Abstract
Geothermal systems in Yellowstone National Park (YNP) provide an outstanding opportunity to understand the origin and evolution of metabolic processes necessary for life in extreme environments including low pH, high temperature, low oxygen and elevated concentrations of reduced iron. Previous phylogenetic studies of acidic ferric iron mats from YNP have revealed considerable diversity of uncultivated and undescribed archaea. The goal of this study was to obtain replicate de novo genome assemblies for a dominant archaeal population inhabiting acidic iron-oxide mats in YNP. Detailed analysis of conserved ribosomal and informational processing genes indicates that the replicate assemblies represent a new candidate phylum within the domain Archaea referred to here as ‘Geoarchaeota’ or ‘novel archaeal group 1 (NAG1)’. The NAG1 organisms contain pathways necessary for the catabolism of peptides and complex carbohydrates as well as a bacterial-like Form I carbon monoxide dehydrogenase complex likely used for energy conservation. Moreover, this novel population contains genes involved in the metabolism of oxygen including a Type A heme copper oxidase, a bd-type terminal oxidase and a putative oxygen-sensing protoglobin. NAG1 has a variety of unique bacterial-like cofactor biosynthesis and transport genes and a Type3-like CRISPR system. Discovery of NAG1 is critical to our understanding of microbial community structure and function in extant thermophilic iron-oxide mats of YNP, and will provide insight regarding the evolution of Archaea in early Earth environments that may have important analogs active in YNP today.
Similar content being viewed by others
Introduction
Although members of the domain Archaea were actually discovered in the late 18th century (Wolfe, 2006) and the first isolates were obtained in the early 20th century (Harrison and Kennedy, 1922; Barker, 1936), these organisms were not recognized as a separate lineage from the Bacteria and Eukarya until the groundbreaking work focused on the phylogenetics of the 16S rRNA gene (Woese et al., 1976; Woese and Fox, 1977). The newly recognized domain Archaea (Woese et al., 1990) included the methanogens and halophiles together in the phylum Euryarchaeota, and a second major phylum (Crenarchaeota) that included the first cultured thermophiles in the order Sulfolobales (Brock et al., 1972). Since then, considerable progress has resulted in the isolation and subsequent genome sequencing of numerous thermophiles and hyperthermophiles belonging to the Crenarchaeota (Wolfe, 2006).
Characterization of 16S rRNA gene sequences from uncultivated organisms across a myriad of natural environments as well as isolation of key reference organisms has expanded our knowledge of the phylogenetic diversity and evolutionary relationships among members of the domain Archaea. In fact, phylogenetic placement of deeply rooted sequences from Obsidian Pool in Yellowstone National Park (YNP) led to the proposal of a third candidate phylum in the mid-1990s, the Korarchaeota (Barns et al., 1994). Enrichment of Candidatus Korarchaeum cryptofilum and subsequent full-genome analysis provided further evidence that the Korarchaeota represent a separate phylum within the Archaea (Elkins et al., 2008). A fourth phylum (Nanoarchaeota) was described after the discovery and genome analysis of the symbiont species Candidatus Nanoarchaeum equitans (Huber et al., 2002), although placement of the Nanoarchaeota as a new phylum is still debated and is based primarily on a single genome sequence and 16S rRNA sequences (Brochier et al., 2005). More recently, the candidate phylum Thaumarchaeota was proposed (Brochier-Armanet et al., 2008), and detailed analysis of genome sequence from multiple representatives of this lineage provides convincing evidence that this group of organisms is significantly different from other recognized phyla within the Archaea (Spang et al., 2010). Specifically, phylogenetic analysis of housekeeping genes (that is, ribosomal proteins and enzymes involved in translation, replication, cell division and repair) from Nitrosopumilus maritimus, Nitrososphaera gargensis and Candidatus Cenarchaeum symbiosum provides strong evidence for the unique and distinguishing genetic features of members of this phylum. The first isolate from this group was the ammonia-oxidizing, marine organism N. maritimus (Könneke et al., 2005), and this discovery led to numerous enrichment cultures and molecular studies demonstrating the ubiquity of these organisms in various marine settings (Hatzenpichler et al., 2008; Walker et al., 2010; Muller et al., 2010; Blainey et al., 2011). Isolates and enrichment cultures from this phylum now include diverse members from marine, soil and hydrothermal systems (de la Torre et al., 2008; Tourna et al., 2011), and include organisms with novel metabolisms and/or morphologies, such as the ‘giant’ Thaumarchaeota (Muller et al., 2010). The discovery of Thaumarchaeota has resulted in major contributions to our understanding of global C and N cycling and the potential importance of CO2 fixation and nitrification in the open ocean (Ingalls et al., 2006).
High-temperature environments in YNP have yielded a wealth of knowledge regarding the extant diversity, distribution and metabolism of archaea (Barns et al., 1996; Inskeep et al., 2005; Meyer-Dombard et al., 2005; Inskeep et al., 2010). Previous studies of acidic ferric iron mats, in particular, have shown a remarkable diversity of archaea, which corresponds with the combination of unique geochemistry and high temperature of these systems (Inskeep et al., 2005; Kozubal et al., 2008; Inskeep et al., 2010; Kozubal et al., 2012). Prior 16S rRNA gene sequencing of high-temperature (65–80 °C) acidic Fe-oxide mats has revealed the presence of novel archaea, but cultivation of representative(s) from this group has yet to be successful. Here, we report for the first time a thorough analysis and description of four replicated de novo assemblies of a deeply rooted microorganism, which we propose as a representative of a candidate phylum-level lineage (Geoarchaeota) within the domain Archaea. Analysis of metagenome sequence from replicate Fe-oxide mat samples (60–78 °C) reveals a dominant archaeal population with novel strategies for energy conservation and oxygen respiration. Evidence is provided here that these organisms are members of one the earliest currently known lineages of the domain Archaea, and that members of this phylum-level group are limited primarily to thermophilic habitats, especially those containing ferrous Fe and low levels of oxygen.
Materials and methods
Sample collection and metagenome sequencing
DNA for metagenome sequencing was extracted from ∼10 g of iron-oxide microbial mat from 100 Spring Plain (Norris Geyser Basin; YNP; (Easting 523042/Northing 4953337) sampled at two different time points using a FastDNA Spin Kit (MO BIO Laboratories, Carlsbad, CA, USA). Environmental DNA was precipitated with 3.0 M sodium acetate and 100% ethanol. Sanger sequencing of 3-kb random insert pUC18 libraries was used to obtain random shotgun metagenome sequence (average read length ∼800 bp) of the first sample obtained in Fall 2007 (OSPB-1). Further replicates sampled 2.4 year after OSPB-1 (OSPB-2, OSPC and OSPD) were sequenced using 454 titanium pyrosequencing (average read length ∼380 bp; Table 1).
Metagenome sequence analysis and assembly of NAG1
Random shotgun sequence reads were assembled using Celera (JCVI) and Newbler (Roche/454, Basel, Switzerland) assemblers. Assembly statistics (coverage, quality scores and GC content) were compiled by the Joint Genome Institute (JGI; Walnut Creek, CA, USA). All aspects of library construction and sequencing performed at the JGI can be found at http://www.jgi.doe.gov/ including assembly methods, base error and possible mis-assembly corrections. Contigs were further analyzed using nucleotide word frequency (Teeling et al., 2004) principal component analysis (NWF-PCA) to separate novel archaeal group 1 (NAG1) contigs from other species (performed at http://gos.jcvi.org/openAccess/scatterPlotViewer2.html with the following parameters: word size=four, minimum contig size at 2000 bases, and the chop sequence size at 4000 bases as described previously for YNP ferric iron mat systems; Inskeep et al., 2010). Further verification of NAG1 contigs was obtained by plotting coverage vs G+C content for all metagenome contigs, which clearly verified the NAG1 sequences as separate from other populations. A group of contigs exhibiting similar sequence character in NWF-PCA space were further screened using G+C%, contig coverage and blastp identity to improve the de novo assemblies for each replicate sample. For example, the largest NAG1 scaffold (1.36 Mb) was obtained from replicate OSPB-1 and contained the 16S and 23S ribosomal gene sequences as well as the majority of ribosomal proteins for the NAG1 population. Nearly identical clustering occurred for the replicate assemblies (OSPB-2, OSPC and OSPD) resulting in 68, 104 and 52 scaffolds, respectively. Assembled sequence was compared with reference databases (blastp) and top sequence hits were assigned to their nearest phylum; however, as indicated by the phylogenetic placement of this candidate phylum, the NAG1 assemblies demonstrate a lack of identity to currently described reference genomes (>70% of the NAG1 sequence is <50% identical to known references). NAG1 assemblies were compared with four reference genomes from other related archaeal phyla including Ferroplasma acidarmanus fer1, Thermofilum pendens HRK5, N. maritimus SCM1 and Candidatus Korarchaeum cryptofilum OPF8 using NWF-PCA analysis.
Annotation of NAG1 de novo assemblies
Gene models, predicted functions and pathway/subsystem assignments derived from IMG/ER and the rapid annotation using subsystems technology were compared and used to annotate the NAG1 sequence clusters screened using NWF-PCA (Aziz et al., 2008). Cofactor utilization and central metabolic subsystems were manually curated using the SEED subsystem editor (Overbeek et al., 2005) to facilitate mapping genes to known subsystem variants and the Artemis genome browser (Rutherford et al., 2000) to identify gene fragments that corresponded to missed genes belonging to these subsystems. Individual protein sequences were further analyzed from function prediction by alignment with sequences with known biochemical functions. The replicate OSPB-1 de novo assembly is located on the Joint Genome Institute IMG/M website under the IMG submission ID number 2423.
Phylogenetic analysis
All phylogenies were constructed with the MEGA 4.0 software package (Tamura et al., 2007). A total of 20 concatenated protein phylogenetic trees were constructed around 32 full-length ribosomal proteins (Figure 3) as well as single-copy proteins RNA polymerase, DNA polymerase and transcription factors (Supplementary Figure S1). Two different alignment methods (ClustalW and Muscle) and three substitution models (Dayhoff and Jones–Taylor–Thornton, no. of differences and p-distance) were used for both the neighbor-joining and minimum evolution distance methods (Nei and Kumar, 2000). In addition, three substitution models (Poisson, Dayhoff and Jones–Taylor–Thornton) were utilized to construct three maximum likelihood trees for each of the two alignments (Jones et al., 1992; Nei and Kumar, 2000). Finally, the ClustalW and Muscle alignments were utilized to obtain corresponding maximum parsimony trees. Reliability of inferred trees was conducted with the bootstrap test of phylogeny using 1000 bootstraps. Alignments were obtained with default gap penalty parameters and were edited by manual inspection and deletion of large gaps. A consensus tree was constructed for the concatenated protein tree of 32 ribosomal proteins (Figure 3a). Maximum likelihood trees (obtained from Muscle alignments and the Jones–Taylor–Thornton substitution model) are shown for single protein trees and are similar to topologies of consensus trees.
A total of 16 nucleotide 16S/23S phylogenetic trees (Figure 3b) were constructed with two different alignment methods (ClustalW and Muscle), three substitution models (no. of differences, p-distance and maximum composite likelihood) for both neighbor-joining and minimum evolution distance methods. Maximum likelihood (Jukes–Cantor substitution model) and maximum parsimony trees were constructed for both ClustalW and Muscle alignments (other details are as described above). Reliability of inferred trees was conducted with the bootstrap test of phylogeny using 1000 bootstraps. Alignments were obtained with default gap penalty parameters and were edited by manual inspection and deletion of large gaps in the alignments. Results from all treeing methods were utilized to construct a consensus 16S/23S phylogenetic tree.
Fluorescence in situ hybridization
Approximately 10 g of dispersed OSP_B mat was incubated at 75 °C in 160 ml serum bottles for 3 days with 50 ml of OSP source spring water. An 0.5-g aliquot of culture slurry was washed with Nycodenz buffer (Bertin et al., 2011) then fixed with 4% paraformaldehyde at 4 °C for 16 h (Fuchs et al., 2007). Fixed cells were further separated from the solid ferric iron with the nonionic density gradient medium Nycodenz (60% w/v; Sigma-Aldrich D2158; St. Louis, MO, USA) and pelleted at 10 000 g for 30 min (Bertin et al., 2011). The cells were resuspended in 500 μl of 1:1 phosphate-buffered saline:ethanol. 50 μl of this suspension was fixed onto a glass microscope slide and fluorescence in situ hybridization (FISH) performed with Cy5-labeled probe (5′-GAG TTC TTA CCT ATC CGG G-3′; Integrated DNA Technologies, Coralville, IA, USA) specific for the NAG1 16S ribosomal RNA sequence. The Cy5 probe was designed around 16S rRNA class I and II sites and had no homology to 16S rRNA gene sequences of other organisms from OSP Springs or other YNP iron mats (Behrens et al., 2003). The probe sequence was further analyzed using IDT ScitTools OligoAnalyzer 3.0 (Integrated DNA Technologies) to determine possible hairpins, dimers and the melting temperature. Formamide stringency optimization was conducted at concentrations of 0%, 5%, 20% and 35% according to Fuchs et al. (2007), and highest specificity was determined to be 5% based on comparison of fluorescence signal with appropriate Cy5 excitation laser. In addition, fresh OSPB mat was stained with SYBR gold to show cell distribution in a natural sample. Cy5 fluorescence staining was not successful on natural mat samples because of high mineral content. Both Nycodenz preparations and SYBR gold-labeled mat were viewed on a Leica SP5 CSLM (Leica Microsystems GmbH; Wetzlar, Germany) inverted scanning confocal laser microscope.
Results
Metagenome assemblies
De novo ‘genome’ assemblies of a ‘novel archaeal Group 1’ population were generated from random shotgun sequence of community DNA obtained from four separate sampling points (60–75 °C) within the outflow channel of an acidic (pH 3.5) Fe-oxide geothermal spring in YNP (Figure 1a; Table 1). Two replicates were obtained from the same location and temperature (76–78 °C), but sampled over 2 years apart (OSPB-1 and OSPB-2). Additional replicates were sampled downstream at temperatures of 73 °C (OSP-C) and 60 °C (OSP-D) at the same time as OSPB-2 (Figure 1a). Contigs assigned to each replicate de novo assembly have similar G+C content (%) and sequence read coverage that clearly distinguish them from other population clusters in this community (example shown in Supplementary Figure S1). Comparison of NAG1 sequence to reference databases (via blastp or blastn) revealed a consistent pattern (that is, poor sequence similarity to current reference genomes), and is clearly different than the other three to four predominant populations present in these systems (Kozubal et al., 2012). The assemblies were evaluated using NWF-PCA (Teeling et al., 2004), which showed that the sequence content and character (that is, G+C content, codon usage) are nearly identical among the four NAG1 replicates, and that these assemblies differ significantly compared with four representative phyla within the domain Archaea, including those from the Crenarchaeota, Euryarchaeota and Thaumarchaeota (Figure 1b).
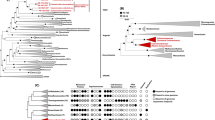
(a) One Hundred Springs Plain Spring (OSP) located within Norris Geyser Basin (YNP). Arrows indicate spring source and sampling points with corresponding temperature and dissolved oxygen values obtained along primary flow path. (b) NWF-PCA of four replicate de novo assemblies for NAG1 (circled, four shades of brown and red) and genomes from T. pendens, Candidatus Korarchaeum cryptofilum, N. maritimus and F. acidarmanus. Axes x, y, z correspond to PC1, PC2 and PC3. (c) Average G+C content (%) of NAG1 scaffolds from replicate OSPB-1 (red circles) plotted as a function of decreasing scaffold length (solid black line, cumulative total sequence length; solid red line, average G+C content; dotted line, indicates first eight scaffolds where consensus sequence is >90%).
The largest assembly of the NAG1 population (OSPB-1) was obtained using Sanger sequencing (3 kb short-insert library, pUC18 vector) and resulted in 1.7 Mb of nonredundant sequence on eight scaffolds (Figure 1c, Table 1). Blastp matches of 1932 protein-coding open reading frames identified in the OSPB-1 assembly reveal a distribution of top hits scattered across nearby phyla within the domain Archaea, as well as a significant number of protein-coding genes that have best matches to references within the domain Bacteria (15%). However, the majority (>90%) of these matches are <60% amino acid (aa) identity to reference genomes. Whole-genome nucleotide alignments of the replicate NAG1 assemblies (OSPB-2, OSPC and OSPD) obtained from 454 titanium pyrosequencing show nearly identical sequence content (>99% nucleotide sequence identity) and synteny with one another (Table 1).
The assembled NAG1 sequence from OSPB-1 (1.7 Mb) likely corresponds to a near-complete consensus genome, based on the finding that this assembly contains the expected number of archaeal RNA polymerase subunits, transcription factors, ribosomal proteins and all genes necessary for tRNA synthesis (Supplementary Table S1). Addition of NAG1-like contigs or scaffolds less than ∼5 kb in size (that is, scaffolds 9–30) did not result in significant increases in nonredundant sequence or functional information; consequently, our analysis is focused on the largest 8 scaffolds from OSPB-1 (Figure 1c) as well as a conservative group of NAG1 scaffolds from replicate assemblies (those contigs that could be clearly defined using NWF-PCA, G+C content and blastp analysis; Table 1). Replicate NAG1 assemblies obtained using titanium pyrosequencing (OSPB-2, OSPC and OSPD) did not reveal or contribute any additional sequence content. Furthermore, each replicate assembly contains one complete and identical copy of 5S, 16S and 23S rRNA genes. Definitive identification of NAG1 cells using FISH probes specific for these 16S rRNA sequences show that NAG1 cells are ∼1 μm diameter cocci and are easily located in Nycodenz cell preparations (Figure 2a). The NAG1 organisms, along with Metallosphaera yellowstonensis, are among the dominant cocci in the Fe-mat samples. Based on metagenome replicates, >80% of the archaeal sequence in OSP_B belongs to NAG1 (∼50–55%), M. yellowstonensis (12–21%) and Acidolobus spp. (10–15%) as determined by three separate phylogenetic methods. All three of these organisms are ∼1 μm diameter cocci (Boyd et al., 2007; Kozubal et al., 2008; this study). Most of the remaining sequences belong to Vulcanisaeta- or Hydrogenobaculum-like spp. (rods and filaments, respectively). SYBR gold staining shows the distribution of these cell morphologies in an undisturbed OSP_B ferric iron mat. Given the successful identification of NAG1 organisms using a FISH probe on cell separations from the same sample, it is likely that a fraction of the cocci identified using a global (SYBR gold) stain of intact iron mat are indeed NAG1 organisms. FISH probes proved to be highly problematic on iron mat samples without sample dispersion and Nycodenz preparation to remove mineral material.
Confocal microscopy of cell preparations and undisturbed Fe-oxide microbial mat. (a) NAG1-specific 16S rRNA FISH of Nycodenz cell preparations (to remove mineral phases) from OSP_B iron mat samples. Cells were identified as ∼1 μm cocci (green). (b). Undisturbed Fe-oxide mat from site OSPB stained with SYBR gold (scale bar=20 μm).
Phylogenetic analysis of NAG1: representative of a candidate phylum in the Archaea
Consensus concatenated trees of 32 ribosomal proteins and 16S/23S nucleotide sequences shared among archaeal species show consistently that the four replicate NAG1 assemblies cluster together as a new lineage between the current phylum Crenarchaeota and candidate phylum Korarchaeota (Figures 3a and b, Supplementary Figure S2). Both the concatenated ribosomal protein and 16S/23S trees are built as consensus trees using 18 and 16 phylogenetic analyses, respectively, and both approaches yield consistent phylogenetic placement of the NAG1 lineage. Several universally conserved housekeeping genes have been utilized previously for phylum-level patterning in the Archaea (Spang et al., 2010), and this same analysis of de novo NAG1 assemblies supports the conclusion that NAG1 organisms are representatives of a phylum-level lineage (Supplementary Figure S2; Supplementary Table S2). Specifically, topoisomerase and translation proteins show a distinct pattern difference in NAG1 populations compared with other Archaea. Topoisomerase IA, IB and IIA were not found in NAG1 consensus assemblies, but the genome(s) do contain a reverse gyrase most similar to Thermococcus onnurineu, a member of the Euryarchaeota. NAG1 organisms contain cdvABC genes for cell division (Bernander and Ettema, 2010), which are phylogenetically distinct and fall between the Crenarchaeota and the Thaumarchaeota. Furthermore, unlike the Crenarchaeota, the NAG1 organisms contain sequences for small and large subunits of DNA polymerase D; the small subunit represents the most deeply rooted deduced PolD sequence currently available in public databases, branching between recognized members of Archaea and Eukarya (Supplementary Figure S1). Phylogenetic analysis of other RNA polymerases places the NAG1 population(s) between the Crenarchaeota and either the Thaumarchaeota, Korarchaeota or Euryarchaeota, depending on the specific subunit used (Supplementary Figure S2). Moreover, maximum parsimony trees suggest that the NAG1 lineage is more deeply rooted than other methods and branches between the Thaumarchaeota and Euryarchaeota (Supplementary Figure S3).
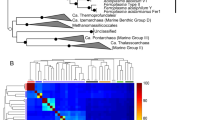
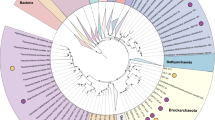
(a) Consensus concatenated phylogenetic protein tree of 32 ribosomal proteins shared by the three domains of life (Giardia intestinalis was used as the outgroup). Tree is a consensus of 18 different phylogenetic methods. Consensus bootstrap values for nodes >900/1000 are noted by solid circles and values >700/1000 are noted by hollow circles. (b) Consensus concatenated 16S/23S rRNA nucleotide tree (Hydrogenobaculum sp. Y04AAS1 was used as the outgroup). Tree is a consensus of 16 different phylogenetic methods. Consensus bootstrap values for nodes >800/1000 are noted by solid circles and values >600/1000 are noted by hollow circles.
Numerous full-length 16S rRNA gene sequences from acidic Fe-oxide mats in YNP have been characterized during the last decade, and many of these belong to the proposed candidate phylum Geoarchaeota. For example, 60% of ∼250 clones from site OSPB are highly similar to (>99.5%) and clade with the 16S rRNA gene sequences obtained from the de novo assemblies (Supplementary Figure S4). This agrees favorably with analysis of random metagenome sequence where ∼55% of the total reads belong to the NAG1 assembly (true for both OSPB-1 Sanger and OSPB-2 pyrosequencing runs). Moreover, 16S rRNA sequences from previous studies of Fe mats in YNP and other extreme environments show at least two major clades within this novel lineage (Supplementary Figure S4). Clade A is represented by over 600 clones from low-pH ferric iron mats from YNP (with the exception of one sequence from Great Boiling Springs, NV, USA) and includes the 16S rRNA sequences of the four replicate de novo assemblies from OSP Spring, which are all 100% identical. Moreover, the majority of environmental clones in clade A are >99.5% similar to the 16S rRNA sequences obtained using independent metagenome sequencing. The absence of sequence entries from other locations indicates that the collective attributes of these Fe(III)-oxide mats are unique, or that similar habitats have not been characterized elsewhere on the planet. The combination of high temperature, ferrous iron, hypoxic to oxic conditions and moderately acidic conditions appears to define the niche of these deeply rooted organisms. The second major group within this candidate phylum (clade B) is represented by ∼25 sequences from a limited number of neutral pH thermophilic systems including Obsidian Pool (YNP). These 16S rRNA sequences are only 81–85% similar (nt) to NAG1 sequences within clade A (Supplementary Figure S4).
Functional analysis of NAG1 genome sequence
Energy conservation and central C metabolism
The de novo NAG1 assemblies provide sequence data necessary for understanding metabolic attributes of these organisms in acidic high-temperature Fe-mats of YNP (Figure 4). Genes coding for chemolithotrophic pathways known to be associated with extreme environments (Inskeep et al., 2010; Kozubal et al., 2011), including the oxidation of ferrous iron, hydrogen, arsenic, sulfur, ammonium or methane, were not found in the NAG1 assemblies. However, the NAG1 populations have two separate loci that encode aerobic carbon monoxide (CO) dehydrogenases and associated maturases (Dobbek et al., 2002; King and Weber, 2007). One locus encodes a ‘Form I’ CO dehydrogenase along with coxFSM and the other contains three coxL ‘Form II’ CO dehydrogenase sequences along with coxDEFG (Supplementary Figures S5, S6). The NAG1 ‘Form I’ CO dehydrogenase contains the active binding site (amino-acid sequence=IAYRCSFR) implicated in aerobic CO oxidation and has similar gene arrangement and sequence similarity to the known CO utilizer Thermomicrobium roseum (Wu et al., 2009; 68%, 73% and 47% amino-acid identity for CoxS, CoxM and CoxL subunits, respectively). Although the ‘Form II’ CO oxidases may be involved in CO oxidation, they have been shown to have a broader substrate range (King and Weber, 2007).
Summary of NAG1 metabolic attributes based on replicate de novo assemblies. (bd, bd-type oxygen reductase; CO, aerobic CO dehydrogenase CoxSML; ED, Entner–Doudoroff; EM, Embden–Meyerhof; HCO, heme copper oxygen reductase complex IV of the respiratory chain (NAG1 has subunits I, II and III of a Type A HCO); I, NADH dehydrogenase complex I of the respiratory chain; II, succinate dehydrogenase Complex II).
The NAG1 assemblies contain a number of protein-coding genes important in the external acquisition and subsequent degradation of amino acids and peptides. For example, genes involved in branched-chain amino acid, dipeptide and oligopeptide uptake ABC transport systems, a CstA peptide transporter, aminotransferases, peptidases and proteases all support the hypothesis that NAG1 populations acquire and degrade protein as a carbon and energy source. The assemblies contain genes for a variety of related transporters that include branched chain amino acids (livFGKHM), dipeptides (dppABCDF), oligopeptides (appABCDF) and peptides (cstA; Figure 4). In addition, NAG1 assemblies contain a glycoside hydrolase (EC 3.2.1.23) for hydrolysis of glycosidic bonds as well as other genes necessary for glycogen and starch metabolism (Figure 4). It is unclear whether the polysaccharide degradation potential observed in the genome sequence of NAG1 assemblies relates to storage and utilization of glycogen and/or hydrolysis of complex carbohydrates found in the environment.
The absence of a Type 1 4-hydroxybutyryl-CoA dehydratase and bifunctional acetyl/propionyl-CoA carboxylase suggests that NAG1 populations do not fix CO2 through the 3-hydroxypropionate/4-hydroxybutyrate pathway observed in the Crenarchaeota and Thaumarchaeota (Berg et al., 2007). No other obvious modes of CO or CO2 fixation were noted in replicate NAG1 assemblies, based on genes coding for key marker enzymes of currently recognized autotrophic pathways (Berg et al., 2010). Although NAG1 has a gene coding for a Type II 4-hydroxybutyryl-CoA dehydratase, it is not known whether this protein is involved in CO2 fixation. Consequently, the organism may either obtain carbon from autotrophic organisms in the spring (for example, M. yellowstonensis, Hydrogenobaculum spp.) or from exogenous sources. The gene content of NAG1 suggests metabolism of simple carbon compounds such as peptides and fatty acids and/or degradation of more complex organic substrates. Other NAG1 genes involved in central C metabolism were annotated manually including those required for the Embden–Meyerhof pathway and gluconeogenesis. A unique feature of the NAG1 Embden–Meyerhof pathway is the presence of a glucokinase gene that is similar to an atypical member of the glucose-phosphorylating enzymes initially characterized in Sulfolobus tokodaii (Nishimasu et al., 2006). NAG1 contains all genes required for both the phosphorylating and non-phosphorylating branches of the archaeal Entner–Doudoroff pathway. In addition, NAG1 has all genes necessary for the oxidative pentose phosphate pathway, which includes an F420-dependent (FGD), glucose-6-phosphate (G6P) dehydrogenase (that may also be linked with oxidative stress—see below) and a bacterial-like 6-phospho-gluconate dehydrogenase. Furthermore, NAG1 has the archaeal-specific ribulose monophosphate pathway and components of the non-oxidative pentose phosphate pathway including a ribose-5-phosphate isomerase and a transketolase.
Respiratory complexes, oxygen dependence and oxidative stress
NAG1 sequence contains both a Type A heme copper oxidase (HCO) and a bd ubiquinol oxidase complex. The NAG1 HCO (subunit I) is distinct from bacterial Type A HCOs and forms a separate branch near the base of a subunit I protein tree (Supplementary Figure S7) between members of the Thaumarchaeota and all other archaea with Type A HCOs. In addition, the terminal oxidase complex contains three subunits and exhibits operon structure and gene content considerably similar to that found in mitochondrial terminal oxidases (along with thaumarchaeal sequences). The operon coding for the Rieske and cytochrome b components of electron transport chain complex III was found in NAG1 assemblies, but neither cytochrome C1 nor CbsA are located on this operon, and are found elsewhere in the genome. This structure is more typical of a euryarchaeotal-like complex III. Genes coding for proteins involved in anaerobic respiration were not found in NAG1 assemblies (for example, respiration on ferric Fe, arsenate, nitrate, sulfate and/or sulfur). Although NAG1 contains an alcohol dehydrogenase and certain components of the subsystem resulting in acetyl-CoA fermentation to butyrate, it does not appear to contain all the proteins necessary for mixed-acid fermentation. NAG1 contains an FGD G6P dehydrogenase, which is most similar to a functionally characterized enzyme in Mycobacterium smegmatis (Hasan et al., 2010) and is another example of specific NAG1 gene content that is more ‘bacterial-like’. G6P is believed to be a source of reducing power in M. smegmatis for reduction of reactive oxygen species with FGD having an active role in ensuring high amounts of G6P. The NAG1 FGD homolog is part of a two gene operon with a superoxide dismutase (sodA), which further suggests a role for the FGD in oxidative stress.
The NAG1 assemblies also contain a globin protein (169 aa) with a clear protoglobin loop and heme domain (pfam 11563). Moreover, the NAG1 sequence is the most deeply rooted entry of all known globins/protoglobins (Supplementary Figure S8). The protein sequence is related to characterized protoglobins in Aeropyrum pernix, which suggests possible involvement in binding O2, CO and NO (Freitas et al., 2004). Other genes encoding proteins for heavy metal detoxification and reduction of oxidative stress are found in the NAG1 assembly and include mercuric reductase (merA), arsenic resistance (arsABC), a superoxide dismutase (sodA) and multiple peroxiredoxins. However, the ars genes were found distributed throughout the genome(s) and are not part of a single operon.
Cofactor biosynthesis
The NAG1 population has necessary pathways for synthesis of thioredoxin, pyridoxine, riboflavin, flavine adenine dinucleotide, cobalamin, siroheme, lipoic acid and salvage of cobalamin and niacin (Figure 4), several of which differ from those commonly found in archaea. Specifically, NAG1 appears to synthesize thiozole from glycine rather than tyrosine and has thiO rather than thiH, which would make NAG1 the only known archaeon with this ‘bacterial-like’ pathway. NAG1 also has a ‘bacterial-like’ SUF biosynthesis system as the sole pathway for FeS synthesis, including sufA and sufS, which have been found exclusively in bacteria. NAG1 contains all genes necessary for the biosynthesis of coenzyme F420 and has 10 putative FGD enzymes in addition to the (G6P) dehydrogenase mentioned above including four luciferase monooxygenase-like proteins, four FGD N(5),N(10)-methylenetetrahydro-methanopterin reductases, a F420 hydrogenase/dehydrogenase beta subunit and an nicotinamide adenine dinucleotide phosphate oxidase-dependent F420 reductase.
NAG1 does not contain any biotin synthesis or salvage genes, and contains no genes coding for enzymes that require this cofactor. Biotin is considered an essential vitamin throughout the domains Bacteria and Eukarya as it is utilized as a carboxyl carrier in pathways that mediate fatty acid synthesis, branched-chain amino-acid catabolism and gluconeogenesis. Moreover, the Sulfolobales, some Desulfurococcales and possibly one member of the Thermoproteales (T. pendens) utilize biotin-dependent carboxylases for the 3-hydroxypropionate/4-hydroxybutyrate CO2 fixation pathway, but these enzymes are not required for central carbon metabolism and lipid biosynthesis (Berg et al., 2010). Interestingly, genes encoding biotin-dependent enzymes are not found in the genomes of other archaea including the Korarchaeota, Thermoplasmatales and Acidilobus spp. Consequently, including the candidate phylum Geoarchaeota, representatives of four archaeal phyla do not contain biotin-dependent enzymes.
Heme is predicted to be required by NAG1 organisms (for example, subunit I HCOs), but identifiable heme uptake genes were not identified. However, NAG1 ‘genomes’ have the early hemALBCD heme synthesis genes found in most archaea (missing in the Thermococcales, Nanoarchaeota, Korarchaeota and some Crenarchaeota; Storbeck et al., 2010). Interestingly, NAG1 also contains hemEH (only found in the bacteria), but is missing hemFG necessary for a complete bacterial-like heme synthesis pathway. Furthermore, genes for a proposed alternative archaeal heme synthesis pathway (SUMT, PC2-DH, nirD, nirH and nirJ12) are not found in the NAG1 assemblies suggesting heme biosynthesis via an unknown alternative pathway (Storbeck et al., 2010).
Evidence for viral defense
The NAG1 population from OSPB-1 contains a unique CRISPR (Clustered Regularly Interspaced Short Palindromic Repeats) region with several putative CRISPR protein-coding genes and a CRISPR array containing 19 direct repeats (DR=5′-CTTAAACTCAGAAGAGGATTGAAAG)-3′ with associated spacer regions (1217 total nucleotides). The putative Csm3-like (RAMP) proteins and a 896 amino acid hypothetical Cas10 polymerase are found upstream of the CRISPR array with similar gene arrangement to T. pendens, Metallosphaera sedula and S. tokodaii (Supplementary Figure S9). However, unlike these organisms, homologs to Cas1, Cas2 and Cas4 proteins were not found in NAG1 assemblies. Additional hypothetical CRISPR-associated genes were found on a separate locus of NAG1 including Cas6, a Cas3 helix DNA-binding region, a herA helicase and a nuclease. Although the herA helicase and nuclease are not similar to any known cas sequences, they may have similar function to Cas3 proteins. The CRISPR-associated proteins in NAG1 suggest a Type III-polymerase system without Cas1 and Cas2 (Makarova et al., 2011).
Definitive CRISPR/Cas regions were not found in the replicate assemblies from OSPB-2, OSPC and OSPD, sampled 2.4 year after OSPB-1. Although four homologous sequences to the OSPB-1 direct repeats were found in the replicate assemblies (OSPB-2, OSPC and OSPD) obtained using pyro-sequencing, they were not identified as complete CRISPRs, and this may be due to poorer assembly of shorter read lengths from pyro-sequencing. Further studies may clarify the temporal nature and identification of CRISPR direct repeats, variable spacer regions and/or other CRISPR/Cas proteins. The presence of a unique CRISPR/Cas region within the largest NAG1 assembly (OSPB-1) is strong evidence that this population is a host to viruses not yet characterized. Moreover, variable spacer regions found within the NAG1 assemblies did not match any gene content from publically available viral sequences or gene content found within metagenome assemblies.
Discussion
Here, we describe for the first time de novo assemblies of a novel archaea, which we propose belongs in a candidate phylum Geoarchaeota (from ‘Geo’, Gr., meaning ‘Earth’). The large majority of currently described 16S rRNA gene sequences from this lineage were obtained from acidic ferric Fe mats in YNP geothermal springs. These moderately acidic, high-temperature, high-ferrous Fe and hypoxic to oxic conditions appear to define the niche of NAG1 organisms. The NAG1 population is one of four to five predominant community members and represents ∼20–55% of the total sequence reads in OSP Spring, depending on the exact sample location and temperature (60–78 °C). Other community members present in the acidic Fe mats of OSP spring include organisms most closely related to M. yellowstonensis Acidilobus spp., Vulcanisaeta spp. and Hydrogenobaculum spp. (Aquificales), as well as other less abundant, uncharacterized archaea and bacteria (Kozubal et al., 2012).
Phylogenetic analysis (Figure 3; Supplementary Figure S2) and phylum-level patterning of specific marker genes (Supplementary Table S2; Spang et al., 2010) provide strong evidence that the NAG1 population is a representative of a new phylum, with phylogenetic placement between the Euryarchaeota, Crenarchaeota and Thaumarchaeota as well as the Eukarya. However, given that the de novo assemblies of NAG1 may not constitute a complete genome, missing genes may be located in gaps between contigs, although this is unlikely in cases where genes are missing from conserved operons. Additional examples of genes that are not shared with the Crenarchaeota include the prevalence of coenzyme FGD enzymes. The assembly proteins (cofHG) for coenzyme F-420 are not found in any crenarchaeal genomes, but are prevalent in methanogens and thaumarchaea.
The discovery of the NAG1 lineage contributes to our understanding of the evolution of archaea and their metabolic interactions with oxygen. Specifically, the NAG1 contains the most deeply rooted protoglobin known to date (Supplementary Figure S8), and appears to provide an evolutionary linkage to higher globins such as hemoglobin that are thought to have evolved from an ancestral protoglobin (Moens et al., 1996; Freitas et al., 2005; Nardini et al., 2008). Protoglobins from the obligately aerobic hyperthermophile A. pernix and the strictly anaerobic methanogen Methanosarcina acetivorans have been found to bind molecular oxygen, nitric oxide and CO, potentially protecting the organisms from nitrosative and oxidative stress (Freitas et al., 2004). Consequently, protoglobins have been found in both aerobic and anaerobic organisms. Possible functions of the NAG1 protoglobin include CO transfer to the CoxSML complex, a sensor for CO in the environment, or O2 transfer to a terminal oxidase in low O2 environments. The NAG1 organisms also contain two terminal oxidase complexes including an advanced Type A HCO and a bd-ubiquinol oxidase, which suggests that these organisms can potentially metabolize O2 under different conditions (that is, oxic/hypoxic).
The divergence of the Euryarchaeota from the Crenarchaeota is thought to have occurred ∼3.2–3.6 Ga, whereas the split between the Thermoproteales and Sulfolobales/Desulfurococcales was hypothesized to occur ∼2.3–2.7 Ga (Blank, 2009). The divergence of the NAG1 lineage would have occurred between these two periods, likely before ‘the Great Oxidation Event’ ∼2.45 Ga (Bekker et al., 2004) and perhaps before the evolution of oxygenic photosynthesis at ∼2.7 Ga (Barley et al., 2005). Phylogenetic analysis of the NAG1 HCO shows the deduced protein to be among the most deeply rooted Type A HCOs in the Archaea (Supplementary Figure S7), and may have been important in the divergence of other archaeal Type A HCOs in high temperature, slightly acidic ferrous iron systems, before the proposed rise in oxygen (>10−5 present atmospheric levels; Bekker et al., 2004; Barley et al., 2005).
Organisms within the NAG1 lineage are important in many of the moderately acidic Fe-rich mats studied in YNP (as inferred from 16S rRNA gene distribution) and may exhibit unique biochemical signatures that, if preserved in the fossil record, would provide additional tools for understanding the importance of this habitat type in Earth’s evolutionary history. Archaeal lipid biomarkers have generated significant interest as a tool for determining the presence of specific archaea in different paleobiological contexts. Analysis of NAG1 genes for isoprenoid biosynthesis has revealed a farnesylgeranyl diphosphate synthase with a chain-length determination region identical to that of A. pernix, which is one of only a very few archaeal species to produce C25 archaeols exclusively (Supplementary Figure S10; Matsumi et al., 2011). Consequently, the genetic evidence suggests that NAG1 organisms may also produce C25 archaeols, and this may prove to be a unique biomarker for examining various Fe(III) formations with paleobiological significance, especially given that members of the Sulfolobales produce C40 caldarchaeols via an alternate mevalonate pathway.
Although it is not yet known whether any useful biomarkers specific to the NAG1 lineage can be utilized with confidence, this is a high priority for future work that could answer whether NAG1-like populations were important in other Fe oxide formations preserved in the rock record. Further genetic, phylogenetic and structural analysis of proteins produced by NAG1-like organisms or other members of this lineage will provide additional insight regarding the exact phylogenetic placement of the candidate phylum Geoarchaeota, as well as possible evolutionary linkages to both domains Bacteria and Eukarya. Moreover, efforts to isolate relevant members of this lineage should now have a higher probability of success using genomic information to customize growth conditions.
References
Aziz RK, Bartels D, Best AA, DeJongh M, Disz T, Edwards RA et al (2008). The RAST Server: rapid annotations using subsystems technology. BMC Genomics 9: 75.
Barker HA . (1936). Studies upon the methane-producing bacteria. Arch Mikrobiol 7: 420–438.
Barley ME, Bekker A, Krapez B . (2005). Late Archean to early Paleoproterozoic global tectonics, environmental change and the rise of atmospheric oxygen. Earth Planet Sci Lett 238: 156–171.
Barns SM, Fundyga RE, Jeffries MW, Pace NR . (1994). Remarkable archaeal diversity detected in a Yellowstone National Park hot spring environment. Proc Natl Acad Sci USA 91: 1609–1613.
Barns SM, Delwiche CF, Palmer JD, Pace NR . (1996). Perspectives on archaeal diversity, thermophily and monophyly from environmental rRNA sequences. Proc Natl Acad Sci USA 93: 9188–9193.
Behrens S, Rühland C, Inácio J, Huber H, Fonseca A, Spencer-Martins I et al (2003). In situ accessibility of small-subunit rRNA of members of the domains Bacteria, Archaea, and Eucarya to Cy3-labeled oligonucleotide probes. Appl Environ Microbiol 69: 1748–1758.
Bekker A, Holland HD, Wang P-L, Rumble D, Stein HJ, Hannah JL et al (2004). Dating the rise of atmospheric oxygen. Nature 427: 117–120.
Berg IA, Kockelkorn D, Buckel W, Fuchs G . (2007). A 3-hydroxypropionate/4-hydroxybutyrate autotrophic carbon dioxide assimilation pathway in Archaea. Science 318: 1782–1786.
Berg IA, Kockelkorn D, Ramos-Vera WH, Say RF, Zarzycki J, Hügler M et al (2010). Autotrophic carbon fixation in archaea. Nat Rev Microbiol 8: 447–460.
Bernande R, Ettema TJ . (2010). FtsZ-less cell division in archaea and bacteria. Curr Opin Microbiol 13: 747–752.
Bertin PN, Heinrich-Salmeron A, Pelletier E, Goulhen-Chollet F, Arsène-Ploetze F, Gallien S et al (2011). Metabolic diversity among main microorganisms inside an arsenic-rich ecosystem revealed by meta- and proteo-genomics. ISME J 11: 1735–1747.
Blainey PC, Mosier AC, Potanina A, Francis CA, Quake SR . (2011). Genome of a low-salinity ammonia-oxidizing archaeon determined by single-cell and metagenomic analysis. PLoS One 6: e16626.
Blank CE . (2009). Not so old Archaea—the antiquity of biogeochemical processes in the archaeal domain of life. Geobiology 5: 495–514.
Boyd ES, Jackson RA, Encarnacion G, Zahn JA, Beard T, Leavitt WD et al (2007). Isolation, characterization, and ecology of sulfur-respiring crenarchaea inhabiting acid-sulfate-chloride-containing geothermal springs in Yellowstone National Park. Appl Environ Microbiol 73: 6669–6677.
Brochier C, Gribaldo S, Zivanovic Y, Confalonieri F, Forterre P . (2005). Nanoarchaea: representatives of a novel archaeal phylum or a fast-evolving euryarchaeal lineage related to Thermococcales? Genome Biol 6: R42.
Brochier-Armanet C, Boussau B, Gribaldo S, Forterre P . (2008). Mesophilic Crenarchaeota: proposal for a third archaeal phylum, the Thaumarchaeota. Nat Rev Microbiol 6: 252.
Brock TD, Brock KM, Belly RT, Weiss RL . (1972). Sulfolobus: a new genus of sulfur-oxidizing bacteria living at low pH and high temperature. Arch Mikrobiol 84: 54–68.
de la Torre JR, Walker CB, Ingalls AE, Konneke M, Stahl DA . (2008). Cultivation of a thermophilic ammonia oxidizing archaeon synthesizing crenarchaeol. Environ Microbiol 10: 810–818.
Dobbek H, Gremer L, Kiefersauer R, Huber R, Meyer O . (2002). Catalysis at a dinuclear [CuSMo(==O)OH] cluster in a CO dehydrogenase resolved at 1.1-A resolution. Proc Natl Acad Sci USA 99: 15971–15976.
Elkins JG, Podar M, Graham DE, Makarova KS, Wolf Y, Randau L et al (2008). A korarchaeal genome reveals new insights into the evolution of the Archaea. Proc Natl Acad Sci USA 105: 8102–8107.
Freitas TA, Hou S, Dioum EM, Saito JA, Newhouse J, Gonzalez G . (2004). Ancestral hemoglobins in Archaea. Proc Natl Acad Sci USA 101: 6675–6680.
Freitas TA, Saito JA, Hou S, Alam M . (2005). Globin-coupled sensors, protoglobins, and the last universal common ancestor. J Inorg Biochem 99: 23–33.
Fuchs BM, Pernthaler J, Amann R . (2007). Single cell identification by fluorescence in situ hybridization. In: Methods for General and Molecular Microbiology 3rd Edn Reddy CA, Beveridge TJ, Breznak JA, Marzluf G, Schmidt TM, Snyder LR, (eds). ASM Press: Washington, DC, pp 886–896.
Harrison FC, Kennedy ME . (1922). The red discoloration of cured codfish. Trans Roy Soc Can 16: 101–152.
Hasan MR, Rahman M, Jaques S, Purwantini E, Daniels LJ . (2010). Glucose 6-phosphate accumulation in mycobacteria: implications for a novel F420-dependent anti-oxidant defense system. Biol Chem 25: 19135–19144.
Hatzenpichler R, Lebedeva EV, Spieck E, Stoecker K, Richter A, Daims H et al (2008). Moderately thermophilic ammonia-oxidizing crenarchaeote from a hot spring. Proc Natl Acad Sci USA 105: 2134–2139.
Huber H, Hohn MJ, Rachel R, Fuchs T, Wimmer VC, Stetter KO . (2002). A new phylum of Archaea represented by a nanosized hyperthermophilic symbiont. Nature 417: 63–67.
Ingalls AE, Shah SR, Hansman RL, Aluwihare LI, Santos GM, Druffel ER et al (2006). Quantifying archaeal community autotrophy in the mesopelagic ocean using natural radiocarbon. Proc Natl Acad Sci USA 103: 6442–6447.
Inskeep WP, Ackerman GG, Taylor WP, Korf S, Kozubal MA, Macur RE . (2005). On the energetics of chemolithotrophy in nonequilibrium systems: case studies of geothermal springs in Yellowstone National Park. Geobiol 3: 297–320.
Inskeep WP, Rusch DB, Jay ZJ, Herrgard MJ, Kozubal MA, Richardson TH et al (2010). Metagenomes from high-temperature chemotrophic systems reveal geochemical controls on microbial community structure and function. PLoS One 5: e9773.
Jones DT, Taylor WR, Thornton JM . (1992). The rapid generation of mutation data matrices from protein sequences. Comput Appl Biosci 8: 275–282.
King GM, Weber CF . (2007). Distribution, diversity and ecology of aerobic CO-oxidizing bacteria. Nat Rev Microbiol 2: 107–118.
Könneke M, Bernhard AE, de la Torre JR, Walker CB, Waterbury JB, Stahl DA . (2005). Isolation of an autotrophic ammonia-oxidizing marine archaeon. Nature 437: 543–546.
Kozubal M, Macur RE, Korf S, Taylor WP, Ackerman GG, Nagy A et al (2008). Isolation and distribution of a novel iron-oxidizing crenarchaeon from acidic geothermal springs in Yellowstone National Park. Appl Environ Microbiol 74: 942–949.
Kozubal MA, Dlakic M, Macur RE, Inskeep WP . (2011). Terminal oxidase diversity and function in ‘Metallosphaera yellowstonensis’: gene expression and protein modeling suggest mechanisms of Fe(II) oxidation in the Sulfolobales. Appl Environ Microbiol 77: 1844–1853.
Kozubal MA, Macur RE, Jay ZJ, Beam JP, Malfatti SA, Tringe SG et al (2012). Microbial iron cycling in acidic geothermal springs of Yellowstone National Park: Integrating molecular surveys, geochemical processes and isolation of novel Fe-active microorganisms. Front Microbio 3: 109 (in press).
Makarova KS, Aravind L, Wolf YI, Koonin EV . (2011). Unification of Cas protein families and a simple scenario for the origin and evolution of CRISPR-Cas systems. Biol Direct 6: 38.
Matsumi R, Atomi H, Driessen AJ, van der Oost J . (2011). Isoprenoid biosynthesis in Archaea—biochemical and evolutionary implications. Res Microbiol 1: 39–52.
Meyer-Dombard DR, Shock EL, Amend JP . (2005). Archaeal and bacterial communities in geochemically diverse hot springs of Yellowstone National Park, USA. Geobiol 3: 211–227.
Moens L, Vanfleteren J, Van de Peer Y, Peeters K, Kapp O, Czeluzniak J et al (1996). Globins in nonvertebrate species: dispersal by horizontal gene transfer and evolution of the structure-function relationships. Mol Biol Evol 13: 324–333.
Muller F, Brissac T, Le Bris N, Felbeck H, Gros O . (2010). First description of giant Archaea (Thaumarchaeota) associated with putative bacterial ectosymbionts in a sulfidic marine habitat. Environ Microbiol 8: 2371–2383.
Nardini M, Pesce A, Thijs L, Saito JA, Dewilde S, Alam M et al (2008). Archaeal protoglobin structure indicates new ligand diffusion paths and modulation of haem-reactivity. EMBO Rep 9: 157–163.
Nei M, Kumar S . (2000) Molecular Evolution and Phylogenetics. Oxford University Press: New York, pp 333.
Nishimasu H, Fushinobu S, Shoun H, Wakagi T . (2006). Identification and characterization of an ATP-dependent hexokinase with broad substrate specificity from the hyperthermophilic archaeon Sulfolobus tokodaii. J Bacteriol 5: 2014–2019.
Overbeek R, Begley T, Butler RM, Choudhuri JV, Chuang HY, Cohoon M et al (2005). The subsystems approach to genome annotation and its use in the project to annotate 1000 genomes. Nucleic Acids Res 33: 5691–5702.
Rutherford K, Parkhill J, Crook J, Horsnell T, Rice P, Rajandream MA et al (2000). Artemis: sequence visualization and annotation. Bioinformatics 16: 944–945.
Spang A, Hatzenpichler R, Brochier-Armanet C, Rattei T, Tischler P, Spieck E et al (2010). Distinct gene set in two different lineages of ammonia-oxidizing archaea supports the phylum Thaumarchaeota. Trends Microbiol 18: 331–340.
Storbeck S, Rolfes S, Raux-Deery E, Warren MJ, Jahn D, Layer G . (2010). A novel pathway for the biosynthesis of heme in Archaea: genome-based bioinformatic predictions and experimental evidence. Archaea 13: 175050.
Tamura K, Dudley J, Nei M, Kumar S . (2007). MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol Biol Evol 24: 1596–1599.
Teeling H, Meyerdierks A, Bauer M, Amann R, Glöckner FO . (2004). Application of tetranucleotide frequencies for the assignment of genomic fragments. Environ Microbiol 6: 938–947.
Tourna M, Stieglmeier M, Spang A, Könneke M, Schintlmeister A, Urich T et al (2011). Nitrososphaera viennensis, an ammonia oxidizing archaeon from soil. Proc Natl Acad Sci USA 108: 8420–8425.
Walker CB, de la Torre JR, Klotz MG, Urakawa H, Pinel N, Arp DJ et al (2010). Nitrosopumilus maritimus genome reveals unique mechanisms for nitrification and autotrophy in globally distributed marine crenarchaea. Proc Natl Acad Sci USA 107: 8818–8823.
Woese CR, Sogin M, Stahl D, Lewis BJ, Bonen L . (1976). A comparison of the 16S ribosomal RNAs from mesophilic and thermophilic Bacilli: Some modifications in the Sanger method for RNA sequencing. J Mol Evol 7: 197–213.
Woese CR, Fox GE . (1977). Phylogenetic structure of the prokaryotic domains: the primary kingdoms. Proc Natl Acad Sci USA 74: 5088–5090.
Woese CR, Kandler O, Wheelis ML . (1990). Towards a natural system of organisms. Proposal for the domains Archaea, Bacteria, and Eucarya. Proc Natl Acad Sci USA 87: 4576–4579.
Wolfe RS . (2006). The Archaea: a personal overview of the formative years. In: The Prokaryotes: A Handbook on the Biology of Bacteria 3rd edition Dworkin M, Flakow S, Rosenberg E, Karl-Heinz S, Stackenbrandt E, (eds) Springer: Heidelberg, pp 1–8.
Wu D, Raymond J, Wu M, Chatterji S, Ren Q, Graham JE et al (2009). Complete genome sequence of the aerobic CO-oxidizing thermophile Thermomicrobium roseum. PLoS One 4: e4207.
Acknowledgements
Authors from MSU appreciate support from the Department of Energy (DOE)-Joint Genome Institute Community Sequencing Program (CSP 787081), NASA Exobiology (via the Thermal Biology Institute, MSU), NSF IGERT (0654336), DOE-Pacific Northwest National Laboratory (subcontract no. 112443), the Montana Agricultural Experiment Station (911300), B Pitts (Center for Biofilm Engineering) for assistance and training in confocal microscopy, and C Hendrix and T Olliff for permitting this work in YNP (Permit YELL-2007–2008-SCI-5068). The work conducted by the Joint Genome Institute (DE-AC02-05CH11231) and the Pacific Northwest National Laboratory (Foundational Scientific Focus Area) is supported by the Genomic Science Program, Office of Biological and Environmental Research, US DOE.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Kozubal, M., Romine, M., Jennings, R. et al. Geoarchaeota: a new candidate phylum in the Archaea from high-temperature acidic iron mats in Yellowstone National Park. ISME J 7, 622–634 (2013). https://doi.org/10.1038/ismej.2012.132
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2012.132
Keywords
This article is cited by
-
Hyperthermophile diversity microbes in the Calientes geothermal field, Tacna, Peru
Brazilian Journal of Microbiology (2023)
-
Global patterns of diversity and metabolism of microbial communities in deep-sea hydrothermal vent deposits
Microbiome (2022)
-
Culexarchaeia, a novel archaeal class of anaerobic generalists inhabiting geothermal environments
ISME Communications (2022)
-
Brockarchaeota, a novel archaeal phylum with unique and versatile carbon cycling pathways
Nature Communications (2021)
-
A standardized archaeal taxonomy for the Genome Taxonomy Database
Nature Microbiology (2021)