Abstract
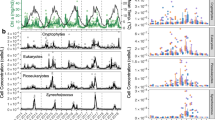
Monterey Bay, CA is an Eastern boundary upwelling system that is nitrogen limited much of the year. In order to resolve population dynamics of microorganisms important for nutrient cycling in this region, we deployed the Environmental Sample Processor with quantitative PCR assays targeting both ribosomal RNA genes and functional genes for subclades of cyanobacteria (Synechococcus) and ammonia-oxidizing Archaea (Thaumarchaeota) populations. Results showed a strong correlation between Thaumarchaea abundances and nitrate during the spring upwelling but not the fall sampling period. In relatively stratified fall waters, the Thaumarchaeota community reached higher numbers than in the spring, and an unexpected positive correlation with chlorophyll concentration was observed. Further, we detected drops in Synechococcus abundance that occurred on short (that is, daily) time scales. Upwelling intensity and blooms of eukaryotic phytoplankton strongly influenced Synechococcus distributions in the spring and fall, revealing what appear to be the environmental limitations of Synechococcus populations in this region. Each of these findings has implications for Monterey Bay biogeochemistry. High-resolution sampling provides a better-resolved framework within which to observe changes in the plankton community. We conclude that controls on these ecosystems change on smaller scales than are routinely assessed, and that more predictable trends will be uncovered if they are evaluated within seasonal (monthly), rather than on annual or interannual scales.
Similar content being viewed by others
Introduction
Ocean time series are essential for accurate predictions of climate scenarios in this age of anthropogenic change (Doney, 1999). Marine biological time series have proven invaluable for uncovering major oceanographic trends that would not have been observed otherwise (Karl et al., 1995; Chavez, 1999; Karl, 1999, 2007; McGowan et al., 1998; Chavez et al., 2003). Although monthly time series are essential for providing data on annual and decadal patterns, the factors controlling changes in microbial populations are known to occur on shorter time, and smaller space scales (that is, from seconds to months, depending on the process: Karl and Dore, 2001; Johnson et al., 2009). However, sustaining high-frequency assessments of microbial populations is generally not possible given logistical and practical constraints, particularly for remote locations. With few exceptions, acquisition of detailed information concerning microbial distributions and activities is typically limited to ship-based surveys. The reliance on traditional ship-based sampling has driven the development and testing of new biological sensors (Paul et al., 2007; Shade et al., 2009; Scholin, 2010).
The Environmental Sample Processor (ESP) is among this new generation of sensors. Modeled as an ‘ecogenomic sensor’ (Scholin, 2010), the instrument provides for subsurface, hands-off collection of discrete water samples, particle concentration and application of various molecular biological analyses including quantitative PCR (Preston et al., 2011) as well as DNA and protein probe arrays for detection of a variety of organisms, genes and metabolites (Goffredi et al., 2006; Haywood et al., 2007; Greenfield et al., 2008; Doucette et al., 2009; Preston et al., 2009; Scholin et al., 2009; Scholin, 2010). At present, the instrument can perform these types of assays with daily resolution and in real-time, and transmit data to a remote location. The ESP can also archive samples for more in-depth analyses upon instrument retrieval (Scholin et al., 2009; Ottesen et al., 2011).
Monterey Bay, CA serves as an optimal location for a high-resolution time series study, as it has been historically monitored by the Monterey Bay Aquarium Research Institute (MBARI) time series (Pennington and Chavez, 2000), and is a region of enormous biogeochemical interest with strong seasonality in environmental conditions (Breaker and Broenkow, 1994). Productivity in Monterey Bay is nitrogen limited for much of the year (Johnson et al., 2006). Upwelling is the primary source of nitrate, and remineralization within the surface waters adds to the nitrogen pool during productive and stratified periods (Dugdale and Goering, 1967; Johnson et al., 2006). Eukaryotic and some cyanobacterial phytoplankton can assimilate nitrate as well as ammonia, and compete with each other for limiting nitrogen in the bay.
The Thaumarchaeota (or group I Archaea, formerly the Crenarchaeota (Brochier-Armanet et al., 2008; Spang et al., 2010)) also have an important role in the nitrogen cycle in coastal marine systems by oxidizing ammonia to nitrite (Könneke et al., 2005), thus altering the form of nitrogen available for photosynthesis and other processes in upwelled waters and deeper in the photic zone where they reside (Mincer et al., 2007; Santoro et al., 2010; Preston et al., 2011). The Thaumarchaeota have widespread global distributions, and have been estimated to comprise over 20% of marine picoplankton (Karner et al., 2001). These organisms are key nitrifiers in Monterey Bay (Francis et al., 2005; Mincer et al., 2007; Santoro et al., 2010) and maintain large, but temporally variable populations in coastal California waters (Massana et al., 1997; Murray et al., 1999; Mincer et al., 2007).
The cyanobacteria are significant contributors to marine primary production (Goericke and Welschmeyer, 1993; Liu et al., 1997) and perform many essential functions to maintain the health of food webs (Wilhelm and Trick, 1994; Zehr et al., 2001). The cyanobacterial genus Synechococcus can assimilate a diversity of nitrogen compounds, including ammonium and nitrate, and reside in relatively nutrient-rich waters at coastal sites throughout the world (Scanlan and West, 2002). The Synechococcus populations in coastal California waters are genetically diverse (Toledo and Palenik, 1997; Paerl et al., 2008; Palenik et al., 2009) and strain abundances vary over the course of the year (Tai and Palenik, 2009; Paerl et al., in press).
Despite their important roles in marine nutrient cycling, Thaumarchaeota and Synechococcus population dynamics in Monterey Bay are not well resolved. In an effort to better understand the controls on these organisms’ distributions, we deployed an ESP in the spring and fall months of 2009 in shallow waters at the MBARI's coastal time series station M0. This study applies qPCR assays to detect both general groups of organisms as well as specific subclades to determine trends in microbial distribution relative to environmental change. Physical, chemical and biological parameters were measured over 57 days, along with the corresponding changes in the prokaryotic communities using the ESP. Thaumarchaeota were detected with a general 16S rRNA assay (Suzuki et al., 2000) and subclades were enumerated with a functional gene assay targeting the ammonia monooxygenase α-subunit gene for the marine shallow water ecotype (amoA, group A; Mosier and Francis, 2011). Synechococcus were detected with a comprehensive rbcL assay targeting the ribulose-1,5-bisphosphate carboxylase gene that accounts for 89% total Synechococcus in Monterey Bay when compared with flow cytometry-derived cell counts (Preston et al., 2011; Paerl et al., in press), while Synechococcus subclades were detected with a functional gene assay coding for the assimilatory nitrate reductase gene (narB) and targeting subclades C C1 and D C2, associated with Synechococcus clades I and IV, respectively, from cultured isolates (Paerl et al., 2011).
The utility of the ESP for quantifying microbial population dynamics in Monterey Bay has been demonstrated by Preston et al. (2011), where microbes with predictable dynamics were quantified during the upwelling season, for technical validation of the ESP technology. Here it was shown that Thaumarchaeota abundances increase with upwelling, as expected based on their deeper distributions. Aside from this proof-of-concept field study, their abundance near shore has not been addressed, and it is generally assumed based on previous findings offshore (Mincer et al., 2007; Santoro et al., 2010) that the Thaumarchaea are scarce, if at all present in shallow waters, and that any distribution to shallow waters is determined by upwelling and mixing alone. Paerl et al. (in press) have shown that Synechococcus populations are abundant for most of the year near the coast, with population decreases corresponding with times of intense upwelling, and a fairly consistent presence in surface water otherwise. With daily sampling, Preston et al. (2011) further resolved the period of Synechococcus appearance in coastal waters, at the relaxation of upwelling. Little is known, however, regarding the drivers of specific Synechococcus subclade distributions over the rest of the year, nor whether their populations are consistently present when upwelling intensity is low. This study sought to obtain higher resolution of the abundance of both Thaumarchaeal and Synechococcus populations and subclades, within and between seasons. The resulting data illustrate a highly dynamic microbial community reflective of regional scale physical processes, such as upwelling, as well as localized processes in stable conditions, where we posit that inter-organismal interactions have a determinant role in dictating microbial distributions.
Materials and methods
Instruments were deployed in Monterey Bay, CA at Station M0 (36.83N, 121.90W) on a mooring as described previously (Doucette et al., 2009). ESP deployments occurred in the spring (from 14 May to 11 June) and fall (from 24 September to 28 October) of 2009, and included an attached Conductivity, Temperature, Depth Sensor (CTD: Seabird SBE 16+CTD, Bellevue, WA, USA) with fluorometer (Turner Cyclops-7, Turner Designs, Sunnyvale, CA, USA) and transmissometer (WetLABS Cstar, WET Labs, Philomath, OR, USA). An in situ ultraviolet spectrophotometer (Johnson and Coletti, 2002) was also coupled to the ESP for in situ nitrate analyses. Data were recorded every 12 min from accompanying sensors. Data were transmitted to shore in real time over a radio modem. The instrument was deployed slightly deeper in the spring (10–11.4 m) than the fall (5.8–8.8 m).
Instrument preparation
To prepare the ESP, tubing was cleaned and reagents were loaded as described (Preston et al., 2011). Quantitative PCR assays were loaded onto the microfluidic block for the following genes: Thaumarchaeota 16S rRNA (which amplifies all ammonia-oxidizing Archaea known; Preston et al., 2011), and amoA water column group A (which targets all known amoA group A sequences from a Monterey Bay database, representing 825 sequences from a total of 9634): Mosier and Francis, 2011), Synechococcus RuBisCO (rbcL, targets all cultivated Synechococcus clades I–IV and all known rbcL sequences in Monterey Bay (Paerl et al., in press), narB group C_C1* and group D_C2 (Paerl et al., 2011). The narB and rbcL assays were the same: each 30 μl reaction contained 1 × Accuprime Supermix II (Invitrogen, Carlsbad, CA, USA), 0.333 μM primers, and 2 μM probe. The amoA group A assay contained 1 × Accuprime, 2.5 mM additional MgCl2, 0.5 μM forward primer, and 0.333 μM reverse primer. 16S rRNA amplification and cycling conditions were as described previously (Preston et al., 2011). For enumeration, the deployed reagents were used on the microfluidic block of the ESP to create duplicate standard curves from 10-fold linearized plasmid standard dilutions ranging from 102 to 105 or 106 gene copies per reaction. Amplification plots were analyzed and standard curves generated in Excel. Quantitative PCR assay cross-reactivities were low or non-existent (published in Mosier and Francis, 2011; Preston et al., 2011; Paerl et al., in press), and detection limits on the microfluidic block were 10–20 copies per ml for Synechococcus targets and 10–50 copies per ml for Thaumarchaeal targets. qPCR standard curve information for the spring deployment are reported elsewhere (Preston et al., 2011). For the fall deployment, copy numbers for the standards in each assay were linear with cycle threshold (Ct). Linear regressions for all assays had r2 values of >0.98, and amplification efficiencies were 90–110% with the exception of the narB group C C1 assay, which had an efficiency of 88.6%. Analyses of this target group are discussed with consideration of the implications of this lower reaction efficiency. The DNA extraction efficiency in the ESP microfluidic block was 98–104% compared with parallel extractions performed in the lab with a Qiagen DNeasy Kit (Qiagen, Valencia, CA, USA).
ESP operations included 24 discrete sampling events in which particles retained on 0.22 μm pore size filters were subjected to qPCR analyses, and an equal number of discrete sampling events where particulates were preserved for later analysis (after Ottesen et al., 2011). The day before each deployment (on 14 May and 23 September), 50 ml of sterile water (Sigma-Aldrich, St Louis, MO, USA) was sampled by the intake of the instrument in order to determine the degree of internal contamination, if any. Negative controls were also run during each deployment (on 17 and 30 May in the spring and 9 and 16 October in the fall), each containing a core system negative control (where a ‘negative lysate’ is created and run through the entire system), and a no-template control (where the PCR is run with the elution water used for DNA extractions in normal operating conditions).
Assay performance during deployment
Though the ESP does not allow standard curve generation for all samples taken for all assays while deployed, internal positive controls give some degree of confidence in target quantification on a per sample basis. For each sample, the degree of PCR inhibition was assessed using a positive control reaction as described in Preston et al., (2011). Samples were only included in our analyses if the Ct of the internal positive control was between 33.5 and 36.5 (spring) or 27.2 and 29.7 (fall). These ranges were within the Ct variability encountered for each PCR module in reactions with water; the template of the internal positive control is included in the primer/probe reagent. None of the no-template control reactions amplified indicating uncontaminated reagents and sufficient cleaning of the PCR microfluidics between the samples.
In core system negative control reactions, amplification occurred for a few of the targets. However, in each case, the Ct values translated to copy numbers that were at least an order of magnitude lower than the sample, which does not affect our interpretations of the trends. The fall deployment allowed interrogation of subpopulations of both Synechococcus and Thaumarchaeota. Subclade dynamics correlated with total clade dynamics, providing support that these groups were adequately quantified. Pearson and Spearman correlation analyses were performed using SigmaPlot 11.2 (Systat Software, Inc., Chicago, IL, USA).
Additional data
Mooring data from station M1 (Pennington and Chavez, 2000) were used to create the water column profiles for Monterey Bay, and chlorophyll satellite images were obtained from Moderate Resolution-Imaging Spectroradiometer on NOAA's Coastwatch/SWFSC Oceanwatch Live Access Server (http://coastwatch.pfeg.noaa.gov).
Results and discussion
Seasonal conditions at station M0
Seasonal upwelling dominated spring conditions at Station M0, leading to two major bloom periods that were clearly distinguished by increased chlorophyll-a fluorescence (Figure 1). During this deployment period, conditions developed from initial intense upwelling to relaxation and eventual stratification of the water column (Figures 1 and 2). The first week of the deployment, when upwelling was sustained, high nitrate concentrations (near 25 μM), high salinity (33.9 PSU) and low temperatures (<11 °C) were observed as expected (Breaker and Broenkow, 1994; Figures 1 and 2). As a reflection of deep-water transport to surface waters (Bolin and Abbott, 1963), chlorophyll-a concentrations were low and constant (below 10 μg l−1) until the relaxation of upwelling around 24 May. The following seven ESP-collected biological samples after this date were from periods of high chlorophyll-a/low transmissivity (until 2 June: black-outlined boxes Figure 3), indicating abundant eukaryotic phytoplankton at station M0 (Green et al., 2003; Green and Sosik, 2004; Behrenfeld and Boss, 2006). These blooms were not adequately captured by satellite imaging (http://coastwatch.pfeg.noaa.gov; Figure 1), but microscopy of Monterey Bay seawater during this period supports this interpretation with the identification of a bloom of Chaetoceros spp. on 27 May (HABMAP: http://cimt2007plankton.wordpress.com/). A tongue of nitrate-rich, low-salinity, offshore water entered the bay displacing the bloom (Preston et al., 2011), and this was followed by increased stratification that persisted through the end of the deployment when nitrate dropped to 5–10 μM, salinity to 33.6 PSU and temperatures raised to 13–14 °C (Figure 1). There was only trace precipitation over the course of the spring deployment.
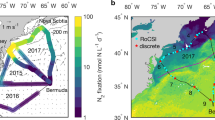
Environmental data for Monterey Bay in (a) spring and (b) fall. First panel includes Moderate Resolution-Imaging Spectroradiometer 8-day composite satellite images of chlorophyll-a for selected dates during each deployment (most were not available for the spring dates). Star in right panel is approximate location of Station M0. Color legends range from purple to red: 18 May 2009: from −0.6 to 2.0, 3 June 2009: from −0.7 to 2.1, 12 June 2009: from −0.5 to 2.3, 1 October 2009: from −0.4 to 2.4, 8 October 2009: from −0.4 to 2.4 and 14 October 2009: from −0.8 to 1.8. Units are log mg m−3. Panels below contain continuous temperature, nitrate, chlorophyll-a and salinity data gathered from in situ ultraviolet spectrophotometer, CTD sensors coupled with the ESP during each deployment. The two bloom periods in the spring are designated with a box from 24 May to 2 June and after 8 June. Arrows point to periods of offshore water influx into the bay.
Abundances of bacterial and archaeal gene copy numbers per ml seawater over time for (a) spring and (b) fall, 2009, plotted on a log scale. Panels below show Monterey Bay water column temperature and salinity characteristics over this period (note the differences in scales between the spring and fall). Dotted line within each of the bottom panels corresponds with the average deployment depth for each season.
Eukaryotic phytoplankton abundances. Percentage transmission vs chlorophyll-a for both spring (squares) and fall (diamonds) periods. Top left of graph indicates high phytoplankton relative to bottom right. The size of the data point within each season corresponds to the time of sampling: the larger the data point the later it is in the deployment. As described in the text, dark outlined points are those depicting upwelling relaxation in the spring (squares, mid-deployment) and of the influx of the offshore water mass in the fall (diamonds, end of deployment).
After an initial unstable phase that lasted 3 days, fall conditions were well-stratified relative to spring, with warmer (12–15 °C), fresher (33.2–33.5 PSU) water, steady physical and chemical conditions and no deep water supply to the surface detected (Figures 1 and 2). Nitrate was less variable in the fall than the spring, with concentrations ranging 8–12 μM with the exception of several transient (hourly) spikes to 17 μM. Chlorophyll-a concentrations increased over time, beginning at an average of 7 μg l−1 (with a range from approximately 3–20 μg l−1) and increasing to an average of 17 μg l−1 (9–38 μg l−1 range; Figures 1 and 2). The ratio of chlorophyll-a to percentage transmission during ESP sampling periods was high, indicating abundant eukaryotic phytoplankton for most of the fall (Figure 3; Green et al., 2003; Green and Sosik, 2004; Behrenfeld and Boss, 2006). Microscopy of Monterey Bay seawater for this period also documented high abundances of eukaryotic phytoplankton, with dinoflagellates (Ceratium spp.) dominating for the majority of the deployment period (sampled at Santa Cruz Wharf on 23 and 29 September; 5, 12 and 19 October: http://cimt2007plankton.wordpress.com/), and Pseudonitszchia-like diatoms observed in high numbers for one of the five samples (on 5 October: http://cimt2007plankton.wordpress.com/). There was only trace precipitation over this period, with the exception of 4.54 inches of rain on 13 October. This rain event was coincident with the onshore influx of a chlorophyll-poor water mass (Figure 3, outlined diamonds).
Microbial population dynamics
Upwelling was the major driver of microbial distributions in the spring, as expected, but population abundances fluctuated despite the relative stability of the water column in stratified fall conditions. Our data lead us to hypothesize that blooms of eukaryotic phytoplankton have an effect on the Thaumarchaeota and Synechococcus clade dynamics over this period.
Thaumarchaeal population dynamics
For much of the spring period, the Thaumarchaeota were abundant in surface waters, based on 16S rRNA gene copy numbers (with a range of undetectable to 6.0 × 104 copies per ml; Preston et al., 2011). A weak negative correlation with chlorophyll-a during this upwelling period likely reflects the transport of chlorophyll-poor, Thaumarchaea-rich deep water to the surface (Pearson's r=−0.448, P=0.037; Spearman's correlation insignificant). Thaumarchaeota abundances were positively correlated with the magnitude of upwelling, using temperature (Pearson's r=−0.636, P=0.001; Spearman's ρ=−0.853, P<0.0001), salinity (Pearson's r=0.558, P=0.007; Spearman's ρ=0.704, P=0.0002) and nitrate as a proxy (Figure 4a; Pearson's r=0.769, P<0.0001; Spearman's ρ=0.831, P<0.0001). Aside from one exception, all quantified genes show correlations with these three upwelling proxies in the spring, demonstrating the impact of upwelling on these microbial populations. Synechococcus, however, did not correlate with nitrate (see below), and the observation that Thaumarchaeal abundances correlate most strongly with nitrate (vs temperature and salinity) may reflect their role in marine nitrification.
Thaumarchaea population dynamics. (a) Bubble plot of Thaumarchaea abundances (that is, 16S rRNA gene copy numbers) in the spring (thick outlined) and fall (no outline) vs nitrate and chlorophyll-a concentrations. Size of bubble corresponds to the population size; brightness corresponds with time (the darker the bubble, the earlier in the deployment). Bubbles with dotted outlines indicate time points corresponding with the period of upwelling relaxation in the spring. (b) Thaumarchaea total population abundances (that is, 16S rRNA gene copy numbers in dark), and amoA water column A gene copy numbers (in light) over the fall period, plotted vs nitrate and chlorophyll.
Thaumarchaeota were more abundant in the fall (16S rRNA gene range of 2.7 × 103 to 1.3 × 106 per ml), and again correlated with cold (Pearson's r=−0.754, P=0.0002; Spearman's ρ=−0.805, P<0.0001), saline waters (Pearson's r=0.625, P=0.004; Spearman's ρ=0.853, P<0.0001), but the connection between Thaumarchaeal abundance and nitrate concentration was not supported in these stratified conditions. Also in contrast to the spring, Thaumarchaea 16S rRNA genes showed a significant positive correlation with chlorophyll-a in the fall (Pearson's r=0.554, P=0.014; Spearman's ρ=0.721, P=0.0003) (Figure 4a). During this time, large Thaumarchaeal populations were sustained in the surface waters and 16S rRNA gene copy numbers grew to become two orders of magnitude more abundant over time (Figure 4a). These data suggest that Thaumarchaeal populations can thrive in surface waters during blooms of eukaryotic phytoplankton. The decrease in abundance that correlates with a drop in chlorophyll towards the end of the deployment may be a reflection of the chlorophyll-poor water mass that moved onshore at this time (which was warmer and less saline, two conditions previously suggested to be unfavorable based on marine Thaumarchaea distributions (Murray et al., 1999; Mincer et al., 2007; Santoro et al., 2010).
In order to determine whether the Thaumarchaea targeted by the 16S rRNA gene primers used in this study have the genetic potential to oxidize ammonia, we correlated their abundance with the abundance of ammonia monooxygenase subunit A functional genes (amoA). There is now considerable evidence for the existence of two distinct clades of marine ammonia-oxidizing Archaea, namely the ‘shallow’ (group A) and ‘deep’ (group B) water column ammonia-oxidizing Archaea (Francis et al., 2005; Hallam et al., 2006; Beman et al., 2008; Mosier and Francis, 2011). It is worth noting that both marine water column amoA subclades would be detected by the described Thaumarchaeal 16S rRNA qPCR assay. In this study, we specifically targeted the shallow (group A) water column ammonia-oxidizing Archaea ecotype, and only during stratification in the fall season. Previous work in the Monterey Bay region shows relative consistency between total amoA gene and marine group I 16S rRNA gene abundances (Santoro et al., 2010), implying that most, if not all, of the Thaumarchaea contain at least one copy of the amoA gene.
Though Thaumarchaeota subclade populations have not been quantified at depths relevant to this study (5–10 m), or near shore within Monterey Bay, trends from past studies in the region (Mincer et al., 2007; Santoro et al., 2010) would imply that the shallow water subclade A might dominate in surface waters at station M0. We find that this subclade is abundant for the duration of the fall deployment (with a range of 2.1 × 103 to 7.5 × 104 copies per ml), but the ratio of amoA, group A to 16S rRNA gene copy numbers ranges from 0.05 to 1.39, with an average of 0.26 (these numbers are not directly comparable to ratios found in past studies, as those enumerated total amoA group A and B abundances rather than that of group A only (Mincer et al., 2007; Santoro et al., 2010). For just six time points, the shallow subclade A makes up >50% of the Thaumarchaea 16S rRNA copy number. This corresponds with the lowest chlorophyll-a concentrations (Figure 4b) after movement of the offshore water mass into the bay, leading to the hypothesis that the shallow subclade is dominant in shallow waters offshore, but that we are missing the amoA diversity (for example, ‘deep’ subclade or otherwise), closer to the coast. Future deployments will measure amoA diversity at this site and target all subclades in stratified fall conditions, in order to determine the relative contribution of each to the total Thaumarchaeal community.
In a previous California coastal time series study, Thaumarchaeal abundances correlated with the characteristics of deeper waters, including low temperature, high salinity and low chlorophyll (Murray et al., 1999). Murray et al. present correlative data from 32 months integrated over the course of the time series, so any potential short-term effects may have been overlooked. Furthermore, though longer in duration than this study, the previous study has monthly temporal resolution, while this study has daily resolution. Despite these differences, we expected the trends to be similar: large Thaumarchaeal abundances in the spring that correlate with upwelling, and low numbers in the fall as a result of increased water column stratification. Although the expected trends were indeed observed in the spring, the higher relative abundance of the Thaumarchaea populations in the fall points to additional unforeseen factors dictating their distributions. Repeated observation of this phenomenon as a result of higher resolution sampling, affords sufficient confidence to investigate the cause behind this finding in the near future.
The high abundance of Thaumarchaea in the surface waters at this coastal station during stratification has far reaching implications. If this correlation between the Thaumarchaea population and chlorophyll-a holds true during nutrient-rich stratified conditions in Monterey Bay, then we would expect near shore regions to experience several periods of Thaumarchaeal dominance annually, perhaps up to 3–4 months per year (Breaker and Broenkow, 1994). Assuming that this population is oxidizing ammonia, it is competing with phytoplankton for this preferred nitrogen source where nitrogen is limiting. Furthermore, the relative decrease in ammonia and increase in nitrite (and therefore nitrate) might sustain growth of larger phytoplankton in the bay (L’Helguen et al., 2008) and thereby contribute to increased carbon export during these periods (Yool et al., 2007).
Synechococcus population dynamics
Over the 2 months (spring and fall) of ESP sampling at station M0, rbcL-based Synechococcus abundance estimates ranged from undetectable to 104 genes per ml. The only correlations in the spring included a positive relationship between Synechococcus abundance and temperature (Pearson's r=0.708, P=0.0002; Spearman's ρ=0.708, P=0.0002), as well as a negative relationship with salinity (Pearson's r=−0.567, P=0.006; Spearman's ρ=−0.758, P<0.0001), likely reflecting the inability of Synechococcus to remain in surface waters during active upwelling. Once upwelling ceased and conditions became more uniform, Synechococcus increased to 2.4 × 103 copies per ml over the course of 3 days (Figures 1 and 2).
We observed no correlation between Synechococcus abundances and any of the environmental parameters measured over the fall period. Though the water column was stratified throughout the fall and there was very little change in the physico-chemical environment, major drops in Synechococcus abundance (from 104 to <10 per ml over 1 day) were observed twice during this time. These crashes coincided with spikes in chlorophyll (Figure 5, time points with thick black dotted outlines), and may be the result of competition or grazing. In a southern California coastal time series, total Synechococcus abundances correlated weakly with temperature, and a weaker negative correlation was observed with phosphate concentration (Tai and Palenik, 2009). It is likely that different parameters control Synechococcus distribution over the course of the year in Monterey Bay. In the future, we hope to deploy more physical and chemical sensors, including phosphate and ammonia sensors, in order to further resolve the environmental context during these time series.
Synechococcus populations and environmental parameters. Bubble plot of Synechococcus total abundances (that is, rbcL copy numbers) in the spring (dark outlined) and fall (no outline) vs nitrate and chlorophyll-a concentrations. Size of bubble corresponds to the population size; brightness corresponds with time (the darker the bubble, the earlier in the deployment). Bubbles with dotted outlines correspond to time points corresponding with the period of upwelling relaxation in the spring. Plus signs correspond with chlorophyll-a and nitrate concentrations sampled by the MBARI Monterey Bay time series at Station M0 over the course of 4 years with 3-week resolution. Dotted box indicates the range of chlorophyll-a and nitrate concentrations where we sampled abundant Synechococcus.
Only a narrow range of chlorophyll-a and nitrate concentrations were sampled during the spring and fall ESP series, representing 18.5% of the range sampled by MBARI's Biological Oceanography Group at station M0 over the course of 4 years (2006–2010; Figure 5). We found that Synechococcus abundances were highest over a predictable range, specifically less than 16 μM nitrate and 15 μg l−1 chlorophyll-a. The only exceptions to this included the period of upwelling relaxation in the spring when conditions were still erratic and Synechococcus was not yet abundant (demarcated in Figure 1a, and corresponding to the few small dashed circles within the box in Figure 5), and a single time point in the fall where chlorophyll concentrations were highest (26.6 μg l−1). It appears that the environmental parameters recorded during our deployments were more extreme than typical conditions at this coastal station, and we may have sampled the upper limits of Synechococcus distribution with respect to nitrate and chlorophyll-a concentration at Station M0. We intend to continue deploying over various times of year in order to sample a higher diversity of conditions, to gather data on Synechococcus distributions under more common circumstances and to test whether the observed trends hold true.
Previous time series in Monterey Bay demonstrates the degree of variation observed for Synechococcus populations over the year, with total abundances ranging from undetectable to 105 copies per ml seawater, and peak abundances occurring over the summer and throughout the fall and winter (Paerl et al., in press). Daily sampling with the ESP further resolves Synechococcus population dynamics, and it is apparent that the population does not maintain continuously high abundances over the summer and fall periods in Monterey Bay.
Synechococcus clades I and IV appear to be common in temperate coastal waters (Paerl et al., in press; Zwirglmaier et al., 2008). In the fall of our time series, narB was used to differentiate between the dominant Synechococcus groups C_C1* (part of clade I) and D_C2 (of clade IV). Previously in Monterey Bay, these groups maintained relatively high abundances, with subgroup D_C2 being slightly more abundant during 2006 and 2008 (Paerl et al., in press). These narB subgroups were also found to be most abundant in the coastal-transition and upwelling zones of the California Current System, and low in abundance in oligotrophic waters further offshore (Paerl et al., 2011).
Unlike average conditions in the previous study, here we find that group C_C1* (clade I) is slightly numerically dominant over clade D_C2. Representative Synechococcus strains from clade I and IV differ in their gene repertoires (Dufresne et al., 2008), so there is the genetic potential for the presence of these different subgroups to have implications for ecosystem function or nitrogen cycling in this region. Both groups are lower in abundance in offshore waters (Figure 6a) and low-salinity waters (Figure 6b). This contradicts what might be expected based on previously described higher distributions of Synechococcus in offshore waters (Paerl et al., in press), though a likely explanation is that these sampled waters were not from CCS transition waters. The relative population sizes of each subclade demonstrate no clear differences with respect to temperature or salinity preferences for either subclade (Figure 6b). Though these were the only subclades quantified, the rbcL copy number remains high when narB groups C_C1* and D_C2 numbers decrease, indicating the presence of additional subclade(s) in offshore waters.
Synechococcus subclade dynamics. (a) Synechococcus clade abundances over time. narB, group D_C2 (black) and group C_C1* (grey) gene copy numbers correspond to the Synechococcus subclades enumerated in the fall (from September to October, 2009). Arrow points to time of offshore water influx. (b) Synechococcus total (filled circles) and subclade (black solid and dotted open circles) abundances with respect to temperature and salinity in the fall.
Conclusions
Owing to high-resolution sampling, we have demonstrated that high Thaumarchaea abundances can be seen in the stratified fall surface waters, and that these abundances can exceed Thaumarchaea numbers transported to surface waters during active upwelling. Such high abundances of ammonia oxidizers likely have significant implications for local nitrogen cycling. If the mass of data we retrieved from the Thaumarchaeaota in the fall series were instead a single data point in 1 month, over the course of a year time series, we may have disregarded it as an outlier, as the conditions corresponding with these high abundances are not conditions that normally coincide with abundant Thaumarchaeota. These data demonstrate that Thaumarchaeota reside with abundant phytoplankton in the surface waters over long periods of time; however, it is currently unknown whether this co-occurrence is typical in stratified waters in Monterey Bay. It is now possible to test this theory by sampling high and low chlorophyll regimes within the same water mass during stratified conditions, using adaptive sampling with robotic instrumentation.
The daily resolution of the ESP also allowed us to demonstrate higher variability in Synechococcus populations in the fall than anticipated based on previous time series with approximately monthly resolution. Due to the high number of samples within this season, we were successful in linking low Synechococcus numbers to high chlorophyll concentration, indicating a possible inverse relationship between bloom conditions and Synechococcus abundances. Here, we have demonstrated the deeper insight gained through high-resolution biological sensing, without the need for continuous on-station ship-time.
The sampling frequencies of most time series programs necessitate a more general interpretation of microbial dynamics and though seasonal trends may be investigated, factors that are important for biogeochemistry are easily obscured by the low likelihood of their continued observation due to intermittent sampling. We have shown that the factors driving Synechococcus and Thaumarchaea population dynamics changed between seasons, and such effects that occur over shorter scales can be easily overlooked when only investigating the dominant drivers on annual or interannual cycles.
References
Behrenfeld M, Boss E . (2006). Beam attenuation and chlorophyll concentration as alternative optical indices of phytoplankton biomass. J Mar Res 64: 431–451.
Beman JM, Popp BN, Francis CA . (2008). Molecular and biogeochemical evidence for ammonia oxidation by marine Crenarchaeota in the Gulf of California. ISME J 2: 429–441.
Bolin R, Abbott D . (1963). Studies on the marine climate and phytoplankton of the central coastal area of California. California Cooperative Oceanic Fisheries InvestigatIons, Progress Report (1963) 9: 23–45.
Breaker L, Broenkow W . (1994). The circulation of Monterey Bay and related processes. Oceanogr Mar Biol Ann Rev 32: 1–64.
Brochier-Armanet C, Boussau B, Gribaldo S, Forterre P . (2008). Mesophilic Crenarchaeota: proposal for a third archaeal phylum, the Thaumarchaeota. Nat Rev Microbiol 6: 245–252.
Chavez FP . (1999). Biological and chemical response of the equatorial Pacific Ocean to the 1997–98 El Niño. Science 286: 2126–2131.
Chavez FP, Ryan J, Lluch-Cota SE, iquen CM . (2003). From anchovies to sardines and back: multidecadal change in the Pacific Ocean. Science 299: 217–221.
Doney SC . (1999). Major challenges confronting marine biogeochemical modeling. Global Biogeochem Cycles 13: 705–714.
Doucette GJ, Mikulski CM, Jones KL, King KL, Greenfield DI, Marin R et al. (2009). Remote, subsurface detection of the algal toxin domoic acid onboard the environmental sample processor: assay development and field trials. Harmful Algae 8: 880–888.
Dufresne A, Ostrowski M, Scanlan DJ, Garczarek L, Mazard S, Palenik BP et al. (2008). Unraveling the genomic mosaic of a ubiquitous genus of marine cyanobacteria. Genome Biol 9: R90.
Dugdale RC, Goering JJ . (1967). Uptake of new and regenerated forms of nitrogen in primary productivity. Limnol Oceanogr 12: 196–206.
Francis CA, Roberts KJ, Beman JM, Santoro AE, Oakley BB . (2005). Ubiquity and diversity of ammonia-oxidizing archaea in water columns and sediments of the ocean. Proc Natl Acad Sci USA 102: 14683–14688.
Goericke R, Welschmeyer NA . (1993). The marine prochlorophyte Prochlorococcus contributes significantly to phytoplankton biomass and primary production in the Sargasso Sea. Deep Sea Res Oceanogr Res Pap 40: 2283–2294.
Goffredi SK, Jones WJ, Scholin CA, Marin III R, Vrijenhoek RC . (2006). Molecular detection of marine invertebrate larvae. Mar Biotechnol (NY) 8: 149–160.
Green RE, Sosik HM, Olson RJ . (2003). Contributions of Phytoplankton and other particles to inherent optical properties in New England continental shelf waters. Limnol Oceanogr 48: 2377–2391.
Green RE, Sosik HM . (2004). Analysis of apparent optical properties and ocean color models using measurements of seawater constituents in New England continental shelf surface waters. J Geophys Res 109: C03026.
Greenfield DI, Marin R, Doucette GJ, Mikulski C, Jones K, Jensen S et al. (2008). Field applications of the second-generation environmental sample processor (ESP) for remote detection of harmful algae: 2006–2007. Limnol Oceanogr Methods 6: 667–679.
Hallam SJ, Mincer TJ, Schleper C, Preston CM, Roberts K, Richardson PM . (2006). Pathways of carbon assimilation and ammonia oxidation suggested by environmental genomic analyses of marine Crenarchaeota. PLoS Biol 4: e95.
Haywood AJ, Scholin CA, Marin III R, Steidinger KA, Heil C, Ray J . (2007). Molecular detection of the brevetoxin-producing dinoflagellate Karenia brevis and closely related species using rRNA-targeted probes and a semiautomated sandwich hybridization assay. J Phycol 43: 1271–1286.
Johnson KS, Berelson WM, Boss ES, Chase Z, Claustre H, Emerson SR et al. (2009). Observing biogeochemical cycles t global scales with profiling floats and gliders. Oceanography 22: 216–225.
Johnson K, Coletti L, Chavez F . (2006). Diel nitrate cycles observed with in situ sensors predict monthly and annual new production. Deep Sea Res Oceanogr Res Pap 53: 561–573.
Johnson KS, Coletti LJ . (2002). In situ ultraviolet spectrophotometry for high resolution and long-term monitoring of nitrate, bromide and bisulfide in the ocean. Deep Sea Res Oceanogr Res Pap 49: 1291–1305.
Karl DM, Letelier R, Hebel D, Tupas L, Dore J, Christian J et al. (1995). Ecosystem changes in the North Pacific subtropical gyre attributed to the 1991–92 El Nino. Nature 373: 230–234.
Karl DM . (1999). Minireviews: a sea of change: biogeochemical variability in the North Pacific subtropical gyre. Ecosystems 2: 181–214.
Karl DM, Dore JE . (2001). Microbial ecology at sea: sampling, subsampling, and incubation considerations. Methods Microbiol 30: 13–27.
Karl DM . (2007). Microbial oceanography: paradigms, processes and promise. Nat Rev Microbiol 5: 759–769.
Karner MB, DeLong EF, Karl DM . (2001). Archaeal dominance in the mesopelagic zone of the Pacific Ocean. Nature 409: 507–510.
Könneke M, Bernhard AE, de la Torre JR, Walker CB, Waterbury JB, Stahl DA . (2005). Isolation of an autotrophic ammonia-oxidizing marine archaeon. Nature 437: 543–546.
L’Helguen S, Maguer J-F, Caradec J . (2008). Inhibition kinetics of nitrate uptake by ammonium in size-fractionated oceanic phytoplankton communities: implications for new production and f-ratio estimates. J Plankton Res 30: 1179–1188.
Liu HB, Nolla HA, Campbell L . (1997). Prochlorococcus growth rate and contribution to primary production in the equatorial and subtropical North Pacific Ocean. Aquat Microbial Ecol 12: 39–47.
Massana R, Murray A, Preston C, DeLong E . (1997). Vertical distribution and phylogenetic characterization of marine planktonic Archaea in the Santa Barbara channel. Appl Environ Microbiol 63: 50–56.
McGowan JA, Cayan DR, Dorman LM . (1998). Climate-ocean variability and ecosystem response in the Northeast Pacific. Science 281: 210–217.
Mincer TJ, Church MJ, Taylor LT, Preston C, Karl DM, DeLong EF . (2007). Quantitative distribution of presumptive archaeal and bacterial nitrifiers in Monterey Bay and the North Pacific Subtropical Gyre. Environ Microbiol 9: 1162–1175.
Mosier AC, Francis CA . (2011). Determining the distribution of marine and coastal ammonia-oxidizing archaea and bacteria using a quantitative approach. In: Martin GK (ed). Methods in Enzymology. Academic Press: New York, pp 205–221.
Murray AE, Blakis A, Massana R, Strawzewski S, Passow U, Alldredge A et al. (1999). A time series assessment of planktonic archaeal variability in the Santa Barbara Channel. Aquat Microbial Ecol 20: 129–145.
Ottesen EA, Marin III R, Preston CM, Young CR, Ryan JP, Scholin CA et al. (2011). Metatranscriptomic analysis of autonomously collected and preserved marine bacterioplankton. ISME J; e-pub ahead of print 30 June 2011; doi: 10.1038/ismej.2011.70.
Paerl RW, Foster RA, Jenkins BD, Montoya JP, Zehr JP . (2008). Phylogenetic diversity of cyanobacterial narB genes from various marine habitats. Environ Microbiol 10: 3377–3387.
Paerl RW, Johnson KS, Welsh RM, Worden AZ, Chavez FP, Zehr JP . (2011). Differential distributions of Synechococcus subgroups across the California current system. Front Microbiol 2: 59.
Paerl RW, Turk KA, Beinart RA, Chavez FP, Zehr JP . (in press). Seasonal change in the abundance of Synechococcus and multiple distinct phylotypes in Monterey determined by rbcL and narB quantitive PCR. Environ Microbiol.
Palenik B, Ren Q, Tai V, Paulsen IT . (2009). Coastal Synechococcus metagenome reveals major roles for horizontal gene transfer and plasmids in population diversity. Environ Microbiol 11: 349–359.
Paul J, Scholin C, Van Den Engh G, Perry MJ . (2007). In situ instrumentation. Oceanography 20: 70–78.
Pennington JT, Chavez FP . (2000). Seasonal fluctuations of temperature, salinity, nitrate, chlorophyll and primary production at station H3/M1 over 1989–1996 in Monterey Bay, California. Deep Sea Res Topical Stud Oceanogr 47: 947–973.
Preston C, Harris A, Ryan JP, Roman B, Marin R, Jensen S et al. (2011). Application of quantitative PCR on a coastal mooring. PLOS One 6: e22522.
Preston CM, Marin R, Jensen SD, Feldman J, Birch JM, Massion EI et al. (2009). Near real-time, autonomous detection of marine bacterioplankton on a coastal mooring in Monterey Bay, California, using rRNA-targeted DNA probes. Environ Microbiol 11: 1168–1180.
Santoro AE, Casciotti KL, Francis CA . (2010). Activity, abundance and diversity of nitrifying archaea and bacteria in the central California Current. Environ Microbiol 12: 1989–2006.
Scanlan DJ, West NJ . (2002). Molecular ecology of the marine cyanobacterial genera Prochlorococcus and Synechococcus. Fems Microbiol Ecol 40: 1–12.
Scholin C, Doucette G, Jensen S, Roman B, Pargett D, Marin R et al. (2009). Remote detection of marine microbes, small Invertebrates, harmful algae, and biotoxins using the environmental sample processor (Esp). Oceanography 22: 158–167.
Scholin C . (2010). What are ‘ecogenomic sensors?’ A review and thoughts for the future. Ocean Sci 6: 51–60.
Shade A, Carey CC, Kara E, Bertilsson S, McMahon KD, Smith MC . (2009). Can the black box be cracked? The augmentation of microbial ecology by high-resolution, automated sensing technologies. ISME J 3: 881–888.
Spang A, Hatzenpichler R, Brochier-Armanet C, Rattei T, Tischler P, Spieck E et al. (2010). Distinct gene set in two different lineages of ammonia-oxidizing archaea supports the phylum Thaumarchaeota. Trends Microbiol 18: 331–340.
Suzuki MT, Taylor LT, DeLong EF . (2000). Quantitative analysis of small-subunit rRNA genes in mixed microbial populations via 5'-nuclease assays. Appl Environ Microbiol 66: 4605–4614.
Tai V, Palenik B . (2009). Temporal variation of Synechococcus clades at a coastal Pacific Ocean monitoring site. ISME J 3: 903–915.
Toledo G, Palenik B . (1997). Synechococcus diversity in the California current as seen by RNA polymerase (rpoC1) gene sequences of isolated strains. Appl Environ Microbiol 63: 4298–4303.
Wilhelm SW, Trick CG . (1994). Iron-limited growth of Cyanobacteria - multiple siderophore production is a common response. Limnol Oceanogr 39: 1979–1984.
Yool A, Martin AP, Fernandez C, Clark DR . (2007). The significance of nitrification for oceanic new production. Nature 447: 999–1002.
Zehr JP, Waterbury JB, Turner PJ, Montoya JP, Omoregie E, Steward GF et al. (2001). Unicellular cyanobacteria fix N-2 in the subtropical North Pacific Ocean. Nature 412: 635–638.
Zwirglmaier K, Jardillier L, Ostrowski M, Mazard S, Garczarek L, Vaulot D et al. (2008). Global phylogeography of marine Synechococcus and Prochlorococcus reveals a distinct partitioning of lineages among oceanic biomes. Environ Microbiol 10: 147–161.
Acknowledgements
This work was partially funded by the MEGAMER facility grant by the Gordon and Betty Moore Foundation and the NSF Center for Microbial Oceanography, Research and Education (C-MORE) (to JPZ). Development and application of the ESP has been funded in part by grants (to CAS) from the David and Lucille Packard Foundation (through funds allocated to MBARI), NSF, NASA Astrobiology, and the Gordon and Betty Moore Foundation. Work was also supported in part by an EPA STAR Graduate Fellowship and Stanford University DARE Fellowship (to ACM), as well a National Science Foundation Grant (OCE-0825363) to CAF. We thank the science and engineering technicians and machinists at MBARI (in particular Cheri Everlove and Roman Marin III) for their invaluable help and dedication toward instrument development, and the crew of the R/V Zephyr for their support and expertise during field operations. Thanks also to the MBARI Biological Oceanography Group, especially Reiko Michisaki for help with MBARI mooring data. Raphael Kudela and lab, John Ryan, Heather Kerkering, Tom Wadsworth, Tracy Cote and the NOAA CoastWatch Program and NASA's Goddard Space Flight Center, OceanColor Web provided and aided in analyses. Thanks to Jim Birch for help with manuscript preparations.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Robidart, J., Preston, C., Paerl, R. et al. Seasonal Synechococcus and Thaumarchaeal population dynamics examined with high resolution with remote in situ instrumentation. ISME J 6, 513–523 (2012). https://doi.org/10.1038/ismej.2011.127
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2011.127
Keywords
This article is cited by
-
Spatio-temporal patterns of Synechococcus oligotypes in Moroccan lagoonal environments
Scientific Reports (2023)
-
Role of Bacterial Community Composition as a Driver of the Small-Sized Phytoplankton Community Structure in a Productive Coastal System
Microbial Ecology (2023)
-
Marine archaeal dynamics and interactions with the microbial community over 5 years from surface to seafloor
The ISME Journal (2017)
-
Marine microbial community dynamics and their ecological interpretation
Nature Reviews Microbiology (2015)
-
Single-taxon field measurements of bacterial gene regulation controlling DMSP fate
The ISME Journal (2015)