Abstract
How reproducibly microbial populations assemble in the wild remains poorly understood. Here, we assess evidence for ecological specialization and predictability of fine-scale population structure and habitat association in coastal ocean Vibrionaceae across years. We compare Vibrionaceae lifestyles in the bacterioplankton (combinations of free-living, particle, or zooplankton associations) measured using the same sampling scheme in 2006 and 2009 to assess whether the same groups show the same environmental association year after year. This reveals complex dynamics with populations falling primarily into two categories: (i) nearly equally represented in each of the two samplings and (ii) highly skewed, often to an extent that they appear exclusive to one or the other sampling times. Importantly, populations recovered at the same abundance in both samplings occupied highly similar habitats suggesting predictable and robust environmental association while skewed abundances of some populations may be triggered by shifts in ecological conditions. The latter is supported by difference in the composition of large eukaryotic plankton between years, with samples in 2006 being dominated by copepods, and those in 2009 by diatoms. Overall, the comparison supports highly predictable population-habitat linkage but highlights the fact that complex, and often unmeasured, environmental dynamics in habitat occurrence may have strong effects on population dynamics.
Similar content being viewed by others
Introduction
Surveys of microbial diversity have consistently highlighted the vast number of genetically different microbes in environmental samples. In ocean water, for example, thousands of distinct ribotypes (cells characterized by different rRNA sequences) can co-occur, a considerable fraction of which at very low frequency (for example, Acinas et al., 2004; Sogin et al., 2006; and Rusch et al., 2007). Moreover, metagenomic comparison has revealed similar metabolic functions in phylogenetically distant microbes (Venter et al., 2004; Howard et al., 2008; Mou et al., 2008) suggesting that these might be, to a high degree, functionally interchangeable. Taken together, these results pose serious challenges for interpretation of community structure and dynamics (Polz et al., 2006). A key question is to what extent microbes are optimized for, and therefore linked to, specific ecological opportunities and to what extent they tend to act as ecological generalists. If microbes display a high degree of ecological specialization, then community assembly may be largely deterministic, and, importantly, structure–function relationships would become predictable at the level of bacterial taxonomy. This is because taxa, recognizable by some type of marker genes, would represent populations that have been competitively optimized to occupy a defined niche. Alternatively, communities may assemble from pools of generalists, strongly influenced by stochastic processes (dispersal, founder effects, bottlenecks) and allowing for frequent invasions. In this case, predictability would be much more challenging and only possible at the level of genetic diversity representing metabolic function.
Although numerous studies have established differential distribution of microbes among samples, what types of community assembly mechanisms cause observed differences in organismal composition remains poorly understood (Hughes-Martiny et al., 2006; Green et al., 2008). In fact, several recent studies have reached different conclusions. For ocean water and alpine soils, repeated sampling has revealed temporally and spatially recurrent patterns (that is, autocorrelations) of microbial taxa that were statistically significant and predictable from biotic and abiotic factors (Fuhrman et al., 2006; King et al., 2010; Gilbert et al., 2012). Such findings can be interpreted as supporting low ecological redundancy and high niche fidelity. Similar conclusions were reached by following genotypic clusters in the development of acid mine drainage biofilms (Denef et al., 2010). However, experimental addition of model organic compounds to marine samples triggered a taxonomically widespread response, which was taken as evidence for dominance of generalist taxa (Mou et al., 2008). Consistent with this observation, some modeling-based studies have de-emphasized niche-based processes (Sloan et al., 2006; Ofiteru et al., 2010). Instead, they explained community assembly by (near) neutral processes, which explain the composition of communities by stochastic birth-death and immigration processes.
One of the central questions in this debate on community assembly mechanisms is how conserved habitat specialization is for closely related genotypes. To address this question, it is necessary to distinguish microbial habitats within samples and assess the phylogenetic bounds of associated ecologically differentiated populations, both of which remain difficult. Microbial diversity is typically assessed using the conserved rRNA marker genes, for which universal cutoffs are chosen to delineate taxa, and samples are collected at scales, which comprise many different microbial habitats. As a consequence, ecological associations are measured as correlations of operational taxonomic units with macroecological features, such as nutrient concentrations and temperature. However, comparative genomics has revealed large gene content diversity even within identical ribotypes (Tettelin et al., 2008), questioning whether fine-scale ecological associations beyond combinations of macroecological features can evolve. This is because horizontal gene transfer can spread ecologically adaptive genes and alleles among otherwise unrelated genomes (Doolittle and Papke, 2006; Retchless and Lawrence, 2007), and recent analysis has shown that this process can happen on ecological time scales (Boucher et al., 2011; Smillie et al., 2011). Hence, we reason that this debate on community assembly mechanisms will benefit from a system that, akin to macroecology, allows assessment of whether ecologically coherent populations exist that reproducibly associate with physically defined habitats.
Over the past years, we have developed Vibrionaceae bacteria in the coastal ocean as a model for population biology and ecology. Our approach has been to (i) sample hundreds of individuals from fine-scale environmental fractions, (ii) establish high-resolution phylogenetic relationships by multilocus sequence analysis and (iii) test for ecological association using a mathematical model (AdaptML) (Hunt et al., 2008a; Preheim et al., 2011a, 2011b). Populations are identified as groups of related strains sharing a characteristic distribution among environmental samples; these distributions are referred to as ‘projected habitats’ (or ‘habitats’ for short) based on the rationale that they allow differentiation of the true habitats/niches if these are differentially apportioned among samples (Hunt et al., 2008a). To date, we have shown that bacteria of the family Vibrionaceae partition resources in the coastal ocean by differential distribution among the free-living and associated (with suspended organic particles and zooplankton) fractions of bacterioplankton (Hunt et al., 2008a; Preheim et al., 2011a, 2011b). Sampling during different seasons has revealed strong temporal differentiation with the same ‘habitat’ type often occupied by season-specific populations (Hunt et al., 2008a; Preheim et al., 2011a). Moreover, two studies carried out 1 year apart but targeting different types of samples from coastal water have suggested population re-occurrence (Preheim et al., 2011b); however, the evidence remains indirect since no sampling scheme has been reproduced in a manner to ascertain to what extent populations occupy the same ‘habitats’.
Here, we ask to what extent fine-scale population structure and habitat association of Vibrionaceae in coastal ocean water is reproduced in the same season (and presumably overall similar ecological conditions) in different years, as evidence for ecological specialization and predictability. In 2009, we repeated a sampling scheme first carried out in 2006 to differentiate Vibrionaceae lifestyles in the bacterioplankton (combinations of free-living, particle, or zooplankton associations) (Hunt et al., 2008a). Overall, the comparison supports highly predictable population-habitat linkage but highlights that often unmeasured, fine-scale temporal and spatial dynamics, such as shifts in eukaryotic plankton, may have strong effects on population dynamics.
Materials and methods
Environmental sampling and strain isolation
To compare the population structure of Vibrionaceae, we repeated in the fall of 2009 (October 2 and 6) a comprehensive survey carried out 3 years earlier (6 September 2006) (Hunt et al., 2008a). In both years, water samples were collected at high tide from the mouth of the Plum Island Estuary, Ipswich, MA, USA, and thus represent coastal ocean rather than estuarine water. Water temperature was 16 °C in 2006 and 13.5 °C on both days in 2009.
To differentiate free-living and particle/organism-associated bacterial lifestyles, we isolated strains from four size fractions of sequentially filtered seawater. The largest size fraction (⩾63 μm) captures bacterial cells primarily associated with larger phytoplankton and zooplankton; however, this size fraction can also contain some detrital particles. The fraction between 63 and 5 μm is enriched in different types of organic particles and smaller phytoplankton and protozoa (for simplicity hereafter ‘large particle fraction’ since bacterial attachment to these organism is expected to be minor compared with organic particles), while the fraction between 5 and 1 μm may contain large bacterial cells or those attached to small particles (‘small particle fraction’); however, this fraction also provides a firm buffer between obviously attached and free-living cells, which were obtained from the 1- to 0.2-μm fraction.
For the largest size fraction, 100 l samples were passed through a 63-μm mesh, in two and four replicates in 2006 and 2009, respectively. The mesh was rinsed with sterile seawater and the contents washed into 50 ml conical tubes. All other size fractions were derived from 63 μm pre-filtered water, collected in four replicate 4 l Nalgene bottles. These were transported in a cooler to the laboratory where further processing commenced within an hour of collection.
In the laboratory, all ⩾63 μm samples were homogenized using a tissue grinder (VWR Scientific, Radnor, PA, USA), vortexed for 20 min at low speed. The homogenates were diluted 10-fold to 10 000-fold and cells concentrated onto 0.2 μm pore size filters. The <63-μm water samples were sequentially filtered through 5, 1 and 0.2 μm pore size filters, where the 63–5 and 5–1 μm size fractions were collected using gravity filtration to avoid breakdown of fragile particles. For these, filtration was repeated with sterile seawater to further remove cells unattached to particles. Subsequently, all filters were placed into 50 ml conical tubes containing 45 ml sterile seawater and vortexed for 20 min at low speed to break up particles and resuspend bacterial cells. Supernatants were used for isolation of Vibrionaceae by concentrating serial dilutions onto 0.2 μm Supor-200 filters (Pall, Prot Washington, NY, USA) using gentle vacuum pressure. These filters were then placed onto Agar plates containing Vibrio selective Thiosulfate Citrate Bile Salts Sucrose media (BD Difco, Franklin Lakes, NJ, USA) with 2% NaCl (marine TCBS) and incubated at room temperature for 1–3 days. Single colonies were picked and re-streaked three times alternating Tryptic Soy Broth (TSB) (BD Bacto, Franklin Lakes, NJ, USA) with 2% NaCl and marine TCBS media to obtain pure strains.
Isolate characterization by gene sequencing
For preparation of DNA for gene sequencing, isolates were grown in Marine Broth 2216 (BD Difco) for 2–3 days at room temperature with shaking. DNA was released from liquid cultures with Lyse-N-Go (Thermo Fisher Scientific, Rockford, IL, USA). A 600-bp fragment of the hsp60 genes was amplified using H279 and H280 primers (Goh et al., 1996) using the following PCR conditions: 94 °C for 3 min; 30 cycles of 94 °C for 1 min, 37 °C for 1 min, 72 °C for 1 min; 72 °C for 5 min. All sequencing was by the Sanger method at the Josephine Bay Paul Center of the Marine Biological Laboratory in Woods Hole, MA, USA. Raw sequences were trimmed and verified using Sequencher (Gene Codes Corp., Ann Arbor, MI, USA). Sequences (541 bp in length) were aligned with ClustalW (Jeanmougin et al., 1998). The alignment was manually curated using MacClade (Sinauer Associates, Sunderland, MA, USA).
Hsp60 sequences were assigned to populations identified in 2006 based on multilocus sequence analysis as described in Preheim et al. (2011b) and Supplementary Table 1. Taxonomic placement of unassigned populations was done by phylogenetic analysis of 16S rRNA gene sequences. Genes were PCR amplified from representative strains using the 27f and 1492r primers (Lane, 1991). Sequencing was performed with the same primer pair at the Bay Paul Center at the Marine Biological Laboratory. A sequence similarity search was performed against type strain sequences in the Ribosomal Database Project (RDP) employing the Seqmatch tool (Cole et al., 2009). All sequences were aligned with CLUSTAL W (Jeanmougin et al., 1998) and non-overlapping sequences concatenated (final length ∼1200 bp). The alignment was manually refined and served as input for calculation of a maximum likelihood tree using MEGA 5 (Tamura et al., 2011). Percentages of sequence similarity shared with the nearest relatives on the tree were calculated by the EMBOSS Needle tool (available at http://www.ebi.ac.uk/Tools/) and are given in Supplementary Table 1.
Phylogenetic analysis
A maximum likelihood gene tree for all hsp60 gene sequences was constructed using PhyML v.2.4.5 (Guindon and Gascuel, 2003) with the following parameter settings: a likelihood ratio test supported the use of the GTR substitution model (Preheim et al., 2011a); PhyML estimated the transition/transversion ratio and the proportion of invariable nucleotide sites; four gamma categories were used; a BIONJ tree served as starting tree; tree topology, branch lengths and rate parameters were optimized by PhyML. Strains with identical hsp60 loci and isolated from the same sample were included only once in the phylogeny to prevent AdaptML (see below) from identifying clonal expansions as likely habitats.
Population prediction and identification of ‘projected habitat’
We applied an empirical model, AdaptML (Hunt et al., 2008a), to estimate population structure of Vibrionaceae. AdaptML is a maximum likelihood method that employs a hidden Markov model to learn ‘projected habitats’ (distribution patterns among environmental categories) and ecologically cohesive ‘populations’ (groups of related strains sharing the same projected habitat). In this case, the ecological data were the size fractions from which the strains were isolated and phylogenetic information was provided by the hsp60 gene tree with E. coli as an outgroup. A squared distance threshold of 0.025 was used to merge habitats with similar environmental distribution patterns during AdaptML’s iterative model fitting step. Although AdaptML predicts population membership for each isolate, we restricted our analysis to populations that (i) passed a post hoc empirical significance threshold (P-value<0.01) (Hunt et al., 2008a) and (ii) consisted of at least 20 isolates The analysis was re-run 100 times with the same parameters to verify habitat predictions were robust. Model predictions and ecological data for strains were visualized with iTOL (Letunic and Bork, 2007; Figure 1a).
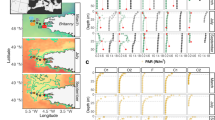
Phylogeny and ecological associations of Vibrionaceae populations inferred by AdaptML. (a) Maximum likelihood tree based on partial sequences of hsp60 genes. Inner and outer rings show the size fraction and year of isolation, respectively. Populations, which contained at least 10 strains in one of the years and passed a post hoc empirical significance threshold (P-value<0.01), are shaded where ochre=nearly equal among years and cross-hatched black and blue=skewed toward 2006 and 2009, respectively. Projected habitats are displayed by colored circles whose colors reflect trends in distributions among the different size fractions. Taxonomic assignment to numbered populations are as follows: #1, Vibrio aestuarianus; #2, V. ordalii; #3, Enterovibrio calviensis; #4, Enterovibrio norvegicus; #5, V. breoganii; #6, V. crassostreae; #7, Vibrio sp. F10; #8, Vibrio sp. F12; #9, V. splendidus; #10, V. kanaloae; #11 and #12, V. tasmaniensis; #13, V. gigantis; #14, V. cyclotrophicus. (b) Ultrametric tree summarizing the distribution of each population among size fractions. (c) Characteristic distribution patterns over size fractions (‘projected habitats’) inferred by AdaptML.
For visualization of the distribution among size fractions of all significant populations in Figure 1b, one representative strain was selected from each population to create an ultrametric tree using the Analysis of Phylogenetics and Evolution (Paradis et al., 2004) package in R programming language (http://www.r-project.org/). Because slightly different numbers of strains were obtained from the different size fractions, the distributions were normalized to the average number of strains per size fraction (349) (Figure 1b).
We did not provide sampling year information to AdaptML, but we manually curated the phylogenetic results where this temporal information was clearly relevant. Two deep-branching clusters were grouped into a single population by AdaptML, but were treated separately (populations #1 and #2) because of their distinct phylogeny and the fact that #1 was overlapping between years, whereas #2 was observed in 2006 only. We also note that, previously, populations #6, #9, #11 and #14 were listed as members of V. splendidus (Hunt et al., 2008a); however, since then taxonomic revision of populations predicted in our studies has shown that only population #9 is congruent with V. splendidus (though we note that there is disagreement among taxonomists; Le Roux et al., 2009; Preheim et al., 2011b; Supplementary Table 1).
Similarity of ecological associations
In addition to AdaptML analysis, we tested whether bacterial populations reproducibly associated with the same size fractions using Fisher’s exact test. We evaluated the null hypothesis that strains associated with congruent populations from 2006 and 2009 possess the same distribution patterns among environmental categories in the 2 years. Only populations that included at least 10 strains in each sampling were compared, and their distributions were normalized to the average size of shared populations per size fraction in each year (56 in 2006 and 94.5 in 2009). A 2 × 4 (years × habitats) contingency table was created for each population based on the normalized data. Fisher’s exact test was computed in the R programming package (http://www.r-project.org/). The criterion for significance was set to P<0.05.
Population-specific genes for V. cyclotrophicus
In the previous analysis of population structure of planktonic Vibrionaceae, two nascent populations of V. cyclotrophicus were identified based on their distinct ecological associations (group II.A and II.B of population #15 in Hunt et al., 2008a). A set of five loci was shown to be present in sub-population II.A but absent in II.B: MSHA biogenesis proteins mshN and mshF (flex1 and flex2); a probable maltose O-acetyltransferase (flex3); a probable glycosyltransferase (flex4); and a putative intercellular adhesion protein (flex5) (Shapiro et al., 2012). Representatives of the two V. cyclotrophicus sub-populations recovered in this study were tested for the presence and absence of these five genes, using diagnostic PCR tests with specific primers according to Shapiro et al. (2012) (Table 1). Phusion High-Fidelity DNA Polymerase (New England Biolobs, Ipswich, MA, USA) was applied with the following PCR conditions: 95 °C for 3 min; 30 cycles of 95 °C for 30 s, 50 °C for 30 s, 72 °C for 15 s.
Analysis of eukaryotic plankton
The composition of larger eukaryote plankton was analyzed by PCR amplification, cloning and sequencing of the mitochondrial cytochrome c oxidase subunit I gene (cox1). DNA was extracted using Puregene DNA Isolation Kit (Qiagen, Valencia, CA, USA) from 0.2 μm filters, which were obtained when concentrating the ⩾63 μm size seawater fractions of 2006 and 2009. The filters were kept frozen until the time of DNA extraction. Concentrations of DNA were measured using a NanoDrop spectrophotometer (Nanodrop Technologies, Wilmington, DE, USA). Equal amounts of template DNA from each replicate sample in each year were pooled. An ∼710-bp fragment of cox1 gene was amplified with universal eukaryote primers LCO1490 and HCO2198 (Folmer et al., 1994) using Phusion High-Fidelity DNA Polymerase (New England Biolabs). PCR was performed with the following parameters: 95 °C for 3 min, and 30 cycles of 95 °C for 30 s, 37 °C for 30 s, 72 °C for 15 s. Single DNA bands were excised from 1.5% agarose gels and were extracted with NucleoSpin Extract II Kit (Macherey-Nagel, Bethlehem, PA, USA). Purified PCR products were ligated into the pJET1.2/blunt cloning vector using CloneJET PCR Cloning Kit (Thermo Scientific, Rockford, IL, USA) according to manufacturer’s protocol. Ligation products were transformed into E. coli competent cells, which were then plated on LB-ampicillin media and were grown overnight at 37 °C. Single colonies were randomly picked into LB-ampicillin liquid broth and were grown with shaking overnight. Liquid cultures were heat shocked (95 °C for 10 min) to release template DNA. The cox1 gene inserts were reamplified employing vector-specific primers. Sequencing was performed with LCO1490 and HCO2198 primers at the Bay Paul Center at the Marine Biological Laboratory. Roughly 100 gene sequences were analyzed from each year. Taxonomic identification was according to data in the NCBI nr database (www.ncbi.nlm.nih.gov) using the BLAST algorithm (Altschul et al., 1990). A maximum likelihood gene tree was constructed as described for the phylogenetic analysis of Vibrionaceae strains.
Results and discussion
The combined analysis of 1396 Vibrionaceae isolates from samples obtained in the fall of 2006 and 2009 resulted in 14 significant populations (Figures 1a and b), whose environmental associations can be summarized as four distinct characteristic distributions (‘projected habitats’ and thereafter ‘habitat’ for simplicity) (Figure 1c). Importantly, most populations are represented in both 2006 and 2009. The five populations occupying habitat HA are predicted to be predominantly free-living in the water column: Vibrio aesturianius (#1), V. ordalii (#2), Enterovibrio calviensis-like (#3), V. tasmaniensis (#12) and V. gigantis (#13) are all highly enriched in the smallest size fraction. All other ‘habitats’ show a large portion of strains associated with different size classes of particles (Figure 1c). Habitat HB appears nearly equally distributed in all size fractions except the largest (⩾63 μm), zooplankton (and larger phytoplankton) enriched fraction. The three populations (E. norvegicus, #4; V. splendidus, #9; V. tasmaniensis, #11) occupying habitat HB may therefore be free-living or associated with organic particles and/or phytoplankton that pass through 63 μm filters. Whether these populations are interpreted as predominantly free-living or particle-attached depends on whether the bacterial cells isolated from the 1–5 μm fraction are actively growing (and therefore large) or attached to very small particles. On the other hand, lifestyles that unambiguously include attached lifestyles are displayed by habitats HC and HD, which are predominantly associated with large organic particles or zooplankton and phytoplankton. For example, for population Vibrio sp. F-10 (#7), which has previously been shown (by isolation from handpicked specimens) to be predominantly associated with living zooplankton (Preheim et al., 2011a), the analysis confirms strong bias toward the largest fraction. Although populations summarized by these two habitats are clearly attached, the observed relative frequencies among size fractions differ substantially among populations (Figure 1b). This indicates that these populations colonize different types of organisms and particles (with different distributions among the size fractions). Overall, the similarity in predicted habitats for the 2006 and 2009 samples give confidence in similar ecology of these Vibrionaceae populations.
The 14 populations fall mainly into two categories: relative abundance nearly equal between 2006 and 2009 samples, or skewed toward either 2006 or 2009. Five populations were in the first category while in the second, two populations (V. breoganii, #5 and V. cyclotrophicus, #14) had moderately differing (3.1- to 6.4-fold) but high abundances in both samplings and the remaining seven were nearly absent from either one of the samplings (that is, <10 isolates) (Figures 1a and 2). By this combined analysis of the data, 11 and 10 populations were sufficiently abundant to be considered represented in 2006 and 2009 samples, respectively; however, the overall number of predicted populations is an underestimate of the actual number because of our restriction to populations with >10 members in at least one of the years.
Distributions patterns among size fractions of populations recovered in both 2006 and 2009. Population numbers and color legend as in Figure 1.
The most striking feature of the populations appearing at similar abundance in both years is that their environmental associations also appear highly similar. Inspection of the relative habitat frequency distribution of isolates of V. aesturianius (#1), V. crassostreae (#6), Vibrio sp. F-10 (#7), V. splendidus (#9) and V. tasmaniensis (#11) suggests matching association with the different size fractions in both years (Figure 2). This visual impression is confirmed by a Fisher’s exact test with the null hypothesis that the relative proportion of populations among size fractions is independent of sample year. This null hypothesis cannot be rejected for all populations shared at similar abundances and is hence consistent with the visual impression of indistinguishable distributions (Table 2). Although V. breoganii (#5), one of the two populations with moderately skewed abundances showed significantly different distribution among size fractions in the two years (Table 2), its ecology may nonetheless be similar. A moderate association with the 1–5 μm fraction was not seen in 2006, but in both years the plankton-enriched and large particle fractions were dominant, and the particle-associated fraction reaches nearly equal representation if the 1–5 μm fraction is interpreted as cells attached to small particles (Figure 2). Regardless, V. breoganii pursued a predominantly attached lifestyle in both years so that overall, six of the seven shared populations show robust and predictable ‘habitat’ associations.
V. cyclotrophicus (#14) was the only population present in both years with qualitatively different distribution among the size fractions between years (Figures 1 and 2). In 2006, it occupied primarily the largest size fraction, indicating a possible association with larger organisms such as zooplankton (Hunt et al., 2008a). In contrast, the population recovered in 2009 behaved like a generalist, occupying all size fractions with near equal frequency (Figure 2). Because >6-fold more strains were recovered in 2009, V. cyclotrophicus may have responded to a shift in ecological conditions.
To shed more light on these differences in V. cyclotrophicus occurrence, we determined whether there was a concurrent shift in the large, eukaryotic plankton community between years. We amplified, sequenced and taxonomically binned the mitochondrial marker gene cox1 and compared the relative frequency of different sequence types in the largest size fraction. This showed that the 2006 sequences were exclusively composed of the calanoid copepod Acartia tonsa while in 2009 only few such sequences were recovered (Figure 3). Instead, the samples were dominated by a centric diatom related at ∼90% nucleotide identity to the large (80–130 μm) species Ditylum brightwelli. Hence, the habitat shift and population expansion of V. cyclotrophicus coincided with a shift in relative composition of eukaryotic plankton.
Based on these observations, we hypothesize that V. cyclotrophicus might undergo a life cycle of attachment to larger organisms during less favorable conditions and then detaching to engage in active growth in the water column in response to increased carbon availability during algal (for example, diatom) blooms. In support, we have recently observed that V. cyclotrophicus isolates are unusual among Vibrio populations in that they are chemotactic toward diatom exudates (Chien, Ahmed, Stocker and Polz, unpublished). Unicellular algae can excrete large amounts of photosynthate (Bertilsson and Jones, 2003) creating a microzone of enriched carbon substrates around the cell. Chemotaxis toward this ‘phycosphere’ (Bell and Mitchell, 1972) may enable bacteria to take advantage of nutrient enrichment around algal cells (Stocker et al., 2008). Recent analysis of monthly samples over a 6-year observation period in the English channel have revealed a very large bloom event of an unidentified Vibrio sp. following a diatom bloom (Gilbert et al., 2012; Larsen et al., 2012). Moreover, V. cholerae, which is believed to be primarily attached to zooplankton (Huq et al., 1990), was also shown to respond to algal blooms by active growth in the free-living state (Worden et al., 2006). Additionally, V. cyclotrophicus may use algal exudates as a cue to hone in on diatoms for attachment, which might be mediated by chitin contained in the diatom cell wall (Durkin et al., 2009). Vibrios, in general, possess the ability to metabolize chitin (Hunt et al., 2008b), and V. cyclotrophicus, in particular, encodes in its flexible genome an MSHA pilus (Shapiro et al., 2012), previously implicated in attachment to chitinous surfaces in V. cholerae (Watnick et al., 1999). Although this pilus might also suggest an association with zooplankton (such as copepods), we note that members of V. cyclotrophicus were nearly absent on hand-picked copepod specimens in a recent study (Preheim et al., 2011b). It will therefore be interesting to explore whether these bacteria discriminate among different types of zooplankton and phytoplankton for attachment.
A further line of evidence for high predictability of population occurrence is that two very recently diverged sister populations of V. cyclotrophicus were also recovered as ecologically distinct populations in this study. This population split was originally described in Hunt et al. (2008a) as group II.A (large particle associated) and II.B (small particle) of population #15 and was recently characterized by sequencing of 20 genomes (13 and 7 isolates, respectively) to reconstruct microevolutionary events accompanying ecological specialization (Shapiro et al., 2012). This showed that the populations remain so closely related that they are indistinguishable in 16S rRNA loci and share >99% average nucleotide identity across the core genome making them one of the most closely related but ecologically differentiated groups of bacteria yet reported (Shapiro et al., 2012). Here, we also detected the sister population II.B of V. cyclotrophicus II.A (population #14 in Figure 1) in initial AdaptML runs with a habitat prediction for population II.B matching the one originally reported (Hunt et al., 2008a). However, because our procedure for conservative estimation of habitat associations includes removal of isolates with identical hsp60 gene sequences to avoid prediction of association by chance sampling of clonal expansions, the size of population II.B dropped below AdaptML’s threshold for population prediction. To nonetheless test whether the two sister populations of V. cyclotrophicus detected in 2006 also occurred in the same habitats in 2009, we developed a PCR assay relying on 5 genes that were shown by genomic comparison among 20 isolates from the 2006 samples to be highly enriched in the flexible genome of population II.A but nearly absent in population II.B. The results confirm a similar pattern in the populations recovered in 2009 with the genes nearly universally present and absent in the corresponding populations II.A and II.B, respectively (Table 3). This therefore suggests that even for these extremely closely related groups of strains, ecological differentiation is reproducible and that habitats are occupied by genetically similar populations.
For the seven populations whose abundance was so skewed that they were nearly exclusive to either one of the samples, the dynamics remain elusive. This is because sampling of relatively few isolates from microbial communities containing high diversity of lognormally distributed taxa, can add a stochastic element to observations of population structure. It is, however, noteworthy that four of these skewed populations were assigned a free-living ‘habitat’ (Figure 1). They may thus have, as suggested for V. cyclotrophicus, more than one alternate habitat and bloom by expanding into the free-living state under certain conditions. Whether this was due to shifts in eukaryotic plankton or other water parameters remains unknown. Indeed, chemical parameters differed considerably among samples and the water temperature was 16 °C in 2006 and 13.5 °C on both days in 2009 (Supplementary Table 2), and Vibrio abundance can be strongly correlated with temperature (Heidelberg et al., 2002; Randa et al., 2004; Thompson et al., 2004; Hsieh et al., 2008).
Overall, our results show that when populations are recovered at similar relative abundances in consecutive years, the habitat predictions are robust and reproducible, suggesting that similar stages of defined population cycles or lifestyles were sampled (for example, bloom versus attachment to a specific host). On the other hand, a considerable fraction of populations was at low frequency or undetectable in one of the samples. This may indicate that these populations occasionally bloom, but are otherwise rare as has been shown for other bacterioplankton populations, which can experience population expansions on relatively short timescales (Rehnstam et al., 1993; Rieman et al., 2000; Piccini et al., 2006; Gilbert et al., 2012). Such blooms of specific populations while the remainder of the community remains constant are likely triggered by specific physical, chemical and/or biotic conditions and are thus consistent with fine-scale structuring and predictable association with what we here term ‘habitats’ (that is, combinations of specific parameters). Alternatively, observation of sporadic population occurrence may simply be due to the stochastic nature of sampling but may also be driven by dispersal rather than specific growth within the sampled environment. The latter conclusion was recently reached for vibrios associated with different body parts (respiratory and gastrointestinal tract) of mussels and crabs (Preheim et al., 2011a). Notably, many of the same populations that appear specifically associated in the water column assemble neutrally in these animals due to feeding dynamics. To clarify the relative importance of different factors in population assembly, more frequent temporal sampling will be needed to determine how finely structured and predictable biological interdependencies might be.
The considerations above highlight the importance of sampling at relevant environmental scales as has also been pointed out for analysis of plant communities where both phylogenetic and spatial scale of observation can influence the interpretation of community assembly (Cavender-Bares et al., 2009). At fine-graded spatial and phylogenetic scales, communities appear assembled according to niche partitioning while decreasing resolution changes interpretation of community assembly mechanisms first to neutrality and finally to habitat filtering (Cavender-Bares et al., 2009). The latter predicts that communities contain more closely related species than expected by chance (phylogenetic clustering) because their close relationship ensures that they share many traits that might enhance their survival in a given environment (Horner-Devine and Bohannan, 2006). These considerations are relevant for microbial communities since they are typically sampled at the ‘bucket’ scale, corresponding more to ecosystem rather than to habitat scales, and with phylogenetic markers (for example, 16S rRNA genes) that have relatively coarse genotypic resolution. The latter is particularly important for microbes since horizontal gene transfer can spread adaptive genes rapidly so that slowly evolving marker genes may not provide sufficient resolution to differentiate at the population level (Doolittle and Papke, 2006; Polz et al., 2006). We therefore suggest that fine spatial and, in particular for aquatic microbes, temporal scale comparative sampling will be important to clarify whether bacterioplankton generally assembles with high niche fidelity and thus according to ‘rules.’
References
Acinas SG, Klepac-Ceraj V, Hunt DE, Pharino C, Ceraj I, Distel DL, Polz MF . (2004). Fine-scale phylogenetic architecture of a complex bacterial community. Nature 430: 551–554.
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ . (1990). Basic local alignment search tool. J Mol Biol 215: 403–410.
Bell W, Mitchell R . (1972). Chemotactic and growth responses of marine bacteria to algal extracellular products. Biol Bull 143: 265–277.
Bertilsson S, Jones JG . (2003). Supply of dissolved organic matter to aquatic ecosystems: autochthonous sources. In: Findlay SEG, Sinsabaugh RL (eds). Aquatic Ecosystems: Interactivity of Dissolved Organic Matter. Academic Press: San Diego, pp 3–24.
Boucher Y, Cordero OX, Takemura A, Hunt DE, Schliep K, Bapteste E, Lopez P, Tarr CL, Polz MF . (2011). Local mobile gene pools rapidly cross species boundaries to create endemicity within global Vibrio cholerae populations. MBio 2: e00335–e00310.
Cavender-Bares J, Kozak KH, Fine PVA, Kembel SW . (2009). The merging of community ecology and phylogenetic biology. Ecol Lett 12: 693–715.
Cole JR, Wang Q, Cardenas E, Fish J, Chai B, Farris RJ, Kulam-Syed-Hohideen AS, McGarrell DM, Marsh T, Garrity GM, Tiedje JM . (2009). The Ribosomal Database Project: improved alignments and new tools for rRNA analysis. Nucleic Acids Res 37: D141–D145.
Denef VJ, Mueller RS, Banfield JF . (2010). AMD biofilms: using model communities to study microbial evolution and ecological complexity in nature. ISME J 4: 599–610.
Doolittle WF, Papke RT . (2006). Genomics and the bacterial species problem. Genome Biol 7: 116.
Durkin CA, Mock T, Armbrust EV . (2009). Chitin in diatoms and its association with the cell wall. Eukaryot Cell 8: 1038–1050.
Folmer O, Black M, Hoeh W, Lutz R, Vrijenhoek R . (1994). DNA primers for amplification of mitochondrial cytochrome c oxidase subunit I from diverse metazoan invertebrates. Mol Mar Biol Biotechnol 3: 294–299.
Fuhrman JA, Hewson I, Schwalbach MS, Steele JA, Brown JV, Naeem S . (2006). Annually reoccurring bacterial communities are predictable from ocean conditions. Proc Natl Acad Sci USA 103: 13104–13109.
Gilbert JA, Steele JA, Caporaso JG, Steinbruck L, Reeder J, Temperton B, Huse S, McHardy AC, Knight R, Joint I, Somerfield P, Fuhrman JA, Field D . (2012). Defining seasonal marine microbial community dynamics. ISME J 6: 298–308.
Goh SH, Potter S, Wood JO, Hemmingsen SM, Reynolds RP, Chow AW . (1996). HSP60 gene sequences as universal targets for microbial species identification: Studies with coagulase-negative staphylococci. J Clin Microbiol 34: 818–823.
Green JL, Bohannan BJ, Whitaker RJ . (2008). Microbial biogeography: from taxonomy to traits. Science 320: 1039–1043.
Guindon S, Gascuel O . (2003). A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol 52: 696–704.
Heidelberg JF, Heidelberg KB, Colwell RR . (2002). Seasonality of Chesapeake Bay bacterioplankton species. Appl Environ Microbiol 68: 5488–5497.
Horner-Devine MC, Bohannan BJM . (2006). Phylogenetic clustering and overdispersion in bacterial communities. Ecology 87: S100–S108.
Howard EC, Sun S, Biers FJ, Moran MA . (2008). Abundant and diverse bacteria involved in DMSP degradation in marine surface waters. Environ Microbiol 10: 397–410.
Hsieh JL, Fries JS, Noble RT . (2008). Dynamics and predictive modeling of Vibrio spp. in the Neuse River Estuary, North Carolina, USA. Environ Microbiol 10: 57–64.
Hughes-Martiny JB, Bohannan BJM, Brown JH, Colwell RK, Fuhrman JA, Green JL, Horner-Divine MC, Kane M, Krumins JA, Kuske CR, Morin PJ, Naeem S, Øvreås L, Reysenbach A-L, Smith VH, Staley JT . (2006). Microbial biogeography: putting microorganisms on the map. Nat Rev Microbiol 4: 102–112.
Hunt DE, David LD, Gevers D, Preheim SP, Alm EJ, Polz MF . (2008a). Resource partitioning and sympatric differentiation among closely related bacterioplankton. Science 320: 1081–1085.
Hunt DE, Gevers D, Vahora NM, Polz MF . (2008b). Conservation of the chitin utilization pathway in the Vibrionaceae. Appl Environ Microbiol 74: 44–51.
Huq A, Colwell RR, Rahman R, Ali A, Chowdhury MAR, Parveen S, Sack DA, Russek-Cohen E . (1990). Detection of Vibrio cholerae O1 in the aquatic environment by fluorescent-monoclonal antibodies and culture methods. Appl Environ Microbiol 56: 2370–2373.
Jeanmougin F, Thompson JD, Gouy M, Higgins DG, Gibson TJ . (1998). Multiple sequence alignment with ClustalX. Trends Biochem Sci 10: 403–405.
King AJ, Freeman KR, McCormick KF, Lynch RC, Lozupone C, Knight R, Schmidt SK . (2010). Biogeography and habitat modelling of high-alpine bacteria. Nat Commun 1: 53.
Lane DJ . (1991). 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M, (eds). Nucleic Acid Techniques in Bacterial Systematics. Wiley & Sons: Chichester, pp 115–175.
Larsen PE, Field D, Gilbert JA . (2012). Predicting bacterial community assemblages using an artificial neural network approach. Nat Methods 9: 621–625.
Le Roux F, Zouine M, Chakroun N, Binesse J, Saulnier D, Bouchier C, Zidane N, Ma L, Rusniok C, Lajus A, Buchrieser C, Médigue C, Polz MF, Mazel D . (2009). Genome sequence of Vibrio splendidus: an abundant marine species with a large genotypic diversity. Environ Microbiol 11: 1959–1970.
Letunic I, Bork P . (2007). Interactive Tree of Life (iTOL): an online tool for phylogenetic tree display and annotation. Bioinformatics 23: 127–128.
Mou X, Sun S, Edwards RA, Hodson RE, Moran MA . (2008). Bacterial carbon processing by generalist species in the coastal ocean. Nature 451: 708–711.
Ofiteru ID, Lunn M, Curtis TP, Wells GF, Criddle CS, Francis CA, Sloan WT . (2010). Combined niche and neutral effects in a microbial wastewater treatment community. Proc Natl Acad Sci US A 107: 15345–15350.
Paradis E, Claude J, Strimmer K . (2004). APE: Analyses of phylogenetics and evolution in R language. Bioinformatics 20: 289–290.
Piccini C, Conde D, Alonso C, Sommaruga R, Pernthaler J . (2006). Blooms of single bacterial species in a coastal lagoon of the southwestern Atlantic Ocean. Appl Environ Microbiol 72: 6560–6568.
Polz MF, Hunt DE, Preheim SP, Weinreich DM . (2006). Patterns and mechanisms of genetic and phenotypic differentiation in marine microbes. Philos Trans R Soc Lond B 361: 2009–2021.
Preheim SP, Boucher Y, Wildschutte H, David LA, Veneziano D, Alm EJ, Polz MF . (2011a). Metapopulation structure of Vibrionaceae among coastal marine invertebrates. Environ Microbiol 13: 265–275.
Preheim SP, Timberlake S, Polz MF . (2011b). Merging taxonomy with ecological population prediction: a case study of Vibrionaceae. Appl Environ Microbiol 77: 7195–7206.
Randa MA, Polz MF, Lim E . (2004). Effects of temperature and salinity of Vibrio vulnificus population dynamics as assessed by quantitative PCR. Appl Environ Microbiol 70: 5469–5476.
Rehnstam A-S, Bäckman S, Smith DC, Azam F, Hagström Å . (1993). Blooms of sequence-specific culturable bacteria in the sea. FEMS Microbiol Ecol 102: 161–166.
Retchless AC, Lawrence JG . (2007). Temporal fragmentation of speciation in bacteria. Science 317: 1093–1096.
Rieman L, Stewrd GF, Azam F . (2000). Dynamics of bacterial community composition and activity during a mesocosm diatom bloom. Appl Environ Microbiol 66: 578–587.
Rusch DB, Halpern AL, Sutton G, Heidelberg KB, Williamson S, Yooseph S et al. (2007). The Sorcerer II Global Ocean Sampling expedition: Northwest Atlantic through Eastern Tropical Pacific. PLoS Biol 5: 398–431.
Shapiro BJ, Friedman J, Cordero OX, Preheim SP, Timberlake SC, Szabo G, Polz MF, Alm EJ . (2012). Population genomics of early events in the ecological differentiation of bacteria. Science 336: 48–51.
Sloan WT, Lunn M, Woodcock S, Head IM, Nee S, Curtis TP . (2006). Quantifying the roles of immigration and chance in shaping prokaryote community structure. Environ Microbiol 8: 732–740.
Smillie CS, Smith MB, Friedman J, Cordero OX, David LA, Alm EJ . (2011). Ecology drives a global network of gene exchange connecting the human microbiome. Nature 480: 241–244.
Sogin ML, Morrison HG, Huber JA, Welch DM, Huse SM, Neal PR, Arrieta JM, Herndl GJ . (2006). Microbial diversity in the deep sea and the underexplored "rare biosphere". Proc Natl Acad Sci USA 103: 12115–12120.
Stocker R, Seymour JR, Samandani A, Hunt DE, Polz MF . (2008). Rapic chemotactic response enables marine bacteria to exploit ephemeral microscale nutrient patches. Proc Natl Acad Sci USA 105: 4209–4214.
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S . (2011). MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28: 2731–2739.
Tettelin H, Riley D, Cattuto C, Medini D . (2008). Comparative genomics: the bacterial pan-genome. Curr Opin Microbiol 12: 472–477.
Thompson JR, Randa MA, Marcelino LA, Tomita A, Lim EL, Polz MF . (2004). Diversity and dynamics of a North Atlantic coastal Vibrio community. Appl Environ Microbiol 70: 4103–4110.
Venter CJ, Remington K, Heidelberg JF, Halpern AL, Rusch D, Eisen JA et al. (2004). Environmental genome shotgun sequncing of the Sargasso Sea. Science 304: 66–74.
Watnick PI, Fullner KJ, Kolter R . (1999). A role for the mannose-sensitive hemagglutinin in biofilm formation by Vibrio cholerae El Tor. J Bacteriol 181: 3606–3609.
Worden AZ, Seidel M, Smriga S, Wick A, Malfatti F, Bartlett D, Azam F . (2006). Trophic regulation of V. cholerae in coastal marine waters. Environ Microbiol 8: 21–29.
Acknowledgements
This work was supported by grants from the National Science Foundation Evolutionary Ecology program (Evolutionary Processes), and the National Science Foundation and National Institutes of Health co-sponsored Woods Hole Center for Oceans and Human Health, the Moore Foundation and the Department of Energy. GS was supported by a stipend from the Rosztoczy Foundation.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Szabo, G., Preheim, S., Kauffman, K. et al. Reproducibility of Vibrionaceae population structure in coastal bacterioplankton. ISME J 7, 509–519 (2013). https://doi.org/10.1038/ismej.2012.134
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2012.134
Keywords
This article is cited by
-
Resolving the structure of phage–bacteria interactions in the context of natural diversity
Nature Communications (2022)
-
Viruses of the Nahant Collection, characterization of 251 marine Vibrionaceae viruses
Scientific Data (2018)
-
Ancestral gene acquisition as the key to virulence potential in environmental Vibrio populations
The ISME Journal (2018)
-
High resolution time series reveals cohesive but short-lived communities in coastal plankton
Nature Communications (2018)
-
Vibrio crassostreae, a benign oyster colonizer turned into a pathogen after plasmid acquisition
The ISME Journal (2017)