, Canbäck B et al (2012). The ectomycorrhizal fungus Paxillus involutus converts organic matter in plant litter using a trimmed brown-rot mechanism involving Fenton chemistry. Environ Microbiol 14: 1477–1487.
Schimel JP, Bennett J . (2004). Nitrogen mineralization: challenges of a changing paradigm. Ecology 85: 591–602.
Talbot JM, Allison SD, Treseder KK . (2008). Decomposer in disguise: mycorrhizal fungi as regulators of soil C dynamics in ecosystems under global change. Funct Ecol 22: 955–963.
Talbot JM, Treseder KK . (2010). Controls over mycorrhizal uptake of organic nitrogen. Pedobiologia 53: 169–179.
Talbot JM, Bruns TD, Smith DP, Branco S, Glassman SI, Erlandsona S et al (2013). Independent roles of ectomycorrhizal and saprotrophic communities in soil organic matter decomposition. Soil Biol Biochem 57: 282–291.
Vanden Wymelenberg A, Gaskell J, Mozuch M, Sabat G, Ralph J, Skyba O et al (2010). Comprataive transcriptomics and secretome analysis of the wood decay fungi Postia placenta and Phanerochaete chrysosporium. Appl Environ Microbiol 76: 3599–3610.
Zhang XZ, Sathitsuskanoh N, Zhang YHP . (2010). Glycoside hydrolase family 9 processive endoglucanase from Clostridium phytofermentans: Heterologous expression, characterization, and synergy with family 48 cellobiohydrolase. Bioresource Technol 101: 5534–5538.
Zhu H, Dancik BP, Higginbotham KO . (1994). Regulation of extracellular proteinase production in an ectomycorrhizal fungus Hebeloma crustuliniforme. Mycologia 86: 227–234.
Acknowledgements
The work was supported by Grants from the Swedish Research Council (VR), the strategic research program Biodiversity and Ecosystem Services in a Changing Climate (BECC), the Danish Agency for Science and Technology and the Research Foundation—Flanders (FWO).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Rineau, F., Shah, F., Smits, M. et al. Carbon availability triggers the decomposition of plant litter and assimilation of nitrogen by an ectomycorrhizal fungus. ISME J 7, 2010–2022 (2013). https://doi.org/10.1038/ismej.2013.91
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2013.91
Keywords
This article is cited by
-
Changes in Soil Particulate Organic Carbon and Their Response to Changing Environments on the Tibetan Plateau, Mongolian Plateau, and Loess Plateau, China
Journal of Soil Science and Plant Nutrition (2023)
-
Biodegradation potential of indigenous litter dwelling ligninolytic fungi on agricultural wastes
Bulletin of the National Research Centre (2020)
-
Large-scale genome sequencing of mycorrhizal fungi provides insights into the early evolution of symbiotic traits
Nature Communications (2020)
-
The soil organic matter decomposition mechanisms in ectomycorrhizal fungi are tuned for liberating soil organic nitrogen
The ISME Journal (2019)
-
Evolutionary dynamics of host specialization in wood-decay fungi
BMC Evolutionary Biology (2018)