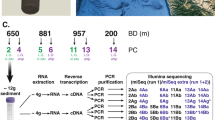
Abstract
Members of the highly diverse Miscellaneous Crenarchaeotal Group (MCG) are globally distributed in various marine and continental habitats. In this study, we applied a polyphasic approach (rRNA slot blot hybridization, quantitative PCR (qPCR) and catalyzed reporter deposition FISH) using newly developed probes and primers for the in situ detection and quantification of MCG crenarchaeota in diverse types of marine sediments and microbial mats. In general, abundance of MCG (cocci, 0.4 μm) relative to other archaea was highest (12–100%) in anoxic, low-energy environments characterized by deeper sulfate depletion and lower microbial respiration rates (P=0.06 for slot blot and P=0.05 for qPCR). When studied in high depth resolution in the White Oak River estuary and Hydrate Ridge methane seeps, changes in MCG abundance relative to total archaea and MCG phylogenetic composition did not correlate with changes in sulfate reduction or methane oxidation with depth. In addition, MCG abundance did not vary significantly (P>0.1) between seep sites (with high rates of methanotrophy) and non-seep sites (with low rates of methanotrophy). This suggests that MCG are likely not methanotrophs. MCG crenarchaeota are highly diverse and contain 17 subgroups, with a range of intragroup similarity of 82 to 94%. This high diversity and widespread distribution in subsurface sediments indicates that this group is globally important in sedimentary processes.
Similar content being viewed by others
Introduction
The archaeal phylum Crenarchaeota was originally considered to comprise extremophiles. Cultivated strains were thermophilic or hyperthermophilic organisms utilizing sulfur for energy metabolism (Burggraf et al., 1997). Then high numbers of pelagic crenarchaeota were discovered in the marine water column, indicating the presence of mesophilic or psychrophilic species (DeLong, 1992; Fuhrman et al., 1992). Current estimates of pelagic crenarchaeota suggest they are abundant with up to 1.3 × 1028 cells in the ocean (Karner et al., 2001). Mesophilic pelagic crenarchaeota have been assigned to Marine Group I, a sister group of thermophilic crenarchaeota. Cultivated strains are autotrophic ammonium-oxidizing archaea (Könneke et al., 2005; Tourna et al., 2011). A third archaeal phylum, the Thaumarchaeota, was proposed for these organisms because of their distinct phylogeny and physiology (Brochier-Armanet et al., 2008).
The Miscellaneous Crenarchaeotal Group (MCG), has only been defined by 16S rRNA sequences and is distinct from the cultured crenarchaeota of the Thermoprotei and Thaumarchaeota (Inagaki et al., 2003). The sequences delineating the MCG were described by Inagaki et al. (2003), but the same clade had been observed before and either called the Terrestrial Miscellaneous Crenarchaeotic Group (Takai et al., 2001) or Group 1.3 (Jurgens et al., 2000). Members of the MCG have been hypothesized to be numerically and ecologically important in oceanic deep subsurface sediments because they often dominate archaeal clone libraries using 16S rRNA genes (Parkes et al., 2005) as well as 16S rRNA (Biddle et al., 2006), which is often rapidly degraded in inactive cells and may reflect the active population (Felske et al., 1996).
Currently, the carbon and energy sources for the MCG are unknown. Biddle et al. (2006) showed that total archaea in MCG-rich sediments are heterotrophic, using organic carbon derived from degradation of fossil organic matter. To explain energetic and isotopic discrepancies, they hypothesized that MCG might perform ‘dissimilatory’ methane oxidation. However, so far the distribution of MCG crenarchaeota relative to methane and sulfate availability in the environment remains unknown. DNA fosmids containing 16S rRNA gene sequences for MCG and functional genes have been identified, but lack information about the carbon and energy sources for MCG (Meng et al., 2009; Li et al., 2012).
In this study, we examined the total diversity of published MCG sequences to identify phylogenetic subgroups of this highly diverse group and identify potential biogeographical trends. We used this comprehensive MCG database to design primers and probes to be used in a polyphasic approach, including rRNA slot blot hybridization, DNA-based quantitative PCR, and CARD-FISH, to investigate the distribution of MCG in eleven different habitats. We provide a protocol optimized for in situ visualization and enumeration of MCG cells. As a specific case study, we compared MCG abundance and taxonomic composition with depth in the White Oak River estuary, North Carolina and Hydrate Ridge methane seeps.
Materials and methods
Sample descriptions
Eleven sites were chosen for quantification of MCG on the basis of available biogeochemical and microbial diversity data (Table 1).
MCG phylogenetic analysis
The ARB/SILVA 106 non-redundant 16S rRNA database (Pruesse et al., 2007) was supplemented with sequences from the Parc database for a total of 4720 MCG sequences (for details see Supplementary Information). A 50% conserved filter considering 804 well-aligned columns between E. coli positions 86 and 935 was applied in a neighbor-joining, RAxML, and PHYML analyses for tree reconstruction using only sequences longer than 940 bp (1276 sequences). Partial sequences were added to the tree by parsimony criteria without allowing changes in the general tree topology. MCG subgroup designations were made when either previous publications had defined a subgroup, or subgroups with >50 sequences were monophyletic with both distance and maximum likelihood methods.
Probe/primer development
An alignment of 615 MCG sequences from ARB/SILVA release 92 (September 2007) was used to design primers specific for the marine-derived MCGs (235 sequences total) using ARB. Sequences, coverage and specificity of probes and primers are listed in Table 2 and Supplementary Table S1.
Total nucleic acids
DNA and RNA were extracted from 1–6 g of frozen sediments using pH8-buffered phenol and bead beating based on protocols described previously (Stahl et al., 1988; MacGregor et al., 1997) with slight modifications. Detailed protocols are provided in the Supplementary Information.
Quantitative PCR
qPCR for 16S rRNA genes was performed as described previously (Lloyd et al., 2011). Total archaeal 16S rRNA genes were quantified using primers ARCH915F and ARC1059R, or ARCH806F and ARCH915R; total bacterial using BAC340F and BAC515R; and total MCG using MCG528F and MCG732R or primers MCG410F and MCG528R (Supplementary Table S1). Standards were made from plasmids containing inserts of environmental 16S cDNA from White Oak River estuary sediments. 69Earc46 (accession no. JN605175) was used for archaea and MCG, and 70c2Aarc103 (accession no. JN616271) was used for bacteria. Plasmids containing non-target 16S rRNA sequences were used as negative controls. R-squared values for qPCR standard curves were between 0.993 and 0.999. No low melting point primer dimers were detected. For each core and extraction method, samples were diluted until the Cp decreased log-linearly with sample dilution, indicating the absence of inhibition effects at 10-fold dilution. Amplification efficiencies (listed in Supplementary Table S2) were determined for each individual environmental DNA sample. In addition, the qPCR products of three samples were cloned and sequenced to check for amplification of non-target DNAs (Supplementary Table S3).
Slot blot hybridization
Approximately 10–100 ng of RNA was blotted onto nylon membranes (MagnaCharge Membrane; GE Water & Process Tech., Trevose, PA, USA) in triplicate and hybridized with 33P-labeled oligonucleotides as described previously (Stahl et al., 1988). The probes and dissociation temperatures used in this study are given in Supplementary Table S1; reference rRNA are given in the Supplementary Methods. Hybridization intensity was measured with a blot imager Typhoon 9400 (GE Healthcare, München, Germany) and analyzed with ImageQuant software (GE Healthcare).
Catalyzed reporter deposition fluorescence in situ hybridization (CARD-FISH)
Sediment samples were fixed in paraformaldehyde, sonicated and filtered onto a polycarbonate filter as previously described (Knittel et al., 2003). Cells were embedded in 0.1% low melting point agarose (NuSieve GTG Agarose, Cambrex Bio Science Rockland Inc., Rockland, ME, USA) and air-dried. Inactivation of endogenous peroxidases and permeabilization of cell walls were done by incubating the filters in 0.01 M HCl with 0.15% H2O2 for 10 min at room temperature. CARD-FISH and subsequent staining with 4',6-diamidino-2-phenylindole (DAPI) followed a previous published protocol (Pernthaler et al., 2002). For dual CARD-FISH, the protocol was repeated on the same filters using a second probe and other fluorescently labeled tyramides after inactivation of peroxidases of initial hybridization as described above. The given CARD-FISH counts are means calculated from 10 to 150 randomly chosen microscopic fields corresponding to 100–800 total DAPI-stained cells. Probe sequences and formamide concentrations required for specific hybridization are given in Supplementary Table S1. The specificity of new MCG probes was evaluated by Clone-FISH (Schramm et al., 2002).
cDNA clone libraries
Total RNA was reverse-transcribed and amplified with general 16S rRNA-targeted archaeal primers, checked for DNA co-extraction, and sequenced as described previously (Lloyd et al., 2011). Pintail (Ashelford et al., 2006) and Chimera Slayer (Haas et al., 2011) were implemented in Mothur v.1.19.4 (Schloss et al., 2009) using the ARB/SILVA Ref106 database, to remove 16S rRNA sequences that were likely chimeric. Further chimeras were identified manually in Pintail. Chimeric sequences were 10% overall. Sequences were clustered into operational taxonomic units (OTUs) based on 97% similarity.
Amplicon sequencing
DNA from 0.24–0.27 mbsf extracted from Core December 06, White Oak River estuary sediments, was amplified using tagged primers for pyrosequencing of the V6 region, ARCH958F and ARCH1048R. Primer details and amplification conditions (http://vamps.mbl.edu/resources/faq.php#tags) were according to those of the Josephine Bay Paul Center (Woods Hole, MA, USA) by the International Census of Marine Microbes project. The protocol to prepare PCR amplicons from archaea was followed as described in Huber et al. (2007). Data processing was followed as described in Huse et al. (2007). The VAMPS pipeline was used to designate MCG amplicons after manual curation of the archaeal reference database. Further analysis of MCG tags was performed using CLUSTAL alignments and OTU selection in Mothur.
Nucleotide sequence accession numbers
Representative MCG sequences obtained from qPCR products and sequences from cDNA libraries have been deposited in GenBank under accession numbers HE584637 to HE584657 and JN605005 to JN605175. Amplicon sequencing data were deposited in GenBank SRA archive under accession number SRX011653.
Results
MCG phylogeny
MCG diversity in our 4720 sequence database was great, with some 16S rRNA similarity values as low as 76% between the most distant sequences. A total of 17 subgroups of MCGs (MCG-1–MCG-17) were supported in both distance and maximum likelihood inferences. These included the previously named MCG-1 through MCG-4 (Sørensen and Teske, 2006), C3 (Inagaki et al., 2006) that we renamed MCG-15, and pMARA-4 (Nercessian et al., 2005) that we renamed MCG-16 for consistency (Figure 1). However, these previously named subgroups covered only 21% of the MCG database. Our new subgroups increase the coverage to 91%. A few subgroups from the literature were not explicitly included in the 17 subgroup designations because they are subsets of some of the other MCG subgroups. Subgroup PM-1 fell within the MCG-2, PM-2 fell within the MCG-4, PM-3 through PM-5 fell within the MCG-1, PM-6 fell within the MCG-3, PM-7 fell within the MCG-8 in some treeing methods and PM-8 fell within the MCG-17 (Webster et al., 2006). pMARA-5 (Nercessian et al., 2005) fell within MCG-17. MBGC (Vetriani et al., 1999) fell within the MCG-8. Group 1.3b (Ochsenreiter et al., 2003) fell within the MCG-6. Groups MCG-13 through MCG-17 were stable, but their branching order varied between treeing methods, so we presented them as a multifurcation. In general, there is high intra-subgroup diversity (Table 2), with most subgroups containing ⩽92% intragroup similarity. Similarities ranged from 82% for MCG-15 to 94% for MCG-5a and MCG-2. MCG-5 split into two adjacent groups each with >90% intragroup similarity in the maximum likelihood trees, so these were designated groups MCG-5a and MCG-5b. The smallest subcluster MCG-2 is restricted to sequences retrieved from marine habitats, whereas all others are mixed and contain sequences from marine, terrestrial and limnic habitats. For an overview of MCG habitats see Supplementary Table S4.
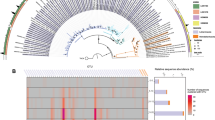
RaxML phylogenetic tree based on 16S rRNA genes showing MCG crenarchaeotal subgroups MCG-1–MCG-17. Subgroup designations follow those of Sørensen and Teske (2006) for MCG-1 through MCG-4, Nercessian et al. (2005) for MCG-16 (formerly pMARA-4), and Inagaki et al. (2006) for MCG-15 (formerly C3). MCG-13 through MCG-17 multifurcate because branching order was inconsistent between tree models. Sequences with no color were not assigned to a subgroup. Black bars indicate the total number of sequences from our MCG database within that subgroup, with the highest value of 1160 for MCG-6 and lowest value of 51 for MCG-5a. White Oak River estuary cDNA sequences (97% OTUs) are labeled with red or blue bars for methane oxidation or methane production zones, respectively. The first two numbers in the clone names are the depth of that sequence, followed by A for core July 05-1, B for July 05-2, E for December 06 and G for July 08-2. Additional membership in each OTU is listed after the initial clone name. Terrestrial Hot Springs Crenarchaeotal Group was used as outgroup and individual clone names are listed in Supplementary Tables S4 and S5. The web-based Interactive Tree of Life was used for tree and data set visualization (Letunic and Bork, 2011).
Design and evaluation of MCG-specific oligonucleotide probes
The design of new probes always aims at a compromise of good target group coverage and high specificity. Owing to the great MCG intragroup diversity it was impossible to target all members of MCG perfectly. The deeply branching subclusters MCG-13–MCG-17, in particular, are not well covered by the four newly developed probes and primers MCG410, MCG493, MCG528 and MCG732. In silico tests revealed a MCG-1–MCG-12 group coverage of 87% for MCG528, 79% for MCG493, 44% for MCG410 and 27% for MCG732 (Table 2). The two probes/primers with the highest group coverage, MCG493 and MCG528, are highly specific having only three and four outgroup hits within the Archaea and none within the Bacteria. Although both probes showed bright signals in Clone-FISH, we recommend the use of probe MCG493 for in situ detection because of a much brighter signal in environmental samples. In slot blots, use of probe MCG493 resulted in easily visualized slot blot signals (Supplementary Figure S1).
In qPCR, all newly designed MCG-targeted primers amplified the 16S rRNA fragment from MCG, and showed no amplification after 40 cycles for another deeply branching crenarchaeotal group (MBGB), ANME-2a, ANME-2c, ANME-1b, Thermoplasmatales, Desulfobacteraceae, Gammaproteobacteria, Cytophagales or Alphaprotobacteria. Amplification resulted in a single, sharp peak, although a small shoulder indicates possible small contribution from non-target sequences. However, sequencing of qPCR products resulted only in MCG sequences, which were also quite diverse indicating that multiple MCG subgroups are indeed amplified by the primers (Supplementary Table S3).
Quantification of MCG in several types of sediments
Members of the MCG were quantified by rRNA slot blot hybridization, qPCR and CARD-FISH in 11 different marine seep and non-seep sediments and microbial mats. Except for oxic surface sediments from Svalbard and most parts of anoxic methanotrophic microbial mats from the Black Sea, MCG rRNA could be detected in all types of habitats (Table 3). Highest MCG rRNA amounts were found in seep sediments from station 19-2 at Hydrate Ridge with 178 ng g−1 at a depth of 4–5 cm. This was accompanied by the highest total archaeal rRNA amount detected in the sediments investigated. Relative values of MCG calculated as a percentage of total archaeal rRNA varied greatly between sites (Figure 2), but was significantly higher in seep and non-seep sediments with sulfate penetration >0.15 mbsf (1.6%±1.2 vs 28.0%±27.1; P=0.06, 90% CI, based on a two-tailed heteroscedastic t-test, where values for all depths in a core were averaged so that each core contributed one value, reducing skewed weighting towards cores with many measurements). Samples with higher potential for exposure to oxygen such as those from hypersaline microbial mats in the Arabian Gulf, and the surface of non-advective sediments from Svalbard (<0.01 mbsf) had lower MCG/archaea ratios (3.9%±9.4) than the deep sulfate penetration sediments (P=0.08, 90% CI).
Relative abundance of MCG crenarchaeota compared with total archaea as determined for different marine habitats by rRNA slot blot hybridization (black bars), qPCR (white bars), and CARD-FISH (gray bars). Colors indicate samples from anoxic methanotrophic microbial mats (peach) or oxic surface sediments (pink) or sediments with sulfate depletion depths of <0.15 mbsf (yellow) or >0.15 mbsf (blue). Dots indicate samples with no MCG detected with slot blot (black) or qPCR (white); other samples with no visible data were not analyzed with that method. When a depth range is listed, values were averaged over adjacent depths. All individual measurements listed in Table 3.
qPCR data conformed with slot blot hybridization (Table 3, Figure 2). As with total rRNA, the percentage of 16S rRNA gene copy numbers relative to that of all archaea was significantly higher in samples with sulfate penetration >0.15 mbsf (1.0%±0.8 vs 67.9%±54.7; P=0.05, 95% CI). Likewise, samples with likely oxygen exposure were also low (0.4%±0.3; P=0.05, 95% CI). In two non-seep habitats MCG were exceptionally abundant: at ODP site 1229 (99%, Peru Margin) and in the White Oak River estuary (98–121%). Values above 100% can be explained best by a reduced binding efficiency or insufficient coverage of used general archaeal primers. Absolute values of MCG abundance were not significantly different between seep vs non-seep sites (slot blot P=0.47, qPCR P=0.15, and CARDFISH P=0.16).
As a third method for MCG quantification we adapted the standard CARD-FISH protocol (Pernthaler et al., 2002) for in situ detection of MCG cells. The permeabilization of the cell walls turned out to be the most crucial for visualization of MCG. Archaeal cell walls are usually permeabilized either by a treatment with 10–15 μg ml−1 proteinase K for 1–5 min, 0.5% SDS for 10 min at room temperature or with 60 U ml−1achromopeptidase at 37 °C (Teira et al., 2004; Knittel and Boetius, 2009; Labrenz et al., 2010). However, none of these methods was successful for the visualization of MCG. Instead, we used 0.01 M HCl for 10 min that resulted in bright signals. Higher HCl concentrations (> 0.1 M) did not increase the fraction of hybridized cells, but rather caused a visible disintegration of cells.
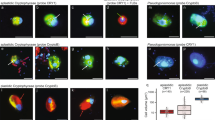
Hot spots of MCG as identified by slot blots, qPCR or 16S rRNA gene libraries were selected for CARD-FISH. About 90% of the MCG cells were small and coccoid (Figure 3) with a cell size of 0.4–0.5 μm. This is a comparable size to that reported for their sister group Marine Benthic Group B (MBGB) crenarchaeota (0.2–0.4 μm) (Knittel et al., 2005). A minor part of the MCG community had a much larger cell size with up to 1 μm in diameter and might reflect the observed large MCG intragroup 16S rRNA sequence diversity. In most cases MCG were detected as single cells but they were also found in aggregates of 2–5 cells. DAPI signal of MCG was clearly visible in most habitats except for all ODP sites investigated where DAPI staining did not work at all. To corroborate the identification of detected MCG cells, dual hybridizations were performed with probe MCG493 and the general archaeal probe ARCH915. MCG493 signals were always co-localized with ARCH915 signals (Figure 3I).
Single cells of MCG crenarchaeota in subsurface sediments from the sulfate–methane transition zone of the White Oak River estuary (0.4 mbsf; a–f) and ODP site 1227 at Peru Margin (5H3, 37.38 mbsf; g–i), visualized by CARD-FISH. a and d show DAPI staining (blue); other panels show the corresponding FISH signals obtained by dual hybridization with the general archaeal probe ARCH915 (b, e, g; red) and MCG-specific probe MCG493 (c, f, h; green). i shows an overly of g and h. Arrows point to MCG cell signals. Scale bars, 5 μm.
In White Oak River sediments 1.2 × 108 cells cm−3 were detected at shallow depth of 0.075 mbsf (12% of total cell counts, 22% of total Archaea). In the subsurface at 0.405 mbsf, the absolute MCG abundance was comparably high but the relative fraction of MCG increased to 30% and 60% of total archaea. In other deep sulfate penetration sediments, MCG were as dominant as in White Oak River sediments; they contributed 18–20% of total Archaea in Janssand intertidal sand flats, 15–74% at ODP sites 1245 and 1250 at Hydrate Ridge, 22% at Peru Margin ODP site 1229, 100% at Peru Basin ODP site 1231 and 48% at eastern equatorial Pacific ODP site 1225. The two latter sites showed very low microbial activities and low organic matter content. In deep sediments (4.65 mbsf) from the crater center of Haakon Mosby mud volcano, a site which is strongly dominated by MCG 16S rRNA sequences (Lösekann-Behrens, Boetius & Knittel, unpublished), MCG cells accounted for 16% of total Archaea.
MCG depth distribution
In order to explore changes in MCG abundance over depth, we examined the White Oak River estuary cores (sulfate depleted at 0.40 mbsf) and site 19-2 from Hydrate Ridge (sulfate depleted at 0.07 mbsf, Figure 4). In White Oak River core Jul 08-1 qPCRs using duplicate primer sets gave roughly equal 16S rRNA gene copies (Figures 4a and b), suggesting low or equal primer bias. At the division between the horizon dominated by sulfate reduction and methane oxidation and that dominated by methanogenesis DNA concentrations for archaea, bacteria or MCG did not change perceptibly. At and below 0.045 mbsf, total archaeal DNA copies matched MCG DNA copies for all four primer sets. Above, at 0.015 mbsf, however, MCG 16S rRNA gene copy numbers were much lower than those of archaea. Bacterial DNA copies were consistently higher than archaeal and MCG DNA copies with the gap narrowing gradually with depth. Large differences in absolute values between July 08-1 and July 08-2 correspond to differences in Cp values, whereas Cp values for standards were nearly identical, suggesting much lower DNA yields during extraction of July 08-2. An interesting contrast to the lack of vertical zonation of MCG in White Oak River estuarine sediments was Hydrate Ridge, where quantities and activities of MCG and total archaea peaked around 0.05 mbsf near the base of the sulfate reduction zone, as shown in both qPCR and RNA slot blot data (Figure 4c). As MCG do not increase relative to archaea, and the archaeal peak is far too large to be caused by the MCG increase alone, this peak is most likely due to an increase in total ANME biomass supported either directly or indirectly by very high rates of anaerobic methane oxidation at this site (Treude et al., 2003).
qPCR of 16S rDNA for White Oak River estuary cores July 08-1 (a) and Jul 08-2 (b) and Hydrate Ridge core 19-2 (c). Primer sets used were for total archaea (ARCH915F-ARCH1059R, filled red squares, and ARCH806F-ARCH915R, open red squares), MCG (MCG528F-MCG732R, filled blue circles and MCG410F-MCG528R, open blue circles) or total bacteria (BAC340R-BAC515R, filled black diamonds). All replicate measurements at each depth are shown. In addition, rRNA concentrations of MCG (blue x’s) and total archaea (red x’s) measured with slot blot hybridization for Hydrate Ridge core 19-2 are shown in c. Sulfate (filled triangles) and methane (open squares or modeled line) concentrations for White Oak River estuary cores July 08-1 (d) and July 08-2 (e) and for Hydrate Ridge (f); (originally published in Lloyd et al. (2011), and in Treude et al., 2003). Dashed horizontal lines are the depth above which there is net anaerobic oxidation of methane (AOM) and below which there is net methane production (MP; Lloyd et al., 2011).
Given a lack of vertical zonation in total MCG abundance in the White Oak River estuary, we investigated possible changes in MCG phylogenetic composition with depth. MCG comprised the majority (44–86%) of 16S rRNA cDNA sequences from White Oak River estuary sediments (Figure 5). Other archaeal sequences included the uncultured groups ANME-1 (Hinrichs et al., 1999), Marine Benthic Groups (MBGA, B and D Vetriani et al., 1999), Deep Sea Euryarchaeotal Group (DSEG; Takai et al., 2001), or Terrestrial Hot Springs Crenarchaeotic Group (THSCG Takai and Horikoshi, 1999) (Supplementary Figure S2). MCG had more OTUs than all other archaea in the sediments combined (92 OTUs vs 79 OTUs, respectively, at 97% similarity) and covered 13 out of the 17 subgroups (Figure 1, Supplementary Figure S1). The general composition and sequence abundance did not change much with depth or season (Supplementary Table S5). No clear phylogenetic separation was observed for sequences from the methane oxidation and production zones (Figure 1, red and blue dots, respectively). No seasonality was observed with MCG types, as sequences from the winter core (December 06) were distributed throughout the MCG tree along with those from summer cores (July 05-2, July 08-1 and July 08-2).
Phylogenetic groupings of all 16S rRNA cDNA clones and 16S rRNA gene V6 amplicon tag sequences for White Oak River estuary sediments that derived from (a) methane oxidation or (b) methane production zone. Numbers at the right side of bars are the total number of sequences in the library. Abbreviations and references for phylogenetic groups are listed in results section.
Amplicon pyrosequencing supported the dominance of MCG within the archaea with 86% of 13 250 sequences (Figure 5). It is surprising that the cDNA clone libraries agreed so well with the amplicon pyrosequencing, as one targets RNA and the other targets DNA, different primers were used, and the V6 amplicon library was 150 times greater than the biggest cDNA clone library. The only difference was that the amplicon data contained sequences for MBGA and the Methanobacteriales, which were presumably in lower numbers in the sediments and therefore missed in the clone libraries. After error processing, which includes clustering that may hide some alpha diversity, 1192 unique MCG sequences were detected. Of these, 790 were singletons and only 19 were seen 100 or more times. These 19 sequences made up 57% of all tags seen within the archaeal White Oak River data set, truly a significant portion of the archaea in this sediment. The diversity seen within these tags was also high. Tags represented over 10 times were aligned and OTUs were calculated. Within these 99 tags, there were 99 OTUs calculated at 99% similarity, 73 OTUs at 98% similarity and 51 OTUs still remained at 97% similarity.
Quantification of total bacterial rRNA
On the basis of the size of MCG rRNA fraction, two seep habitats (Gulf of Mexico site 87, Hydrate Ridge ODP site 1245D) and three non-seep habitats (Janssand, White Oak River, Peru ODP sites 1227A and 1229D) were selected for quantification of total bacterial 16S rRNA. All habitats were characterized by deep sulfate penetration. Surface sediment from Janssand sand flat was strongly dominated by bacterial rRNA (97–98% bacterial rRNA: 2–3% archaeal rRNA) at a depth of 0–0.4 mbsf, but by archaeal rRNA in a deeper sediment horizon at 4.9 mbsf (19% bacterial rRNA: 81% archaeal rRNA). The ratio of bacterial to archaeal rRNA percentages was comparably low for the other subsurface sediments investigated, that is, 14%:86% at ODP site 1245 at Hydrate Ridge, 24%:76% at ODP site 1227, 45%:55% at ODP site 1229 and 34%:66% at White Oak River (Table 3, Figure 6).
Slot blot-based determination of the ratio archaea:bacteria 16S rRNA in selected surface and subsurface sediments. The sum of detected archaeal and bacterial 16S rRNA was set as 100% of prokaryotic rRNA. The archaeal fraction is further resolved in the column that shows the proportion of MCG and other, yet unidentified archaea.
Discussion
When looking at all MCG sequences available in public databases, MCG crenarchaeota appear very diverse and widespread. MCG are present in terrestrial hot springs (Barns et al., 1996), deep oceanic subsurface sediments (Parkes et al., 2005), deep terrestrial subsurface (Inagaki et al., 2003), continental shelf sediments (Vetriani et al., 1998), ancient marine sapropels (Coolen et al., 2002), petroleum-contaminated soil (Kasai et al., 2005), termite guts (Friedrich et al., 2001), mud volcanoes (Heijs et al., 2007), methane hydrate-containing marine sediments (Inagaki et al., 2006), landfill leachate (Huang et al., 2003), anaerobic wastewater reactors (Collins et al., 2005), sulfidic springs (Elshahed et al., 2003), brackish lakes (Hershberger et al., 1996) and coastal salt marshes (Castro et al., 2004). Only 28 out of 4720 MCG sequences were retrieved from potentially oxic habitats (for example, database releases EU370096 and FJ560325). This broad distribution is reflected in a large MCG intragroup diversity with 16S rRNA gene similarity values as low as 76%, which is within the range proposed to define prokaryotic phyla (Yarza et al., 2010). The diversity within most of our subgroups is in the proposed cutoff of 87.7% minimum level for family boundaries (Yarza et al., 2010). It is possible that this evolutionary diversity is also reflected in a high degree of metabolic diversity.
MCG crenarchaeota constituted a major part of archaea in sediments with low respiration rates and deep sulfate penetration
Thaumarchaeota account for up to 40% of total microbes in meso- and bathypelagic deep ocean waters (Karner et al., 2001). In analogy, we present the first quantitative data for the dominance of MCG crenarchaeota in anoxic marine sediments. We used a combination of three methods that are based on different target molecules, that is, DNA and RNA, to provide robust environmental data for MCG independent of their individual methodological constrains. All methods pointed to a higher relative abundance of MCG to total microbes in sediments at sites where sulfate penetrates at least 10 cm.
Cold seeps with high methane fluxes and high microbial respiration rates are characterized by low sulfate penetration depths. Among the cold seep sites with shallow sulfate depletion investigated here, the four sites from Hydrate Ridge and site 156 from the Gulf of Mexico showed the lowest MCG abundance concurrent with extremely high rates of AOM and sulfate reduction (Boetius et al., 2000; Treude et al., 2003; Orcutt et al., 2010). An exception were 4.6 m-deep sediments from sites 371 and 372 at HMMV which showed a high MCG abundance together with the lowest microbial activity due to high fluid flow rates repressing sulfate penetration (DeBeer et al., 2006; Niemann et al., 2006).
Sediments from ‘low-energy’ environments or from cold seeps with low methane fluxes and low microbial respiration rates are characterized by deep sulfate penetration. In all non-seep, low-energy habitats, MCG made up a high proportion of MCG relative to total prokaryotes, ranging from 12% at Janssand to 45% at Peru Margin ODP 1229D or even up to 100% at White Oak River. This finding was independent of water depths, temperature or total carbon contents (organic-poor ODP sites 1225 and 1231 vs organic-rich ODP sites 1227 and 1229). Therefore, it appears that MCG dominate the archaeal population in low activity subsurface sediments. At cold seeps with deep sulfate penetration such as Hydrate Ridge site ODP1245 and ODP1250 as well as Gulf of Mexico site 87, relative abundance of MCG was comparably high (15–74% of archaea) as for non-seep sediments with deep sulfate penetration. An exception was HMMV seep site 336 that showed the lowest relative percentages of MCG of all CARD-FISH counts (3% of archaea). However, the deep sulfate penetration depth at this special site is not the result of low microbial respiration rates, but instead is driven by active pumping of sulfate into the core by Siboglinidae tubeworms (DeBeer et al., 2006).
No support for an association of MCG with the methane cycle
Data obtained in this study do not support the hypothesis that MCG could be anaerobic methanotrophs (Biddle et al., 2006). MCG distribution is not proportional to any geochemical gradients, and specifically, does not co-vary with energy availability from methane and sulfate. We did not find evidence for changes of MCG activity with depth based on rRNA content per MCG cell (range 1–117 fg per cell, calculated based on rRNA slot blot and CARD-FISH). When the White Oak River was studied with a finer depth resolution, this trend was upheld. Similar to many diffusion-driven marine sediments, those of the White Oak River estuary are stratified with respect to dominant microbial electron acceptor. Thus, if MCG crenarchaeota depend on a known terminal electron acceptor either directly or indirectly through microbial syntrophy, MCG presence, abundance or activity would be expected to change downcore. However, we found no evidence for such a coupling with depth. In contrast, sediments from Hydrate Ridge site 19-2 had a peak in MCG abundance (qPCR) and activity (slot blot) coinciding with the bottom of the sulfate reduction zone where highest methane oxidizing activity is presumed to occur (Knittel et al., 2003). Nevertheless, in agreement with White Oak River sediments, the percent MCG of total archaea had no depth trend. Further support for a lack of vertical zonation of MCG came from the lack of MCG phylogenetic composition changes with depth in White Oak River sediments. Secondly, a comparison of MCG cell numbers in seep (for example, Hydrate Ridge, Gulf of Mexico) vs non-seep (for example, North Sea, Peru, White Oak River estuary) sediments did not show remarkable differences, whereas numbers of methanotrophs change dramatically across all the sites investigated (see Knittel and Boetius, 2009). The lack of vertical zonation together with similar cell numbers of MCG at seeps and non-seep sediments suggest that MCG are anaerobes, as they maintain a high abundance well-below oxygen penetration, but do not gain energy from AOM and sulfate is likely not an electron acceptor. This, together with a widespread distribution of MCG, could indicate that they obtain carbon and energy from substrate pools and energy sources that vary little with sediment depth and between habitats. Options for such metabolisms include the slow fermentative degradation of different types of organic matter. MCG are likely to access substrates that are physically or chemically recalcitrant. This conclusion is in agreement with findings from previous literature (Biddle et al., 2006; Lipp et al., 2008; Takano et al., 2010; Webster et al., 2010; Llirós et al., 2011).
Conclusions
The high diversity and high abundance of MCG crenarchaeota in seep and non-seep sediments suggest that they are globally important components of the anaerobic sedimentary microbial community. However, the question of what carbon and energy sources sustain MCG is still open. The lack of abrupt changes in MCG abundance or phylogenetic composition at the downcore transition from zones with sulfate reduction/methane oxidation to those with methane production as well as the lack of difference in MCG cell numbers at seeps vs non-seeps suggests that the MCG community is not active in methane or sulfur cycling. This is supported by a much higher relative abundance of MCG compared with other prokaryotes in low-energy environments. It is, therefore, likely that MCG are anaerobic heterotrophs and either access a wide variety of fermentative substrates or are linked to degradation of refractory organic matter, potentially using other electron acceptors than sulfate. This possible contribution to post-depositional carbon remineralization makes MCG a promising target for future culturing and ecophysiological research.
References
Ashelford K, Chuzhanova NA, Fry JC, Jones AJ, Weightman AJ . (2006). New screening software shows that most recent large 16S rRNA gene clone libraries contain chimeras. App Environ Microbiol 72: 5734–5741.
Barns SM, Delwiche CF, Palmer JD, Pace NR . (1996). Perspectives on archaeal diversity, thermophily and monophyly from environmental rRNA sequences. Proc Natl Acad Sci USA 93: 9188–9193.
Biddle JF, Lipp JS, Lever MA, Lloyd KG, Sørensen KB, Anderson R et al. (2006). Heterotrophic Archaea dominate sedimentary subsurface ecosystems off Peru. Proc Natl Acad Sci USA 103: 3846–3851.
Boetius A, Ravenschlag K, Schubert C, Rickert D, Widdel F, Gieseke A et al. (2000). A marine microbial consortium apparently mediating anaerobic oxidation of methane. Nature 407: 623–626.
Brochier-Armanet C, Boussau B, Gribaldo S, Forterre P . (2008). Mesophilic crenarchaeota: proposal for a third archaeal phylum, the Thaumarchaeota. Nat Rev Microbiol 6: 245–252.
Burggraf S, Huber H, Stetter KO . (1997). Reclassification of the crenarchaeal orders and families in accordance with 16S rRNA sequence data. Int J Syst Bacteriol 47: 657–660.
Castro H, Ogram A, Reddy KR . (2004). Phylogenetic characterization of methanogenic assemblages in eutrophic and oligotrophic areas of the Florida Everglades. Appl Environ Microbiol 70: 6559–6568.
Collins G, O’Connor L, Mahony T, Gieseke A, de Beer D, O’Flaherty V . (2005). Distribution, localization, and phylogeny of abundant populations of Crenarchaeota in anaerobic granular sludge. Appl Environ Microbiol 71: 7523–7527.
Coolen MJL, Cypionka H, Sass AM, Sass H, Overmann J . (2002). Ongoing modification of Mediterranean Pleistocene sapropels mediated by prokaryotes. Science 296: 2407–2410.
DeBeer D, Sauter E, Niemann H, Kaul N, Foucher J-P, Witte U et al. (2006). In situ fluxes and zonation of microbial activity in surface sediments of the Haakon Mosby mud volcano. Limnol Oceanogr 51: 1315–1331.
DeLong EF . (1992). Archaea in coastal marine environments. Proc Natl Acad Sci USA 89: 5685–5689.
D’Hondt S, Jorgensen BB, Miller DJ, Batzke A, Blake R, Cragg BA et al. (2004). Distributions of microbial activities in deep subseafloor sediments. Science 306: 2216–2221.
Elshahed MS, Senko JM, Najar FZ, Kenton SM, Roe BA, Dewers TA et al. (2003). Bacterial diversity and sulfur cycling in a mesophilic sulfide-rich spring. Appl Environ Microbiol 69: 5609–5621.
Felske A, Engelen B, Nübel U, Backhaus H . (1996). Direct ribosome isolation from soil to extract bacterial rRNA for community analysis. Appl Environ Microbiol 62: 4162–4167.
Friedrich MW, Schmitt-Wagner D, Lueders T, Brune A . (2001). Axial differences in community structure of Crenarchaeota and Euryarchaeota in the highly compartmentalized gut of hte soil-feeding termite Cubitermes orthognathus. Appl Environ Microbiol 67: 4880–4890.
Fuhrman JA, McCallum K, Davis AA . (1992). Novel major archaebacterial group from marine plankton. Nature 356: 148–149.
Gittel A, Mußmann M, Sass H, Cypionka H, Könneke M . (2008). Identity and abundance of active sulfate-reducing bacteria in deep tidal flat sediments determined by directed cultivation and CARD-FISH analysis. Environ Microbiol 10: 2645–2658.
Haas BJ, Gevers D, Earl AM, Feldgarden M, Ward DV, Giannoukos G . (2011). Chimeric 16S rRNA sequence formation and detection in Sanger and 454-pyrosequenced PCR amplicons. Genome Res 21: 494–504.
Heijs SK, Haese RR, van der Wielen P, Forney LJ, van Elsas JD . (2007). Use of 16S rRNA gene based clone libraries to assess microbial communities potentially involved in anaerobic methane oxidation in a Mediterranean cold seep. Microbial Ecol 53: 384–398.
Hershberger KL, Barns SM, Reysenbach AL, Dawson SC, Pace NR . (1996). Wide diversity of Crenarchaeota. Nature 384: 420.
Hinrichs KU, Hayes JM, Sylva SP, Brewer PG, DeLong EF . (1999). Methane-consuming archaebacteria in marine sediments. Nature 398: 802–805.
Huang L-N, Chen Y-Q, Zhou H, Luo S, Lan C-Y, Qu L-H . (2003). Characterization of methanogenic Archaea in the leachate of a closed municipal solid waste landfill. FEMS Microbiol Ecol 46: 171–177.
Huber JA, Welch DBM, Morrison HG, Huse SM, Neal PR, Butterfield DA et al. (2007). Microbial population structures in the deep marine biosphere. Science 318: 97–100.
Huse S, Huber J, Morrison H, Sogin M, Welch D . (2007). Accuracy and quality of massively parallel DNA pyrosequencing. Genome Biol 8: R143.
Inagaki F, Nunoura T, Nakagawa S, Teske A, Lever M, Lauer A et al. (2006). Biogeographical distribution and diversity of microbes in methane hydrate-bearing deep marine sediments on the Pacific Ocean Margin. Proc Natl Acad Sci USA 103: 2815–2820.
Inagaki F, Suzuki M, Takai K, Oida H, Sakamoto T, Aoki K et al. (2003). Microbial communities associated with geological horizons in coastal subseafloor sediments from the Sea of Okhotsk. Appl Environ Microbiol 69: 7224–7235.
Jørgensen BB, Dunker R, Grünke S, Røy H . (2010). Filamentous sulfur bacteria, Beggiatoa spp., in arctic marine sediments (Svalbard, 79°N). FEMS Microbiol Ecol 73: 500–513.
Jurgens G, Glöckner F-O, Amann R, Saano A, Montonen L, Likolammi M et al. (2000). Identification of novel Archaea in bacterioplankton of a boreal forest lake by phylogenetic analysis and fluorescent in situ hybridization. FEMS Microbiol Ecol 34: 45–56.
Karner MB, DeLong EF, Karl DM . (2001). Archaeal dominance in the mesopelagic zone of the Pacific Ocean. Nature 409: 507–510.
Kasai Y, Takahata Y, Hoak T, Watanabe K . (2005). Physiological and molecular characterization of a microbial community established in unsaturated, petroleum-contaminated soil. Environ Microbiol 7: 806–818.
Knittel K, Boetius A . (2009). Anaerobic oxidation of methane: progress with an unknown process. Ann Rev Microbiol 63: 311–334.
Knittel K, Boetius A, Lemke A, Eilers H, Lochte K, Pfannkuche O et al. (2003). Activity, distribution, and diversity of sulfate reducers and other bacteria in sediments above gas hydrate (Cascadia margin, Oregon). Geomicrobiol J 20: 269–294.
Knittel K, Lösekann T, Boetius A, Kort R, Amann R . (2005). Diversity and distribution of methanotrophic archaea at cold seeps. Appl Environ Microbiol 71: 467–479.
Kohls K, Abed RMM, Polerecky L, Weber M, De Beer D . (2010). Halotaxis of cyanobacteria in an intertidal hypersaline microbial mat. Environ Microbiol 12: 567–575.
Könneke M, Bernhard AE, de la Torre JR, Walker CB, Waterbury JB, Stahl DA . (2005). Isolation of an autotrophic ammonia-oxidizing marine archaeon. Nature 437: 543–546.
Labrenz M, Sintes E, Toetzke F, Zumsteg A, Herndl GJ, Seidler M et al. (2010). Relevance of a crenarchaeotal subcluster related to Candidatus Nitrosopumilus maritimus to ammonia oxidation in the suboxic zone of the central Baltic Sea. ISME J 4: 1496–1508.
Letunic I, Bork P . (2011). Interactive tree of life v2: online annotation and display of phylogenetic trees made easy. Nucleic Acids Res 39: W475–W478.
Li P-Y, Xie B-B, Zhang X-Y, Qin Q-L, Dang H-Y, Wang X-M et al. (2012). Genetic structure of three fosmid-fragments encoding 16S rRNA genes of the Miscellaneous Crenarchaeotic Group (MCG): implications for physiology and evolution of marine sedimentary archaea. Environ Microbiol 14: 467–479.
Lipp JS, Morono Y, Inagaki F, Hinrichs K-U . (2008). Significant contribution of Archaea to extant biomass in marine subsurface sediments. Nature 454: 991–994.
Llirós M, Alonso-Sáez L, Gich F, Plasencia A, Auguet O, Casamayor EO et al. (2011). Active bacteria and archaea cells fixing bicarbonate in the dark along the water column of a stratified eutrophic lagoon. FEMS Microbiol Ecol 77: 370–384.
Lloyd KG, Alperin MJ, Teske A . (2011). Environmental evidence for net methane production and oxidation in putative ANaerobic MEthanotrophic (ANME) archaea. Environ Microbiol 13: 2548–2564.
MacGregor BJ, Moser DP, Alm EW, Nealson KH, Stahl DA . (1997). Crenarchaeota in Lake Michigan sediment. Appl Environ Microbiol 63: 1178–1181.
Meng J, Wang F, Wang F, Zheng Y, Peng X, Zhou H et al. (2009). An uncultivated crenarchaeota contains functional bacteriochlorophyll a synthase. ISME J 3: 106–116.
Nercessian O, Bienvenu N, Moreira D, Prieur D, Jeanthon C . (2005). Diversity of functional genes of methanogens, methanotrophs and sulfate reducers in deep-sea hydrothermal environments. Environ Microbiol 7: 118–132.
Niemann H, Lösekann T, DeBeer D, Elvert M, Nadalig T, Knittel K et al. (2006). Novel microbial communities of the Haakon Mosby mud volcano and their role as a methane sink. Nature 443: 854–858.
Ochsenreiter T, Selezi D, Quaiser A, Bonch-Osmolovskaya L, Schleper C . (2003). Diversity and abundance of Crenarchaeota in terrestrial habitats studied by 16S RNA surveys and real time PCR. Environ Microbiol 5: 787–797.
Orcutt B, Joye SB, Kleindienst S, Knittel K, Ramette A, Reitz A et al. (2010). Impact of natural oil and higher hydrocarbons on microbial diversity, distribution, and activity in Gulf of Mexico cold-seep sediments. Deep-Sea Res II 57: 2008–2021.
Orcutt BN . (2007). Anaerobic oxidation of methane in cold seeps and gas hydrates: Responsible microorganisms, rates of activity and interactions with other processes. Ph.D. dissertation University of Georgia, Athens.
Parkes RJ, Webster G, Cragg BA, Weightman AJ, Newberry CJ, Ferdelman TG et al. (2005). Deep sub-seafloor prokaryotes stimulated at interfaces over geological time. Nature 436: 390–394.
Pernthaler A, Pernthaler J, Amann R . (2002). Fluorescence in situ hybridization and catalyzed reporter deposition for the identification of marine bacteria. Appl Environ Microbiol 68: 3094–3101.
Pruesse E, Quast C, Knittel K, Fuchs B, Ludwig W, Peplies J et al. (2007). SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res 35: 7188–7196.
Ravenschlag K, Sahm K, Knoblauch C, Jørgensen BB, Amann R . (2000). Community structure, cellular rRNA content and activity of sulfate-reducing bacteria in marine Arctic sediments. Appl Environ Microbiol 66: 3592–3602.
Rossel PE, Lipp JS, Fredricks HF, Arnds J, Boetius A, Elvert M et al. (2008). Intact polar lipids of anaerobic methanotrophic archaea and associated bacteria. Org Geochem 39: 992–999.
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB et al. (2009). Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75: 7537–7541.
Schramm A, Fuchs BM, Nielsen JL, Tonolla M, Stahl DA . (2002). Fluorescence in situ hybridization of 16S rRNA gene clones (Clone-FISH) for probe validation and screening of clone libraries. Environ Microbiol 4: 713–720.
Sørensen KB, Teske A . (2006). Stratified communities of active archaea in deep marine subsurface sediments. Appl Environ Microbiol 72: 4596–4603.
Stahl DA, Flesher B, Mansfield HR, Montgomery L . (1988). Use of phylogenetically based hybridization probes for studies of ruminal microbial ecology. Appl Environ Microbiol 54: 1079–1084.
Takai K, Horikoshi K . (1999). Genetic diversity of archaea in deep-sea hydrothermal vent environments. Genetics 152: 1285–1297.
Takai K, Moser DP, DeFlaun M, Onstott TC, Fredrickson JK . (2001). Archaeal diversity in waters from deep South African gold mines. Appl Environ Microbiol 67: 5750–5760.
Takano Y, Chikaraishi Y, Ogawa NO, Nomaki H, Morono Y, Inagaki F et al. (2010). Sedimentary membrane lipids recycled by deep-sea benthic archaea. Nat Geosci 3: 858–861.
Teira E, Reinthaler T, Pernthaler A, Pernthaler J, Herndl GJ . (2004). Combining catalyzed reporter deposition-fluorescence in situ hybridization and microautoradiography to detect substrate utilization by bacteria and archaea in the deep ocean. Appl Environ Microbiol 70: 4411–4414.
Tourna M, Stieglmeier M, Spang A, Könneke M, Schintlmeister A, Urich T et al. (2011). Nitrososphaera viennensis, an ammonia oxidizing archaeon from soil. Proc Natl Acad Sci USA 108: 8420–8425.
Tréhu AM, Bohrmann G, Rack FR, Torres ME . Proc. ODP, Init. Repts., 204. College Station, TX: Ocean Drilling Program doi:10.2973/odp.proc.ir.204.2003. (2003).
Treude T, Boetius A, Knittel K, Wallmann K, Jørgensen BB . (2003). Anaerobic oxidation of methane above gas hydrates at Hydrate Ridge, NE Pacific Ocean. Marine Ecol Prog Ser 264: 1–14.
Van Gaever S, Raes M, Pasotti F, Vanreusel A . (2010). Spatial scale and habitat-dependent diversity patterns in nematode communities in three seepage related sites along the Norwegian Sea margin. Mar Ecol 31: 77.
Vetriani C, Jannasch HW, MacGregor BJ, Stahl DA, Reysenbach AL . (1999). Population structure and phylogenetic characterization of marine benthic archaea in deep-sea sediments. Appl Environ Microbiol 65: 4375–4384.
Vetriani C, Reysenbach A-L, Doré J . (1998). Recovery and phylogenetic analysis of archaeal rRNA sequences from continental shelf sediments. FEMS Microbiol Lett 161: 83–88.
Webster G, Parkes RJ, Cragg BA, Newberry CJ, Weightman AJ, Fry JC . (2006). Prokaryotic community composition and biogeochemical processes in deep subseafloor sediments from the Peru Margin. FEMS Microbiol Ecol 58: 65–85.
Webster G, Rinna J, Roussel EG, Fry JC, Weightman AJ, Parkes RJ . (2010). Prokaryotic functional diversity in different biogeochemical depth zones in tidal sediments of the Severn Estuary, UK, revealed by stable-isotope probing. FEMS Microbiol Ecol 72: 179–197.
Yarza P, Ludwig W, Euzéby J, Amann R, Schleifer K-H, Glöckner FO et al. (2010). Update of the All-Species Living Tree Project based on 16S and 23S rRNA sequence analyses. Syst Appl Microbiol 33: 291–299.
Acknowledgements
We greatly acknowledge Antje Boetius for fruitful discussions and for providing samples from expeditions with the research vessels SONNE, POLARSTERN, POURQUOI PAS?, Tina Treude for samples from POSEIDON and Kai-Uwe Hinrichs and Katharina Kohls for sediments from ODP sites and hypersaline mats, respectively. Samples were taken in the framework of the GEOTECHNOLOGIEN programs MUMM I and II (grants 03G0554A and 03G0608A) funded by the German Ministry of Education and Research (BMBF) and the German Research Foundation, by the EU 6th FP HERMES, and by the Ocean Drilling Program (Leg 201 and 204) funded by the National Science Foundation (NSF) and participating countries under the Joint Oceanographic Institutions. Dirk Rickert is acknowledged for sulfate measurements in Hydrate Ridge sediments. Howard Mendlovitz, Daniel Albert, John Biddle, Andrew Steen, and Luke McKay helped with core acquisition and sectioning. Niculina Musat and Barbara MacGregor are acknowledged for helpful discussions about RNA extraction and slot blot hybridization. Financial support for K Kubo came from the German Academic Exchange Service (DAAD), for KGL from the US Environmental Protection Agency Star Fellowship, the Danish National Research Foundation. JFB was supported by a NASA NPP Fellowship administered by ORAU; and AT was supported from NOAA-NIUST and the NASA Astrobiology Institute for 16S clone library sequencing, and the Keck Foundation for V6-tag sequencing. Further support was provided by the Max Planck Society.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Supplementary Information accompanies the paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Kubo, K., Lloyd, K., F Biddle, J. et al. Archaea of the Miscellaneous Crenarchaeotal Group are abundant, diverse and widespread in marine sediments. ISME J 6, 1949–1965 (2012). https://doi.org/10.1038/ismej.2012.37
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2012.37
Keywords
This article is cited by
-
Thiobacillus as a key player for biofilm formation in oligotrophic groundwaters of the Fennoscandian Shield
npj Biofilms and Microbiomes (2023)
-
Microbial degradation of various types of dissolved organic matter in aquatic ecosystems and its influencing factors
Science China Earth Sciences (2023)
-
Microbial communities in the deep-sea sediments of the South São Paulo Plateau, Southwestern Atlantic Ocean
International Microbiology (2023)
-
Determination of carbon-fixing potential of Bathyarchaeota in marine sediment by DNA stable isotope probing analysis
Science China Earth Sciences (2023)
-
Conserved and lineage-specific hypothetical proteins may have played a central role in the rise and diversification of major archaeal groups
BMC Biology (2022)