Abstract
We investigated the mechanisms of osmoadaptation in the order Halobacteriales, with special emphasis on Haladaptatus paucihalophilus, known for its ability to survive in low salinities. H. paucihalophilus genome contained genes for trehalose synthesis (trehalose-6-phosphate synthase/trehalose-6-phosphatase (OtsAB pathway) and trehalose glycosyl-transferring synthase pathway), as well as for glycine betaine uptake (BCCT family of secondary transporters and QAT family of ABC transporters). H. paucihalophilus cells synthesized and accumulated ∼1.97–3.72 μmol per mg protein of trehalose in a defined medium, with its levels decreasing with increasing salinities. When exogenously supplied, glycine betaine accumulated intracellularly with its levels increasing at higher salinities. RT-PCR analysis strongly suggested that H. paucihalophilus utilizes the OtsAB pathway for trehalose synthesis. Out of 83 Halobacteriales genomes publicly available, genes encoding the OtsAB pathway and glycine betaine BCCT family transporters were identified in 38 and 60 genomes, respectively. Trehalose (or its sulfonated derivative) production and glycine betaine uptake, or lack thereof, were experimentally verified in 17 different Halobacteriales species. Phylogenetic analysis suggested that trehalose synthesis is an ancestral trait within the Halobacteriales, with its absence in specific lineages reflecting the occurrence of gene loss events during Halobacteriales evolution. Analysis of multiple culture-independent survey data sets demonstrated the preference of trehalose-producing genera to saline and low salinity habitats, and the dominance of genera lacking trehalose production capabilities in permanently hypersaline habitats. This study demonstrates that, contrary to current assumptions, compatible solutes production and uptake represent a common mechanism of osmoadaptation within the Halobacteriales.
Similar content being viewed by others
Introduction
Members of the halophilic archaea (order Halobacteriales) are characterized by an absolute salt requirement and conspicuous growth in hypersaline habitats where they often represent the majority of microbial biomass. To survive in high salinity environments, the Halobacteriales maintain an intracellular osmotic pressure that is equal to or higher than that of the surrounding environment (Oren, 2013). In general, maintenance of high intracellular osmotic pressure by halophilic and halotolerant microorganisms could be achieved either by the uptake and accumulation of high concentrations of inorganic ions such as K+ (salting-in strategy) or by synthesis and/or uptake of highly soluble organic solutes that do not interfere with intracellular enzymatic activities and cellular processes (compatible solute strategy) (Grant, 2004).
Within the Halobacteriales, utilization of salting-in strategy for osmoadaptation has been demonstrated in multiple model species such as Halobacterium salinarum, Haloarcula marismortui, Haloferax volcanii, Haloferax mediterranei, Haloferax gibbonsii, Halorubrum saccharovorum and Halorubrum trapanicum (Christian and Waltho, 1962; Ginzburg et al., 1970; Lanyi and Silverman, 1972; Matheson et al., 1976; Pérez-Fillol and Rodríguez-Valera, 1986; Mojica et al., 1997; Oren et al., 2002a). These studies reported high levels of intracellular K+ (1.9–5.5 M) in all examined species. Intracellular K+ levels were shown to be dependent on the extracellular salt concentrations, with its levels increasing with the increase of the total salinity of the growth media (Pérez-Fillol and Rodríguez-Valera, 1986). This salting-in strategy for osmoadaptation is more energetically favorable when compared with the alternative compatible solute accumulation strategy, as the latter requires the synthesis and accumulation of increasing amounts of organic solutes at higher salinities. However, the permanently high levels of intracellular K+ trigger multiple structural and metabolic adaptations that limit the metabolic flexibility and ecological range of microorganisms utilizing this strategy for osmoadaptation (Oren, 2013).
In addition to their prevalence in hypersaline habitats, members of the Halobacteriales are also consistently observed in saline (10–17% NaCl) environments as a fraction of a more complex halophilic and halotolerant prokaryotic and eukaryotic population (Bowman et al., 2000; Perreault et al., 2007; Stivaletta et al., 2011; Youssef et al., 2012; Xiao et al., 2013). Further, members of the Halobacteriales have also been identified as a minor constituent of the microbial population in low-salinity environments including sea water (Ventosa et al., 1984), seawater aquarium (Inoue et al., 2011), anaerobic springs (Elshahed et al., 2004), low salinity salterns (Mani et al., 2012), coastal salt marshes (Munson et al., 1997, Purdy et al., 2004), and hydrothermal (Takai et al., 2001) and geothermal (Ellis et al., 2008) vents, where pockets of salinity occasionally develop. Therefore, in such habitats, the Halobacteriales must cope with extended periods of low salinities and/or constant fluctuations in salinity levels, conditions that are not conducive to microorganisms solely dependent on a salting-in strategy for osmoadaptation.
Prior investigations of osmoadaptive strategies within the Halobacteriales have been conducted on isolates obtained from permanently hypersaline environments, for example, Halobacterium salinarum strain NRC 34002, Haloferax mediterranei and Haloarcula marismortui (isolated from the dead sea (Oren et al., 1990)). Here, we decided to investigate the potential for alternative mechanisms of osmoadaptation in multiple Halobacteriales genera, with special emphasis on Haladaptatus paucihalophilus, known for its capability to grow in low and fluctuating salinities and survive extended incubations in distilled water (Savage et al., 2007). While few prior studies on the production (Nicolaus et al., 1988; Desmarais et al., 1997; Goh et al., 2011) and uptake (Kokoeva et al., 2002) of compatible solutes in the Halobacteriales have been published, detailed understanding of the extent of distribution, genetic basis and ecological significance of the process is lacking. We demonstrate that multiple Halobacteriales genera are capable of trehalose or 2-sulfotrehalose production, and that the uptake of glycine betaine is a widely utilized process for osmoadaptation in the Halobacteriales. These results suggest that the osmoadaptive strategies of the Halobacteriales are more complex than previously implied from prior studies on model Halobacteriales strains.
Materials and methods
Archaeal strains and growth media
H. paucihalophilus strain DX253T (DSM18195) was isolated (Savage et al., 2007) and maintained in our laboratory. All other strains were either kindly provided by Dr Marc T Facciotti (University of California Davis) or purchased from the German Collection of Microorganisms (DSMZ, Braunschweig, Germany). H. paucihalophilus was grown in a defined media similar in composition to DSMZ 1125 but with no yeast extract and with pyruvate rather than sucrose as a carbon source. This was done to ensure that any compatible solutes detected in cell extracts are synthesized de novo. All other strains were cultured in their DSMZ media at three different salt concentrations (Supplementary Table S1).
Genomic sequencing
The genome of H. paucihalophilus strain DX253T was sequenced using the service of a commercial sequencing provider (Engencore, now Selah Genomics, Columbia, SC, USA) using 454 FLX technology. The genome was assembled using Newbler assembler (Branford, CT, USA) into 32 Contigs (N50=267, 250 bp, N90=1 00 497 bp). Gene calling, annotation and metabolic construction were conducted using the Department of Energy Integrated Microbial Genomes (IMG) platform (Markowitz et al., 2012).
Analytical methods
Identification of and quantification of K+ and compatible solutes in cell-free extracts
Details on the methods for identification and quantifications of ions and compatible solutes are presented as Supplementary Text. In brief, the presence and identity of compatible solutes in ethanolic cell-free extracts were determined using 1H nuclear magnetic resonance spectroscopy (1H-NMR) and high performance liquid chromatography (HPLC). Quantification of various compatible solutes and ions was done in perchloric acid cell-free extracts (Supplementary Text). Trehalose was determined enzymatically using using a trehalase assay kit (Megazyme, Wicklow, Ireland). To differentiate trehalose from 2-sulfotrehalose, which is also detectable using this kit, we devised a fast method to detect the presence of a 2′-O-sulfonate group based on the action of abalone snail sulfatase (Sigma-Aldrich, St Louis, MO, USA), shown to be specific for cleaving the 2′-O-sulfonate group of trehalose (Uzawa et al., 2004), followed by quantification of the released inorganic sulfate turbidimetrically (Lundquist et al., 1980) (Supplementary Text). Glycine betaine was quantified using a colorimetric assay for quaternary ammonium compounds, as described before (Grieve and Grattan, 1983). Intracellular K+ levels were quantified in cell-free extracts of various Halobacteriales species spectophotmetrically (Takagi et al., 1981). Intracellular free amino acids in cell-free extracts were assayed for free amino acids using the services of the Protein Chemistry Lab of Texas A&M University (Supplementary Text)
RNA extraction and gene expression
Cell pellets from late exponential phase cultures were solubilized in TRI Reagent Solution (Ambion, Carlsbad, CA, USA) and processed according to the manufacturer’s instruction. Contaminating DNA was removed from all RNA preparations using RQ1 DNase (Promega, Madison, WI, USA). DNA-free RNA was then precipitated using Lithium Chloride Precipitation Solution (Ambion) according to the manufacturer’s instructions to remove any contaminating small RNAs. Total RNA reverse transcription (cDNA construction) was conducted on 1 μg of RNA preparations using Superscript III first-strand synthesis kit (Invitrogen, Carlsbad, CA, USA) with random hexamers (Invitrogen). Reverse transcription qPCR (RT-qPCR) was conducted on the cDNA obtained using primers shown in Supplementary Table S2. Levels of expression of different genes were compared with glyceraldehyde-3-phosphate dehydrogenase gene expression (Supplementary Text).
Phylogenetic analysis
RNA polymerase β-subunit (RpoB′) protein sequences from publicly available Halobacteriales genomes (n=82) were aligned in ClustalX (Larkin et al., 2007) and used to construct a maximum likelihood tree in Mega (Tamura et al., 2011) using a WAG model with Γ distribution of rates among sites as previously suggested (Minegishi et al., 2010). Halobacteriales OtsA (trehalose-6-phosphate synthase) and OtsB (trehalose-6-phosphatase) protein sequences, as well as those from reference bacterial and archaeal taxa were also aligned using ClustalX, and the best protein substitution model was predicted in Mega (Tamura et al., 2011). The predicted model was applied in maximum likelihood tree construction and support values were obtained using 100 bootstrapping events.
Ecological distribution of otsAB-harboring genera
We examined the correlation between the presence/absence of trehalose synthesis otsAB genes in a genome and the respective salt minimum, maximum and optimum concentrations using χ2 Contingency tables for correlation (Plackett, 1983) (see Supplementary Text for details). To study the correlation between otsAB presence and the ecological distribution of various genera, we re-examined the relative abundance of various Halobacteriales genera in 13 different environments with salinities ranging from permanently hypersaline (n=9), saline and fluctuating salinity habitats (where salinity is usually <250 g l−1 and exhibits fluctuation; n=3), and mostly non-saline with only occasional development of temporal and spatial saline pockets (n=1, Zodletone spring). Relative abundances of Halobacteriales genera with and without the OtsAB system in their genome were compared between various groups of environments using Student’s t-test. Values were considered significantly different if the P-value associated with the t-test was <0.0038 (equivalent to an α value of 0.05 corrected for 13 pairwise comparisons using the Bonferroni correction). The relative abundances values obtained were also used in a principal-component analysis to evaluate the differences in genera composition between the environments using the R statistical package (R Development Core Team, 2011).
GenBank accession number
The genome of H. paucihalophilus has been deposited in Genbank under the accession number AEMG00000000.
Results
General features and osmoadaptive capabilities inferred from H. paucihalophilus genome
The general features of H. paucihalophilus genome are shown in Supplementary Table S3. The genome and predicted proteome displayed several features characteristic of the Halobacteriales such as high GC content (61.8%), a higher proportion of acidic residues (Glu+Asp=15.95%), lower proportion of basic residues (Lys+Arg=8.79%), lower proportion of cysteine residues (0.73%) and lower proportion of large hydrophobic residues (Ile+Leu+Met+Phe=18.98%). The predicted proteome also showed a low average isoelectric point (pI=5.12), with the pI of cytoplasmic proteins (average pI=4.69) significantly lower (Student’s t-test P-value=2.4E−205) than the pI of membrane proteins (average pI=6.88) (Supplementary Table S3, S4).
We queried the H. paucihalophilus genome for genes putatively involved in osmoadaptation. As expected, the genome contained all genes necessary for Na+ extrusion, K+ uptake and chloride homeostasis (Supplementary Text). More importantly, the genome contained genes necessary for the synthesis of trehalose. Out of the five mechanisms known for trehalose biosynthesis (Avonce et al., 2006), two were identified in the genome: the trehalose-6-phosphate synthase/trehalose-6-phosphatase (OtsA/B) pathway and the trehalose glycosyl-transferring synthase pathway (TreT) (Table 1). The genome contained two copies of the trehalose-6-phosphate synthase gene (otsA), one copy of trehalose-6-phosphatase (otsB) clustered with one of the otsA genes and one copy of trehalose glycosyl-transferring synthase TreT (Table 2). No genes for the synthesis of other compatible solutes such as glycine betaine or ectoine were identified. However, the genome contained genes necessary for the uptake of glycine betaine as evident by the presence of genes encoding the two major betaine transporter families: The Betaine/Carnitine/Choline Transporter (BCCT) Family of secondary transporters (TC 2.A.15), and the Quaternary Amine Uptake Transporter (QAT) Family of ABC transporters (TC 3.A.1.12) (Table 1).
Trehalose biosynthesis and glycine betaine uptake by H. paucihalophilus
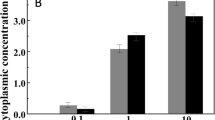
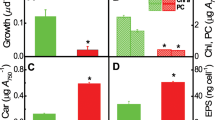
Trehalose was identified using 1H-NMR in cell-free extracts of H. paucihalophilus cells grown on a defined yeast extract-free medium with pyruvate as the sole carbon source (Figure 1). Trehalose was differentiated from 2-sulfotrehalose previously identified in the cell-free extracts of several Halobacteriales spp. (Desmarais et al., 1997) by differences in the 1H chemical shifts at the C-2 position. The trehalose spectrum shows similar resonances for C-2 and C-2′ of ∼3.63–3.64 p.p.m. (Figure 1, blue), whereas 2-sulfotrehalose shows a downfield shift for one of the C-2 positions, due to the presence of the sulfontate group, to 4.35 p.p.m. (Desmarais et al., 1997). Further, the absence of 2-sulfotrehalose was confirmed by the lack of free inorganic sulfate accumulation after treatment of cell-free extracts with C-2-specific abalone snail sulfatase, using Natronococcus occultus, Natrialba magadii and Natronobacterium gregoryi as positive controls. No evidence for the accumulation of other polyols (for example, glycerol), zwitterions (for example, glycine betaine) or free amino acids was observed using 1H-NMR scans or HPLC. Intracellular trehalose concentrations in H. paucihalophilus cells were highest when grown at lower salinities, and progressively decreased in cultures grown at higher salinities (from 3.72 μmol per mg protein of trehalose when grown at 75 g l−1 salinity to 1.97 μmol per mg protein of trehalose when grown at 275 g l−1 salinity) (Figure 2a). Intracellular K+ levels in H. paucihalophilus were ∼17 times that measured in E. Coli K12 cells grown in LB media (Figure 2c), but only a fraction (0.55 times at 125 g l−1 salinity to 0.28 times at 275 g l−1 salinity) of the intracellular K+ concentration measured in Halobacterium salinarum strain NRC-1. Interestingly intracellular K+ levels in H. paucihalophilus were relatively stable at all salinities examined, as opposed to the salinity-dependent increase in intracellular K+ concentrations observed in Halobacterium salinarum (Figure 2a; Pérez-Fillol and Rodríguez-Valera (1986)).
Trehalose (-<>-), glycine betaine (-Δ-) and intracellular potassium levels (□) in H. paucihalophilus grown in defined medium (a) and in the presence of 1% betaine (b). Comparison of H. paucihalophilus intracellular potassium levels (▪) to those in Halobacterium salinarum strain NRC-1 ( ) and E. coli (□) is shown in (c). Intracellular K+ are levels are determined only at 10 g l−1 for E. coli due to its inability to grow at higher salinities.
) and E. coli (□) is shown in (c). Intracellular K+ are levels are determined only at 10 g l−1 for E. coli due to its inability to grow at higher salinities.
No glycine betaine was observed in cell-free extracts of H. paucihalophilus grown in a defined media (Figure 2a). However, when exogenously supplied (10 g l−1), cells of H. paucihalophilus accumulated large amounts of glycine betaine intracellularly in a salinity-dependent pattern (0.9 μmol per mg protein of betaine when grown at 75 g l−1 salinity to 3.21 μmol per mg protein of betaine when grown at 275 g l−1 salinity; Figure 2b). Overall patterns of intracellular K+ and trehalose accumulation in the presence of glycine betaine (Figure 2b) were comparable to those observed in defined media (Figure 2a). Although trehalose levels were slightly lower in presence of glycine betaine, these differences were only significant at higher salinities (175 and 225 g l−1, Student’s t-test P-value=0.017 and 0.007, respectively).
Transcriptional analysis of trehalose synthesis genes in H. paucihalophilus
We used RT-qPCR to quantify gene expression levels of all four genes putatively involved in trehalose biosynthesis in H. paucihalophilus. Expression levels of the complete OtsAB system genes (otsA2 and otsB genes) were 3–4 orders of magnitude higher than that of otsA1 or treT genes (Table 2). Further, the pattern of gene expression of the complete OtsAB system mirrored the pattern of intracellular trehalose accumulation observed at various salinities, where gene expression was highest at lower salinity (2016–4688 times that of glyceraldehyde-3-phosphate expression at 75 g l−1 salinity) and progressively decreased at higher salinities (to 0.74–2.84 times that of glyceraldehyde-3-phosphate expression at 275 g l−1 salinity) (Table 3). These results strongly suggest the involvement of the complete OtsAB system in trehalose synthesis in H. paucihalophilus.
Genes mediating trehalose synthesis and betaine uptake are widely distributed in Halobacteriales genomes
Out of 83 genomes belonging to 28 different Halobacteriales genera examined, a complete OtsAB system was identified in 39 different species comprising 17 different genera (Haladaptatus, Halalkalicoccus, Halosimplex, Halococcus, Halovivax, Halobiforma, Haloterrigena, Halobacterium, Halogranum, Halosarcina, Halopiger, Natrialba, Natronobacterium, Natronococcus, Natronorubrum, Natronolimnobius and Natrinema; Table 3). The presence or absence of a complete OtsAB system was a defining characteristic in all genera: within any Halobacteriales genus; all species either possessed or lacked a complete OtsAB system. Additional systems of trehalose production (trehalose glycosyl-transferring synthase (TreT), trehalose synthase (TreS) and trehalose phosphorylase (TreP) but not maltooligosyl-trehalose synthase/maltooligosyl-trehalose trehalohydrolase (TreYZ)) were sporadically distributed in few of the sequenced genomes (Table 3).
Genes for glycine betaine synthesis from glycine were not identified in any of the Halobacteriales genomes examined. However, genes encoding representatives of the BCCT family of betaine transporter were identified in 60 Halobacteriales genomes and genes encoding the complete ABC betaine transporter were identified in 13 genomes (Table 3). Three distinct patterns for the distribution of glycine betaine uptake genes were identified: (1) genera or species with no evidences for betaine transport (for example, members of the genera Halobiforma, Halostagnicola, Haloplanus, Halobaculum, and Halomicrobium and multiple species within the genera Haloferax, Halorubrum, Halorhabdus, Halobacterium, Natronomonas, Natrialba and Natrinema; (2) genera or species with only the BCCT transporters (for example, genera Halobacterium, Halalkalicoccus, Haloterrigena, Haloarcula, Halosimplex, Halovivax, Haloquadratum, Halogeometricum, Halogranum, Halopiger, Natronobacterium and Natronolimnobius, and multiple species within the genera Halococcus, Halorhabdus, Haloferax, Halorubrum, Natronomnas, Natrialba, Natronorubrum, Natronococcus and Natrinema) and (3) genera or species with representatives of both families of transporters (for example, genera Haladaptatus and Halosarcina and various species within the genera Halococcus, Haloferax, Natronorubrum, Natronococcus and Natrinema) (Table 3).
Experimental evaluation of trehalose/2-sulfotrehalose production and glycine uptake within the Halobacteriales
We quantified the intracellular levels of trehalose and glycine betaine in 17 different Halobacteriales species grown at different salinities that are close to their low, optimum and high salt concentrations (Table 3 and Supplementary Table S5). Trehalose production was observed in the examined otsAB-harboring species Halovivax ruber, Halococcus saccharolyticus, Halosarcina pallida, Halogranum rubrum, Haloterrigena turkmenica and Natronorubrum tibetense, whereas 2-sulfotrehalose production was observed in the examined otsAB-harboring species Natrialba magadii, Natronococcus occultus, Natronobacterium gregoryi, Natrinema pellirubrum and Halalkalicoccus jeotgali. No trehalose was produced in examined species lacking the OtsAB system (Haloferax mediterranei, Halorhabdus utahenesis, Halogeometricum borinquense, Halostagnicola larsenii, Halorubrum lacusprofundi and Halobacterium salinarum NRC-1) (Table 3). Similar to patterns observed in H. paucihalophilus, trehalose or 2-sulfotrehalose concentrations were highest at lower salinities (with the exception of Halovivax ruber, Halosarcina pallida and Natrinema pellirubrum) (Supplementary Table S5).
Several patterns of glycine betaine accumulation were observed amongst the examined species. Strains with no genomic evidences of both betaine uptake systems did not accumulate betaine (for example, Halostagnicola larsenii). Strains with genomic evidence of both families of betaine uptake accumulated betaine (for example, Halococcus saccharolyticus, Halosarcina pallida and Haloterrigena turkmenica). Within strains harboring only the BCCT family of betaine uptake transporters, some species did not accumulate betaine (Halalkalicoccus jeotgali, Halovivax ruber, Natrialba magadii and Natronobacterium gregoryii), whereas others did (Halorhabdus utahenesis, Haloferax medeterranei, Halogeometricum borinquense, Halorubrum lacusprofundi, Natrinema pellirubrum, Natronococcus occultus and Halogranum rubrum) (Table 3). Intracellular glycine betaine levels in all glycine betaine-accumulating species examined were salinity-dependent (Supplementary Table S5). While these results do not exclusively establish the role of either system in glycine betaine uptake, they suggest that (1) the BCCT system can function in some strains as the sole mediator of glycine betaine uptake and (2) the presence of a BCCT family representative in a genome does not necessarily translate into betaine uptake capabilities.
Intracellular K+ levels in trehalose- and 2-sulfotrehalose-producing strains ranged between 2.1 and 9.6 μmol per mg protein and, in general, were fairly constant at various levels of salinities examined (Supplementary Table S5, with the exception of Halosarcina pallida). On the other hand, intracellular K+ levels in genera incapable of trehalose or 2-sulfotrehalose production were significantly higher (4.19–29.7 μmol per mg protein) (Student’s t-test P-value=1.57E−6), and in general, increased with the increase of media salinity (Supplementary Table S5).
Phylogeny and distribution of Halobacteriales OtsAB system genes
Halobacteriales OtsA- and OtsB-predicted protein sequences showed high level of homology (average sequence identities of 67.59% and 60.76% for OtsA and OtsB, respectively). Within OtsA and OtsB protein trees, Halobacteriales proteins formed a coherent, deep-branching cluster that was only distantly related to sequences from other bacterial and archaeal phyla (Figures 3a and b). Further, Halobacteriales OtsA and OtsB phylogenies mirrored those inferred from reference 16S rRNA and RpoB′ trees (Figure 4) (Walsh et al., 2004; Enache et al., 2007; Minegishi et al., 2010). Collectively, these patterns suggest that trehalose production is an ancient trait within the Halobacteriales rather than a recent acquisition through horizontal gene transfer.
Maximum likelihood phylogenetic tree inferred from: (a) trehalose-6-phosphate synthase (OtsA) and (b) trehalose-6-phosphatase (OtsB) protein sequence alignments for members of Halobacteriaceae as well as other bacterial and archaeal species. The optimal model used as predicted by Mega was WAG+Γ with γ shape parameter=1.03, and proportion of invariable sites (I=0.016). Bootstraps values (100 replicates) are shown for the nodes with >50 bootstrap support. The tree was rooted using OtsA/B protein sequence of Schizosaccharomyces japonicum. Numbers in parenthesis represent the number of taxa comprising each wedge.
Maximum likelihood phylogenetic tree inferred from RNA polymerase β-subunit (RpoB′) protein sequence alignment for members of the Halobacteriales. The optimal model used was WAG+Γ as suggested by Minegishi et al. (2010). Bootstraps values (100 replicates) are shown for the nodes with more than 50 bootstrap support. × indicates putative incidences of gene loss events during Halobacteriales evolution. Numbers in parenthesis represent the number of taxa comprising each wedge.
We further examined the occurrence pattern of the ostAB genes within the Halobacteriales by constructing an RpoB′ tree of all species with sequenced genomes (Figure 4). We identified a distinct distribution pattern of the ostAB genes at the supragenus level. This pattern suggests the occurrence of several gene loss events during Halobacteriales evolution. otsAB genes were present in all members of Halobacteriales Clade I (as defined by Walsh et al. (2004), Enache et al. (2007) and Minegishi et al. (2010), including the genera Halovivax, Natronobacterium, Halobiforma, Natronorubrum, Haloterrigena, Halopiger, Natronolimnobius, Natrinema, Natrialba and Natronococcus), as well as the genera Halococcus, Haladaptatus, Halalkalicoccus and Halosimplex. On the other hand, otsAB genes were absent in all members of Halobacteriales clade II (as defined by Walsh et al. (2004), Enache et al. (2007), Minegishi et al. (2010), including the genera Haloferax, Halorubrum, Halogeometricum, Haloquadratum, Haloplanus and Halobaculum), the Halorhabdus–Halomicrobium–Haloarcula clade, as well as within the genera Halobacterium and Natronomonas.
Ecological distribution of Halobacteriales genera with and without OtsAB system
Comparison of the minimum, optimum and maximum salt requirements between various Halobacteriales species revealed that otsAB-harboring genera have lower salt optima than those lacking the system (3.2 M NaCl for OtsAB harboring genera versus 3.8 M NaCl for OtsAB lacking genera, P=0.0372, albeit weak, Cramer’s V value=0.37). No similar significant correlation was identified between otsAB occurrence and salt minima or maxima of various Halobacteriales species.
We quantified the relative abundances of Halobacteriales genera with and without OtsAB system in relation to the total Halobacteriales community with published genomes (referred to as ‘sequenced Halobacteriales community’ thereafter) in previously published data sets from 13 different environments with salinities ranging from permanently hypersaline (n=9), saline and fluctuating salinity habitats (where salinity is usually <250 g l−1, exhibits fluctuation and the Halobacteriales community is usually a fraction of a more diverse prokaryotic community) (n=3) and mostly non-saline with only occasional development of temporal and spatial pockets (n=1, Zodletone spring). OtsAB-lacking genera represented the majority of the sequenced Halobacteriales community in permanently hypersaline habitats such as the Dead Sea and Great Salt lake (87.9% and 85.7% of sequenced Halobacteriales communities, respectively) and different crystallizer ponds examined (100% of sequenced Halobacteriales community, Table 4). On the other hand, the saline and fluctuating salinity habitats examined had a population of both OtsAB-harboring (44.5–55.1% of the sequenced Halobacteriales community) and OtsAB-lacking genera. Finally, Zodletone spring, an environment that is mostly non-saline with only occasional development of temporal and spatial pockets, has the largest proportion of OtsAB-harboring genera (67.6% of the sequenced Halobacteriales community). Principal component analysis conducted using the relative abundances of Halobacteriales genera with and without OtsAB system in relation to the sequenced Halobacteriales community indicated that the two groups of environments (hypersaline versus saline or fluctuating salinity) are divergent with respect to the abundances of OtsAB-lacking and OtsAB-harboring genera (Figure 5).
Discussion
In this study, we demonstrate that the following: (1) trehalose and 2-sulfotrehalose biosynthetic capabilities are widely spread in multiple Halobacteriales genera; (2) otsAB genes in H. paucihalophilus are highly expressed and their expression patterns broadly correspond to trehalose production patterns at various salinities; (3) a phylogenetic distinction exists between otsAB-harboring versus otsAB-lacking Halobacteriales taxa at the genus and supragenus levels; (4) the predominance of otsAB-lacking genera in permanently hypersaline habitats and (5) the occurrence of glycine betaine uptake as an additional osmoadaptive mechanism in the Halobacteriales.
Osmoadaptive landscape in trehalose- and 2-sulfotrehalose-synthesizing Halobacteriales
Quantification of trehalose and 2-sulfotrehalose in 17 different species (Figure 2, Supplementary Table S5) demonstrates that, in general, while these compounds are produced at all examined salinities, their concentrations are highest at lower media salinities. These production patterns are consistent with our understanding of the energetic costs of compatible solutes biosynthesis (Oren, 1999), with prior studies that documented trehalose importance as a compatible solute at lower salinities (for example in Chromohalobacter israelenis (Regeva et al., 1990)) and the fact that within specific lineages such as Cyanobacteria, the utilization of trehalose as an osmoprotectant appears to be more prevalent in strains with relatively lower salt tolerance (Hagemann, 2011).
The observed trehalose/2-sulfotrehalose production patterns, coupled to the near-constant levels of K+ observed at various salinities in the majority of species examined, argues for the need to employ additional osmoadaptive mechanisms when these genera encounter higher salinities. We argue that glycine betaine uptake represents an important mechanism for osmoadaptation in trehalose- and 2 sulfotrehalose-producing genera at higher salinities. Genes for glycine betaine uptake were identified in 36 out of 39 otsAB-harboring genomes, and glycine betaine uptake was experimentally verified in 7 out of 11 trehalose/2 sulfotrehalose-producing species (Table 3). The production of additional yet-unidentified compatible solutes cannot be ruled out, although genomic and experimental analysis failed to identify the production of any of the commonly produced compatible solutes (Roberts, 2005) within examined species.
2-sulfotrehalose, rather than trehalose, was identified as the osmoadaptive agent in multiple Halobacteriales species (Supplementary Table S5). It appears that 2-sulfotrehalose producers exhibit a distinct phylogenetic affiliation and pH preference when compared with trehalose producers. 2-sulfotrehalose producers identified belonged to Halobacteriales Clade II, with the notable exception of Haloalkalicoccus jeotgali. Further, the majority of 2-sulfotrehalose producers identified in this as well as in a prior study (Desmarais et al., 1997) are haloalkaliphiles, with the exception of the two neutrophilic species Natrinema pellirubrum and Halalkaicoccus jeotgali. The rationale for the modification of a non-reducing neutral sugar to an anionic derivative, and whether such modification confers any advantage in alkalisaline conditions is not yet entirely clear. It is interesting to note that orthologs for the well described and only known enzyme specific for trehalose sulfonation (Mougous et al., 2004) were not identified in Halobacteriales genomes examined, and hence an alternative, yet unknown, mechanism must be involved in the process.
Overall, trehalose- and 2-sulfotrehalose-producing lineages had lower intracellular K+ levels when compared with genera incapable of trehalose/2-sulfotrehalose production. However, these levels are significantly higher than those observed in moderately halophilic heterotrophs and Cyanobacteria that are dependent on compatible solute production for osmoadaptation (Ventosa and Oren, 1996; Hagemann, 2011). Therefore, these relatively higher intracellular K+ levels in trehalose- and 2-sulfotrehalose-producing Halobacteriales lineages imply that K+ accumulation still has an integral role in osmoadaptation within these lineages, and that trehalose/2-sulfotrehalose-producing Halobacteriales should not be regarded as solely dependent on compatible solute production and uptake mechanisms for osmoadaptation. In addition, trehalose/2-sulfotrehalose-producing Halobacteriales, similar to other Halobacteriales lineages, have an acidic proteome with a high concentration of Asp and Glu residues (∼17%) and a low predicted pI (4.76±0.12). Hence, the high levels of intracellular K+ in trehalose/2-sulfotrehalose-producing Halobacteriales also function to stabilize the acidic proteome via interactions with the negatively charged acidic protein surface required for increasing the protein hydration shell and preventing unfolding or salting out (Oren and Mana, 2002).
Osmoadaptive landscape in genera lacking OtsAB system
On the other hand, 11 out of 28 genera with sequenced genomes examined lack the genes necessary for trehalose biosynthesis, an observation that was experimentally verified in 6 representative strains. Within this group, a salting-in strategy depending on the accumulation of high levels of intracellular K+ in a salinity-dependent manner appears to be the primary mechanism for osmoadaptation. As such, our results are in agreement with prior studies on osmoadaptive mechanisms in the Halobacteriales (Christian and Waltho, 1962; Ginzburg et al., 1970; Lanyi and Silverman, 1972; Matheson et al., 1976; Pérez-Fillol and Rodríguez-Valera, 1986; Mojica et al., 1997; Oren et al., 2002b), all of which have been conducted on strains that lack trehalose production capabilities (Halobacterium salinarum, Haloarcula marismortui, Haloferax volcanii, Haloferax mediterranei, Haloferax gibbonsii, Halorubrum saccharovorum and Halorubrum trapanicum). However, in addition to the well-established role of intracellular K+ accumulation, this study demonstrates that uptake of the compatible solute glycine betaine is a common process in otsAB-lacking genera. Several bacterial and eukaryotic ineages capable of synthesizing glycine betaine could thrive in hypersaline (>25% NaCl) habitats, for example Cyanobacteria (Oren, 2010) and Dunaliella species (Oren, 2005), and hence this mechanism could have environmental relevance in specific habitats. Although glycine betaine uptake has been reported in Halobacterium salinarum sp. strain NRC-1 (Kokoeva et al., 2002), the overall role of this process in Halobacteriales osmodaptation has largely been unexplored.
Evolutionary and ecological dichotomy between trehalose-/2sulfotrehalose-producing versus non-producing Halobacteriales
Analysis of the phylogeny and distribution of otsAB genes revealed valuable insights into the evolution of osmoadaptive mechanisms, and the implications of such events on the ecological distribution of various Halobacteriales lineages. Phylogenetic analysis indicated that trehalose biosynthesis is an ancient trait that has subsequently been lost in some lineages during Halobacteriales evolution. We argue that the loss of otsAB genes was associated with the spread of Halobacteriales into hypersaline habitats, as this negative selection pressure event would alleviate the energetic costs associated with the synthesis of high level of trehalose to cope with the external hypersaline environments. The resulting energetic benefit of this mutation outweighs its anticipated drawback: the loss of osmoadaptive flexibility, as the benefit of such flexibility is highly diminished in permanently hypersaline habitats. Further, this loss of osmoadaptive flexibility could possibly be compensated for by the capability to uptake glycine betaine from the environment. Indeed, our analysis of Halobacteriales community in multiple habitats of various salinities demonstrated a clear preference and dominance of genera incapable of trehalose production in permanently hypersaline habitats.
In conclusion, this study highlights the role of compatible solute biosynthesis and uptake in the Halobacteriales, and provides ecological and evolutionary context to the observed variations and complexities of osmoadaptive strategies. The complexity and multiplicity of osmoadaptive mechanisms in the Halobacteriales reported here, as opposed to the prior view of sole dependence on a single solute (K+ ions) for osmoadaptation, raise multiple yet-unanswered questions regarding the kinetics of compatible solutes production, as well as the relative contribution, interaction and regulation of various mechanisms in the overall process of osmoadaptation in the Halobacteriales.
Accession codes
References
Avonce N, Mendoza-Vargas A, Morett E, Iturriaga G . (2006). Insights on the evolution of trehalose biosynthesis. BMC Evol Biol 6: 109.
Baati H, Guermazi S, Amdouni R, Gharsallah N, Sghir A, Ammar E . (2008). Prokaryotic diversity of a Tunisian multipond solar saltern. Extremophiles 12: 505–518.
Bidle K, Amadio W, Oliveira P, Paulish T, Sharron H, Earnest C . (2005). A phylogenetic analysis of Haloarchaea found in a solar saltern. Bios 76: 89–96.
Bodaker I, Sharon I, Suzuki MT, Feingersch R, Shmoish M, Andreishcheva E et al. (2009). Comparative community genomics in the Dead Sea: an increasingly extreme environment. ISME J 4: 399–407.
Bowman JP, McCammon SA, Rea SM, McMeekin TA . (2000). The microbial composition of three limnologically disparate hypersaline Antarctic lakes. FEMS Microbiol Lett 183: 81–88.
Burns DG, Camakaris HM, Janssen PH, Dyall-Smith ML . (2004). Combined use of cultivation-dependent and cultivation-independent methods indicates that members of most Haloarchaeal groups in an Australian crystallizer pond are cultivable. Appl Environ Microbiol 70: 5258–5265.
Christian JH, Waltho JA . (1962). Solute concentrations within cells of halophilic and non-halophilic bacteria. Biochim Biophys Acta 65: 506–508.
Desmarais D, Jablonski PE, Fedarko NS, Roberts MF . (1997). 2-Sulfotrehalose, a novel osmolyte in haloalkaliphilic archaea. J Bacteriol 179: 3146–3153.
Ellis DG, Bizzoco RW, Kelley ST . (2008). Halophilic Archaea determined from geothermal steam vent aerosols. Environ Microbiol 10: 1582–1590.
Elshahed MS, Najar FZ, Roe BA, Oren A, Dewers TA, Krumholz LR . (2004). Survey of archaeal diversity reveals an abundance of halophilic Archaea in a low-salt, sulfide- and sulfur-rich spring. Appl Environ Microbiol 70: 2230–2239.
Enache M, Takashi I, Fukushima T, Usami R, Dumitru L, Kamekura M . (2007). Phylogenetic relationships within the family Halobacteriaceae inferred from rpoB’ gene and protein sequences. Int J Syst Evol Microbiol 57: 2289–2295.
Ginzburg M, Sachs L, Ginzburg BZ . (1970). Ion metabolism in a Halobacterium: I. Influence of age of culture on intracellular concentrations. J Gen Physiol 55: 187–207.
Goh F, Jeon YJ, Barrow K, Neilan BA, Burns BP . (2011). Osmoadaptive strategies of the archaeon Halococcus hamelinensis Isolated from a hypersaline stromatolite environment. Astrobiology 11: 529–536.
Grant WD . (2004). Life at low water activity. Philos Trans Roy Soc B 359: 1249–1267.
Grieve CM, Grattan SR . (1983). Rapid assay for determination of water soluble quaternary ammonium compounds. Plant Soil 70: 303–307.
Hagemann M . (2011). Molecular biology of cyanobacterial salt acclimation. FEMS Microbiol Rev 35: 87–123.
Inoue K, Itoh T, Ohkuma M, Kogure K . (2011). Halomarina oriensis gen. nov., sp. nov., a halophilic archaeon isolated from a seawater aquarium. Int J Syst Evol Microbiol 61: 942–846.
Kokoeva MV, Storch KF, Klein C, Oesterhelt D . (2002). A novel mode of sensory transduction in archaea: binding protein-mediated chemotaxis towards osmoprotectants and amino acids. EMBO J 15: 2312–2322.
Lanyi JK, Silverman MP . (1972). The state of binding of intracellular K+in Halobacterium cutirubrum. Can J Microbiol 18: 993–995.
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H et al. (2007). Clustal W and Clustal X version 2.0. Bioinformatics 23: 2947–2948.
Lundquist P, Mårtensson J, Sörbo B, Ohman S . (1980). Turbidimetry of inorganic sulfate, ester sulfate, and total sulfur in urine. Clin Chem 26: 1178–1181.
Mani K, Salgaonkar B, Braganca J . (2012). Culturable halophilic archaea at the initial and crystallization stages of salt production in a natural solar saltern of Goa, India. Aquat Biosyst 8: 15.
Markowitz VM, Chen I-MA, Palaniappan K, Chu K, Szeto E, Grechkin Y et al. (2012). IMG: the integrated microbial genomes database and comparative analysis system. Nucleic Acid Res 40: D115–D122.
Matheson AT, Sprott GD, McDonald IJ, Tessier H . (1976). Some properties of an unidentified halophile: growth characteristics, internal salt concentration, and morphology. Can J Microbiol 22: 780–786.
Minegishi H, Kamekura M, Itoh T, Echigo A, Usami R, Hashimoto T . (2010). Further refinement of the phylogeny of the Halobacteriaceae based on the full-length RNA polymerase subunit B' (rpoB') gene. Int J Syst Evol Microbiol 60: 2398–2408.
Mojica FJ, Cisneros E, Ferrer C, Rodríguez-Valera F, Juez G . (1997). Osmotically induced response in representatives of halophilic prokaryotes: the bacterium Halomonas elongata and the archaeon Haloferax volcanii. J Bacteriol 179: 5471–5481.
Mougous JD, Petzold CJ, Senaratne RH, Lee DH, Akey DL, Lin FL et al. (2004). Identification, function and structure of the mycobacterial sulfotransferase that initiates sulfolipid-1 biosynthesis. Nat Struct Mol Biol 11: 721–729.
Munson MA, Nedwell DB, Embley TM . (1997). Phylogenetic diversity of Archaea in sediment samples from a coastal salt marsh. Appl Environ Microbiol 63: 4729–4733.
Nicolaus B, Gambacorta A, Basso AL, Riccio R, Rosa MD, Grant WD . (1988). Trehalose in Archaebacteria. Syst Appl Microbiol 10: 215–217.
Oh D, Porter K, Russ B, Burns D, Dyall-Smith M . (2010). Diversity of Haloquadratum and other haloarchaea in three, geographically distant, Australian saltern crystallizer ponds. Extremophiles 14: 161–169.
Oren A . (1999). Bioenergetic aspects of halophilism. Microbiol Mol Biol Rev 63: 334–348.
Oren A . (2005). A hundred years of Dunaliella research: 1905–2005. Saline Syst 1: 2.
Oren A . (2010). Mats of filamentous and unicellular Cyanobacteria in hypersaline environments. In: Seckbach J, Oren A (eds). Microbial Mats. Springer: Netherlands, pp 387–400.
Oren A . (2013). Life at high salt concentrations. The Prokaryotes—Prokaryotic Communities and Ecophysiology (4th edn). Springer: Berlin-Heidelburg.
Oren A, Ginzburg M, Ginzburg BZ, Hochstein LI, Volcani BE . (1990). Haloarcula marismortui (Volcani) sp. nov., nom. rev., an extremely halophilic bacterium from the Dead Sea. Intl J Syst Bacteriol 40: 209–210.
Oren A, Heldal M, Norland S, Galinski E . (2002a). Intracellular ion and organic solute concentrations of the extremely halophilic bacterium Salinibacter ruber. Extremophiles 6: 491–498.
Oren A, Mana L . (2002b). Amino acid composition of bulk protein and salt relationships of selected enzymes of Salinibacter ruber, an extremely halophilic bacterium. Extremophiles 6: 217–223.
Perreault NN, Andersen DT, Pollard WH, Greer CW, Whyte LG . (2007). Characterization of the prokaryotic diversity in cold saline perennial springs of the Canadian high arctic. Appl Environ Microbiol 73: 1532–1543.
Plackett RL . (1983). Karl Pearson and the chi-squared test. Int Statist Rev 51: 59–72.
Podell S, Ugalde JA, Narasingarao P, Banfield JF, Heidelberg KB, Allen EE . (2013). Assembly-driven community genomics of a hypersaline microbial ecosystem. PLoS One 8: e61692.
Purdy KJ, Cresswell-Maynard TD, Nedwell DB, McGenity TJ, Grant WD, Timmis KN et al. (2004). Isolation of haloarchaea that grow at low salinities. Environ Microbiol 6: 591–595.
Pérez-Fillol M, Rodríguez-Valera F . (1986). Potassium ion accumulation in cells of different halobacteria. Microbiologia (Madrid, Spain) 2: 73–80.
R Development Core Team (2011) R: A Language and Environment for Statistical Computing. Reference Index., 2.13 (2nd edn) R Foundation for Statistical Computing: Vienna, Austria.
Regeva R, Peria I, Gilboa H, Avi-Dor Y . (1990). 13C NMR study of the interrelation between synthesis and uptake of compatible solutes in two moderately halophilic eubacteria: Bacterium Ba1 and Vibro costicola. Arch Biochem Biophys 278: 106–112.
Roberts M . (2005). Organic compatible solutes of halotolerant and halophilic microorganisms. Saline Syst 1: 5.
Savage KN, Krumholz LR, Oren A, Elshahed MS . (2007). Haladaptatus paucihalophilus gen. nov., sp. nov., a halophilic archaeon isolated from a low-salt, sulfide-rich spring. Int J Syst Evol Microbiol 57: 19–24.
Stivaletta N, Barbieri R, Cevenini F, López-García P . (2011). Physicochemical conditions and microbial diversity associated with the evaporite deposits in the Laguna de la Piedra (Salar de Atacama, Chile). Geomicrobiol J 28: 83–95.
Takagi M, Nakamura H, Sanui Y, Ueno K . (1981). Spectrophotometric determination of sodium by ion-pair extraction with crown ether complexes and monoanionic dyes. Analyt Chim Acta 126: 185–190.
Takai K, Komatsu T, Inagaki F, Horikoshi F . (2001). Distribution of archaea in a black smoker chimney structure. Appl Environ Microbiol 67: 3618–3629.
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S . (2011). MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28: 2731–2739.
Uzawa H, Nishida Y, Sasaki K, Nagatsuka T, Hiramatsu H, Kobayashi K . (2004). Sulfatase-catalyzed assembly of regioselectively O-sulfonated p-nitrophenyl α-d-gluco- and α-d-mannopyranosides. Carbohyd Res 339: 1597–1602.
Ventosa A, Oren A . (1996). Halobacterium salinarum nom. corrig., a name to replace Halobacterium salinarium (Elazari-Volcani) and to include Halobacterium halobium and Halobacterium cutirubrum. Int J Syst Bacteriol 46: 347–347.
Ventosa A, Rodriguez-Valera F, Poindexter JS, Reznikoff WS . (1984). Selection for moderately halophilic bacteria by gradual salinity increases. Can J Microbiol 30: 1279–1282.
Walsh DA, Bapteste E, Kamekura M, Doolittle WF . (2004). Evolution of the RNA polymerase B′ subunit gene (rpoB′) in Halobacteriales: a complementary molecular marker to the SSU rRNA gene. Mol Biol Evol 21: 2340–2351.
Walsh DA, Papke RT, Doolittle WF . (2005). Archaeal diversity along a soil salinity gradient prone to disturbance. Environ Microbiol 7: 1655–1666.
Xiao W, Wang Z-G, Wang Y-X, Schneegurt MA, Li Z-Y, Lai Y-H et al. (2013). Comparative molecular analysis of the prokaryotic diversity of two salt mine soils in southwest China. J Basic Microbiol doi:10.1002/jobm.201200200.
Youssef NH, Ashlock-Savage KN, Elshahed MS . (2012). Phylogenetic diversities and community structure of members of the extremely halophilic Archaea (order Halobacteriales) in multiple saline sediment habitats. Appl Environ Microbiol 78: 1332–1344.
Acknowledgements
We thank Ratnakar Deole and Audra Liggenstoffer for technical assistance, Margret Eastman for NMR analysis and Ramy Aziz for helpful discussions. This work was supported by the National Science Foundation Microbial Observatories Program (grant EF0801858).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Youssef, N., Savage-Ashlock, K., McCully, A. et al. Trehalose/2-sulfotrehalose biosynthesis and glycine-betaine uptake are widely spread mechanisms for osmoadaptation in the Halobacteriales. ISME J 8, 636–649 (2014). https://doi.org/10.1038/ismej.2013.165
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2013.165
Keywords
This article is cited by
-
Assessment of diversity of archaeal communities in Algerian chott
Extremophiles (2023)
-
Physiological and genomic insights into abiotic stress of halophilic archaeon Natrinema altunense 4.1R isolated from a saline ecosystem of Tunisian desert
Genetica (2023)
-
Hly176B, a low-salt tolerant halolysin from the haloarchaeon Haloarchaeobius sp. FL176
World Journal of Microbiology and Biotechnology (2023)
-
Asgard archaea in saline environments
Extremophiles (2022)
-
Revealing the salinity adaptation mechanism in halotolerant bacterium Egicoccus halophilus EGI 80432T by physiological analysis and comparative transcriptomics
Applied Microbiology and Biotechnology (2021)