Abstract
Understanding the relationship between prokaryotic traits and phylogeny is important for predicting and modeling ecological processes. Microbial extracellular enzymes have a pivotal role in nutrient cycling and the decomposition of organic matter, yet little is known about the phylogenetic distribution of genes encoding these enzymes. In this study, we analyzed 3058 annotated prokaryotic genomes to determine which taxa have the genetic potential to produce alkaline phosphatase, chitinase and β-N-acetyl-glucosaminidase enzymes. We then evaluated the relationship between the genetic potential for enzyme production and 16S rRNA phylogeny using the consenTRAIT algorithm, which calculated the phylogenetic depth and corresponding 16S rRNA sequence identity of clades of potential enzyme producers. Nearly half (49.2%) of the genomes analyzed were found to be capable of extracellular enzyme production, and these were non-randomly distributed across most prokaryotic phyla. On average, clades of potential enzyme-producing organisms had a maximum phylogenetic depth of 0.008004–0.009780, though individual clades varied broadly in both size and depth. These values correspond to a minimum 16S rRNA sequence identity of 98.04–98.40%. The distribution pattern we found is an indication of microdiversity, the occurrence of ecologically or physiologically distinct populations within phylogenetically related groups. Additionally, we found positive correlations among the genes encoding different extracellular enzymes. Our results suggest that the capacity to produce extracellular enzymes varies at relatively fine-scale phylogenetic resolution. This variation is consistent with other traits that require a small number of genes and provides insight into the relationship between taxonomy and traits that may be useful for predicting ecological function.
Similar content being viewed by others
Introduction
A major goal and challenge in microbial ecology is to link ecologically important metabolic functions to prokaryotic identity. Progress toward this goal has been hindered not only by the resistance of many microbes to traditional culture methods (Rappé and Giovannoni, 2003), but also by the inherent difficulty in attributing a microbially mediated process, measured in a complex environmental sample, to a specific taxon. Identifying the potential of specific organisms to contribute to an ecological process is valuable for making functional predictions and informing ecological models. Trait-based models have already shown promise for predicting ecosystem dynamics such as community structure (Follows et al., 2007) and decomposition (Allison, 2012), but would benefit from improved characterization of taxon-trait relationships. Traits related to resource acquisition are of particular interest, as these may be involved in both responding to and affecting ecosystem processes (Lavorel and Garnier, 2002).
The metabolic machinery of microorganisms contributes to ecosystem processes, including nutrient cycling and decomposition of organic matter. Decomposition by prokaryotes is initiated by the activity of extracellular enzymes that are secreted outside the cell, attached to the cell wall or located in the periplasm. These enzymes function to acquire energy and resources from organic matter for cellular growth, while catalyzing important transformations in the carbon (C), nitrogen (N) and phosphorus (P) cycles (Chrost and Siuda, 2002). Specific enzymes are produced to target the principal reservoirs of C, N and P in the environment. For example, alkaline phosphatases (APs) are broad specificity enzymes that release inorganic P (in the form of a phosphate group) from nucleic acids, phospholipids and other phosphate esters (Torriani, 1960; Sebastian and Ammerman, 2009, 2011). AP serves as the primary means by which prokaryotes hydrolyze P from organic material for uptake and use (Karl, 2000). Likewise, chitinases (CHIs) are produced to access the C and N resources contained in chitin, the second most abundant biopolymer on Earth next to cellulose (Jolles and Muzzarelli, 1999). Complete hydrolysis of chitin to monomer units also requires degradation by β-N-acetyl-glucosaminidase (NAG) (Gooday, 1990). The basics of biosynthesis, structure, catalytic properties and genetic regulation for extracellular enzymes are well known for only a few prokaryotes (for example, Bradshaw et al., 1981; Bassler et al., 1991; Jolles and Muzzarelli, 1999). Additionally, the range of taxa that are capable of producing specific extracellular enzymes remains largely unknown (Arnosti, 2011).
Our understanding of the metabolic capabilities of prokaryotic taxa has advanced significantly due to the sequencing of microbial genomes. For example, genomic analysis has allowed unprecedented insight into the environmental resource adaptation and ecological function of several key lineages, including Prochlorococcus (Rocap et al., 2003; Martiny et al., 2006), Synechococcus (Palenik et al., 2006), Roseobacter (Newton et al., 2010) and Escherichia coli (Luo et al., 2011). Since genome sequences represent the complete genetic repertoire of potential functions available to an organism, including strategies available for resource acquisition, the increasing availability of sequenced prokaryotic genomes holds much promise for clarifying taxon-trait relationships (Fraser et al., 2000; Ward and Klota, 2011).
Several previous studies have reported that traits related to resource use are associated with specific taxa or ecotypes at very fine-scale phylogenetic resolution (⩾97% 16S rRNA sequence identity). This has been demonstrated for resource traits such as particle colonization (Hunt et al., 2008), light adaptation (Moore et al., 1998; West and Scanlan, 1999; Johnson et al., 2006; Becraft et al., 2011) and nutrient use (Jaspers and Overmann, 2004; Martiny et al., 2006; Bhaya et al., 2007; Choudhary and Johri, 2011). This fine-scale association appears to hold true across the prokaryotic domains for genetically simple carbon use traits (Martiny et al., 2012); however, it is unknown whether extracellular enzyme traits also follow this pattern.
The objectives of the present study were two-fold: (1) to determine which prokaryotic taxa have the genetic potential to produce extracellular enzymes; and (2) to evaluate the linkages between that genetic potential and phylogeny, as defined by the 16S ribosomal RNA gene sequence. We used sequenced prokaryotic genomes to identify taxa with the genetic potential to produce AP, CHI and NAG enzymes. We then evaluated the linkages between that genetic potential and phylogeny by examining the size and average 16S rRNA sequence identity of clades capable of AP, CHI or NAG production. We hypothesized that vertical inheritance should result in a non-random distribution of enzyme-positive genotypes among prokaryotic taxa. If the genetic potential to produce extracellular enzymes is highly conserved, then enzyme-positive genotypes should be shared within large, deep clades (Figure 1a). In this case, broad changes in the taxonomic composition of a community may influence functioning associated with the trait. More likely, the genetic potential to produce extracellular enzymes is less conserved (more labile) and enzyme-positive genotypes should be found within smaller, closely related clades (Figure 1b). Changes in community composition would be unlikely to have a big impact on the functioning of such a dispersed trait. If phylogenetic conservation is related to genetic complexity as predicted by Martiny et al. (2012), then enzyme-positive genotypes should be associated with small clades (Figure 1b) since extracellular enzyme production is commonly encoded by a few genes (see references in Supplementary Tables S1–S3). Alternatively, the traits under investigation may be randomly associated with prokaryotic taxa (Figure 1c), possibly resulting from frequent gene gain/loss, rapid convergent evolution or horizontal gene transfer that obscure vertical inheritance (Doolittle, 1999; Gogarten et al., 2002; Snel et al., 2002; Boucher et al., 2003).
Hypothetical illustration depicting possible scenarios for the phylogenetic distribution and conservation of enzyme traits. Circles denote genomes with the genetic potential to produce extracellular enzymes (‘enzyme-positive genotypes’). Shaded branches highlight the phylogenetic depth (τD) of clades of enzyme-positive genotypes. Case (a) represents a strong correlation between phylogeny and enzyme trait, resulting in a large, deeply branching clade of enzyme-positive genotypes with relatively low 16S rRNA sequence identity (strong phylogenetic conservation). The correlation between phylogeny and enzyme trait is more labile in case (b), resulting in smaller clades of enzyme-positive genotypes with higher 16S rRNA sequence identity. Case (c) represents a random association between phylogeny and enzyme trait (no phylogenetic conservation).
We also hypothesized that the genetic potential to produce CHI and NAG enzymes should be positively correlated, thereby allowing the complete hydrolysis of chitin substrates by an individual organism. We used correlation analyses to test for associations between CHI- and NAG-positive genotypes within the data set.
Materials and methods
Identification of enzyme protein families
To target protein families with confirmed enzyme function, we conducted a literature search for empirically characterized amino acid sequences of prokaryotic AP (EC 3.1.3.1), CHI (EC 3.2.1.14) and NAG (EC 3.2.1.52). These enzymes were selected based on their ecological relevance and reasonably well-defined function/nomenclature. In all, 15–17 amino acid sequences from at least 11 different prokaryotic species were used as representatives for each enzyme (Supplementary Tables S1–S3). We then used the approach recently described by Martiny et al. (2012). The amino acid sequences were used to query the SEED database (Overbeek et al., 2005; http://seed-viewer.theseed.org) for the matching protein family identified by a unique ‘FIGfam’ number. SEED’s FIGfams are sets of isofunctional proteins that are homologous along the full length of the amino acid sequence (Meyer et al., 2009). Using FIGfams is a conservative method of identifying functional proteins, but provides an advantage over BLAST analysis since each FIGfam may encompass several genes coding for the same function. In total, 15 AP query sequences from 15 different species matched three FIGfams (FIG000766, FIG021520 and FIG024565). Seventeen CHI query sequences from 11 different species matched two FIGfams (FIG001347 and FIG004885). Sixteen NAG query sequences from 12 different species matched five FIGfams, but only three were used in our analysis (FIG001088, FIG003166 and FIG149633) because two corresponded to a different enzyme commission number (excluded FIG010408 and FIG048839). For each identified FIGfam, we compiled a list of prokaryotic genomes that had a protein sequence associated with the FIGfam function using the SEED API (Disz et al., 2010; http://www.theseed.org/servers/). This approach generated a matrix relating each genome to its corresponding genotype for AP, CHI and NAG. Genomes with the genetic potential to produce AP, CHI and/or NAG enzymes will be referred to as ‘enzyme-positive genotypes,’ whereas genomes in which the targeted protein families were not detected will be called ‘negative genotypes.’ We recognize that negative genomes may still contain genes for (non-homologous) proteins of the same or similar function that do not correspond to a defined FIGfam.
16S rRNA phylogenetic tree construction
We then generated a 16S rRNA phylogeny of all the annotated prokaryotic genomes available from SEED as of 5 December 2011. The organism name (derived from the NCBI species taxonomy ID) was used to extract the corresponding aligned 16S rRNA sequence from the Silva database (Release 108; Pruesse et al., 2007; http://beta.arb-silva.de/). Four eukaryotic large ribosomal subunit sequences were added to the alignment as outgroups (Arabidopsis thaliana, GenBank accession no. AX059457; Saccharomyces cerevisiae, AB278124; Aspergillus niger, EU884135; and Candida albicans, M60302). To account for uncertainty in the phylogeny, the sequence alignment was bootstrap sampled to generate 100 data sets using the SEQBOOT program from the PHYLIP software package (Felsenstein, 2005; http://evolution.genetics.washington.edu/phylip.html). The pairwise genetic distances between 16S rRNA sequences in each of the 100 data sets were measured with the DNADIST program utilizing the F84 model of nucleotide substitution (Felsenstein and Churchill, 1996). Phylogenetic trees were inferred from the square distance matrix of each data set with the neighbor-joining algorithm implemented by the NEIGHBOR program and randomized input order for the sequences. An additional distance matrix and neighbor-joining tree were constructed from the non-bootstrapped sequence alignment using the same methods and were used for visualization in iTOL (Letunic and Bork, 2007, 2011; http://itol.embl.de/) and to calculate phylogenetic-independent contrasts (PICs).
Phylogenetic conservation analysis
To evaluate the relationship between enzyme genotype and 16S rRNA phylogeny, we used the consenTRAIT algorithm (Martiny et al., 2012). The algorithm identified the root node of clades of enzyme-positive genotypes in which 90% of the terminal descendants shared the genotype for the enzyme of interest. Within each enzyme-positive clade, consenTRAIT calculated the average consensus sequence distance (d) between the root node (R) and the terminal node (S) for all clade members (m). If an enzyme-positive genotype did not have any neighbors that shared the trait (‘singleton’), then the average consensus sequence distance was calculated as half the branch length to the nearest neighbor. Trait depth, τD, was then calculated as the average of d values for n clades in the phylogeny:

This calculation was repeated for each bootstrapped tree and then averaged across the set of 100 trees to obtain the number of singletons, number of enzyme-positive clades, size of enzyme-positive clades, and τD for each enzyme studied. Bootstrap sampling with replacement approximates a normal distribution around a single sequence alignment, so analyzing the set of 100 bootstrapped trees allowed us to account for variation in the phylogeny. The calculated τD represents the average sequence difference within an enzyme-positive clade. τD was multiplied by two, and then subtracted from 1 as a measure of sequence identity of organisms within an enzyme-positive clade, and is comparable to a cutoff for defining operational taxonomic units.
To determine whether enzyme genotype and phylogeny were significantly non-randomly associated, we compared the observed size and trait depth (τD) of enzyme-positive clades to the same values calculated after randomizing the genome-genotype associations 1000 times (10 times for each of the 100 bootstrapped trees). The reported P-value is the fraction of randomizations that had a clade size or τD greater than or equal to that of the observed data. We considered enzyme genotype and phylogeny to be significantly associated (‘phylogenetically conserved’) for P-values <0.05.
Trait correlations
For all correlational analyses, the genetic potential to produce a specific enzyme was represented as ‘1’ if present or ‘0’ if absent within each genome. We examined associations between enzyme-positive genotypes specific for AP, CHI or NAG production first without considering phylogeny by calculating the Pearson’s product-moment correlation and Spearman’s rank correlation for all pairwise trait combinations using the ‘Hmisc’ package in R (Harrell, 2012; R v2.15.1, R Core Team, 2012; http://www.R-project.org/). Additionally, we calculated co-occurrence frequencies for directional pairwise combinations of enzyme-positive genotypes to account for the differences in total abundance of each enzyme in the data set. For example, we calculated the fraction of NAG-positive genotypes that also contained CHI. We then used the analysis of traits (AOT) function in the Phylocom software package (v4.2; Webb et al., 2008, 2011; http://phylodiversity.net/phylocom/) to calculate PICs that test for associations between specific enzyme-positive genotypes while correcting for the non-independence of related taxa (Felsenstein, 1985; Garland et al., 1992). Independent contrasts are calculated by AOT as the difference between the mean trait values of the bifurcating descendants at each internal node. The AOT function can be used to contrast two binary traits because the proportion of taxa possessing each trait is a continuous value between 0 and 1 for each clade. Contrasts were calculated for all pairwise trait combinations using the phylogeny generated from the non-bootstrapped sequence alignment. Before analysis, we used the FigTree program (v1.3.1; http://tree.bio.ed.ac.uk/software/figtree/) to transform all branch lengths into proportions to account for any zero values. We tested for significance of the resulting Pearson correlation coefficient using a table of critical values (Sokal and Rohlf, 1995).
The full output of AOT also includes a test of phylogenetic signal, which represents the degree to which each trait is conserved across the phylogeny (Blomberg and Garland, 2002). We used this calculation to validate the results of the consenTRAIT analysis. Phylocom uses the variance of PICs to calculate the phylogenetic signal similar to Blomberg and Garland’s K statistic (Blomberg and Garland, 2002; Blomberg et al., 2003; Webb et al., 2011) and determines the significance by comparing the observed PIC mean to a distribution of PIC means generated from 1000 randomizations of trait values across the tips of the phylogeny.
Results
Taxonomic composition of sequenced prokaryotic genomes
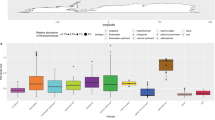
We analyzed 3058 prokaryotic genomes, including 30 phyla of Bacteria and Archaea, with 1–1312 genomes per phylum (Figure 2a). Proteobacteria (42.9% of genomes analyzed) was the most represented phylum in the data set. Other phyla that were disproportionately represented in the data set included Firmicutes (28.9%), Actinobacteria (10.0%) and Bacteroidetes (4.8%). Total representation from these abundant phyla accounted for 86.6% of the data set. The remaining phyla in the data set all contained <100 genomes each, and more than half of those contained <10 genomes each.
Occurrence of the genetic potential to produce AP, CHI and/or NAG enzymes (‘enzyme-positive genotypes’) among prokaryotic taxa. (a) Log abundance of genomes by phylum. (b) Proportion of enzyme-positive genotypes within each phylum. (c–e) Proportion of enzyme-positive genotypes specific for AP, CHI and NAG enzyme production.
Taxonomic distribution of enzyme-positive genotypes
Almost half of the 3058 genomes (1504, 49.2%) were identified as enzyme positive for AP, CHI, NAG or some combination (Figures 2b and 3). NAG-positive genotypes were most common (detected in 1127 genomes or 36.9% of all genomes analyzed), followed by AP- and CHI-positive genotypes (976, 31.9% and 461, 15.1%, respectively). Enzyme-positive genotypes were detected in all phyla except Korarchaeota, Thaumarchaeota, Aquificae, Chlamydiae, Chrysiogenetes and Fibrobacteres. Multi-enzyme genotypes capable of producing all three enzymes were found in the Acidobacteria, Actinobacteria, Bacteroidetes, Chloroflexi, Cyanobacteria, Dictyoglomi, Firmicutes, Proteobacteria, Spirochaetes, Theromotogae and Verrucomicrobia groups. AP- and NAG-positive genotypes were most broadly distributed across prokaryotic taxa and were detected in 22 and 19 phyla, respectively. CHI-positive genotypes were more narrowly distributed, and were found in only 12 of the 30 phyla.
Distribution of prokaryotic genomes with the genetic potential for AP, CHI or NAG enzyme production across a neighbor-joining phylogeny of 16S rRNA sequences. The red inner ring shows AP-positive genotypes, the blue middle ring shows CHI-positive genotypes and the green outer ring shows NAG-positive genotypes. Gray bars represent enzyme-negative genotypes. Highlighted clades depict (1) Enterococcus, (2) Burkholderia, (3) Vibrio and (4) Escherichia.
Although enzyme-positive genotypes were distributed across all the major groups of prokaryotes, the pattern of distribution within most groups was variable. For example, genomes within Escherichia (n=153) and Enterococcus (n=75) genera showed notable variation in the genetic potential to produce all three enzymes (Figure 3). This pattern is striking given the high rRNA relatedness (>99% identity) of the genomes within Escherichia. By contrast, enzyme genotype was consistent within few taxonomic groups, including Burkholderia (n=57) species for all enzymes studied, and Vibrio (n=60) species for CHI and NAG only.
Phylogenetic conservation
Clades of enzyme-positive genotypes varied broadly in size and relatedness (Figure 4a; Supplementary Figures S1a and S2a). The mean clade size for enzyme-positive genotypes was 1.74–2.02 genomes per clade (Table 1). For all three enzymes studied, randomizing the assignment of enzyme-positive genotypes across the phylogeny resulted in a significantly higher occurrence of singletons and smaller clade size than the observed data (Table 1; Figure 4b, Supplementary Figures S1b and S2b; P<0.001). Additionally, the mean trait depth (τD) for all three enzymes was also significantly different from the null (randomized) distribution. τD within clades ranged from 0.008004 16S rRNA distance for CHI to 0.009780 16S rRNA distance for AP (Table 1; Figure 4c; Supplementary Figures S1c and S2c; P<0.012). This corresponds to a 16S sequence identity of 98.40% for CHI and 98.04% for AP. Supporting this, we also detected a significant phylogenetic signal for all three enzymes (P<0.001 for AP and P=0.001 for CHI and NAG). Thus, based on the larger than random clade size, trait depth and phylogenetic signal, our analysis suggests that the enzyme-positive genotypes were non-randomly associated with the phylogeny.
Phylogenetic conservation of AP-positive genotypes. (a) Clade size range and relationship to mean trait depth, τD (average branch length from the root node to terminal nodes within enzyme-positive clades). (b) Frequency distribution of mean clade size of randomized data. The red bar represents the observed mean clade size, P<0.001. (c) Frequency distribution of τD of randomized data. The observed τD (red bar) is significantly different from the null distribution (P<0.001). The color reproduction of this figure is available on the ISME Journal online.
Trait correlations
We next analyzed the correlations between enzyme traits. Both Pearson and Spearman correlation tests supported significant (P<0.001, n=3058) associations between all pairwise enzyme combinations. AP- and NAG-positive genotypes, which were each individually more abundant in the data set than CHI-positive genotypes, were also most highly correlated (r=0.419). CHI-NAG and AP-CHI genotype combinations were slightly less correlated (r=0.322 and 0.302, respectively). These association patterns remained similar and significant (P<0.001, n=3058) even after considering phylogeny using independent contrasts. AP- and NAG-positive genotypes were still most correlated (r=0.474), followed by CHI-NAG (r=0.354) and AP-CHI (r=0.207).
Evaluating the frequency of directional combinations of enzyme-positive genotypes revealed a different association pattern. As we hypothesized, a high proportion of CHI-positive genotypes (73.75%) were also positive for NAG, regardless of the genotype for AP. By contrast, only 30.17% of NAG-positive genotypes were also positive for CHI. These results indicated that genomes with the genetic potential to produce CHI enzymes were mostly a subset of those capable of producing NAG enzymes.
Discussion
Our objectives were to determine which prokaryotic taxa have the genetic potential to produce AP, CHI and/or NAG enzymes (‘enzyme-positive genotypes’), and to evaluate the linkages between that genetic potential and phylogeny for sequenced prokaryotic genomes. Consistent with our hypothesis, the genetic potential to produce AP, CHI and/or NAG enzymes showed a significantly non-random association with phylogeny when measured by both the size (Figure 4b; Supplementary Figures S1b and S2b; P<0.001) and average phylogenetic relatedness (Figure 4c; Supplementary Figures S1c and S2c; P<0.012) of clades containing enzyme-positive genotypes. Although this finding suggests that vertical inheritance was generally important for the distribution of these traits in prokaryotes, the minimum phylogenetic relatedness within enzyme-positive clades was high, on average >98% 16S rRNA sequence identity (Table 1).
The relatedness value of >98% is closer to the species demarcation threshold suggested for predicting phenotypic potential (Konstantinidis et al., 2006a, 2006b) than the common threshold for distinguishing prokaryotic species (Stackebrandt and Goebel, 1994; Petti, 2007). This function could be inferred for operational taxonomic units defined by a 98% sequence identity cutoff or higher. However, the sequence identity values presented here should be interpreted with some caution. The high percentage of singletons (enzyme-positive genotypes whose nearest neighbors did not share the trait) for each enzyme is likely a consequence of under sampling prokaryotic diversity. When these singletons were excluded as clades in the phylogenetic conservation analysis, the average relatedness within enzyme-positive clades decreased (AP=96.9%, CHI=98.0% and NAG=96.7%). The number of singletons may change as we continue to sample more of the prokaryotic diversity in nature, though it is unclear how trait associations may be affected. Sampling biases in prokaryotic genome sequencing likely also influenced the variation in individual clade size and phylogenetic relatedness (Figure 4a; Supplementary Figures S1a and S2a), such that large clades with high relatedness are the result of deep sampling of a few taxonomic groups in the data set (for example, E. coli). Despite these biases, the fine-scale association that we detected between enzyme genotype and phylogeny is evidence of ‘microdiversity,’ the occurrence of ecologically or physiologically distinct populations within phylogenetically related groups (Moore et al., 1998).
Microdiversity is a well-documented phenomenon among prokaryotes, particularly for traits related to resource use. Prochlorococcus isolates have been divided into ecotypes based on exploitation of light resources, despite being >97% similar in ribosomal identity (Moore et al., 1998; West and Scanlan, 1999). Martiny et al. (2006) also found that the gene content for phosphate acquisition was not congruent with rRNA phylogeny for members of Prochlorococcus. Likewise, extremely close relatives of hot spring Synechococcus have been shown to differ in their adaptations to light levels (Becraft et al., 2011) as well as phosphorus and nitrogen use pathways (Bhaya et al., 2007). Isolates of Pseudomonas, Acinetobacter (Choudhary and Johri, 2011) and Brevundimonas (Jaspers and Overmann, 2004), which had identical 16S rRNA gene sequences, were also found to occupy different ecological niches, with each using a unique combination of carbon substrates. Additionally, Hunt et al. (2008) demonstrated that Vibrio isolates could be resolved into ecologically distinct populations based on resource partitioning within the water column. These examples represent cases of local adaptation or niche specialization, which can be the first step in the process of ecological speciation. Likewise, the microdiversity detected for AP-, CHI- and NAG-positive genotypes may provide evidence for the importance of extracellular enzymes in the origins of ecological diversity.
Although not tested here, we speculate that several ecological processes could contribute to microdiversity in the ability to use phosphate esters and chitin among prokaryotic taxa. It is likely that gene content reflects differential adaptation to environmental conditions even among closely related organisms. Extracellular enzymes are commonly produced by prokaryotes in low-nutrient environments to access the resources trapped in high molecular weight compounds (Torriani, 1960; Münster and Chrost, 1990; Amon and Benner, 1996; Delpin and Goodman, 2009). As such, microdiversity in enzyme genotype may occur if individuals are adapted to contrasting nutrient regimes. For example, nitrogen transporter genes differed between coastal (high nutrient) and open ocean (low nutrient) isolates of Synechococcus (Palenik et al., 2006) and phosphate acquisition genes varied among strains of Prochlorococcus in relation to the specific nutrient availability where the strains were isolated (Martiny et al., 2006). These patterns of variation were reflected in the distribution of AP-positive genotypes among Cyanobacteria in our data set (Figure 3), and may be the result of frequent local resource adaptation. Ideally, environmental data collected from an organism’s isolation source could be used to test for correlations with particular traits. We suspect that the taxonomic distribution of AP-, CHI- and NAG-positive genotypes may be related to nutrient supply; however, these data were not available for the sequenced genomes used in this study. Fortunately, the field of microbial ecology is experiencing a shift toward more meticulous measurement and reporting of contextual environmental data (for example, Yilmaz et al., 2011).
Microdiversity in enzyme genotype may also be associated with lifestyle strategy. Commensal or pathogenic strains may have greater access to more labile nutrients than their free-living counterparts, alleviating the necessity for extracellular enzymes. We notably did not detect the ability to use phosphate esters or chitin within Fibrobacteres, a gut symbiont (Ransom-Jones et al., 2012), or Chlamydiae, an obligate intracellular pathogen (Horn, 2008). Recently, Luo et al. (2011) found that environmental isolates of E. coli shared a set of genes important for resource acquisition that were absent from enteric isolates. Likewise, the enteric strains shared a set of genes involved in the transport and use of several labile nutrients, which were absent from the environmental isolates. We found that members within Escherichia varied in their genetic potential to produce all three enzymes (Figure 3). This variation may be evidence that they are changing their ecology or even speciating more often than other lineages, such as Burkholderia, which was relatively coherent for enzyme genotype (Figure 3).
Finally, horizontal gene transfer may also result in important ecological differences between closely related organisms (Welch et al., 2002). The extent to which horizontal gene transfer may occur for resource acquisition machinery among prokaryotes is unclear. Previous work suggests that horizontal gene transfer may significantly impact the evolution of chitin-degrading enzymes (Garcia-Vallve et al., 1999). However, adaptive transfer of genes is limited to those that can be transferred as a functional unit containing a complement of genes that are involved in processing a single resource molecule (Wiedenbeck and Cohan, 2011). Structural or physiological incompatibilities may further inhibit the horizontal transfer of genes between individuals (Cohan and Koeppel, 2008).
The fine-scale association between enzyme genotype and phylogeny is consistent with the relatively simple genetic structure underlying the production of extracellular enzymes. These enzyme systems involve few genes/operons that may allow enzyme function to be rapidly gained or lost though evolutionary processes including horizontal gene transfer. Martiny et al. (2012) found similar association values for carbon usage traits, which ranged from 96.6% to 97.8% 16S rRNA sequence identity within positive clades, depending on the data set analyzed. By contrast, more complex traits such as oxygenic photosynthesis or sulfate reduction involve many more genes and show phylogenetic conservation at much deeper taxonomic levels (80% and 92.2% mean 16S rRNA sequence identity, respectively; Martiny et al., 2012). Other traits not directly related to resource use, such as rRNA operon copy number (Rastogi et al., 2009) and host adaptation (Ettema and Andersson, 2009), may also be conserved at deeper taxonomic levels.
We predicted that the genetic potential to produce CHI and NAG enzymes should be positively correlated, thereby allowing the complete hydrolysis of chitin substrates by an individual organism. In support of our hypothesis, we found a significant association between the ability to hydrolyze chitin and use the end products of hydrolysis, regardless of whether we corrected for shared evolutionary history. Allocation to extracellular enzymes represents a significant investment of carbon and nitrogen, and cells should be under selection to regulate production and maximize substrate use (Allison et al., 2007, 2011). Most of the Vibrio genomes in our data set were capable of complete chitin hydrolysis, which is consistent with their known ecology. Vibrios can be found in a range of aquatic habitats and have one of the most well-studied chitinolytic systems of any prokaryote (Keyhani and Roseman, 1999; Xibing Li and Roseman, 2004; Pruzzo et al., 2008). While a high proportion of CHI-positive genotypes (73.75%) were also positive for NAG, we detected potential for CHI production without the potential for NAG production in a few groups within the Firmicutes phylum (Figure 3). For these strains, CHI production may be primarily involved in virulence (Larsen et al., 2010) or biofilm formation (Tirumalai and Prakash, 2011) instead of resource acquisition. Alternatively, we may not have identified all NAG-positive genotypes in our analysis. By contrast, a majority (69.83%) of NAG-positive genomes appeared to lack the genetic potential for CHI production. We suspect that this may be related to the numerous biological and environmental sources of NAG substrates aside from the products of CHI activity, causing a relatively low correlation between the two enzymes. We also detected significant positive correlations between AP- and CHI-positive genotypes as well as AP- and NAG-positive genotypes, suggesting that some prokaryotes may function as general ‘enzyme producers,’ with the ability to produce a suite of extracellular enzymes. Furthermore, the similarity of the raw correlations with the PIC analysis indicates that these traits are correlated independent of phylogeny, and that there may be selection for the traits to co-occur.
Identifying the appropriate level of phylogenetic relatedness for a trait of interest is critical for defining ecologically coherent units and predicting/interpreting community function. Our study shows that extracellular enzyme genes are not randomly distributed throughout the prokaryotic phylogeny, but the average level of phylogenetic conservation for enzyme genes is less than the level typically used to define bacterial operational taxonomic units based on 16S markers. Consequently, the microdiversity associated with enzyme traits could mask correlations between enzyme activity and community composition (for example, Frossard et al., 2012) if operational taxonomic units are defined below ∼98% 16S rRNA sequence identity. Trait-based techniques such as GeoChip (He et al., 2007), which allow for directed assessment of functional genes and processes, may be more appropriate for investigating the ecological consequences of phylogenetically fine-scale microbial traits than inferring function based on phylogenetic marker genes. Continued efforts by microbial ecologists to identify the levels of phylogenetic conservation of metabolic functions among prokaryotes will enhance our ability to predict microbial impacts on ecosystem processes.
Accession codes
References
Alam M, Nikaidou N, Tanaka H, Watanabe T . (1995). Cloning and sequencing of chiC gene of Bacillus circulans WL-12 and relationship of its product to some other chitinases and chitinase-like proteins. J Ferment Bioeng 80: 454–461.
Allison SD . (2012). A trait-based approach for modelling microbial litter decomposition. Ecol Lett 15: 1058–1070.
Allison SD, Gartner TB, Holland K, Weintraub M, Sinsabaugh RL . (2007). Soil enzymes: Linking proteomics and ecological process. In Hurst CJ, Garland JL, Mills AL, (eds). Manual of Environmental Microbiolgy 3rd Edition. ASM Press: Washington, DC, pp 704–711.
Allison SD, Weintraub MN, Gartner TB, Waldrop MP . (2011). Evolutionary-economic principles as regulators of soil enzyme production and ecosystem function. In Shukla G, Varma A, (eds). Soil Enzymology. Springer-Verlag: Berlin, Heidelberg, pp 229–243.
Amon RMW, Benner R . (1996). Bacterial utilization of different size classes of dissolved organic matter. Limnol Oceanogr 41: 41–51.
Arnosti C . (2011). Microbial extracellular enzymes and the marine carbon cycle. Annu Rev Mar Sci 3: 401–425.
Asgeirsson B, Andrésson OS . (2001). Primary structure of cold-adapted alkaline phosphatase from a Vibrio sp. as deduced from the nucleotide gene sequence. Biochim Biophys Acta 1549: 99–111.
Au S, Roy KL, von Tigerstrom RG . (1991). Nucleotide sequence and characterization of the gene for secreted alkaline phosphatase from Lysobacter enzymogenes. J Bacteriol 173: 4551–4557.
Bassler BL, Yu C, Lee YC, Roseman S . (1991). Chitin utilization by marine bacteria: degradation and catabolism of chitin oligosaccharides by Vibrio furnissii. J Biol Chem 266: 24276–24286.
Becraft ED, Cohan FM, Kühl M, Jensen SI, Ward DM . (2011). Fine-scale distribution patterns of Synechococcus ecological diversity in microbial mats of Mushroom Spring, Yellowstone National Park. Appl Environ Microbiol 77: 7689–7697.
Bhaya D, Grossman AR, Steunou A-S, Khuri N, Cohan FM, Hamamura N et al (2007). Population level functional diversity in a microbial community revealed by comparative genomic and metagenomic analyses. ISME J 1: 703–713.
Blaak H, Schnellmann J, Walter S, Henrissat B, Schrempf H . (1993). Characteristics of an exochitinase from Streptomyces olivaceoviridis, its corresponding gene, putative protein domains and relationship to other chitinases. Eur J Biochem 214: 659–669.
Blomberg SP, Garland T . (2002). Tempo and mode in evolution: phylogenetic inertia, adaptation and comparative methods. J Evol Biol 15: 899–910.
Blomberg SP, Garland T, Ives AR . (2003). Testing for phylogenetic signal in comparative data: behavioral traits are more labile. Evolution 57: 717–745.
Boucher Y, Douady CJ, Papke RT, Walsh DA, Boudreau MER, Nesbø CL et al (2003). Lateral gene transfer and the origins of prokaryotic groups. Annu Rev Genet 37: 283–328.
Bradshaw RA, Cancedda F, Ericsson LH, Neumann PA, Piccoli SP, Schlesinger MJ et al (1981). Amino acid sequence of Escherichia coli alkaline phosphatase. Proc Natl Acad Sci USA 78: 3473–3477.
Chang CN, Kuang W-J, Chen EY . (1986). Nucleotide sequence of the alkaline phosphatase gene of Escherichia coli. Gene 44: 121–125.
Choudhary DK, Johri BN . (2011). Ecological significance of microdiversity: coexistence among casing soil bacterial strains through allocation of nutritional resource. Indian J Microbiol 51: 8–13.
Chrost RJ, Siuda W . (2002). Ecology of microbial enzymes in lake ecosystems. In Enzymes in the Environment: Activity, Ecology, and Applications Burns RG, Dick RP, (eds). Marcel Dekker, Inc.: New York, pp 35–72.
Cohan FM, Koeppel AF . (2008). The origins of ecological diversity in prokaryotes. Curr Biol 18: R1024–R1034.
Delpin MW, Goodman AE . (2009). Nitrogen regulates chitinase gene expression in a marine bacterium. ISME J 3: 1064–1069.
Disz T, Akhter S, Cuevas D, Olson R, Overbeek R, Vonstein V et al (2010). Accessing the SEED genome databases via web services API: tools for programmers. BMC Bioinformatics 11: 319.
Doolittle WF . (1999). Phylogenetic classification and the universal tree. Science 284: 2124–2128.
Ettema TJG, Andersson SGE . (2009). The alpha-proteobacteria: the Darwin finches of the bacterial world. Biol Lett 5: 429–432.
Felsenstein J . (2005), PHYLIP.
Felsenstein J . (1985). Phylogenies and the comparative method. Am Nat 125: 1–15.
Felsenstein J, Churchill GA . (1996). A Hidden Markov Model approach to variation among sites in rate of evolution. Mol Biol Evol 13: 93–104.
Follows MJ, Dutkiewicz S, Grant S, Chisholm SW . (2007). Emergent biogeography of microbial communities in a model ocean. Science 315: 1843–1846.
Fraser CM, Eisen JA, Salzberg SL . (2000). Microbial genome sequencing. Nature 406: 799–803.
Frossard A, Gerull L, Mutz M, Gessner MO . (2012). Disconnect of microbial structure and function: enzyme activities and bacterial communities in nascent stream corridors. ISME J 6: 680–691.
Fujita M, Tsuchida A, Hirata A, Kobayashi N, Goto K, Osumi K et al (2011). Glycoside hydrolase family 89 alpha-N-acetylglucosaminidase from Clostridium perfringens specifically acts on GlcNAc alpha1,4Gal beta1R at the non-reducing terminus of O-glycans in gastric mucin. J Biol Chem 286: 6479–6489.
Garcia-Vallve S, Palau J, Romeu A . (1999). Horizontal gene transfer in glycosyl hydrolases inferred from codon usage in Escherichia coli and Bacillus subtilis. Mol Biol Evol 16: 1125–1134.
Garland TheodoreJ, Harvey PH, Ives AR . (1992). Procedures for the analysis of comparative data using phylogenetically independent contrasts. Syst Biol 41: 18–32.
Gogarten JP, Doolittle WF, Lawrence JG . (2002). Prokaryotic evolution in light of gene transfer. Mol Biol Evol 19: 2226–2238.
Gomez PF, Ingram LO . (1995). Cloning, sequencing and characterization of the alkaline phosphatase gene (phoD) from Zymomonas mobilis. FEMS Microbiol Lett 125: 237–245.
Gooday GW . (1990). The ecology of chitin degradation. In Marshall KC, (eds). Advances in Microbial Ecology Vol. 11. Springer: New York, pp 387–419.
Harrell FE, with contributions from many other users (2012), Hmisc: Harrell Miscellaneous. R Package version 3.9-3. http://cran.r-project.org/package=Hmisc.
Hauksson JB, Andrésson OS, Ásgeirsson B . (2000). Heat-labile bacterial alkaline phosphatase from a marine Vibrio sp. Enzyme Microb Technol 27: 66–73.
He Z, Gentry TJ, Schadt CW, Wu L, Liebich J, Chong SC et al (2007). GeoChip: a comprehensive microarray for investigating biogeochemical, ecological and environmental processes. ISME J 1: 67–77.
Horn M . (2008). Chlamydiae as symbionts in eukaryotes. Annu Rev Microbiol 62: 113–131.
Howard MB, Ekborg NA, Taylor LE, Weiner RM, Hutcheson SW . (2003). Genomic analysis and initial characterization of the chitinolytic system of Microbulbifer degradans Strain 2-40. J Bacteriol 185: 3352–3360.
Hulett FM, Bookstein C, Jensen K . (1990). Evidence for two structural genes for alkaline phosphatase in Bacillus subtilis. J Bacteriol 172: 735–740.
Hulett FM, Kim EE, Bookstein C, Kapp NV, Edwards CW, Wyckoff HW . (1991). Bacillus subtilis alkaline phosphatases III and IV. Cloning, sequencing, and comparisons of deduced amino acid sequence with Escherichia coli alkaline phosphatase three-dimensional structure. J Biol Chem 266: 1077–1084.
Hunt DE, David LA, Gevers D, Preheim SP, Alm EJ, Polz MF . (2008). Resource partitioning and sympatric differentiation among closely related bacterioplankton. Science 320: 1081–1085.
Jaspers E, Overmann J . (2004). Ecological significance of microdiversity: Identical 16S rRNA gene sequences can be found in bacteria with highly divergent genomes and ecophysiologies. Appl Environ Microbiol 70: 4831–4839.
Johnson ZI, Zinser ER, Coe A, McNulty NP, Woodward EMS, Chisholm SW . (2006). Niche partitioning among Prochlorococcus ecotypes along ocean-scale environmental gradients. Science 311: 1737–1740.
Jolles P, Muzzarelli RAA . (1999). Preface. In Jolles P, Muzzarelli RAA, (eds). Chitin and Chitinases. Birkhauser Verlag: Basel.
Karl DM . (2000). Phosphorus, the staff of life. Nature 406: 31–32.
Keyhani NO, Roseman S . (1999). Physiological aspects of chitin catabolism in marine bacteria. Biochim Biophys Acta 1473: 108–122.
Kim J-W, Peterson T, Bee G, Hulett FM . (1998). Bacillus licheniformis MC14 alkaline phosphatase I gene with an extended COOH-terminus. FEMS Microbiol Lett 159: 47–58.
Kim Y-J, Park T-S, Kim H-K, Kwon S-T . (1997). Purification and characterization of a thermostable alkaline phosphatase produced by Thermus caldophilus GK24. J Biochem Mol Biol 30: 262–268.
Kless H, Sitrit Y, Chet I, Oppenheim AB . (1989). Cloning of the gene coding for chitobiase of Serratia marcescens. MGG 217: 471–473.
Konstantinidis KT, Ramette A, Tiedje JM . (2006a). The bacterial species definition in the genomic era. Philos Trans R Soc Lond B Biol Sci 361: 1929–1940.
Konstantinidis KT, Ramette A, Tiedje JM . (2006b). Toward a more robust assessment of intraspecies diversity, using fewer genetic markers. Appl Environ Microbiol 72: 7286–7293.
Kubota T, Miyamoto K, Yasuda M, Inamori Y, Tsujibo H . (2004). Molecular characterization of an intracellular beta-N-acetylglucosaminidase involved in the chitin degradation system of Streptomyces thermoviolaceus OPC-520. Biosci Biotechnol Biochem 68: 1306–1314.
Lan X, Zhang X, Kodaira R, Zhou Z, Shimosaka M . (2008). Gene cloning, expression, and characterization of a second β-N-acetylglucosaminidase from the chitinolytic bacterium Aeromonas hydrophila Strain SUWA-9. Biosci Biotechnol Biochem 72: 492–498.
Larsen MH, Leisner JJ, Ingmer H . (2010). The chitinolytic activity of Listeria monocytogenes EGD is regulated by carbohydrates but also by the virulence regulator PrfA. Appl Environ Microbiol 76: 6470–6476.
Lavorel S, Garnier E . (2002). Predicting changes in community composition and ecosystem functioning from plant traits: revisiting the Holy Grail. Funct Ecol 16: 545–556.
Letunic I, Bork P . (2007). Interactive Tree Of Life (iTOL): an online tool for phylogenetic tree display and annotation. Bioinformatics 23: 127–128.
Letunic I, Bork P . (2011). Interactive Tree Of Life v2: online annotation and display of phylogenetic trees made easy. Nucleic Acids Res 39: W475–W478.
Li H, Morimoto K, Katagiri N, Kimura T, Sakka K, Lun S et al (2002). A novel beta-N-acetylglucosaminidase of Clostridium paraputrificum M-21 with high activity on chitobiose. Appl Microbiol Biotechnol 60: 420–427.
Li X, Roseman S . (2004). The chitinolytic cascade in Vibrios is regulated by chitin oligosaccharides and a two-component chitin catabolic sensor/kinase. Proc Natl Acad Sci USA 101: 627–631.
Lin J, Xiao X, Zeng X, Wang F . (2006). Expression, characterization and mutagenesis of the gene encoding β-N-acetylglucosaminidase from Aeromonas caviae CB101. Enzyme Microb Technol 38: 765–771.
Luo C, Walk ST, Gordon DM, Feldgarden M, Tiedje JM, Konstantinidis KT . (2011). Genome sequencing of environmental Escherichia coli expands understanding of the ecology and speciation of the model bacterial species. Proc Natl Acad Sci USA 108: 7200–7205.
Majumdar A, Ghatak A, Ghosh RK . (2005). Identification of the gene for the monomeric alkaline phosphatase of Vibrio cholerae serogroup O1 strain. Gene 344: 251–258.
Martiny AC, Coleman ML, Chisholm SW . (2006). Phosphate acquisition genes in Prochlorococcus ecotypes: evidence for genome-wide adaptation. Proc Natl Acad Sci USA 103: 12552–12557.
Martiny AC, Tai APK, Veneziano D, Primeau F, Chisholm SW . (2009). Taxonomic resolution, ecotypes and the biogeography of Prochlorococcus. Environ Microbiol 11: 823–832.
Martiny AC, Treseder KK, Pusch GD . (2012). Phylogenetic conservatism of functional traits in microorganisms. ISME J e-pub ahead of print 13 December 2012 10.1038/ismej.2012.160.
Matsuo Y, Kurita M, Park JK, Tanaka K, Nakagawa T, Kawamukai M et al (1999). Purification, characterization and gene analysis of N-acetylglucosaminidase from Enterobacter sp. G-1. Biosci Biotechnol Biochem 63: 1261–1268.
Mayer C, Vocadlo DJ, Mah M, Rupitz K, Stoll D, Warren RAJ et al (2006). Characterization of a beta-N-acetylhexosaminidase and a beta-N-acetylglucosaminidase/beta-glucosidase from Cellulomonas fimi. FEBS J 273: 2929–2941.
Meyer F, Overbeek R, Rodriguez A . (2009). FIGfams: Yet another set of protein families. Nucleic Acids Res 37: 6643–6654.
Moore LR, Rocap G, Chisholm SW . (1998). Physiology and molecular phylogeny of coexisting Prochlorococcus ecotypes. Nature 393: 464.
Morimoto Kenji YoshimotoM, Karita S, Tetsuya Kimura, Kunio Ohmiya, Kazuo. Sakka . (2007). Characterization of the third chitinase Chi18C of Clostridium paraputrificum M-21. Appl Microbiol Biotechnol 73: 1106–1113.
van Mourik A, Bleumink-Pluym NMC, van Dijk L, van Putten JPM, Wösten MMSM . (2008). Functional analysis of a Campylobacter jejuni alkaline phosphatase secreted via the Tat export machinery. Microbiology 154: 584–592.
Münster U, Chrost RJ . (1990). Origin, composition and microbial utilization of dissolved organic matter. In Overbeck J, Chrost RJ, (eds). Aquatic Microbial Ecology: Biochemical and Molecular Approaches. Springer-Verlag: New York, pp 8–46.
Newton RJ, Griffin LE, Bowles KM, Meile C, Gifford S, Givens CE et al (2010). Genome characteristics of a generalist marine bacterial lineage. ISME J 4: 784–798.
Nilgiriwala KS, Alahari A, Rao AS, Apte SK . (2008). Cloning and overexpression of alkaline phosphatase PhoK from Sphingomonas sp. strain BSAR-1 for bioprecipitation of uranium from alkaline solutions. Appl Environ Microbiol 74: 5516–5523.
Overbeek R, Begley T, Butler RM, Choudhuri JV, Chuang H-Y, Cohoon M et al (2005). The subsystems approach to genome annotation and its use in the project to annotate 1000 genomes. Nucleic Acids Res 33: 5691–5702.
Palenik B, Ren Q, Dupont CL, Myers GS, Heidelberg JF, Badger JH et al (2006). Genome sequence of Synechococcus CC9311: Insights into adaptation to a coastal environment. Proc Natl Acad Sci USA 103: 13555–13559.
Park T, Lee J-H, Kim H-K, Hoe H-S, Kwon S-T . (1999). Nucleotide sequence of the gene for alkaline phosphatase of Thermus caldophilus GK24 and characteristics of the deduced primary structure of the enzyme. FEMS Microbiol Lett 180: 133–139.
Petti CA . (2007). Detection and identification of microorganisms by gene amplification and sequencing. Clin Infect Dis 44: 1108–1114.
Pruesse E, Quast C, Knittel K, Fuchs BM, Ludwig W, Peplies J et al (2007). SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res 35: 7188–7196.
Pruzzo C, Vezzulli L, Colwell RR . (2008). Global impact of Vibrio cholerae interactions with chitin. Environ Microbiol 10: 1400–1410.
R Core Team. (2012), R: A language and environment for statistical computing. R Foundation for Statistical Computing. http://www.r-project.org/.
Ransom-Jones E, Jones DL, McCarthy AJ, McDonald JE . (2012). The Fibrobacteres: an important phylum of cellulose-degrading bacteria. Microb Ecol 63: 267–281.
Rappé MS, Giovannoni SJ . (2003). The uncultured microbial majority. Annu Rev Microbiol 57: 369–394.
Rashid MH, Mori M, Sekiguchi J . (1995). Glucosaminidase of Bacillus subtilis: cloning, regulation, primary structure and biochemical characterization. Microbiology 141: 2391–2404.
Rastogi R, Wu M, DasGupta I, Fox GE . (2009). Visualization of ribosomal RNA operon copy number distribution. BMC Microbiol 9: 208.
Ray JM, Bhaya D, Block MA, Grossman AR . (1991). Isolation, transcription, and inactivation of the gene for an atypical alkaline phosphatase of Synechococcus sp. strain PCC 7942. J Bacteriol 173: 4297–4309.
Rina M, Pozidis C, Mavromatis K, Tzanodaskalaki M, Kokkinidis M, Bouriotis V . (2000). Alkaline phosphatase from the Antarctic strain TAB5: properties and psychrophilic adaptations. Eur J Biochem 267: 1230–1238.
Rocap G, Larimer FW, Lamerdin J, Malfatti S, Chain P, Ahlgren NA et al (2003). Genome divergence in two Prochlorococcus ecotypes reflects oceanic niche differentiation. Nature 424: 1042–1047.
Roy NK, Ghosh RK, Das J . (1982). Monomeric alkaline phosphatase of Vibrio cholerae. J Bacteriol 150: 1033–1039.
Sebastian M, Ammerman JW . (2011). Role of the phosphatase PhoX in the phosphorus metabolism of the marine bacterium Ruegeria pomeroyi DSS-3. Environ Microbiol Rep 3: 535–542.
Sebastian M, Ammerman JW . (2009). The alkaline phosphatase PhoX is more widely distributed in marine bacteria than the classical PhoA. ISME J 3: 563–572.
Senba M, Kashige N, Nakashima Y, Miake F, Watanabe K . (2000). Cloning of the gene of beta-N-acetylglucosaminidase from Lactobacillus casei ATCC 27092 and characterization of the enzyme expressed in Escherichia coli. Biol Pharm Bull 23: 527–531.
Sitrit Y, Vorgias CE, Chet I, Oppenheim AB . (1995). Cloning and primary structure of the chiA gene from Aeromonas caviae. J Bacteriol 177: 4187–4189.
Snel B, Bork P, Huynen MA . (2002). Genomes in flux: the evolution of archaeal and proteobacterial gene content. Genome Res 12: 17–25.
Sokal RR, Rohlf FJ . (1995) Biometry: The Principles and Practice of Statistics in Biological Research 3rd Edition WH Freeman and Company: New York.
Stackebrandt E, Goebel BM . (1994). Taxonomic Note: a place for DNA-DNA reassociation and 16S rRNA sequence analysis in the present species befinition in Bacteriology. Int J Syst Bacteriol 44: 846–849.
Suginta W, Chuenark D, Mizuhara M, Fukamizo T . (2010). Novel β-N-acetylglucosaminidases from Vibrio harveyi 650: cloning, expression, enzymatic properties, and subsite identification. BMC Biochem 11: 40.
Suzuki K, Taiyoji M, Sugawara N, Nikaidou N, Henrissat B, Watanabe T . (1999). The third chitinase gene (chiC) of Serratia marcescens 2170 and the relationship of its product to other bacterial chitinases. Biochem J 343: 587–596.
Svitil AL, Kirchman DL . (1998). A chitin-binding domain in a marine bacterial chitinase and other microbial chitinases: implications for the ecology and evolution of 1,4-beta-glycanases. Microbiology 144: 1299–1308.
Techkarnjanaruk S, Pongpattanakitshote S, Goodman AE . (1997). Use of a promoterless lacZ gene insertion to investigate chitinase gene expression in the marine bacterium Pseudoalteromonas sp. strain S9. Appl Environ Microbiol 63: 2989–2996.
Tews I, Vincentelli R, Vorgias CE . (1996). N-Acetylglucosaminidase (chitobiase) from Serratia marcescens: gene sequence, and protein production and purification in Escherichia coli. Gene 170: 63–67.
Tirumalai PS, Prakash S . (2011). Expression of chitinase and chitin binding proteins (CBP’s) by Listeria monocytogenes J0161 in biofilm and co-culture broths. African J Microbiol Res 5: 5188–5193.
Torriani A . (1960). Influence of inorganic phosphate in the formation of phosphatases by Escherichia coli. Biochim Biophys Acta 38: 460–469.
Tsujibo H, Hatano N, Mikami T, Izumizawa Y, Miyamoto K, Inamori Y . (1998). Cloning, characterization and expression of beta-N-acetylglucosaminidase gene from Streptomyces thermoviolaceus OPC-520(1). Biochim Biophys Acta 1425: 437–440.
Tsujibo Hiroshi, Orikoshi H, Baba N, Miyahara M, Katsushiro Miyamoto, Yasuda M et al (2002). Identification and characterization of the gene cluster involved in chitin degradation in a marine bacterium, Alteromonas sp. strain O-7. Appl Environ Microbiol 68: 263–270.
Ueda M, Kawaguchi T, Arai M . (1994). Molecular cloning and nucleotide sequence of the gene encoding chitinase II from Aeromonas sp. no. 10S-24. J Ferment Bioeng 78: 205–211.
Vaaje-Kolstad G, Bunaes AC, Mathiesen G, Eijsink VGH . (2009). The chitinolytic system of Lactococcus lactis ssp. lactis comprises a nonprocessive chitinase and a chitin-binding protein that promotes the degradation of α- and β-chitin. FEBS J 276: 2402–2415.
Ward NL, Klota MG . (2011). Harnessing the power of microbial genomics for exploring exceptions and shifting perceptions. Front Microbiol 1: 1–3.
Watanabe T, Suzuki K, Oyanagi W, Ohnishi K, Tanaka H . (1990). Gene cloning of chitinase Al from Bacillus circulans WL-12 revealed its evolutionary relationship to Serratia chitinase and to the Type III homology units of fibronectin. J Biol Chem 265: 15659–15665.
Webb CO, Ackerly DD, Kembel S . (2011), Phylocom manual: Software for the analysis of phylogenetic community structure and character evolution..
Webb CO, Ackerly DD, Kembel SW . (2008). Phylocom: software for the analysis of phylogenetic community structure and trait evolution. Bioinformatics 24: 2098–2100.
Welch RA, Burlandt V, Plunkett G, Redford P, Roesch P, Rasko D et al (2002). Extensive mosaic revealed by the complete genome sequence of uropathogenic Escherichia coli. Proc Natl Acad Sci USA 99: 17020–17024.
West NJ, Scanlan DJ . (1999). Niche-partitioning of Prochlorococcus populations in a stratified water column in the Eastern North Atlantic Ocean. Appl Environ Microbiol 65: 2585–2591.
Wiedenbeck J, Cohan FM . (2011). Origins of bacterial diversity through horizontal genetic transfer and adaptation to new ecological niches. FEMS Microbiol Rev 35: 957–976.
Wu J-R, Shien J-H, Shieh HK, Hu C-C, Gong S-R, Chen L-Y et al (2007). Cloning of the gene and characterization of the enzymatic properties of the monomeric alkaline phosphatase (PhoX) from Pasteurella multocida strain X-73. FEMS Microbiol Lett 267: 113–120.
Yilmaz P, Gilbert JA, Knight R, Amaral-Zettler L, Karsch-Mizrachi I, Cochrane G et al (2011). The genomic standards consortium: bringing standards to life for microbial ecology. ISME J 5: 1565–1567.
Zaheer R, Morton R, Proudfoot M, Yakunin A, Finan TM . (2009). Genetic and biochemical properties of an alkaline phosphatase PhoX family protein found in many bacteria. Environ Microbiol 11: 1572–1587.
Zappa B, Rolland J, Flament D, Gueguen Y, Boudrant J, Dietrich J . (2001). Characterization of a highly thermostable alkaline phosphatase from the Euryarchaeon Pyrococcus abyssi. Appl Environ Microbiol 67: 4504–4511.
Acknowledgements
We thank three reviewers, Kevin Thornton, Kathleen Treseder and the UC Irvine Microbial Ecology group for many helpful comments on the manuscript. This work was supported in part by the National Science Foundation Dimensions of Biodiversity program and Graduate STEM Fellows in K-12 Education program.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies this paper on The ISME Journal website
Rights and permissions
About this article
Cite this article
Zimmerman, A., Martiny, A. & Allison, S. Microdiversity of extracellular enzyme genes among sequenced prokaryotic genomes. ISME J 7, 1187–1199 (2013). https://doi.org/10.1038/ismej.2012.176
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2012.176
Keywords
This article is cited by
-
Biogeochemistry of Earth before exoenzymes
Nature Geoscience (2023)
-
Root microbiota confers rice resistance to aluminium toxicity and phosphorus deficiency in acidic soils
Nature Food (2023)
-
Bioaugmentation potential of inoculum derived from anaerobic digestion feedstock for enhanced methane production using water hyacinth
World Journal of Microbiology and Biotechnology (2023)
-
Strategies and Structure Feature of the Aboveground and Belowground Microbial Community Respond to Drought in Wild Rice (Oryza longistaminata)
Rice (2021)
-
Increased microbial expression of organic nitrogen cycling genes in long-term warmed grassland soils
ISME Communications (2021)