Abstract
Here we describe, the longest microbial time-series analyzed to date using high-resolution 16S rRNA tag pyrosequencing of samples taken monthly over 6 years at a temperate marine coastal site off Plymouth, UK. Data treatment effected the estimation of community richness over a 6-year period, whereby 8794 operational taxonomic units (OTUs) were identified using single-linkage preclustering and 21 130 OTUs were identified by denoising the data. The Alphaproteobacteria were the most abundant Class, and the most frequently recorded OTUs were members of the Rickettsiales (SAR 11) and Rhodobacteriales. This near-surface ocean bacterial community showed strong repeatable seasonal patterns, which were defined by winter peaks in diversity across all years. Environmental variables explained far more variation in seasonally predictable bacteria than did data on protists or metazoan biomass. Change in day length alone explains >65% of the variance in community diversity. The results suggested that seasonal changes in environmental variables are more important than trophic interactions. Interestingly, microbial association network analysis showed that correlations in abundance were stronger within bacterial taxa rather than between bacteria and eukaryotes, or between bacteria and environmental variables.
Similar content being viewed by others
Introduction
Only recently with the introduction of molecular techniques satisfactory descriptions of natural microbial assemblages have been generated (Fierer and Jackson, 2006; Rusch et al., 2007; Costello et al., 2009; Caporaso et al., 2011). In this paper, we summarize a 6-year time series of 16S rRNA tag pyrosequencing of samples taken from a long-time series station in the English Channel. The aim was to understand seasonal variability and to try to determine which environmental factors might have the greatest influence on the varying diversity. In contrast to terrestrial environments that are essentially static, the marine environment has the added complication that the dispersion and movement of populations will be driven by hydrography. This adds to difficulties of interpretation of results, particularly if the sampling design is Eulurian (a fixed site) rather than Lagrangian (moving with the water flow). The Western English Channel has been studied intensively for more than 100 years (Southward et al., 2005), and this wealth of data provide a robust context with which to explore temporal microbiological complexity. Inferences can be drawn regarding how bacterioplankton assemblages may potentially interact with the environment as well as with specific groups of organisms.
Previous efforts to determine which factors might affect microbial communities have largely focused on the relative importance of temperature and nutrient concentrations (Cullen, 1991; Kirchman et al., 1995; Morris et al., 2005; Fuhrman et al., 2006; Fuhrman, 2009; Gilbert et al., 2009). These are obvious candidates because of the strong effect of temperature on biological processes (Nedwell and Rutter, 1994) and the fact that nutrient availability can drive niche structure through resource partitioning (Church, 2009). Of greatest relevance to the present study is the recent demonstration that bacterioplankton diversity followed a latitudinal gradient, with maximum potential richness being primarily driven by temperature, with many other factors modulating an intricate network of richness at any particular temperature (Fuhrman et al., 2008).
The aim of the current study was to further characterize seasonal patterns of bacterioplankton diversity in the Western English Channel, beyond an initial 1-year study by Gilbert et al. (2009). Using these data, we tested three competing alternative hypotheses about potential drivers of diversity patterns, namely whether the observed seasonal patterns correlate with (1) varying concentrations of inorganic nutrients, (2) annual water–temperature cycle or (3) the population structure of the eukaryotic phytoplankton and zooplankton. The null hypothesis was that the seasonal patterns in microbial community composition in the Western English Channel showed no relationship with any of the physical or biological factors measured in this study.
Materials and methods
Sampling, DNA extraction, 16S rDNA V6 amplification and pyrosequencing
Seawater samples were collected on 72 instances from January 2003 to December 2008, from the L4 sampling site (50° 15.00′ N, 4° 13.02′) of the Western Channel Observatory (http://www.westernchannelobservatory.org.uk). Sampling, extraction, amplification, and sequencing protocols and environmental parameter analysis were performed simultaneously on the same samples as described previously by Gilbert et al. (2009); extensive information can be found in Supplementary Information (Supplementary Tables S1–S3). Bacterial diversity was examined in the context of the broad range of biotic and abiotic variables that are routinely measured at the Observatory. These included phytoplankton and zooplankton species abundance, the concentrations of ammonia, nitrate+nitrite, phosphate, silicate, total organic carbon and nitrogen, salinity, chlorophyll, photosynthetically active radiation, North Atlantic Oscillation data, day length, primary productivity and temperature. Statistical analyses used the routines of PRIMER (Clarke and Warwick, 2001; Clarke and Gorley, 2006).
Sequence data analysis
All sequence data were treated as reported previously (Gilbert et al., 2010), using the same quality control that included random resampling to standardize the sequencing effort as described below, Sequence data noise reduction using Single-Linkage Preclustering (SLP; Huse et al., 2010) and analysis (sample similarity derived from Bray–Curtis indices weighted on taxon abundance matrices) also followed previous protocols. In addition, several noise reduction strategies such as SLP (Huse et al., 2010) and denoiser (Reeder and Knight, 2010) were compared to examine the impact of pyrosequencing errors on community diversity patterns observed in the data (see Supplementary Figure S1a). It is important to stress that both known and unknown biases associated with these techniques meant that these data could not be seen as quantitative, and hence all analyses are based on relative changes derived through comparison. As the same sequencing and sampling effort was applied to each sample, the operational taxonomic unit (OTU) richness (S) was used as a diversity metric, which showed a 97% correlation to two extrapolative estimators of diversity (Chao1 and Ace) over the 72 samples (Supplementary Figure S1b). Changes in community diversity and relationship to environmental parameters were examined using various nonparametric multivariate methods, discriminant function analysis (DFA), and association networks (see Supplementary Information).
To determine whether microbial communities in the Western English Channel demonstrated seasonal patterns over many years, 747 496 16S rDNA V6 sequences were analyzed, including those previously published for the year 2007 (Gilbert et al., 2009). To compensate for potential overestimation in diversity resulting from pyrosequencing and amplification errors, a clustering technique was used. SLP grouped OTUs at 2% sequence identity and an average-linkage clustering followed, based on pair-wise alignments (Huse et al., 2010), which resulted in 8794 OTUs. To remove sequencing effort bias, each sample was randomly resampled to the smallest individual sample sequencing effort (4505) as described before (Gilbert et al., 2009). This resulted in a total of 4204 OTUs (for all 72 samples combined). Approximately, 53% of the OTUs were represented by only a single sequence (singletons). These results, in terms of relative abundance, were confirmed using a second denoising technique, Denoiser (Reeder and Knight, 2010), which generated greater total richness (21 130 OTUs). However, comparison between Denoiser, SLP and no-denoising/filtering indicated that overall, the same patterns of community diversity were evident with each technique (Supplementary Figure S1). SLP constituted by far the most conservative OTU predictions, and was therefore used for subsequent analysis.
Results
Seasonal variations in diversity and persistence
Bacterioplankton were very diverse at this station and a total of 8794 different OTUs (defined using SLP) over a 6-year period were identified. Figure 1 summarizes the taxonomic identify of all the OTUs sequenced and also gives an indication of the persistence of OTUs in microbial communities at L4 over a 6-year time period. Although this study has shown high diversity of bacterioplankton in the English Channel, as with other studies of natural assemblages, the majority of sequences could not be identified to species. Indeed, only 6 of the 10 most abundant OTUs could be annotated below the level of Class and, of the top 100 most abundant OTUs, only 2% could be identified to the species level. The taxonomic level to which the OTUs could be identified was—Phylum (9%), Class (32%), Order (10%), Family (26%), Genus (21%). This was true using a number of different annotation strategies (that is, GAST (Sogin et al., 2006); BLAST against Greengenes (DeSantis et al., 2006), SILVA (Pruesse et al., 2007) and RDP (Maidak et al., 2001); RDP classifier (Maidak et al., 2001); data not shown, references in Supplementary Information). These results suggest that a large fraction of as-of-yet uncharacterized lineages were present, even among the most abundant taxa, and highlights the difficulties associated with accurate annotation of short read-length tag sequences from hypervariable 16S rRNA regions (Wang et al., 2007; Liu et al., 2008).
Persistence of OTUs in microbial communities at L4 over a 6-year time period. Median OTU abundance, calculated for all time points, over a 6-year period is set proportional to node size on a logarithmic scale. Only OTUs found in at least 5% of the time-series samples (⩾4) are shown. This includes 22.53% of the OTUs, representing 97.48% of the sampled organisms. Node coloring shows the differences in persistence over time, with the color scale from orange (5%), yellow (16%), green (35%), blue (66%), red (100%) reflecting increasing persistence.
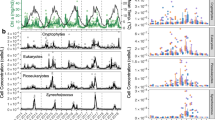
Although there are significant seasonal variations in OTU frequency throughout a 6-year period (Figure 2), there are also strong repeating patterns. As other studies of marine microbial diversity have demonstrated, the Alphaproteobacteria were the most abundant Class. The OTUs most frequently recorded were members of the Rickettsiales and Rhodobacteriales. Other OTUs with high frequency were the Flavobacteriales (Class: Bacteroidetes) and there were also peaks in the Gammaproteobacteria (Vibrionales and Pseudomonadales).
Plot representing the seasonal dynamics (grouped as an average of seasons; Winter: January–March; Spring: April–June; Summer: July–September; Fall: October–December) of taxa grouped at the taxonomic level of Order in the L4 6-year time series. Frequency is recorded based on abundances within a resampled abundance of 4101 sequences per sample. Only Orders whose average frequency peaked above 10% of the resampled community abundance were included.
Alpha diversity of the observed OTUs (S) was relatively constant across the time series, but showed distinct cyclical patterns with maxima in winter and minima in summer (Figure 3). The mean S per time point was 286, with an average minimum of 179 in summer and maximum of 437 in winter. This pattern was further confirmed by permutation-based analysis of variance (of S) for all taxa, and for a range of phyla (Supplementary Table S4). S was most similar when comparing the same time of year, and differences between seasons and among years were both highly significant. Seasonal differences tended to be greater than inter-annual (greater pseudo-F values although there were fewer d.f.). This lack of significant interaction terms suggested that the seasonal cycle was consistent across years. Overall persistence (Figure 1) was linked to abundance; OTUs that were present at more than three time points accounted for 97.48% of the sequences. In total, only 12 OTUs were found at every 1 of the 72 time-points, yet these were exceptionally abundant, comprising ∼35% of all the sequence reads.
Seasonal trends in most abundant bacteria
The two most abundant Orders were Rickettsiales and Rhodobacterales, and they had different seasonal abundances. The Rickettsiales sequences were dominated by the SAR11 clade and tended to peak in winter (Figure 4). At this time, light and primary production were low, and inorganic nutrient concentrations were at their maximum. In contrast, the Rhodobacterales, which were dominated by the Roseobacter clade, tended to peak in Spring and Autumn, when nutrient concentrations were lower yet primary productivity was higher. This is consistent with what is known from single-strain-level studies; SAR11 are considered to be obligate oligotrophs, while the Roseobacter clade contains many genera whose cultured representatives tend to grow in organic nutrient-rich media, and may be likely to respond at times when rates of primary production are higher.
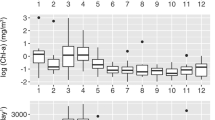
Plot representing the seasonal dynamics of the bacterial Orders, Rickettsiales and Rhodobacterales, and environmental parameters, chlorophyll a and soluble reactive phosphorus (SRP) in the L4 6-year time series. Frequency is recorded based on abundances (abundance of sequences per taxa) within a resampled abundance of 4505 sequences per sample.
Rare taxa may dominate the assemblage
The largest bacterial ‘bloom’ occurred during August 2003, and this constituted a single Vibrio sp., which represented 54% of the sequences. Yet, for the rest of the time series, this taxon was relatively rare, having an abundance of 0–2%. Interestingly, this peak was correlated with an increase in the relative abundance of the diatom, Chaetoceros compressus. This diatom was also typically present at low abundance, between 0.002–0.2% of total phytoplankton biomass (Supplementary Table S2). However, in August 2003, C. compressus accounted for 1.2% of total eukaryotic phytoplankton. Our data do not distinguish between a causal relationship—a specific dependence of a bacterial species on a specific phytoplankton species—and simple co-occurrence, which might be a response to unusual environmental conditions. Certainly at this time point, the highest total organic nitrogen and carbon concentrations, and second highest chlorophyll a concentration were measured in the whole time series between 2003 and 2008 (Supplementary Table S1).
Seasonal succession in the community composition is robust
The dataset of environmental and biological variables was examined to investigate potential relationships between bacterioplankton and the environmental and eukaryotic abundance data. The community composition (rather than richness) was used, after determining whether seasonal patterns in community composition were as robust as those for species richness. Three different subsets of the bacterial OTUs, that is, the most abundant, most common and most variable (see Supplementary Materials) were defined. These definitions were robust across the different denoising strategies (that is, the same OTUs (based on sequence identity, with the same taxonomic inference defined). Using DFA, an eigenvector technique that, in this case, searches out the taxa which are best able to predict the month (Fuhrman et al., 2006), we found that for each subset, the bacterial community could correctly predict the month with 100% accuracy, showed a clear repeating pattern (Figure 5), and was able to explain >60% of the variance in the community structure (Supplementary Table S5).
Annual repeating patterns from the bacterioplankton community sampled monthly from 2003–2008 in the English Channel determined by DFA where the model used the bacterioplankton community to predict the month. Upper row of graphs shows the time-series analysis of the first discriminant function (DFA1) over 72 months. The lower row shows the autocorrelation of the discriminant function with up to a 50-month lag. The lines in the lower row represent correlations with P<0.05.
These patterns for most abundant, common and variable subsets are similar to those reported for similar subsets in a Californian near-surface bacterioplankton time series (Fuhrman et al., 2006), suggesting that seasonal succession patterns of marine surface water bacterial communities in temperate regions may be conserved across different biomes. The Californian study was based on automated ribosomal RNA intergenic spacer analysis fingerprint technology, but the sequence-based annotation provided by this study allowed considerably better predictions for the bacterial taxa contributing most strongly to these signals. In this instance, these were members of the Alphaproteobacteria (for example, SAR11 and Rhodobacteriaciae groups), the Gammaproteobacteria (for example, Pseudomonas, Pseudoalteromonas, and Vibrio groups), the Cyanobacteria, and the Bacteroidetes (for example, Flavobacteriaceae group; Supplementary Table S6).
Seasonal variance in community composition
The relative significance of environmental versus biological factors in describing the seasonal variation in bacterioplankton assemblages was investigated using DFA. DFA, via multiple regression using environmental factors and eukaryotic counts, was used to predict the first discriminant function (DF1) from each subset of the community (that is, most abundant, most common and most variable). Environmental parameters explained 49–91% of the variance in DF1, while eukaryotic variables explained 18–51% of the variance (Supplementary Table S6). This suggests that that the seasonally responsive members of the microbial community were responding to changing environmental factors, while interactions between the bacteria and the eukaryotes may have had a less comprehensive influence. Obviously, as shown for the Vibrio bloom in 2003, this trend is not absolutely uniform, and blooms of rare taxa can be influenced by the presence of eukaryotes. However, as defined by the robust annual cyclicity, the community recovers from these ‘rare-bloom’ events, suggesting an overall bottom-up influence on the community composition and structure. Essentially this suggests that nutrient concentrations, physical parameters and biology all demonstrate significant influence in an extraordinarily complex matrix.
Annual day length cycle explains most of the variability in the seasonal pattern of species diversity
To test whether changes in nutrients or temperature provided the best correlation with changes in community diversity, distance-based linear modelling was used (described in detail in Supplementary Material). This showed that, although a significant fit could be ascribed to a combination of temperature and photosynthetically active radiation and the richness of all OTUs, the most significant fit was always to the annual change in day length (Supplementary Table S7). This was best modelled by a cosine term (DX1) with the peak centered on December 22. When day length (DX1) was combined with serial day (D), it described 66.3% of the variance in OTU richness. However, when examining the phototrophic Cyanobacteria (Supplementary Table S7), the relationship of richness to day length was not always evident, for example, diversity peaked in spring but not in winter, and hence coincided with the lowest annual temperatures at L4. To account for the Cyanobacteria and to significantly improve the fit of our model (δAIC> −2), a second seasonal artificial term centered on March 22 (a sine-derived term—DX2) was added that closely tracked temperature. Also, because most of the taxa show subtle changes in their seasonal cyclicity over these years, it was possible to significantly improve the model further by adding a linear time trend term (D). However, this did not improve the fit for the cyanobacterial community diversity, which was remarkably stable over the 6 years. Strikingly, the Cyanobacteria were unique in that a combination of photosynthetically active radiation, temperature and nitrate/nitrite concentration provided as good a fit as the artificial descriptors (DX1, DX2 and D; Supplementary Table S7). Not unexpectedly, this suggests that, unlike other groups, the species diversity of these primary producers can be well defined by a combination of light availability, nitrogen availability and temperature, reflecting a different set of niches compared with the other potentially heterotrophic bacterioplankton.
Discussion
The repeating cycles in bacterioplankton diversity in this Eulerian study raise the question of whether unique water masses pass through the English Channel, and whether those water masses contain characteristic bacterioplankton assemblages. This is almost certainly not the case as the hydrography of the Western English Channel has been studied extensively (Southward et al., 2005). From the earliest studies in the 1930s using drift-bottles, it was known that there was a strong flow through the English Channel from west to east. Later modelling and observational studies showed the importance of wind over a very wide shelf region (including the North Sea) in determining flow through the Western English Channel (Pingree and Griffiths, 1980). Southerly winds resulted in the greatest net transport of water along the English Channel through the Straits of Dover and into the southern North Sea; westerly winds were less effective.
It has recently been calculated that average residence time at the sampling site is on the order of 2 weeks (Lewis and Allen, 2009), although dispersion occurs continuously. The repeating annual patterns of bacterioplankton demonstrated in this study cannot be due to the repeated intrusion of water mass with an annual periodicity. We do not know how representative these robust annual patterns are of the entire English Channel. It may be that the observed patterns represent seasonal changes in bacterioplankton on the Celtic Sea Shelf, which is advected into the Western English Channel. Given that this advection will largely depend on wind conditions, it seems unlikely that such similar patterns would occur over a 6-year period. Clearly, further sampling on the European Shelf will be required to answer the question of the representativeness of this station.
The relationship between OTU richness and day length is interesting. To the best of our knowledge, this is the only example from a marine dataset where a single variable has such explanatory capacity (66.3% of the variance in OTU richness). There are examples from terrestrial systems; for example, tRFLP analysis identified an r2 value of 0.7 between bacterial community richness and pH (Fierer and Jackson, 2006). Temperature would imply a clear mechanism; we can see no such direct mechanism that result in day length directly controlling bacterioplankton assemblages.
Other environmental factors that could suggest direct mechanisms did have significant relationships. They did not, however, apply to the most common and abundant taxa, but the composition of the most variable taxa could be significantly predicted by nutrient concentrations (NH4+, total organic nitrogen (TON), soluble reactive phosphate, primary production and broad shifts in ocean currents indicated by the North Atlantic Oscillation (Supplementary Table S6). Overall, we conclude that the monthly pattern and response to broad seasonal changes indicate that the most common and most abundant bacterial OTUs have temporally defined niches. In contrast, the most variable OTUs have niches that can be defined temporally as well as by nutrient pulses and changes in currents. Temporal niche structure suggests taxa with a resilient seasonal pattern, for example, SAR11 and Rhodobacteriaciae, although tracking nutrient pulses and currents, are potentially less resilient to changing environmental conditions. However, the relationship is complex, and potentially a function of abundance, commonality and variability, as both SAR11 and Rhodobacteriaciae are in the most abundant, most common and most variable subset.
Interestingly, interactions were strongest within the bacterial and eukaryotic domains rather than between them, and relationships were stronger between bacterial taxa than with environmental variables. Association network analysis was employed in an attempt to deconvolute the complex network of relationships that were driving the observed DFA results. However, this revealed that the strongest correlations exist between bacterial OTUs (whether abundant, common or variable) and, to a slightly lesser extent among eukaryotes, compared with correlations between these two domains or between either bacteria or eukaryotes and environmental factors (Figure 6). Also, the integrity of these relationships was maintained across the three chosen subsets of OTUs (Figure 6). Even among the highly variable OTUs, which might be expected to respond to changing conditions enabling growth from rare to abundant, most significant correlations were still between bacteria (Supplementary Figure S2). Also, at a highly correlated (r>0.7, P<0.001, q<0.0012) level, there were many eukaryotic taxa in a loosely intercorrelated group (Supplementary Figure S2a), but there are still very few specific connections between the eukaryotes and the bacteria. Mostly the bacteria were correlated to one another and to the environmental factors, and the eukaryotes were also connected to one another and the environmental factors. The highly intercorrelated group (Supplementary Figure S2b) was almost completely devoid of eukaryotes, but was connected to an herbivorous, parasitic copepod (Poescilostomatoida), and to the seasonal factor DX1, NO2+NO3, and an interconnected cluster of Gammaproteobacteria, Bacillus and Actinobacteria OTUs.
Broad view of correlation network for the microbial community and the environment at station L4. The network shows strong correlations (r>0.8, P<0.001, q<0.002) between microbial and environmental parameters for the 300 most abundant bacterial taxa (a), the 300 most common bacterial taxa (b), and the 300 most variable bacterial taxa (c). Bacteria are shown in blue, eukaryotes are shown in red and environmental variables are shown in yellow.
Interactions between eukaryotes and bacteria became more apparent when moderate correlations were examined between different subsets of the eukaryotic community and the 300 most abundant bacterial OTUs. Mixotrophic eukaryotes (potential grazers on bacteria) and autotrophic eukaryotes both showed complex interactions with the prokaryotic community (Supplementary Figure S3). Although flagellates (when grouped by size) were correlated to each other (r=0.59, P<0.001, q<0.0012) and, naturally, to the total number of flagellates, only two bacterial OTUs (a single Rhodobactereaceae OTU and a single Cyanobacteria OTU) are correlated to all three groups (Supplementary Figure S3a). There were many bacterial and eukaryotic OTUs, which correlate to two of the flagellate subgroups, and a smaller number, which correlate to only one of the flagellate subgroups. The 5 μm flagellates were negatively correlated to a Betaproteobacterial and a Gammaproteobacterial OTU, and the diatom Paralia sulcata in samples with a 1-month lag, which reflects an increase in those abundant members of the community following a decrease in 5 μm-sized flagellates (Supplementary Figure S3a).
A similar situation was applied to correlations between autotrophic eukaryotes and abundant bacterial OTUs. The diatom, P. sulcata, correlated negatively to the total diatom counts with a 1-month time lag (Supplementary Figure S3b). This may indicate a situation where P. sulcata dominated the diatom community, while the total number of diatoms decreased. These two eukaryotic nodes shared 26 bacterial OTUs that correlated positively to P. sulcata and negatively with a 1-month time lag to the total diatom count (Supplementary Figure S3b). These bacterial OTUs may reflect a community shift indicated by the increase of P. sulcata and the 26 Proteobacteria, Bacteroidetes and Verrucomicrobia when the total number of diatoms decreased. The winter peak seasonal cycle, DX1, also positively correlated to P. sulcata and negatively correlated, with a 1-month lag, to total diatoms in the same way, possibly implying seasonal community succession. There were positive contemporaneous correlations between P. sulcata and NO3+NO2, between silicate and mixed layer depth, and a negative 1-month lagged correlation between the North Atlantic Oscillation and total diatom counts; these results indicate that nutrient concentrations may be drivers of this succession (Supplementary Figure S3b). Interestingly, there were only positive correlations between bacterial OTUs and 2 μm flagellates (Supplementary Figure S3a), even though 2 μm flagellates might be expected to be the major grazers of bacterioplankton. Bacterial OTUs were also positively correlated to total flagellates, total phytoplankton, coccolithophores and Emiliania huxleyi (Supplementary Figure S3b).
Many environmental factors were highly correlated (r>0.7, P<0.001, q<0.0012) with both eukaryotic OTUs and bacterial OTUs, when both the 300 most variable bacteria (Supplementary Figure S4a) and the 300 most common bacteria (Supplementary Figure S4b) were considered. Strikingly, the seasonal index peaking in winter (DX1) was correlated almost exclusively to bacterial OTUs, including Proteobacteria (for example, Alphaproteobacteria, Gammaproteobacteria, Nitrospira), unidentified bacteria, Deferribacteres and Owenweeksia in both the common and variable sub-networks (Supplementary Figure S4). Cladocera and Echinodermata were the only eukaryotes that connected to DX1 and they were negatively correlated with no lag and a 1-month lag, respectively. This suggests that seasonal factors (for example, day length, which is a proxy for DX1) may be more important for the bacterioplankton than for the eukaryotic community. The spring seasonal factor, DX2 was correlated with a 3-month lag to Cladocera (indicating a summer increase in abundance), and was negatively correlated to a Bacteroidetes OTU in the most variable subset (Supplementary Figure S4a).
Positive correlations were widespread in the microbe–environment network. Primary production (monthly average) was correlated to total diatoms, total ciliates, total microzooplankton and a Rhodobacteriaceae OTU (which also correlated to daily primary production and temperature). Daily primary production (ML primary production, calculated from observed chlorophyll values and integrated over the observed mixed layer depth) was also positively correlated to total diatoms, total phytoplankton, total ciliates and echinodermata (Supplementary Figure S4). This suggests that, as productivity and nutrients increased, these bacteria and eukaryotes also increased in abundance, that is, these taxa appear to perform best in a productive system. There was little correlation-based evidence for top-down effects in this system, although this may be a function of a lack of resolution of bacterivores among the eukaryotes or perhaps a limitation of this kind of analysis.
Local similarity analysis, with its ability to see time-lagged correlations, also provided insight into the relationships between environmental factors themselves. Although day length was not correlated to temperature at the 0.7 level, the Winter seasonal cycle (DX1) was negatively correlated to day length with no time delay, and to temperature and primary production with a 1-month time delay (Supplementary Figure S4); that is, day length changed seasonally, followed by a change in temperature. DX1 and day length (which was positively correlated to photosynthetically active radiation and primary production) may be serving as combinatory signals of seasonal environmental change, involving factors such as changes in input of energy into the system. These combinatory variables may more closely map the changes in the whole community of bacterioplankton as well as the individual bacterial OTUs connected to them. NO2+NO3 were highly correlated with soluble reactive phosphate and silicate (Supplementary Figure S4). However, soluble reactive phosphate was correlated only to a Gammaproteobacteria OTU and a Rhodobacteriaciae OTU, while silicate was not highly correlated to any bacteria or eukaryotes. NO2+NO3 was positively correlated to 12 bacterial OTUs, which were also positively correlated to DX1, and there were 10 bacterial taxa that were positively correlated solely to NO2+NO3. The close coupling between these taxa and NO2+NO3 (that is, these taxa were only abundant when there was an increased availability of nitrogen) suggests that these taxa may be seasonally nitrogen limited in this ecosystem.
Regardless of the subset of OTUs (for example, abundant, variable, or common) analysed, each subset was able to predict the month. In addition, each of the networks appeared to identify many of the same connections when we examined 300 taxa from any of the subsets (Figure 6, Supplementary Figures S2–S4). OTUs were ranked differently within each subset, but they produced similar patterns, which were nearly identical at the r>0.8 correlation level (Figure 6). This was partly due to the stability of the bacterioplankton community at L4 and the depth of sampling into this community. It was also an effect of using statistical analyses that require a certain number of occurrences in order to detect a pattern; by design, these analyses would ignore the once-a-decade occurrence, for example, the spike in Vibrio spp. abundance in the summer of 2003. However, comparing these subsets allowed for a better sense of the ecology behind these bacterial OTUs. This is demonstrated most clearly when restricting the correlations to the 50 most common and most variable bacterial taxa, and their relationship to environmental factors (Figure 7). For instance, a SAR11 (Alphaproteobacteria_03_2), although common, changed abundance seasonally (it was the 6th most variable bacterial OTU) and increased in abundance when inorganic nutrient concentrations increased. A Rhizobiales member (Alphaproteobacteria_03_121) that correlated with NO2+NO3 (Figure 7a) was not as variable (Figure 7b), whereas the Deferribacteres member (Deferribacteres_03_12) that correlated with NO2+NO3 (Figure 7b) was not common (Figure 7a), but increased in abundance along with increased NO2+NO3 concentration. Among these observations of common influence, there were also hints at ecological differences between these OTUs. Although some taxa seemed to follow inorganic nutrient concentrations (for example, SAR11 and Deferribacteres), others followed system productivity (for example, Rhodobacteriales) or temperature (Gammaproteobacteria OTUs; Figure 7). These observations, made possible by extended studies of microbial assemblages, will lead to deeper understanding of microbial niches in the ocean and elsewhere.
Sub-networks of highly correlated (r>0.7, P<0.001) variables built around environmental factors from the 50 most common (a) and 50 most variable (b) bacterial OTUs. Interactions between environmental variables and eukaryotic interactions with environmental variables have been removed for clarity. OTU identifications are from http://vampsarchive.mbl.edu/diversity/diversity_old.php. Identifications more specific than the taxonomic order are shown in parentheses. Solid lines represent positive correlations, dashed lines represent negative correlations. Black lines show no time delay while red arrows are delayed by 1 month.
This study has confirmed that strong seasonal patterns occur in this surface water microbial community and that potential drivers of this structure could be identified from the observatory data. Strikingly, the variable with most explanatory power for overall bacterial richness was day length, which appears to be as important for describing temporal community structure in coastal temperate seas as pH is for describing spatial microbial structure in terrestrial ecosystems. This study has highlighted the added value of much longer temporal observations of natural communities. Although the overall community succession was robust, subtle changes in the patterns of individual taxa were observed and were only detectable because of the long (6 years) time series. Examples of different taxa showing different seasonal cycles were SAR11 and Roseobacter, which had nearly exactly opposite peaks in richness. Additionally, blooms of rare OTUs may be linked to changes in eukaryotic species and environmental variables. Seasonal succession in the community composition was robust and the most variable OTUs were best at predicting the time of year. Environmental factors, rather than interactions with eukaryotes, were better at explaining seasonal variance in bacterial community composition. Meanwhile, interactions were strongest within domains rather than between them, and correlative relationships were stronger between taxa than with environmental variables. This may indicate that biological rather than physical factors can be more important in defining the fine-grain community structure. Finally, in making comparisons of the bacterial OTU subsets, a fundamental stability in the community has been shown, which suggests that the robust seasonal cyclicity noted for the alpha- and beta-diversity is also self-evident in the interactions between members of the community.
References
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Lozupone CA, Turnbaugh PJ et al. (2011). Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc Natl Acad Sci USA 108 (Suppl 1): 4516–4522.
Church MJ . (2009). Resource control of bacterial dynamics in the Sea. In: Kirchman DL (ed). Microbial Ecology of the Oceans. Wiley & Sons Inc.: NJ, USA.
Clarke KR, Gorley RN . (2006). PRIMER v6: User Manual/Tutorial. Primer-E Ltd.: Plymouth, UK.
Clarke KR, Warwick RM . (2001). Change in Marine Communities: An Approach to Statistical Analysis and Interpretation, 2nd edn. Primer-E Ltd: Plymouth, UK.
Costello EK, Lauber CL, Hamady M, Fierer N, Gordon JI, Knight R . (2009). Bacterial community variation in human body habitats across space and time. Science 326: 1694–1697.
Cullen JJ . (1991). Hypothesis to explain high-nutrient conditions in the open sea. Limnology Oceanography 36: 1578–1590.
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K et al. (2006). Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72: 5069–5072.
Fierer N, Jackson RB . (2006). The diversity and biogeography of soil bacterial communities. Proc Natl Acad Sci USA 103: 626–631.
Fuhrman JA . (2009). Microbial community structure and its functional implications. Nature 459: 193–199.
Fuhrman JA, Hewson I, Schwalbach MS, Steele JA, Brown MV, Naeem S . (2006). Annually reoccurring bacterial communities are predictable from ocean conditions. Proc Natl Acad Sci USA 103: 13104–13109.
Fuhrman JA, Steele JA, Hewson I, Schwalbach MS, Brown MV, Green JL et al. (2008). A latitudinal diversity gradient in planktonic marine bacteria. Proc Natl Acad Sci USA 105: 7774–7778.
Gilbert JA, Field D, Swift P, Newbold L, Oliver A, Smyth T et al. (2009). The seasonal structure of microbial communities in the Western English Channel. Environ Microbiol 11: 3132–3139.
Gilbert JA, Field D, Swift P, Thomas S, Cummings D, Temperton B et al. (2010). The taxonomic and functional diversity of microbes at a temperate coastal site: a ‘multi-omic’ study of seasonal and diel temporal variation. PLoS One 5: e15545.
Huse SM, Welch DM, Morrison HG, Sogin ML . (2010). Ironing out the wrinkles in the rare biosphere through improved OTU clustering. Environ Microbiol 12: 1889–1898.
Kirchman DL, Rich JH, Barber RT . (1995). Biomass and biomass production of heterotrophic bacteria along 140-degrees-W in the equatorial pacific - effect of temperature on the microbial loop. Deep-Sea Res Part Ii-Topical Stud Oceanography 42: 603–619.
Lewis K, Allen JI . (2009). Validation of a hydrodynamic-ecosystem model simulation with time-series data collected in the Western English Channel. Mar Syst 77: 296–311.
Liu Z, DeSantis TZ, Andersen GL, Knight R . (2008). Accurate taxonomy assignments from 16S rRNA sequences produced by highly parallel pyrosequencers. Nucleic Acids Res 36: e120.
Maidak BL, Cole JR, Lilburn TG, Parker CT, Saxman PR, Farris RJ et al. (2001). The RDP-II (Ribosomal Database Project). Nucleic Acids Res 29: 173–174.
Morris RM, Vergin KL, Cho JC, Rappe MS, Carlson CA, Giovannoni SJ . (2005). Temporal and spatial response of bacterioplankton lineages to annual convective overturn at the Bermuda Atlantic Time-series Study site. Limnology Oceanography 50: 1687–1696.
Nedwell DB, Rutter M . (1994). Influence of temperature on growth-rate and competition between 2 psychrotolerant antarctic bacteria - low-temperature diminishes affinity for substrate uptake. Appl Environ Microbiol 60: 1984–1992.
Pingree RD, Griffiths KD . (1980). Currents driven by a steady uniform wind stress on the shelf seas around the British Isles. Oceanol Acta 3: 227–235.
Pruesse E, Quast C, Knittel K, Fuchs BM, Ludwig WG, Peplies J et al. (2007). SILVA: a comprehensive online resource for quality checked and aligned ribosomal RNA sequence data compatible with ARB. Nucleic Acids Res 35: 7188–7196.
Reeder J, Knight R . (2010). Rapidly denoising pyrosequencing amplicon reads by exploiting rank-abundance distributions. Nat Methods 7: 668–669.
Rusch DB, Halpern AL, Sutton G, Heidelberg KB, Williamson S, Yooseph S et al. (2007). The Sorcerer II Global Ocean Sampling expedition: northwest Atlantic through eastern tropical Pacific. PLoS Biol 5: e77.
Sogin ML, Morrison HG, Huber JA, Mark Welch D, Huse SM, Neal PR et al. (2006). Microbial diversity in the deep sea and the underexplored ‘rare biosphere’. Proc Natl Acad Sci USA 103: 12115–12120.
Southward AJ, Langmead O, Hardman-Mountford NJ, Aiken J, Boalch GT, Dando PR et al. (2005). Long-term oceanographic and ecological research in the Western English Channel. Adv Mar Biol 47: 1–105.
Wang Q, Garrity GM, Tiedje JM, Cole JR . (2007). Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73: 5261–5267.
Acknowledgements
We would like to thank Dr KR Clarke for providing extensive expertize in statistical modelling, and Margaret Hughes for providing the pyrosequencing technical support. All sequencing data and environmental metadata can be found in the INSDC SRA under ERP000118 (http://www.ebi.ac.uk/ena/data/view/ERP000118). This work was supported in part by the US Deptartment of Energy under Contract DE-AC02-06CH11357. JAF and JAS were supported by NSF Grant 0703159 and JAS and SH by the Sloan Foundation (ICoMM).
Author information
Authors and Affiliations
Corresponding author
Additional information
Disclaimer
The submitted manuscript has been created in part by UChicago Argonne, LLC, Operator of Argonne National Laboratory (‘Argonne’). Argonne, a US Department of Energy Office of Science laboratory, is operated under Contract No DE-AC02-06CH11357. The US Government retains for itself, and others acting on its behalf, a paid-up nonexclusive, irrevocable worldwide license in said article to reproduce, prepare derivative works, distribute copies to the public, and perform publicly and display publicly, by or on behalf of the Government.
Supplementary Information accompanies the paper on The ISME Journal website
Supplementary information
Rights and permissions
This work is licensed under the Creative Commons Attribution-NonCommercial-No Derivative Works 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Gilbert, J., Steele, J., Caporaso, J. et al. Defining seasonal marine microbial community dynamics. ISME J 6, 298–308 (2012). https://doi.org/10.1038/ismej.2011.107
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2011.107
Keywords
This article is cited by
-
Microbiome response in an urban river system is dominated by seasonality over wastewater treatment upgrades
Environmental Microbiome (2023)
-
Organic fertilizer and hydrothermal conditions change the distribution of Medicago sativa L. productivity and soil bacterial diversity in coastal saline soil
Chemical and Biological Technologies in Agriculture (2023)
-
Spatio-temporal patterns of Synechococcus oligotypes in Moroccan lagoonal environments
Scientific Reports (2023)
-
Wavelet clustering analysis as a tool for characterizing community structure in the human microbiome
Scientific Reports (2023)
-
Frequency of change determines effectiveness of microbial response strategies
The ISME Journal (2023)