Abstract
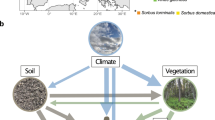
Understanding the interactions among microbial communities, plant communities and soil properties following deforestation could provide insights into the long-term effects of land-use change on ecosystem functions, and may help identify approaches that promote the recovery of degraded sites. We combined high-throughput sequencing of fungal rDNA and molecular barcoding of plant roots to estimate fungal and plant community composition in soil sampled across a chronosequence of deforestation. We found significant effects of land-use change on fungal community composition, which was more closely correlated to plant community composition than to changes in soil properties or geographic distance, providing evidence for strong links between above- and below-ground communities in tropical forests.
Similar content being viewed by others
Main
Land-use change, such as deforestation, is one of the greatest threats to biodiversity worldwide (Thomas et al., 2004), particularly within tropical ecosystems (Sala et al., 2000; Gibson et al., 2011). In tropical ecosystems, the responses of soil fungal biodiversity to land-use change are only beginning to be explored, and responsiveness has been shown in some cases (for example, Fracetto et al., 2013) but not in others (for example, Leal et al., 2009). Fungal communities can be structured by nutrient availability (Rousk et al., 2010) and plant community composition (Carney and Matson, 2006; Peay et al., 2013), suggesting that shifts in soil fungal communities in response to land-use change (Castro et al., 2008; Fracetto et al., 2013) could result from alterations in soil properties and/or plant communities. However, the relative strength of these factors in determining fungal community composition in tropical systems has not been examined.
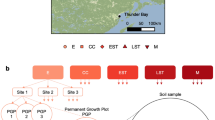
We used an established chronosequence of land use in the Brazilian Amazon rainforest (Rodrigues et al., 2013) that included a primary forest, secondary forest and pasture. We sampled soil using a spatially explicit design within single hectare plots for a total of nine samples per site. To examine changes in fungal community composition we targeted the ITS1 region of the rRNA gene using the fungal-specific primer pair ITS1F-ITS2 with a two-stage PCR designed for paired-end Illumina sequencing. Amplicons were sequenced on an Illumina HiSeq 2000 (Illumina, San Diego, CA, USA) at the University of Oregon, quality-filtered to remove low-quality bases and putative chimeras, and clustered into operational taxonomic units at 97% sequence similarity using the program UCHIME (Edgar, 2010) in the QIIME platform (Caporaso et al., 2010). Plant community composition was determined by direct PCR (Phire Direct PCR, ThermoScientific, Waltham, MA, USA) and Sanger sequencing of the chloroplast trnL intron region (Taberlet et al., 1991, 2007) of roots from the same soil cores collected for microbial community analysis. Root sequences were grouped by plant genera on the basis of top BLAST matches to the GenBank database (Supplementary Table 1). Unique root sequences were deposited in GenBank under accession numbers KF661456–KF661514, and ITS1 sequences were deposited in MG-RAST under ID 4536676.3. Soil properties were determined by quantifying micro- and macronutrients with protocols modified for tropical soil (van Raij et al., 2001; Supplementary Table 2). Analyses were undertaken on matched samples that had sufficient data for fungi, roots and soil properties for a total of 20 samples across the three land uses. A more detailed description of the methods can be found in the Supplementary Material.
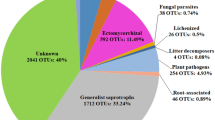
Using an operational taxonomic unit community matrix rarefied to 10 000 sequences per sample, fungal community composition (Bray–Curtis) was compared among the sites using PERMANOVA (Anderson, 2001) and visualized using nonmetric multidimensional scaling. The fungal community composition was significantly different among the three land uses (that is, primary forest, secondary forest and pasture; R2=0.32, F2,17=4.14, P=0.001; Figure 1a), driven mostly by decreased relative abundance of Basidiomycota in the pasture and secondary forest (Supplementary Figure 1). PERMANOVA analysis also showed significant differences in plant community composition (R2=0.41, F2,17=5.92, P=0.001) and measures of soil properties (F2,17=25.6, R2=0.75, P=0.001; Figures 1b and c).
The relative contribution of variation in pairwise geographic distance, environmental distance (Hellinger-transformed Euclidean distance) and plant community dissimilarity (Bray–Curtis) to variation in fungal community composition was determined using multiple regression on distance matrices (Lichstein, 2006). We found that plant community composition was the strongest driver of differences in fungal community composition (Table 1). Although we found significant differences among the sites (Supplementary Table 2), fungal richness was not significantly correlated with plant richness (F=1.68, P=0.21).
Plant identification through targeting of roots within soil cores provides the means to isolate the plant species most likely to interact with the fungal community in a given sampling location. We used this novel approach to characterize plant composition among plots and found that the plant community was a stronger predictor of the fungal community than were soil properties or geographic distance. These results are consistent with those of Mitchell et al. (2010), who showed that plant community composition was a better predictor of microbial community composition than was soil chemistry across a plant successional gradient in a boreal ecosystem. Our findings support those of Peay et al. (2013), who found a significant correlation between plant community composition and fungal community composition in three tropical forests, and further show that fungal community composition is more closely linked to plant community composition than to either environmental variability or geographic distance.
Our results also provide evidence that effects of deforestation on fungal communities are likely mediated in large part by changes in plant communities. Fungal communities in secondary forests were more similar to primary forests than were pasture communities (Figure 1), suggesting that the recovery of fungal communities following pasture abandonment is closely tied to that of plants. A clearer understanding of interactions between plant and fungal communities could prove useful to conservation and restoration biology, as it could identify management strategies that better promote both reforestation and the recovery of microbially mediated ecosystem functions in degraded areas (van der Heijden et al., 2008).
Accession codes
Change history
24 June 2014
This article has been corrected since Advance Online Publication and a corrigendum is also printed in this issue
References
Anderson MJ . (2001). A new method for non-parametric multivariate analysis of variance. Austral Ecol 26: 32–46.
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK et al (2010). QIIME allows analysis of high-throughput community sequencing data. Nat Meth 7: 335–336.
Carney KM, Matson PA . (2006). The influence of tropical plant diversity and composition on soil microbial communities. Microb Ecol 52: 226–238.
Castro AP, Quirino BF, Pappas G Jr, Kurokowa AS, Neto EL, Kruger RH . (2008). Diversity of soil fungal communities of Cerrado and its closely surrounding agriculture fields. Arch Microbiol 190: 129–139.
Edgar RC . (2010). Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26: 2460–2461.
Fracetto GGM, Azevedo LCB, Fracetto FJC, Adreote FD, Lambais MR, Pfenning LH . (2013). Impact of Amazon land use on the community of soil fungi. Sci Agric 70: 59–67.
Gibson L, Lee TM, Koh LP, Brook BW, Gardner TA, Barlow J et al (2011). Primary forests are irreplaceable for sustaining tropical biodiversity. Nature 478: 378–381.
Leal PL, Sturmer SL, Siqueira JO . (2009). Occurrence and diversity of arbuscular mycorrhizal fungi in trap cultures from soils under different land use systems in the Amazon, Brazil. Braz J Microbiol 40: 111–121.
Lichstein JW . (2006). Multiple regression on distance matrices: a multivariate spatial analysis tool. Plant Ecol 188: 117–131.
Mitchell RJ, Hester AJ, Campbell CD, Chapman SJ, Cameron CM, Hewison RL et al (2010). Is vegetation composition or soil chemistry the best predictor of the soil microbial community? Plant Soil 333: 417–430.
Peay KG, Baraloto C, Fine PV . (2013). Strong coupling of plant and fungal community structure across western Amazonian rainforests. ISME J 7: 1852–1861.
Rodrigues JLM, Pellizari VH, Mueller R, Baek K, Jesus EC, Paula FS et al (2013). Conversion of the Amazon rainforest to agriculture results in biotic homogenization of soil bacterial communities. Proc Natl Acad Sci USA 110: 988–993.
Rousk J, Bååth E, Brookes PC, Lauber CL, Lozupone C, Caporaso JG et al (2010). Soil bacterial and fungal communities across a pH gradient in an arable soil. ISME J 4: 1340–1351.
Sala OE, Chapin FS III, Armesto JJ, Berlow E, Bloomfield J, Dirzo R et al (2000). Global biodiversity scenarios for the year 2100. Science 287: 1770–1774.
Taberlet P, Gielly L, Pautou G, Bouvet J . (1991). Universal primers for amplification of three non-coding regions of chloroplast DNA. Plant Mol Biol 17: 1105–1109.
Taberlet P, Coissac E, Pompanon F, Gielly L, Miquel C, Valentini A et al (2007). Power and limitations of the chloroplast trnL (UAA) intron for plant DNA barcoding. Nucleic Acids Res 35: e14.
Thomas CD, Cameron A, Green RE, Bakkenes M, Beaumont LJ, Collingham YC et al (2004). Extinction risk from climate change. Nature 427: 145–148.
van der Heijden MGA, Bardgett RD, van Straalen NM . (2008). The unseen majority: soil microbes as drivers of plant diversity and productivity in terrestrial ecosystems. Ecol Lett 11: 296–310.
van Raij B, Andrade JC, Cantarella H, Quaggio JA . (2001) Análise química para avaliação da fertilidade de solos tropicais. Instituto Agronômico: Campinas, Brazil.
Acknowledgements
We thank the owners of the Fazenda Nova Vida for providing field site access, Vivian Pellizari for logistical support, Wagner Piccinini for field sampling, Jonas Frankel-Bricker and Roo Vandegrift for root amplification, and the helpful suggestions of two anonymous reviewers. Funding was provided by USDA Agriculture and Food Research Initiative Competitive Grant 2009-35319-05186 and by an APS Lewis and Clark Fund for Exploration and Research grant.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary information accompanies this paper on The ISME Journal website
Rights and permissions
About this article
Cite this article
Mueller, R., Paula, F., Mirza, B. et al. Links between plant and fungal communities across a deforestation chronosequence in the Amazon rainforest. ISME J 8, 1548–1550 (2014). https://doi.org/10.1038/ismej.2013.253
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2013.253
Keywords
This article is cited by
-
Response of soil bacteria on habitat-specialization and abundance gradient to different afforestation types
Scientific Reports (2023)
-
The variation of microorganisms and organics during methane production from lignite under an electric field
Biotechnology Letters (2023)
-
Biogeographic responses and niche occupancy of microbial communities following long-term land-use change
Antonie van Leeuwenhoek (2022)
-
Contents of lobetyolin, syringin, and atractylolide III in Codonopsis pilosula are related to dynamic changes of endophytes under drought stress
Chinese Medicine (2021)
-
Erosion reduces soil microbial diversity, network complexity and multifunctionality
The ISME Journal (2021)