Abstract
Microbial metabolic activity occurs at subzero temperatures in permafrost, an environment representing ∼25% of the global soil organic matter. Although much of the observed subzero microbial activity may be due to basal metabolism or macromolecular repair, there is also ample evidence for cellular growth. Unfortunately, most metabolic measurements or culture-based laboratory experiments cannot elucidate the specific microorganisms responsible for metabolic activities in native permafrost, nor, can bulk approaches determine whether different members of the microbial community modulate their responses as a function of changing subzero temperatures. Here, we report on the use of stable isotope probing with 13C-acetate to demonstrate bacterial genome replication in Alaskan permafrost at temperatures of 0 to −20 °C. We found that the majority (80%) of operational taxonomic units detected in permafrost microcosms were active and could synthesize 13C-labeled DNA when supplemented with 13C-acetate at temperatures of 0 to −20 °C during a 6-month incubation. The data indicated that some members of the bacterial community were active across all of the experimental temperatures, whereas many others only synthesized DNA within a narrow subzero temperature range. Phylogenetic analysis of 13C-labeled 16S rRNA genes revealed that the subzero active bacteria were members of the Acidobacteria, Actinobacteria, Chloroflexi, Gemmatimonadetes and Proteobacteria phyla and were distantly related to currently cultivated psychrophiles. These results imply that small subzero temperature changes may lead to changes in the active microbial community, which could have consequences for biogeochemical cycling in permanently frozen systems.
Similar content being viewed by others
Introduction
Arctic and boreal environments cover over 20% of the terrestrial surface, and the northern circumpolar permafrost region contains ∼25% of the world’s total soil organic material (Tarnocai et al., 2009). Increased warming in polar environments is expected to convert these permafrosts into wetlands or thermokarsts, promoting microbial remineralization of stored organic matter as thawing occurs (Schuur et al., 2008). However, many studies dating back to the 1960s (Michener and Elliott, 1964; Larkin and Stokes, 1968) have clearly indicated that microbial life continues below the freezing point, with the first indication of subzero activity in situ coming from Antarctic studies of the cryptoendolithic lichens (Kappen, 1993 (Review)).
Advances in research of subzero microbiology over the past two decades have shown unambiguously that subzero activity occurs. Examples include, but are not limited to the following: measurements of hydrolytic enzyme activity (Gilichinsky et al., 1992); incorporation of isotopic labels/bromodeoxyuridine into macromolecules/metabolites (Carpenter et al., 2000; Rivkina et al., 2000; Christner 2002; Junge et al., 2006; McMahon et al., 2009; Drotz et al., 2010); respiratory evolution of gases (for example, 14CO2) (Gilichinsky et al., 2003; Panikov et al., 2006; Steven et al., 2007; Lacelle et al., 2011); and 14CO2 incorporation into cells (Panikov and Sizova, 2007; Panikov, 2010 (Review)). Unfortunately, these bulk approaches cannot elucidate which specific microorganisms are growing in subzero environments (for example, native permafrost) or whether different members of the microbial community modulate their responses as a function of changing subzero temperatures.
In the current study, we utilized a stable isotope probing (SIP) approach to discern the active bacteria in permafrost (beyond basal metabolism) while in the frozen state. Prior research in Arctic systems has used SIP to monitor active methanotrophs with 13CH4, but the studies were performed at higher incubation temperatures ranging from 4 to 25 °C (Martineau et al., 2010; Liebner et al., 2011; He et al., 2012). Here, we used the SIP method to elucidate how specific members of the bacterial community responded to various subzero incubation temperatures ranging from 0 to −20 °C. SIP was used as it is predicated on the incorporation of 13C carbon by active microorganisms into DNA and the physical separation from the 12C DNA in a cesium chloride density gradient (Radajewski et al., 2000). Therefore, SIP will not detect bacterial cells which are carrying out basal metabolism nor will it provide a signal if the bacteria are performing DNA excision/repair (which usually involves synthesis of 12 bp fragments after damaged base pair removal). Our results indicate that a broad range of permafrost bacteria were able to synthesize DNA between 0 and −20 °C, many of which were only active within a narrow subzero temperature limit. Phylogenetic analysis of 16S rRNA gene clone libraries revealed that the subzero active bacteria were members of the Acidobacteria, Actinobacteria, Chloroflexi, Gemmatimonadetes and the Proteobacteria phyla and were distantly related to currently cultivated psychrophiles. A better understanding of the temperature boundary conditions where Arctic-soil microbial communities can be active should improve efforts to estimate global carbon biogeochemistry, greenhouse gas emission and potential impact on atmospheric radiative balance in high-latitude environments.
Materials and methods
Permafrost SIP microcosms
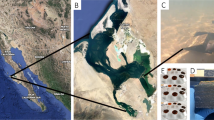
Permafrost cores were collected ∼10 m from the shore of Smith Lake, Fairbanks, Alaska in June 2009. The landscape was characterized by shrub and small trees. The permafrost samples were taken using a two-man portable, gas-powered, coring machine with a drilling inside diameter of 7.5 cm. The thaw depth was about 20 cm at the time of drilling, the active layer of thickness was about 65 cm and the permafrost temperature at a depth of about 1.0 m was ∼−4 to −5 °C. Before drilling, the thawed layer was removed and the drilling started from the frozen surface downwards by more than 1.0 m. After collection, the samples were cooled to −80 °C and shipped to Rutgers University within 24 h, where they arrived frozen, and were stored at −80 °C before the SIP experiments.
Subsamples from the interior of the core (to minimize contamination), ∼30 cm below the base of the active layer, were homogenized using sterilized tools while frozen in a −20 °C walk-in chamber and distributed in 1 g fractions into 2 ml microcentrifuge tubes. The individual microcosms were randomly selected and amended once with 300 μM (50 μl total volume) of either 12C-acetate (controls, n=12) or 13C-acetate (experimental/SIP, n=24). The homogenized soils were briefly (<3 min) placed at 4 °C and vortexed to uniformly distribute the acetate, then immediately transferred to incubators set to either 0 °C, −3 °C, −6 °C, −9 °C, −12 °C or −20 °C for the duration of the experiment. A single control and duplicate experimental incubations for each treatment were halted after 30 and 180 days by flash freezing in liquid nitrogen and samples were stored at −80 °C until processed.
Bacterial community and SIP analysis
Total genomic DNA was purified using a modified phenol/chloroform protocol (Sakano and Kerkhof, 1998) for subsamples of untreated permafrost (maintained at −80 °C), as well as for all SIP microcosm samples incubated at the subzero temperatures. Briefly, five freeze-thaws were followed by a 15 min lysozyme treatment. Samples were then vortexed for ∼15 s with a 1:4 volume of a potassium acetate/acetic acid solution (60% 5 M potassium acetate, 11.5% glacial acetic acid, 28.5% H2O) to suppress the co-purification of humic compounds. This was immediately followed by a 2 × phenol/chloroform extraction. Purified DNA (200–300 ng) was then subjected to cesium chloride (CsCl) isopycnal centrifugation (225 000 g for 48 h) with 100 ng of archaeal (Halobacterium salinarum) 13C-carrier DNA, which has been shown to reduce the amount of 13C-incorporation necessary for signal detection in an SIP experiment (Gallagher et al., 2005). This carrier/SIP approach yields a 12C (top) band and a 13C (bottom) band for all microcosm incubations, representing the resident (12C) and the active (13C) bacterial community, respectively (Figure 1). These 12C and 13C-DNA bands were collected by pipette, rather than fractionating the entire gradient. This approach has been shown to reduce contamination and ensure clean 12C-control amplifications (Padmanabhan et al., 2003; Gallagher et al., 2005; Kerkhof et al., 2011). This does not imply that 12C DNA contamination is not present throughout the gradient. Rather, the lack of detectable amplicon in the 13C-carrier bands of our 12C-control incubations demonstrated that any contamination by 12C bacterial DNA was below our PCR detection limits.
Schematic of SIP/carrier method and TRFLP analysis. Separation of 12C and 13C-DNA in a cesium gradient allows for the fingerprinting of the resident members (blue profile) and active members (red profile) of the bacterial community. The 13C experimental treatment (top path) yields amplification in the carrier band (red peaks), whereas the 12C-control incubations (bottom path) have no PCR/TRFLP signal in the carrier band (red flat-line).
Because reduced metabolic rates under subzero temperature conditions can limit our ability to directly measure growth and the production of biomolecules, all samples were subjected to a two-step PCR 16S rRNA gene amplification (that is, two consecutive PCR reactions). Equal volumes of genomic DNA (5 μl) from either the top (12C) or bottom (13C) bands were added to individual PCR reaction mixtures (50 μl total reaction volume) to allow for the comparison of amplification strength across all treatments and controls. Reaction mixtures contained 20 pmol of the universal bacterial primers 27 Forward (5′-AGAGTTTGATCCTGGCTCAG-3′; labeled with the fluorochrome 6′-FAM (6′-carboxyfluorescein)) and 1100 Reverse (5′-AGGGTTGCGCTCGTTG-3′) per reaction. The PCR amplification parameters were as follows: 94 °C for 5 min and then 94 °C for 30 s, 57 °C for 30 s and 72 °C for 50 s for 25 cycles with a final extension at 72 °C for 7 min using an Applied Biosystems (Foster City, CA, USA) 2720 thermocycler. A small volume of this first round of amplification product (2 μl) was used as a template for the second round of PCR using identical amplification parameters. During the second amplification, aliquots were collected at different cycle numbers (16, 18, 20, 22, 25 and 30 cycles) to determine the lowest cycle number, where either the top or bottom bands yielded a PCR product detected by gel electrophoresis. All 13C-acetate amendments had amplicons at 16 cycles in the carrier band, whereas 12C DNA contamination in the carrier was only apparent in some samples at ⩾20 cycles. Equal masses of fluorescent PCR product (20 ng) amplified at 16 cycles for all microcosms were digested with Mnl I in 20 μl reactions for 6 h at 37 °C. For 12C-acetate controls, the largest volume at 16 cycles used for the 13C-acetate treatments (that is, 5 μl) was digested. The digests were precipitated and resuspended in 19.7 μl of deionized formamide with 0.3 μl of ROX 500 standard and analyzed on an ABI 310 Genetic analyzer with peak detection/quantification using Genescan software (Applied Biosystems). A peak detection limit was calculated as 2 × the average 12C-control bottom-band peak area (Supplementary Table S1). This represented 0.2% of the total average bottom band profile areas of the 13C-amended microcosm samples. Therefore, all peaks that fell below 0.2% of the respective total profile area were excluded from further analysis.
To unambiguously demonstrate 13C-DNA synthesis, the SIP experiments had to meet the following criteria: (a) all 13C-carrier bands (bottom) resulted in little or no amplification/terminal-restriction fragment length polymorphism (TRFLP) signal after incubation with 12C-acetate and (b) the profiles from the 13C-carrier bands incubated with 13C-acetate had to demonstrate either enrichment or suppression of specific TRFLP peaks with respect to their 12C top-band counterparts to rule out contamination. All samples from the 1 month incubations yielded a PCR product from the 12C top bands, whereas the 13C bottom bands yielded either no amplicon or comparable product for both the 12C and 13C-acetate amendments at 20+ cycles of the second step of PCR. However, after 6 months, all treatments amended with 13C-acetate (with the exception of one of the −6 °C biological replicates) yielded PCR products from the 13C-carrier band after 16 cycles of the second step of PCR, whereas all 12C-control treatments did not produce amplicons in the 13C-carrier band under identical conditions (Supplementary Figures S1A–F). To analyze for the enrichment or suppression of different operational taxonomic units (OTUs) we constructed a single composite profile from the replicates and displayed the resident (12C band) and the active (13C band) fingerprints above and below the x axis, respectively, for each temperature incubation using Grapher (Golden Software Inc., Golden, CO, USA; Supplementary Figures S2A–C).
Data analysis
To include all peaks (⩾ detection limit) from all replicates within a treatment, each set of duplicate profiles (for example, 0° replicates A and B, −3° replicates A and B and so on; see Supplementary Figures S1A–F) was first normalized to the highest total community profile area within the pair to compensate for between-run instrument variation. The replicate TRFLP peak areas were then averaged and percent contribution to the overall community profile area was calculated. For assessing the temperature grouping of 13C peaks, the TRFLP peak area data were exported into Excel (Microsoft) and each OTU was categorized according to the presence/absence at all incubation temperatures.
To test whether the observed grouping of OTUs by temperature regime resulted from the analytical/detection level artifacts and the binning approach, we reanalyzed the community profiles utilizing sequentially increasing criteria for peak exclusion. That is, the categorizing/sorting analysis was performed iteratively, excluding peaks that represented less than 0%, 0.2%, 0.5%, 1.0%, 2.0%, 4.0% or 8.0% of the total profile area, respectively (Supplementary Figure S3). Using this approach, highly similar temperature groupings were observed, suggesting that both large and small peaks in the active community profiles demonstrate a distinct temperature pattern.
Clone library generation
Cloning from the 13C-DNA fraction of the −9 °C and −20 °C incubations utilized a TOPO TA cloning kit (Invitrogen, Grand Island, NY, USA). Individual colonies from the libraries (240 clones per temperature treatment) were individually screened by TRFLP to identify partial 16S rRNA genes corresponding to specific TRFLP peaks in the active permafrost-community fingerprints as previously described (Gallagher et al., 2005; Kerkhof et al., 2011). Plasmid preps of recombinant clones corresponding to specific TRFLP peaks were performed using Zyppy Plasmid Miniprep Kits (Zymo, Irvine, CA, USA), and sequenced by the Sanger method (Genewiz, South Plainfield, NJ, USA). Phylogenetic reconstruction began with ∼1100 bp of the permafrost clonal sequences, sequences demonstrating the closest BLAST match (Genbank, NCBI, Bethesda, MD, USA), some known psychrophilic bacteria and representatives of the major bacterial phyla (Supplementary Table S2). A maximum likelihood tree was reconstructed from 784 unambiguously aligned bases and 91 taxa from Genbank or identified in our SIP study using Geneious Pro (Babcock et al., 2007; Drummond et al., 2009).
Results
After 1 and 6 months, independent replicate microcosms from each treatment were destructively sampled and analyzed. After 1 month of incubation, none of the permafrost microcosms unambiguously demonstrated 13C-acetate incorporation (that is, no amplification in 13C bands regardless of treatment or amplification in bottom bands in both 12C and 13C amendments at⩾20 cycle during the second PCR step; see methods). However, after 6 months of incubation, the 13C-DNA synthesis signal was clear and there were no carrier band amplification in 12C controls (for example, Figure 2; for all profiles see Supplementary Figure S1). Overall, 152 different OTUs were detected in the active 13C fractions of the independent microcosms, indicating that a broad range of bacteria were synthesizing DNA at subzero temperatures. Across all subzero temperature treatments, 67% (± 5%) of the 13C peaks were also observed in the 12C fraction. The remaining 33% (±5%) of the 13C peaks were not detectable in the resident, top-band community profile (Supplementary Figure S2). This suggests that these bacteria were present in the resident community but were below our PCR/TRFLP detection limit until stimulated as a result of acetate addition and/or incubation temperature.
Representative control and experimental SIP profiles from the −20 °C permafrost incubations. (a) Control 12C-acetate treatment demonstrating the resident (blue) and active (red) community fingerprints. (b) Experimental 13C-acetate treatment with resident (blue) and active (red) profiles. (c) Composite profiles from the replicate 13C- treatments with the resident profiles (blue line/above axis) and the active profiles (red line/below axis). Specific TRFs that are suppressed (dashed arrows) or enriched (solid arrows) are indicated.
The various 13C-TRFLP peaks (Figure 3, y axis) could be categorized into five groups when expressed as a relative percentage of the overall active community profile (color) plotted against incubation temperatures (Figure 3, x axis). Group 1 consisted of OTUs that synthesized 13C-DNA at 0 to −6 °C and accounted for 14% of all the 13C-labeled TRFLP peaks detected in the study at the various incubation temperatures. Group 2 represented 9% of the 13C-OTUs and only displayed activity at −6 and/or −9 °C. The third group of active microorganisms synthesized DNA at temperatures from −9 to −20 °C and made up 13% of the total 13C-OTUs. A fourth group of active OTUs (15% of all 13C-TRFLP peaks) were observed continuously across the entire range of experimental temperatures (0 to −20 °C). Examples of peaks that occurred in either a narrow temperature range or across all incubation temperatures are readily visualized by examining magnified portions of the raw 13C-profiles (Supplementary Figure S4). Finally, a fifth group of OTUs, representing the largest set of TRFLP peaks in the 13C-profiles (49% of all active OTUs), demonstrated a discontinuous distribution over the incubation temperatures. Many of these continuously/discontinuously distributed 13C-TRFLP peaks likely represent multiple bacterial species with different temperature preferences that are not resolved using our TRFLP approach (see below). The 152 active OTUs detected in this analysis at any given incubation temperature represented 80% of the OTUs observed in untreated permafrost samples stored at −80 °C (Supplementary Figure S5), suggesting that the majority of the Arctic permafrost community is capable of metabolizing acetate (or a product of cross-feeding) at subzero temperatures.
Heat map displaying relative contribution (see color key) of active bacterial OTUs (y axis) in the permafrost-community profiles at the various incubation temperatures (x axis). The OTUs that were cloned and sequenced in the current study are denoted by colored circles on the y axis (also see Figure 4).
To identify the active bacteria in permafrost and assign phylogenetic affiliation to TRFLP peaks, 16S rRNA gene clone libraries were constructed from the 13C-DNA fraction of the −9 °C and −20 °C treatments. Each clone was screened by TRFLP and sequenced as described previously. This approach allows for the matching of nearly full-length 16S rRNA genes (1100 bp) with the corresponding peaks from the 13C-TRFLP profiles (Figures 3 and 4, colored circles). Thirty-eight distinct OTUs were identified among 480 clones (240 per temperature treatment), and a phylogenetic tree was reconstructed using maximum likelihood methods on 784 unambiguously aligned bases of 91 taxa (Figure 4). Many of the active permafrost clones represent deeply branching 16S rRNA phylotypes, distantly related to cultivated psychrophilic strains exhibiting growth at subzero temperatures and to 16S rRNA gene sequences of uncultured bacteria from similar habitats (Supplementary Table S2). The active OTUs were members of diverse phyla, including the Acidobacteria, Actinobacteria, Chloroflexi, Gemmatimonadetes and the Proteobacteria and predominantly represented by the discontinuous terminal restriction fragments (Group 5). In some instances, a Group 5 terminal restriction fragment was represented by two different taxa identified within the clone libraries (for example, terminal restriction fragment 75, 200 and 219), supporting the notion of multiple 16S rRNA genes corresponding to a single terminal restriction fragment peak. Interestingly, none of the clones were affiliated with Firmicutes (for example, Bacillus, Exiguobacterium and Clostridium), which are routinely detected in surveys of permafrost biodiversity (Shi et al., 1997; Gilichinsky et al., 2008). This may be the result of the use of 13C-acetate to discern activity. Alternatively, because spore-forming bacteria may remain in a dormant, non-metabolically active state in subzero environments (Bakermans and Nealson, 2004; Johnson et al., 2007), it is not surprising that Gram+bacteria would not be detected by the clonal libraries derived from the 13C-DNA.
Maximum likelihood phylogenetic tree reconstruction of 16S rRNA genes from the 13C-DNA fraction from the −9 and −20 °C incubations with cultivated psychroactive strains, uncultured phylotypes and other representative species (784 unambiguously aligned bases and 91 taxa). The 13C-labeled TRFs from this study and their respective temperature grouping are indicated by colored circles (see also Figure 3). Source habitat information and GenBank accession numbers of ALL taxa included in the tree construction are presented in Supplementary Table S2.
Discussion
Many recent studies on permafrost bacterial growth and metabolism have focused on temperatures at or above 0 °C, assuming that metabolic activity predominantly occurs when permafrost soils have thawed (Martineau et al., 2010; Liebner et al., 2011; Mackelprang et al., 2011; He et al., 2012; Nicholson et al., 2013). However, the concept that bacterial activity in frozen soils can also have a major role in biogeochemical cycles is changing (Bergholz et al., 2009; McMahon et al., 2009; Vishnivetskaya et al., 2009; Mykytczuk et al., 2012; Tveit et al., 2013). Here, we have demonstrated bacterial growth on 13C-acetate at temperatures down to −20 °C. Our findings of subzero DNA synthesis after 6 months are in agreement with earlier reports indicating microbial growth/metabolism in permafrost within 100–160 days at similar temperature (Rivkina et al., 2000), growth of bacterial isolates at −5 to −15 °C (Bakermans and Nealson, 2004; Mykytczuk et al., 2012, 2013) and acetate incorporation into bacterial lipids in permafrost down to −20 °C (Rivkina et al., 2000). Furthermore, the active bacterial phyla observed in this SIP study (Acidobacteria, Actinobacteria, Chloroflexi, Gemmatimonadetes and many Alphaproteobacteria) have also been shown to produce ribo-tags and mRNA in a metatranscriptome study of the active layer of a Svalbard fen (Tveit et al., 2013). Although comparable results regarding doubling times/temperatures/active microbial phyla have been observed in prior studies, it was surprising to find that certain microorganisms appeared to only synthesize DNA at −9 to −20 °C but not above −6 °C. One possible mechanism to account for the differential temperature response in our experimental microcosms is the change in solute concentrations in the remaining liquid water surrounding the soil particles that occur with decreasing temperatures. Increases in both the production of extracellular polymeric substances (acting as cryoprotectants) and in bacterial abundance under similar subzero temperature conditions have been observed in brine channels from sea ice and frost flowers (Krembs et al., 2002; Collins et al., 2008; Meiners et al., 2008; Bowman and Deming 2010; Krembs et al., 2011). This hypothesis is further supported by recent evidence that permafrost isolates have thermohaline-dependent responses for both polysaccharide and fatty acid composition (Ponder et al., 2005) as well as for gene expression patterns (Mykytczuk et al., 2013). Alternatively, the activity of hypo-psychrophilic bacteria may represent a shift in competition for resources, where opportunistic psychrophiles that are adept at garnering nearly all the organic carbon at near freezing temperatures are inactive below −6 °C. This would allow different members of the psychrophilic community to now incorporate the 13C-labeled carbon and replicate their genomes. Regardless of the underlying mechanisms for the observed growth only between −9 and −20 °C, further studies are needed to replicate and confirm our observations in permafrost.
Interestingly, SIP bacterial genome replication below −6 °C was observed in 22% of the 13C-labeled OTUs detected in this study. If many of the continuously/discontinuously distributed TRFs represent multiple microorganisms (as our phylogenetic analysis suggests) with each OTU active in the same narrow subzero temperature range, the number of bacterial phylotypes capable of growth exclusively below −6 °C could be higher. Recently, a metagenomic study of thawed permafrost suggests that small changes in temperature above freezing can have a profound influence on the metabolic diversity of a permafrost bacterial community (Mackelprang et al., 2011). In this report, we present evidence that a comparable temperature shift in permafrost while in the frozen state (for example, from −10 to −5 °C) may also have the potential to stimulate different bacterial communities that may have divergent metabolic capabilities. These results indicate that it will also be important for future work to determine whether changes in subzero temperatures similarly influence the activity of methanogenic archaea. Unfortunately, the use of archael carrier in our SIP experiments precludes the ability to detect both bacterial and methanogenic 13C-DNA within the same samples.
Combined, these data imply that small temperature changes could have consequences for the degradation of buried organic carbon in permanently frozen systems, similar to those experiencing thaw (Graham et al., 2011; Mackelprang et al., 2011). As the permafrost of today is becoming the active layer of tomorrow, a better understanding of microbial activity while these soils are frozen is clearly necessary. This finding is important as many global-scale land-surface models do not adequately consider the extent of microbial mineralization processes in seasonally frozen soils. Specifically, Schwalm et al. (2010) assessed 22 biosphere models to simulate monthly CO2 exchange in North America and found that output mismatches were greatest in tundra/wetland ecosystems in winter and spring compared with forests/temperate regions in summer and fall. The authors concluded that ‘loss of model skill was linked to insufficient characterization of cold temperature sensitivity of metabolic processes....’. Addressing this critical knowledge gap in subzero activity should improve quantitative models (for example, CO2/CH4 production as a function of subzero temperatures) and predictive understanding of terrestrial ecosystems at high latitudes.
In conclusion, we have demonstrated that some members of the permafrost bacterial community utilize 13C-acetate over a relatively narrow subzero temperature range, whereas others appear to be active over a broad frozen temperature regime. Regardless of the mechanism accounting for this observed subzero activity, this class of ‘hypo-psychrophiles’ denotes a novel component of the permafrost bacterial community. If −20 °C represents a temperature close to the lower temperature limits of life in permafrost, one would expect to observe a large reduction in the number of active community members with a decrease in incubation temperature. However, our SIP data does not demonstrate a significant reduction in active OTUs at colder temperatures (Supplementary Figure S6). Collectively, this implies that DNA synthesis could occur at temperatures much lower than −20 °C, where evidence of respiration or metabolic activity has also been observed (Junge et al., 2006; Panikov et al., 2006; Lacelle et al., 2011).
Finally, our experimental results represent a response to acetate addition, for one permafrost community at a single location and depth with an estimated age of 4.5–5 Kya (Waldrop et al., 2010). Furthermore, our laboratory incubations were done at static temperatures and do not represent the natural progression of temperature change that would be occurring in Arctic environments over the course of years or decades. Questions remain as to whether similar microbial activity patterns are wide spread within permafrost soils, with access to a multitude of natural carbon sources, and how microbial activity might vary with permafrost age. Ultimately, these data support prior findings of biological processes in frozen polar soils and also substantiates the notion that life is currently possible on colder planets (Tsapin and McDonald, 2004), such as Mars, where surficial soils will likely be in a permanently frozen state.
References
Babcock DA, Wawrik B, Paul JH, McGuinness L, Kerkhof LJ . (2007). Rapid screening of a large insert BAC library for specific 16S rRNA genes using TRFLP. J Microbiol Meth 71: 156–161.
Bakermans C, Nealson KH . (2004). Relationship of critical temperature to macromolecular synthesis and growth yield in Psychrobacter cryopegella. J Bacteriol 186: 2340–2345.
Bergholz PW, Bakermans C, Tiedje JM . (2009). Psychrobacter arcticus 273-4 uses resource efficiency and molecular motion adaptations for subzero temperature growth. J Bacteriol 191: 2340–2352.
Bowman JS, Deming JW . (2010). Elevated bacterial abundance and exopolymers in saline frost flowers and implications for atmospheric chemistry and microbial dispersal. Geophys Res Lett 37: doi: 10.1029/2010GL043020.
Carpenter EJ, Lin S, Capone DG . (2000). Bacterial activity in South Pole Snow. Appl Environ Microbiol 66: 4514–4517.
Christner BC . (2002). Incorporation of DNA and protein precursors into macromolecules by bacteria at -15 °C. Appl Environ Microbiol 68: 6435–6438.
Collins RE, Carpenter SD, Deming JW . (2008). Spatial heterogeneity and temporal dynamics of particles, bacteria, and pEPS in Arctic winter sea ice. J Mar Syst 74: 902–917.
Drotz SH, Sparrman T, Nilsson MB, Schleucher J, Oquist MG . (2010). Both catabolic and anabolic heterotrophic microbial activity proceed in frozen soils. Proc Natl Acad Sci USA 107: 21046–21051.
Drummond AJ, Ashton B, Cheung M, Heled J, Dearse M, Moir R et al (2009), Geneious v4.5. Available from http://www.geneious.com.
Gallagher E, McGuinness L, Phelps C, Young LY, Kerkhof LJ . (2005). 13C-carrier DNA shortens the incubation time needed to detect benzoate-utilizing denitrifying bacteria by stable-isotope probing. Appl Environ Microbiol 71: 5192–5196.
Gilichinsky D, Rivkina E, Shcherbakova V, Laurinavichuis K, Tiedje J . (2003). Supercooled water brines within permafrost - An unknown ecological niche for microorganisms: a model for astrobiology. Astrobiology 3: 331–341.
Gilichinsky D, Vishnivetskaya T, Petrova M, Spirina E, Mamykin V, Rivkina E . (2008). Bacteria in permafrost Psychrophiles: from Biodiversity to Biotechnology Margesin R, Schinner F, Marx J-C, Gerday C (eds). Springer-Verlag: Berlin Heidelberg, Germany, pp 83–102.
Gilichinsky DA, Vorobyova EA, Erokhina LG, Fyordorov-Dayvdov DG, Chaikovskaya NR . (1992). Long-term preservation of microbial ecosystems in permafrost. Adv Space Res 12: 255–263.
Graham DE, Wallenstein MD, Vishnivetskaya TA, Waldrop MP, Phelps TJ, Pfiffner SM et al (2011). Microbes in thawing permafrost: the unknown variable in the climate change equation. ISME J 6: 709–712.
He R, Wooller MJ, Pohlman JW, Quensen J, Tiedje JM, Leigh MB . (2012). Shifts in identity and activity of methanotrophs in arctic lake sediments in response to temperature changes. Appl Environ Microbiol 78: 4715–4723.
Johnson SS, Hebsgaard MB, Christensen TR, Mastepanov M, Nielsen R, Munch K et al (2007). Ancient bacteria show evidence of DNA repair. Proc Natl Acad Sci USA 104: 14401–14405.
Junge K, Eicken H, Swanson BD, Deming JW . (2006). Bacterial incorporation of leucine into protein down to −20 °C with evidence for potential activity in sub-eutectic saline ice formations. Cryobiology 52: 417–429.
Kappen L . (1993). Lichens in the Antarctic region. In: Friedmann EI (eds.) Antarctic Microbiology. Wiley-Liss: New York, NY, USA, pp 433–490.
Kerkhof LJ, Williams KH, Long PE . (2011). Phase preference by active, acetate-utilizing bacteria at the rifle, CO integrated field research challenge site. Environ Sci Technol 45: 1250–1256.
Krembs C, Eicken H, Deming JW . (2011). Exopolymer alteration of physical properties of sea ice and implications for ice habitability and biogeochemistry in a warmer Arctic. Proc Natl Acad Sci USA 108: 3653–3658.
Krembs C, Eicken H, Junge K, Deming JW . (2002). High concentrations of exopolymeric substances in Arctic winter sea ice: implications for the polar ocean carbon cycle and cryoprotection of diatoms. Deep-Sea Res Pt I 49: 2163–2181.
Lacelle D, Radtke K, Clark ID, Fisher D, Lauriol B, Utting N et al (2011). Geomicrobiology and occluded O2-CO2-Ar gas analyses provide evidence of microbial respiration in ancient terrestrial ground ice. E&PSL 306: 46–54.
Larkin JM, Stokes JL . (1968). Growth of psychrophilic microorganisms at subzero temperatures. Can J Microbiol 14: 97–101.
Liebner S, Zeyer J, Wagner D, Schubert C, Pfeiffer E-M, Knoblauch C . (2011). Methane oxidation associated with submerged brown mosses reduces methane emissions from Siberian polygonal tundra. J Ecol 99: 914–922.
Mackelprang R, Waldrop MP, DeAngelis KM, David MM, Chavarria KL, Blazewicz SJ et al (2011). Metagenomic analysis of a permafrost microbial community reveals a rapid response to thaw. Nature 480: 368–371.
Martineau C, Whyte LG, Greer CW . (2010). Stable isotope probing analysis of the diversity and activity of methanotrophic bacteria in soils from the Canadian High Arctic. Appl Environ Microbiol 76: 5773–5784.
McMahon SK, Wallenstein MD, Schimel JP . (2009). Microbial growth in Arctic tundra soil at −2 °C. Environ Microbiol Rep 1: 162–166.
Meiners K, Krembs C, Gradinger R . (2008). Exopolymer particles: microbial hotspots of enhanced bacterial activity in Arctic fast ice (Chukchi Sea). Aquat Microb Ecol 52: 195–207.
Michener HD, Elliott RP . (1964). Minimum growth temperatures for food-poisoning, fecal-indicator, and psychrophilic microorganisms. Adv Food Res 13: 349–396.
Mykytczuk NCS, Foote SJ, Omelon CR, Southam G, Greer CW, Whyte LG . (2013). Bacterial growth at −15 °C; molecular insights from the permafrost bacterium Planococcus halocryophilus Or1. ISME J 7: 1211–1226.
Mykytczuk NCS, Wilhelm RC, Whyte LG . (2012). Planococcus halocryophilus sp nov., an extreme sub-zero species from high Arctic permafrost. Int J Syst Evol Microbiol 62: 1937–1944.
Nicholson WL, Krivushin K, Gilichinsky D, Schuerger AC . (2013). Growth of Carnobacterium spp. from permafrost under low pressure, temperature, and anoxic atmosphere has implications for Earth microbes on Mars. Proc Natl Acad Sci USA 110: 666–671.
Padmanabhan P, Padmanabhan S, DeRito C, Gray A, Gannon D, Snape JR et al (2003). Respiration of 13C-labeled substrates added to soil in the field and subsequent 16S rRNA gene analysis of 13C-labeled soil DNA. Appl Environ Microbiol 69: 1614–1622.
Panikov NS . (2010). Microbial activity in frozen soils. In: Margesin R (eds) Permafrost Soils.DOI: 10.1007/978-3-540-69371-0 Springer-Verlag: Berlin Heidelberg, Vol Soil Biology 16: pp 119–148.
Panikov NS, Flanagan PW, Oechel WC, Mastepanov MA, Christensen TR . (2006). Microbial activity in soils frozen to below −39 °C. Soil Biol Biochem 38: 785–794.
Panikov NS, Sizova MV . (2007). Growth kinetics of microorganisms isolated from Alaskan soil and permafrost in solid media frozen down to −35 °C. FEMS Microbiol Ecol 59: 500–512.
Ponder MA, Gilmour SJ, Bergholz PW, Mindock CA, Hollingsworth R, Thomashow MF et al (2005). Characterization of potential stress responses in ancient Siberian permafrost psychroactive bacteria. FEMS Microbiol Ecol 53: 103–115.
Radajewski S, Ineson P, Parekh NR, Murrell JC . (2000). Stable-isotope probing as a tool in microbial ecology. Nature 403: 646–649.
Rivkina EM, Friedmann EI, McKay CP, Gilichinsky DA . (2000). Metabolic activity of permafrost bacteria below the freezing point. Appl Environ Microbiol 66: 3230–3233.
Sakano Y, Kerkhof L . (1998). Assessment of changes in microbial community structure during operation of an ammonia biofilter with molecular tools. Appl Environ Microbiol 64: 4877–4882.
Schuur EAG, Bockheim J, Canadell JG, Euskirchen E, Field CB, Goryachkin SV et al (2008). Vulnerability of permafrost carbon to climate change: Implications for the global carbon cycle. Bioscience 58: 701–714.
Schwalm CR, Williams CA, Schaefer K, Anderson R, Arain MA, Baker I et al (2010). A model-data intercomparison of CO2 exchange across North America: results from the North American Carbon Program site synthesis. J Geophys Res Biogeosci 115: doi:10.1029/2009JG001229.
Shi T, Reeves RH, Gilichinsky DA, Friedmann EI . (1997). Characterization of viable bacteria from Siberian permafrost by 16S rDNA sequencing. Microb Ecol 33: 169–179.
Steven B, Briggs G, McKay CP, Pollard WH, Greer CW, Whyte LG . (2007). Characterization of the microbial diversity in a permafrost sample from the Canadian high Arctic using culture-dependent and culture-independent methods. FEMS Microbiol Ecol 59: 513–523.
Tarnocai C, Canadell JG, Schuur EAG, Kuhry P, Mazhitova G, Zimov S . (2009). Soil organic carbon pools in the northern circumpolar permafrost region. GBioC 23: doi:10.1029/2008GB003327.
Tsapin AI, McDonald GD . (2004). Terrestrial permafrost as a model environment for bioastronomy. Bioastronomy 2002: life among the stars In: Norris RP, Stootman FH (eds) Vol. 213.
Tveit A, Schwacke R, Svenning MM, Urich T . (2013). Organic carbon transformations in high-Arctic peat soils: key functions and microorganisms. ISME J 7: 299–311.
Vishnivetskaya TA, Kathariou S, Tiedje JM . (2009). The exiguobacterium genus: biodiversity and biogeography. Extremophiles 13: 541–555.
Waldrop MP, Wickland KP, White Iii R, Berhe AA, Harden JW, Romanovsky VE . (2010). Molecular investigations into a globally important carbon pool: permafrost-protected carbon in Alaskan soils. Global Change Biol 16: 2543–2554.
Acknowledgements
This study was funded by NSF International Polar Year grant no. 0732956 to NP, TZ, LJK and MMH. We thank Dwayne Elias for helpful discussions. The contributions for this publication include: collected permafrost cores (TZ); designed research (SJT, LJK, MMH and NP); performed research/data collection (SJT, PD and LRM); analyzed data (SJT, LRM, LJK and MMH); and wrote/edited manuscript (SJT, LJK, MMH, LRM, NP, TZ and PD).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Tuorto, S., Darias, P., McGuinness, L. et al. Bacterial genome replication at subzero temperatures in permafrost. ISME J 8, 139–149 (2014). https://doi.org/10.1038/ismej.2013.140
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2013.140
Keywords
This article is cited by
-
Microbial life in 25-m-deep boreholes in ancient permafrost illuminated by metagenomics
Environmental Microbiome (2023)
-
Genomic reconstruction of fossil and living microorganisms in ancient Siberian permafrost
Microbiome (2021)
-
Future threat from the past
Environmental Science and Pollution Research (2021)
-
Cultivation and characterization of snowbound microorganisms from the South Pole
Extremophiles (2021)
-
Bacterial and protozoan dynamics upon thawing and freezing of an active layer permafrost soil
The ISME Journal (2019)