Abstract
The diversity and stability of bacterial communities present in the rhizosphere heavily influence soil and plant quality and ecosystem sustainability. The goal of this study is to understand how ‘Candidatus Liberibacter asiaticus’ (known to cause Huanglongbing, HLB) influences the structure and functional potential of microbial communities associated with the citrus rhizosphere. Clone library sequencing and taxon/group-specific quantitative real-time PCR results showed that ‘Ca. L. asiaticus’ infection restructured the native microbial community associated with citrus rhizosphere. Within the bacterial community, phylum Proteobacteria with various genera typically known as successful rhizosphere colonizers were significantly greater in clone libraries from healthy samples, whereas phylum Acidobacteria, Actinobacteria and Firmicutes, typically more dominant in the bulk soil were higher in ‘Ca. L. asiaticus’-infected samples. A comprehensive functional microarray GeoChip 3.0 was used to determine the effects of ‘Ca. L. asiaticus’ infection on the functional diversity of rhizosphere microbial communities. GeoChip analysis showed that HLB disease has significant effects on various functional guilds of bacteria. Many genes involved in key ecological processes such as nitrogen cycling, carbon fixation, phosphorus utilization, metal homeostasis and resistance were significantly greater in healthy than in the ‘Ca. L. asiaticus’-infected citrus rhizosphere. Our results showed that the microbial community of the ‘Ca. L. asiaticus’-infected citrus rhizosphere has shifted away from using more easily degraded sources of carbon to the more recalcitrant forms. Overall, our study provides evidence that the change in plant physiology mediated by ‘Ca. L. asiaticus’ infection could elicit shifts in the composition and functional potential of rhizosphere microbial communities. In the long term, these fluctuations might have important implications for the productivity and sustainability of citrus-producing agro-ecosystems.
Similar content being viewed by others
Introduction
Plants are the key primary producers in most terrestrial ecosystems and generally exploit soils for resources using complex root systems (Dennis et al., 2010). The root exudates allow the maintenance of a dynamic and nutrient-rich niche around the root–soil interface called the rhizosphere (Curl and Truelove, 1986; DeAngelis et al., 2009). The diversity of nutrients and plant secondary metabolites present in the exudates allows enrichment of specific taxonomic or functional groups of bacteria in the rhizosphere (Bais et al., 2006; Haichar et al., 2008; Dennis et al., 2010). Microbial populations in the rhizosphere are immersed in a framework of interactions known to affect plant fitness and soil quality (Nannipieri et al., 2003; Lugtenberg and Kamilova, 2009). They are involved in fundamental activities that ensure the stability and productivity of both agricultural and natural ecosystems. These microbial populations benefit plant growth by increasing the availability of mineral nutrients, production of phytohormones, degradation of phytotoxic compounds and suppression of pathogens (Kent and Triplett, 2002; Bais et al., 2006; Lugtenberg and Kamilova, 2009). Microbes residing in the rhizosphere also have key roles in ecosystems and influence a large number of important processes, including nutrient acquisition, nitrogen cycling, carbon cycling and soil formation (Nannipieri et al., 2003; Fitter et al., 2005; Cardon and Whitbeck, 2007). These considerations demonstrate the importance of studying the structural and functional properties of rhizosphere microbial communities and the ways in which various factors influence bacterial diversity and microbial activity. Nevertheless, much of the basic information regarding the community structure of rhizosphere microbial community, their principal functions, their relative stability and the forces that govern their continuity is still lacking.
Owing to the interest in basic knowledge of plant disease and the economic importance of plant disease control, a vast amount of research has been dedicated to understanding the basis of the specificity of plant pathogen interactions, how plants defend themselves from infection by pathogens and the role of plant beneficial bacteria in the control of plant pathogens (Montesinos et al., 2002; Lugtenberg and Kamilova, 2009). Cheatham et al. (2009) have stated that the effect of plant disease goes beyond yield and pointed towards the scarcity of reports dealing with the potential of plant diseases to affect various ecological processes and services. It has been postulated that disruption of multitrophic interactions in a stable ecosystem under the influence of invading phytopathogens will cause community reorganization and changes in the local feedback interactions (van der Putten et al., 2007). However, there is a paucity of knowledge on the extent to which such community shifts may occur, on the dynamics of changes and on the putative effects regarding the functioning of ecosystems. In addition, establishing linkages between microbial diversity and ecosystem functions is even more challenging (Nannipieri et al., 2003; Fitter et al., 2005).
A few studies have examined the effects of plant pathogens on the structure of plant-associated bacterial communities (McSpadden Gardener and Weller, 2001; Yang et al., 2001; Araújo et al., 2002; Reiter et al., 2002; Trivedi et al., 2010, 2011); however, the implication of shifts in diversity on ecosystem functions are not well understood. The long-term sustainability of ecosystem productivity requires detailed knowledge on its biodiversity coupled to profound understanding of its functioning. Determining the functional role of complex and dynamic microbial communities is challenging because of our inability to culture a vast majority of microorganisms (Amann et al., 1995). Functional gene microarrays that compare many orthologous genes controlling biogeochemical processes have recently emerged as a way to examine functional genes across a broad range of microorganisms and functions simultaneously (Zhou, 2003; He et al., 2007, 2010a). For example, GeoChip 3.0 targets several thousand genes from diverse species and groups involved in various functions (He et al., 2010a; Zhou et al., 2010). This array allows for a detailed analysis of the functional profiles of microbial communities and is ideal for understanding how these profile changes respond to environmental perturbations (He et al., 2010a, 2010b; Zhou et al., 2010).
Huanglongbing (HLB) is the most devastating disease of citrus in many areas of the world (Asia, North America, South America and Africa). HLB can debilitate the productive capacity of citrus trees with losses of 30–100% reported (Aubert, 1992). HLB is associated with ‘Candidatus Liberibacter spp.’, which are Gram-negative, phloem-limited bacteria (Bové, 2006). The pathogens are transmitted by psyllids Diaphorina citri in Asia, North America and South America and by Trioza erytreae in Africa (Halbert and Manjunath, 2004). HLB pathogens could also be transmitted by HLB-diseased plant materials used for propagation of nursery plants. Candidatus Liberibacter asiaticus (Ca. L. asiaticus) was found in Florida in 2005 and spread to all citrus-growing counties in the state by 2007. HLB was also found in Louisiana in June 2008, and in South Carolina and Georgia in 2009. HLB has not been found in other citrus-growing regions in the United States, such as Texas, California and Arizona. However, D. citri Kuwayama, the vector of ‘Ca. L. asiaticus’, was reported in Texas in 2001 and rapidly spread throughout the state by 2006. Psyllids were also found in California and Mexico in 2009. Importantly, HLB was confirmed in Mexico in December 2009. The infected trees are in the states of Jalisco and Nayarit, which are ∼750 miles away from the Mexican border with California. With the presence of vectors in every citrus-growing state, HLB poses immediate threat to the whole US citrus industry (Gottwald, 2010). Our previous studies have shown that HLB not only affects plant health but also manifests profound effects on the structure and composition of the bacterial community associated with citrus (Sagaram et al., 2009; Trivedi et al., 2010, 2011). As the diversity and stability of the plant-associated bacterial communities heavily influence soil and plant production and ecosystem processes (Nannipieri et al., 2003), fluctuations in the bacterial diversity could have serious implications in the agro-ecosystem sustainability.
The goal of this study was to understand the consequences on microbial community composition and their associated functions under stress caused by the invading phytopathogen. It is hypothesized that the introduction of exogeneous bacteria (phytopathogen) into natural ecosystems may perturb the stability of the microbial community, thus affecting their ecosystem functions. To test this hypothesis, we used citrus- ‘Ca. L. asiaticus’ as a host-pathogen model to evaluate fluctuations in the diversity, composition, structure and functional potential of rhizosphere microbial communities in response to phytopathogen infection. Community composition of the rhizosphere and bulk soil was accessed by clone library construction and sequencing; quantitative real-time PCR (qPCR) was used to provide quantitative data on changes in the numbers of different bacterial taxa and functional groups; and GeoChip 3.0 was used to determine the effect of disease incidence on the functional diversity of rhizosphere microbial communities. This study would advance our understanding of factors that shape the abundances and diversity of species, how these are influenced by various biotic and abiotic factors and what the consequences are for ecological processes and properties.
Materials and methods
Site description and sample processing
Hamlin oranges (Citrus sinensis) trees (three each) were identified as ‘Ca. L. asiaticus’ infected or healthy based on visual symptoms in a grove at Devils garden, Florida. Overall, the sampled infected trees showed foliar symptoms in approximately 15–20% of the canopy indicating a moderate stage of disease infection. Leaf and root samples were collected in triplicate from each tree and brought to the laboratory in a cooler with ice in November 2009 and processed immediately. Soil samples were also collected from regions few meters away from trees (bulk soil samples). The soil type was classified as spodosols (sandy mineral soil, pH 6.8) with low organic content and water-holding capacity. Each rhizosphere sample consisted of the total root system with tightly adhering sediment of each individual plant. HLB-infected citrus had less fine roots as compared with healthy citrus but clear symptoms of root damage and root death were not observed. Roots were shaken gently to remove the loosely adhering soil. Rhizosphere soil was carefully removed from fine roots by gently scraping adhering soil using fine forceps. Special care was taken to detach adhering soil without damaging fine root networks, and considerable effort was made to remove root hairs and fragments from soil samples.
DNA isolation and purification
DNA from root and leaf samples was extracted using the Wizard Genomic DNA purification kit (Promega Corp., Madison, WI, USA) following the protocol for isolating genomic DNA from the plant tissue. The DNA pellet was dried in Vacufuge (Eppendorf, Westbury, NY, USA) for 15 min and dissolved in 100 μl of DNA rehydration solution (Promega Corp.). DNA from soil samples was extracted using the MoBio UltraClean Soil DNA isolation kit (MoBio Laboratories, Solana Beach, CA, USA) as per the manufacturer's instructions. Soil DNA (23 kb) was further purified on 0.8% agarose gel and recovered using the GFX PCR DNA and Gel Band Purification Kit (Amersham Biosciences, Piscataway, NJ, USA) as per the manufacturer's instructions, and eluted in 50 μl ddH2O, after which samples from the same tree were pooled.
Detection of ‘Ca. L. asiaticus’
PCR using primers A2 and J5 was performed to confirm the presence of ‘Ca. L. asiaticus’ in the leaf and root samples (Supplementary Table S1). All PCRs in this study were performed in DNAEngine Peltier thermal cycler (Bio-Rad Laboratories, Hercules, CA, USA). Amplification of DNA was determined by electrophoresis on 1.2% agarose gels for about 30–45 min and visualized by ethidium bromide staining.
All qPCR assays were performed in a 96-well plate using an ABI Prism 7500 Sequence detection system (Applied Biosystems, Foster City, CA, USA). Primer/probe set CQULA04F-CQULAP10P-CQULA04R was used to target the β-operon region of ‘Ca. L. asiaticus’, and qPCR reactions were performed according to the conditions described in Supplementary Table S1. Each individual sample was replicated 4 times on a 96-well plate and the whole reaction was repeated twice to verify the consistency of the method. Results were analyzed using ABI Prism software. Raw data were analyzed using the default settings (threshold=0.2) of the software. The standard equation developed by Trivedi et al. (2009) was used to convert individual Ct values into bacterial population as genome equivalents per gram of samples.
PCR, rRNA gene clone library and sequencing
Primers 27f/1492r were used to amplify the 16S rRNA gene fragments of bacteria under PCR conditions as described in Supplementary Table S1. PCR products were immediately cloned in a pCR2.1-TOPOTA cloning vector and transformed into chemically competent Escherichia coli TOP10 cells (Invitrogen, Carlsbad, CA, USA). Transformed cells were plated on Luria–Bertani agar plates with ampicillin (100 μg l−1) and kanamycin (10 μg l−1). Plates were incubated overnight at 37 °C and then stored at 4 °C for 24 h. Transformants were screened by blue/white colony selection and individual white colonies were inoculated into 96-well culture blocks (Eppendorf) containing 1 ml of freezing medium (Luria–Bertani broth with 10% (v/v) anhydrous glycerol, 25 μg l−1 ampicillin and 12.5 μg l−1 kanamycin) per well. The blocks were incubated at 37 °C for 16 h on a rotary shaker at 200 r.p.m. A 150 ml aliquot of grown cultures was then transferred to sterile 96-well microtitration plates (Corning Inc., New York, NY, USA) for sequencing. Plates were sealed with aluminum seal tape (Eppendorf), and both blocks and plates were stored at −80 °C. Sequencing was performed using the M13 (−20) forward primer at the sequencing facility of the Interdisciplinary Center for Biotechnology Research at the University of Florida.
Sequence analysis
Sequences were analyzed for orientation and detection of non-16S rDNA sequences using OrientationChecker (http://www.bioinformatics-toolkit.org/Squirrel) (Ashelford et al., 2006). The presence of chimeras was assessed using MALLARD (http://www.bioinformatics-toolkit.org/Mallard) (Ashelford et al., 2006). A sequence identified at the 99.9% and 99% cutoffs or suspected at the 99.9% cutoff as a chimeric sequence was discarded. The remaining sequences were aligned using CLUSTAL W (http://www.clustal.org) (Thompson et al., 1994). On the basis of the alignment, a distance matrix was constructed using the DNADIST program from PHYLIP version 3.66 (http://evolution.genetics.washington.edu/phylip.html) with default parameters. The resulting matrices were run in DOTUR (http://www.plantpath.wisc.edu/fac/joh/dotur) (Schloss and Handelsman, 2005) to generate diversity indexes. The default DOTUR settings were used with threshold values of 97% sequence identity for the definition of operational taxonomic units (OTUs). Library coverage was calculated with the nonparametric estimator C (Good, 1953). The reciprocal of Simpson's index (1/D) was used as a measure of diversity to evaluate the level of dominance in a community (Zhou et al., 2002). The number of shared OTUs between libraries was calculated using SONS (www.mothur.org) (Schloss and Handelsman, 2006). UniFrac (Lozupone and Knight, 2005) was applied to examine the differences between clone libraries. A tree file generated by CLUSTAL W (Thompson et al., 1994) and an environment file, which links a file to a library, were uploaded to UniFrac (Lozupone and Knight, 2005). Principal-coordinate analysis (PCA) was performed using UniFrac with the abundance-weighted option (Lozupone and Knight, 2005). The diversity of the clone library was also investigated by rarefaction analysis. Rarefaction curves were calculated using the freeware program aRarefactWin (http://www.uga.edu/strata/software/anRareReadme) (Holland, 1998).
Phylogenetic analysis
The phylogenetic composition of the sequences in each library was evaluated using the Classifier program of RDP-II release 10 (http://rdp.cme.msu.edu) Wang et al., 2007), with confidence levels of 80%. BLASTN (http://www.ncbi.nlm.nih.gov/blast/Blast.cgi?PAGE=Nucleotides) was also used to classify clones and to identify the closest relatives in the GenBank database. For phylogenetic analysis, sequences were aligned using the CLUSTAL W program (Thompson et al., 1994). The neighbor-joining method was used for building the trees. The PHYLIP format tree output was obtained using the bootstrapping procedure; 1000 bootstrap trials were used. The trees were constructed using TreeView software (PHYLIP).
Accession numbers of nucleotide sequences
Nucleotide sequences of 16S rRNA genes for clone libraries have been deposited in the NCBI database under accession numbers shown in Table 1.
Taxon/genus-specific qPCR
Quantitative real-time PCR quantifications for Acidobacteria, Actinobacteria, Firmicutes, Alphaproteobacteria, Bacteroidetes, Betaproteobacteria, Pseudomonas spp., Bacillus spp., Burkholderia spp. and total bacteria were performed using primers and cycling conditions described in Supplementary Table S1. qPCR reactions were carried out on extracted soil DNA from different samples using Absolute qPCR SYBR green mixes (Qiagen Inc., Valencia, CA, USA) on an ABI Prism 7500 Sequence detection system. The relative fractional abundance for each of the groups was calculated by determining the measured copy number of each group-specific qPCR assay and the ‘total bacteria’ assay.
Quantification of functional communities involved in nitrogen cycling
Real-time PCR quantifications of genes encoding the key enzymes of ammonia oxidation (amo encoding the ammonia monooxygenase in both AOB and AOA), nitrate reduction (narG encoding the membrane-bound nitrate reductase) and denitrification (nirK and nirS encoding the cd1 and copper nitrite reductase; nosZ encoding the nitrous oxide reductase) were used to estimate the density of functional communities involved in nitrogen cycling by using primers and conditions described in Supplementary Table S1.
Preparation of standard curve for qPCR
Standard curves for real-time PCR assays were developed by PCR amplifying the respective genes by their specific primers. PCR products were purified using a PCR cleanup kit (Axygen Bioscience, Union City, CA, USA) and cloned into the pGEM-T Easy Vector (Promega Corp.). The resulting ligation mix was transformed into E. coli JM109 competent cells (Promega Corp.) following the manufacturer's instructions. Plasmids used as standards for quantitative analyses were extracted from the correct insert clones of each target gene and sent for sequencing. The plasmid DNA concentration was determined on a NanoDrop ND-1000 spectrophotometer (NanoDrop Technologies Inc., Wilmington, DE, USA), and copy numbers of target genes were calculated directly from the concentration of the extracted plasmid DNA. Tenfold serial dilutions (108–101 copies per μl) of the plasmid DNA were subjected to a qPCR assay in triplicate to generate an external standard curve and to check the amplification efficiency. Standard curve regression coefficients were consistently above 0.99, and melt curve analysis verified a single amplicon per reaction in all the cases. Samples and standards were assessed in at least two different runs to confirm reproducibility of the quantification. Target copy numbers for each reaction were calculated from the standard curve and were used to ascertain the number of copies per μg of DNA. In the analysis of qPCR data, Student's t-tests were used to determine the significance of pairs of values of mean±s.d.
GeoChip analysis
A whole community genome amplification was used to generate ∼3.0 μg of DNA with 50 ng purified DNA as the template using the TempliPhi Kit (GE Healthcare, Piscataway, NJ, USA) following the manufacturer's instructions. To improve amplification efficiency, single-strand-binding protein (267 ng μl−1) and spermidine (0.1 mM) were added to the reaction mix (Wu et al., 2006). Reactions were then incubated at 30 °C for 3 h and stopped by heating the mixtures at 65 °C for 10 min. Template labeling was performed with the fluorescent dye Cy-5 using random priming as described by He et al. (2010a, 2010b). Labeled DNA was purified using the QIA quick purification kit (Qiagen Inc.) according to the manufacturer's instructions, measured on a NanoDrop ND-1000 spectrophotometer and then dried down in a SpeedVac (ThermoSavant, Milford, MA, USA) at 45 °C for 45 min. The labeled target was resuspended in 120 μl hybridization solution containing 50% formamide, 3 × saline-sodium citrate (SSC) buffer, 10 μg of unlabeled herring sperm DNA (Promega Corp.) and 0.1% SDS, and the mix was denatured at 95 °C for 5 min and kept at 50 °C until it was deposited directly onto a microarray. Hybridizations were performed using TECAN Hybridization Station HS4800 Pro (TECAN, US, Durham, NC, USA) according to the manufacturer's recommended method. After washing and drying, the microarray was scanned using a Scan Array Express Microarray Scanner (Perkin-Elmer, Boston, MA, USA) at 633 nm using a laser power of 90% and a photomultiplier tube gain of 75%. Images were then processed by ImaGene 6.0 software (BioDiscovery, El Segundo, CA, USA) in which a grid of individual circles defining the location of each DNA spot on the array was superimposed on the image to designate each fluorescent spot to be quantified. Hybridization spots with a signal-noise ratio ⩾2.0 (He and Zhou, 2008) were kept for further analysis. Normalization among slides was performed based on the mean signals of all spots. Outliers of a gene within technical replicate slides (>2 s.d.) were removed, and gene detection was considered positive when a positive hybridization signal was obtained for at least 33.3% spots targeting the gene of all replicates.
The effects of ‘Ca. L. asiaticus’ infection on functional microbial communities, microbial processes, and environmental parameters were analyzed by computing the response ratio using the formula described by Luo et al. (2006). For the response ratio analysis, the total abundance of each gene category or family was simply the sum of the normalized intensity for the gene category or family. Ordination analyses were further performed using PC-ORD for Windows (McCune and Mefford, 1999) and confirmed by CANOCO 4.5 for Windows (Biometris—Plant Research International, Wageningen, The Netherlands). Detrended correspondence analysis was used to determine the overall functional changes in the microbial communities. Heatmaps were generated using a pairwise centroid linkage hierarchical clustering algorithm from the software program Cluster (http://rana.lbl.gov) (Eisen et al., 1998) and visualized the clustering results using the TreeView software program (see the above URL).
Results
Detection of ‘Ca. L. asiaticus’
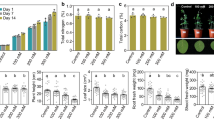
Conventional PCR using primers A2-J5, which target the 16S rRNA genes of ‘Ca. L. asiaticus’, showed a band of ∼703 bp in the infected leaf and root samples, which was not detected in healthy samples (Figure 1a). Sometimes lower levels of ‘Ca. L. asiaticus’ infection are not detected by conventional PCR. We further performed qPCR analysis using primer-probe combination CQULA04F-CQULAP10P-CQULA04R (targeting the β-operon region of ‘Ca. L. asiaticus’) to confirm and quantify the numbers of HLB pathogens in the samples (Figure 1b). qPCR results showed the absence of ‘Ca. L. asiaticus’ in both the leaf and the root samples of asymptomatic samples. The number of ‘Ca. L. asiaticus’ ranged from 2.2 to 4.3 × 107 and 2.5 to 3.4 × 106 genome equivalents per gram of tissue leaf and root samples, respectively. We also tested rhizosphere soil samples to detect the presence of ‘Ca. L. asiaticus’ using both conventional and qPCR methods. As expected, ‘Ca. L. asiaticus’ was not detected in rhizosphere soil samples of either HLB-infected or healthy citrus.
Detection of ‘Ca. L. asiaticus’ in root and leaf samples of HLB symptomatic and healthy citrus. (a) Agarose gel electrophoresis of PCR products amplified using primers A2-J5 targeting 16S rRNA gene of ‘Ca. L. asiaticus’. (b) Quantification of ‘Ca. L. asiaticus’ (genome equivalents per gram of samples) by qPCR using primer-probe combination CQULA04FCQULAP10-CQULA04R targeting the β-operon region of ‘Ca. L. asiaticus’. Symptomatic IT1–IT3; healthy HT1–HT3; M: DNA molecular weight size markers.
Clone library analysis of the effect of ‘Ca. L. asiaticus’ infection on the structure of bacterial community associated with citrus rhizosphere
Statistics of clone libraries
16S rRNA gene clone libraries were constructed from the bulk soil and rhizosphere samples of ‘Ca. L. asiaticus’-infected or healthy citrus. Out of a total of 1344 clones, 45 were low-quality sequences and 74 were identified as chimeric, which were removed from the final analysis. Rarefaction curves of 16S rRNA gene clone libraries of bulk soil and rhizosphere-associated bacteria from ‘Ca. L. asiaticus’-infected and healthy citrus are presented in Supplementary Figure S1. Rarefaction curves did not reach an asymptote indicating insufficient sampling to capture the total diversity of the bacterial community. Although clone library analysis does not provide a complete snapshot of the total bacterial diversity, it provides invaluable information on the most common taxa present in the samples (DeAngelis et al., 2009). Detailed information on the statistics of clone libraries is summarized in Table 1. In pairwise comparisons, the clone library from the rhizosphere of healthy citrus contained significantly higher 16S rRNA gene diversity compared with clone libraries from bulk soil or the ‘Ca. L. asiaticus’-infected citrus rhizosphere, as reflected by the higher Chao1 richness estimations (P⩽0.01). The Chao1 richness estimates were 143.2, 194.23 and 486.2 for the combined clone libraries from bulk soil, ‘Ca. L. asiaticus’-infected and healthy citrus rhizosphere samples, respectively. Similarly, the Shannon–Weiner diversity index of the clone library from healthy samples was significantly greater (P⩽0.01) than that of infected or bulk soil clone library. Qualitative differences in the bacteria from different samples were confirmed by PCA analysis in UniFrac (Figure 2). The UniFrac analysis of cloned sequences separated the rhizosphere of healthy citrus from both the ‘Ca. L. asiaticus’-infected rhizosphere and the bulk soil. A tight cluster of healthy samples distinctly separated from infected samples was observed, suggesting that ‘Ca. L. asiaticus’ infection caused a drastic shift in the community structure of bacteria associated with citrus rhizosphere.
Principal-coordinate analysis for the 16S rRNA gene clone libraries of bulk soil and rhizosphere-associated bacteria in healthy or ‘Ca. L. asiaticus’-infected citrus. The ordination was constructed using UniFrac distances weighted by relative abundances. Each number represents the phylogenetic composition of rhizosphere-associated bacteria in healthy (HT1–HT3) or ‘Ca. L. asiaticus’-infected (IT1–IT3) citrus.
Comparison of phylogenetic diversity in clone libraries of bulk soil and rhizosphere samples of ‘Ca. L. asiaticus’-infected or healthy citrus
Cloned bacterial sequences evidenced differences in the rhizosphere bacterial composition of citrus in response to ‘Ca. L. asiaticus’ infection. A neighbor-joining tree of the 213 cloned OTUs (Figure 3) shows 12 phyla detected: Firmicutes, Bacteroidetes, Proteobacteria, Actinobacteria, Verrucomicrobia, Chloroflexi, Deinococcus-Thermus, Nitrospira, Planctomycetes, Acidobacteria, Gemmatimonadetes and TM7. There were significant differences in the relative abundances of different phyla (different classes for Proteobacteria) between clone libraries of healthy citrus, ‘Ca. L. asiaticus’-infected and bulk soil (Figure 3). The majority of the clones in the healthy library belonged to Proteobacteria (85.2%). The relative abundances of bacteria belonging to Alphaproteobacteria (52.3%), Betaproteobacteria (17.6%) and Gammaproteobacteria (15.3%) were nearly five times greater in clone libraries from healthy citrus as compared with ‘Ca. L. asiaticus’-infected or bulk soil clone libraries. Clone libraries of ‘Ca. L. asiaticus’-infected samples were dominated by bacteria belonging to Firmicutes (25.4%), Actinobacteria (24.0%) and Acidobacteria (23.7%). The relative abundances of each of these phyla were five times greater compared with those of clone libraries from healthy citrus. In the clone library of bulk soil, the majority of the clones belonged to Acidobacteria (35.0%), Firmicutes (28.24%) and Actinobacteria (26.5%). The clone library of the bulk soil consisted of 177 clones belonging to 54 OTUs. The distribution of various OTUs belonging to Proteobacteria, Fimicutes, Actinobacteria and Acidobacteria is presented in Figures 4a–c. Most of the OTUs present in the bulk soil were also observed in the clone library of the ‘Ca. L. asiaticus’-infected citrus rhizosphere.
Heatmap for the relative abundance of operational taxonomic units (OTUs) for 16S rRNA gene clone libraries of bulk soil and citrus rhizosphere-associated bacteria under healthy and ‘Ca. L. asiaticus’-infected conditions. The relative abundance of each OTU in each library is indicated by different colors: yellow (>1%), green (1–2%), orange (2–3%) and red (<3%), with a background of dark blue (0%). The relative abundance for the bacterial groups showing significant differences between clone libraries of bulk soil, ‘Ca. L. asiaticus’-infected and healthy citrus is shown at the right side.
Phylogenetic distribution of operational taxonomic units (OTUs) for (a) Alphaproteobacteria (AP); (b) Gammaproteobacteria (GP) and Betaproteobacteria (BP); and (c) Firmicutes (FIR), Acidobacteria (ACIDO) and Actinobacteria (ACT), of 16S rRNA gene clone libraries of bulk soil and citrus rhizosphere-associated bacteria under healthy (HT1–HT3) and ‘Ca. L. asiaticus’-infected conditions (IT1–IT3). The dendrogram (left) indicates the phylogenetic relationships among the representative sequences of OTUs (defined by 97% identity). The table indicates the relative abundance of clones belonging to each OTU in each library and the results of a BLAST search using the representative sequences.
Clone libraries of ‘Ca. L. asiaticus’-infected and healthy citrus rhizosphere samples consisted of 526 and 521 clones belonging to 124 and 154 OTUs, respectively. In some cases, clearly distinct groups within one genus were also observed. The relatedness of the representative clones of bacterial phyla (class for Proteobacteria) showing significant differences between infected and healthy clone libraries (one each for each taxon found) to their nearest relatives in the NCBI database is shown in Figures 4a–c. The group Alphaproteobacteria consisted of 72 OTUs in clone libraries of healthy samples, whereas only 25 OTUs were found in infected libraries (Figure 4a). In all, 24 OTUs belonging to Gammaproteobacteria were found in clone libraries of healthy samples compared with 11 in infected libraries (Figure 4b). Citrobacter freundii, Enterobacter asburiae and Lysobacter spp. found in all the three clone libraries of healthy samples were not observed in ‘Ca. L. asiaticus’-infected clone libraries. Betaproteobacteria consisted of 16 and 4 OTUs in clone libraries of healthy and infected samples, respectively (Figure 4c). Burkholderia spp. consisted of 7 OTUs. A total of 50 clones were observed in all clone libraries of healthy samples, whereas this was not recovered from infected samples. Firmicutes were represented by 6 and 19 OTUs in the clone libraries of healthy and infected samples, respectively (Figure 4c). Overall, 17 OTUs of Bacillus spp. (a total of 118 clones) were observed in infected clone libraries, whereas these were represented only by 5 OTUs (a total of 8 clones) in clone libraries of healthy samples. Actinobacteria was represented by 9 and 23 OTUs in healthy and infected clone libraries, respectively (Figure 4c). Several OTUs showing similarity of Mycobacterium fortuitum, Actinoplanes globisporus, Asanoa ishikariensis, Microbacterium spp., Conexibacter woesei and Solirubrobacter sp. observed in high frequency in all ‘Ca. L. asiaticus’-infected clone libraries were not observed in healthy libraries. Acidobacteria, the most dominant group of ‘Ca. L. asiaticus’-infected library consisted of a total of 124 clones showing similarity to 9 OTUs in the NCBI database (Figure 4c). In the clone libraries of healthy samples, 5 OTUs of Acidobacteria consisted of a total of 21 clones.
Effects of ‘Ca. L. asiaticus’ infection on rRNA gene abundances of various groups of bacteria in citrus rhizosphere
rRNA gene abundances of total bacteria and specific groups within the total bacterial community was determined using qPCR. The total bacterial population of healthy citrus samples (7.7±1.2 × 109) was significantly greater (at P⩽0.01) than ‘Ca. L. asiaticus’-infected samples (5.5±1.2 × 108) or bulk soil (6.7±1.1 × 107). To determine changes in different groups of bacteria, we have presented our results as fractional copy numbers, which provide a more accurate index of target abundances than the actual copy numbers, as they neutralize the bias produced by the phylotypes with multiple copies of the SSU rRNA gene (Trivedi et al., 2010). Our qPCR results (Supplementary Figure S2) for the numbers of different phylum or genus of bacteria correlate well with trends presented in clone library analysis. Except for Bacteroidetes, qPCR revealed clear differences in the fractional abundances of all groups of bacteria that were observed between ‘Ca. L. asiaticus’-infected and healthy samples from citrus rhizosphere. The proportion of Alphaproteobacteria, Betaproteobacteria, Pseudomonas spp. and Burkholderia spp. was significantly greater in healthy samples. Samples from ‘Ca. L. asiaticus’-infected trees and bulk soil samples revealed greater relative proportion of bacteria belonging to Acidobacteria, Actinobacteria, Firmicutes and Bacillus spp. as compared with the healthy citrus rhizosphere.
As the primary objective of this study was to evaluate ‘Ca. L. asiaticus’-mediated changes in the structure and function of rhizosphere bacterial community of citrus, we did not perform GeoChip 3.0 analysis of bulk soil samples.
Comparison of the functional structure between ‘Ca. L. asiaticus- infected and healthy citrus rhizosphere by GeoChip 3.0
A total of 8763 genes belonging to various functional categories such as antibiotic resistance (472), carbon cycling (1992), energy processes (160), metal resistance (1814), nitrogen cycling (1044), organic remediation (2750), phosphorus utilization (146) and sulfur cycling (385) were detected by GeoChip 3.0. We detected an average of 4659 genes in ‘Ca. L. asiaticus’-infected samples, whereas the average number of genes detected in healthy samples was 6948, revealing a significant difference in the number of genes detected (t-test, P=0.003). Both the Shannon–Wiener (H′) and Simpson's reciprocal 1/D diversity indices indicate higher levels of functional diversity in healthy samples than that seen in ‘Ca. L. asiaticus’-infected samples (Table 2). Both sets of samples exhibited similar Shannon–Wiener's evenness (Table 2). Gene overlap between the samples was also calculated (Table 2). Pairwise comparisons of the genes detected indicated that the proportion of overlapped genes correlated with healthy or ‘Ca. L. asiaticus’-infected citrus rhizosphere samples. Higher percentages of shared genes were observed within similar individual samples. For example, samples HT1 and HT2 (both healthy samples) and IT1 and IT2 (both diseased samples) shared 72.80% and 70.8% of genes, respectively. Between ‘Ca. L. asiaticus’-infected and healthy citrus rhizosphere samples, a lower percentage of genes (51.2%) were shared. All samples have a lower percentage of unique genes.
Cluster analysis of the functional genes identified for all six samples showed two distinct clusters for ‘Ca. L. asiaticus’-infected and healthy samples from citrus rhizosphere (Figure 5a). Detrended correspondence analysis of all detected genes was also used to examine overall functional structure changes in the microbial communities. Detrended correspondence analysis is an ordination technique that uses detrending to remove the arch effect typical in correspondence analysis (Hill and Gauch, 1980). Detrended correspondence analysis revealed that the microbial community structure was significantly altered because of ‘Ca. L. asiaticus’ infection, as indicated by separate clustering between HLB diseased or healthy samples from citrus rhizosphere (Figure 5b). To determine whether the functional structures of ‘Ca. L. asiaticus’-infected and healthy samples from citrus rhizosphere were significantly different, a two-way crossed analysis of similarity was also performed. The analysis of similarity revealed a significant difference between the microbial functional structure of ‘Ca. L. asiaticus’-infected and healthy citrus rhizosphere samples (P⩽0.01).
Multivariate analysis of functional genes associated with the rhizosphere of ‘Ca. L. asiaticus’-infected or healthy citrus. (a) Cluster analysis; and (b) detrended correspondence analysis (DCA) of the functional genes detected in ‘Ca. L. asiaticus’-infected (IT1–IT3) or healthy (HT1–HT3) citrus rhizosphere.
As nitrogen, carbon and phosphorus dynamics are of major importance in the ecosystem processes and biogeochemistry, the genes involved in the cycling of these three elements were examined in more detail.
Nitrogen cycling genes
Overall, the healthy samples have a considerably greater number of detected genes involved in nitrogen cycling as compared with ‘Ca. L. asiaticus’-infected samples, indicating that the functional characteristics of microbial communities of citrus rhizosphere involved in nitrogen cycling were significantly altered by HLB disease (Figure 6). GeoChip data correlated well with the qPCR analysis of various genes involved in nitrogen cycling. A significant reduction in the abundance of genes encoding various key enzymes in nitrogen cycling was observed both in GeoChip and in qPCR analyses (Figure 6).
Relative changes of the detected genes involved in N cycling in ‘Ca. L. asiaticus’-infected samples from citrus rhizosphere. The signal intensity for each gene detected was normalized by all detected gene sequences using the mean. The percentage of a functional gene in a bracket represents the reduction in the signal intensity of the particular gene in ‘Ca. L. asiaticus’ samples compared with healthy samples (bold represent significant differences at P<0.05). Gray-colored genes were not targeted by this GeoChip or not detected in those samples. The bar graphs show the abundances of various genes involved in nitrogen cycling as revealed by qPCR analysis.
A substantial number (358) of genes involved in nitrogen fixation (nifH) was detected, and the abundance of these was significantly higher (P⩽0.01) in healthy than in ‘Ca. L. asiaticus’ citrus rhizosphere samples. The majority of genes detected in total samples were identified as uncultured or unidentified bacterial species followed by Proteobacteria (11.4%), Firmicutes (5.3%), Cyanobacteria (2.7%) and Spirochetes (1.3%). In addition, 18 methanogenic Euryachaeota genes were also detected. Healthy samples have a higher number of nifH genes (297) than do ‘Ca. L. asiaticus’-infected samples (192). Denitrification involves the reduction of nitrate to nitrite, nitric oxide, nitrous oxide and finally nitrogen. GeoChip 3.0 analysis detected four different enzymes involved in denitrification including nitrate reductase (nar), nitrite reductase (nir), nitric oxide reductase (nor) and nitrous oxide reductase (nos). A significant reduction (P⩽0.01) in the abundance of all of the denitrification genes was observed in response to ‘Ca. L. asiaticus’ infection. Overall, 428 genes involved in denitrification were identified in citrus rhizosphere. The number of denitrification genes in healthy samples (401) was significantly higher than in ‘Ca. L. asiaticus’-infected samples (124). The abundance of two key enzymes involved in ammonification, glutamate dehydrogenase (gdh) and urease (ureC) was higher in healthy samples. Ammonia monooxygenase (amo) is a microbial enzyme that catalyzes the oxidation of ammonia to hydroxylamine, the first step in nitrification; subsequently, the enzyme hydroxylamine oxidoreductase (hao) catalyzes the reduction of hydroxylamine to nitrite. GeoChip 3.0 detected only one hao gene which belonged to an uncultured bacterium and was common to all the samples. The majority (60%) of the 20 amo genes detected belonged to Proteobacteria and their abundance was significantly lower (P⩽0.01) in ‘Ca. L. asiaticus’-infected samples. A reduction of 38.4 and 17.2% in the ‘Ca. L. asiaticus’-infected samples was observed in the abundance of dissimilatory nitrogen-reducing enzymes, nitrate reductase (napA) and c-type cytochrome nitrate reductase (nrfA), respectively.
Carbon cycling genes
Five pathways for autotrophic CO2 fixation have been identified so far (Berg et al., 2007), and GeoChip 3.0. contains probes for the genes encoding CO2 fixation enzymes from four pathways: ribulose-l,5-bisphosphate carboxylase/oxygenase (Rubisco) for the Calvin cycle, carbon monoxide dehydrogenase for the reductive acetyl-CoA pathway, PCC/ACC (propionyl-CoA/acetyl-CoA carboxylase) for the 3-hydroxypropionate/malyl- CoA cycle and ATP citrate lyase (AclB) for the reductive acetyl-CoA pathway. The PCC/ACC and Rubisco pathways were identified to be dominant in the citrus-producing agro-ecosystems, whereas only three genes involved in the AclB pathway were detected. A total of 208, 227 and 98 genes were detected for PCC/ACC, Rubisco and carbon monoxide dehydrogenase pathways, respectively. Gene abundances measured by their signal intensities were significantly higher in healthy than in ‘Ca. L. asiaticus’-infected samples (Supplementary Figures 3a–c).
‘Ca. L. asiaticus’ infection had a differential effect on genes involved in carbon degradation (Figure 7). A significant reduction in the abundance of genes involved in the degradation of relatively labile carbon (for example, starch and cellulose), was observed in ‘Ca. L. asiaticus’-infected samples. These include genes encoding enzymes such as α-amylase, pullulanase, glycoamylase, arabinofuranosidase, cellobiase and xylanase. ‘Ca. L. asiaticus’-infected samples have a higher abundance of genes involved in the degradation of recalcitrant carbon, including those encoding glyoxal oxidase, lignin peroxydase and phenol oxidase. There was no statistical difference between the signal intensities of genes involved in cellulose and aromatics degradation in ‘Ca. L. asiaticus’-infected or healthy samples from citrus rhizosphere.
Normalized signal intensity of the detected key gene families involved in carbon degradation in rhizosphere of ‘Ca. L. asiaticus’-infected or healthy citrus. Signal intensities were the sum of detected individual gene sequences for each functional gene, averaged among rhizosphere samples. The complexity of carbon is presented in order from labile to recalcitrant. Error bars represent s.e. *P<0.10.
Phosphorus cycling genes
GeoChip 3.0 targets three enzymes involved in phosphorus utilization, exophosphate (Ppx) for inorganic degradation, polyphosphate kinase (Ppk) for polyphosphate biosynthesis in prokaryotes and phytase for phytate degradation. No significant differences in abundance were observed for Ppk and phytase genes. The total signal intensity of Ppx genes was significantly greater (P⩽0.01) in healthy than in ‘Ca. L. asiaticus’-infected samples. Nearly 60% of the total 97 Ppx genes detected in this study belonged to phylum Proteobacteria. A significant reduction in the number of bacteria possessing Ppx genes was observed in ‘Ca. L. asiaticus’-infected samples (Figure 8). The number of Ppx genes was 84 and 46 in healthy and ‘Ca. L. asiaticus’-infected samples, respectively.
Hierarchical cluster analysis of exophosphate (Ppx) genes in rhizosphere of ‘Ca. L. asiaticus’-infected (IT1–IT3) or healthy (HT1–HT3) citrus. Genes that were present in at least three samples were used for cluster analysis. Results were generated in CLUSTER and visualized using TreeView. Red indicates signal intensities above background, whereas black indicates signal intensities below background. Brighter red coloring indicates higher signal intensities.
Effect of ‘Ca. L. asiaticus’ infection on gene densities of enzymes involved in nitrogen cycling
Our results demonstrate that ‘Ca. L. asiaticus’ infection influences the size of microbial guilds responsible for nitrogen cycling (Figure 6), which was estimated by qPCR quantification for genes encoding the main catalytic enzymes or biosynthetic loci. These effects were significant when gene copy numbers were expressed both as copy numbers ng−1 of extracted DNA (Figure 6) or copy number per gram of soil (data not shown). In general, our qPCR results showed that ‘Ca. L. asiaticus’ infection tends to significantly lower the size of microbial communities involved in various functions.
Discussion
The plant rhizosphere is a dynamic environment in which many factors may affect the structure and species composition of the microbial communities that colonize the roots (Kent and Triplett, 2002; Bais et al., 2006). Microbial communities of the rhizosphere control many belowground processes critical to ecosystem functioning, through their influence on decomposition of organic matter, nutrient cycling and creation of soil structure (Nannipieri et al., 2003; Cardon and Whitbeck, 2007). Shifts in soil microbial community composition and abundance, in turn can significantly influence the dynamics of these processes. It has been postulated that changes in the host physiology due to pathogen infection can have differential influence on rhizosphere community composition and their associated functions (van der Putten et al., 2007). At present, ecosystem simulation models do not include microbial composition, and often neither explicitly considers the effects of phytopathogen infection on microbial activities nor the interactions between diverse microbial processes (Bardgett et al., 2008). In this study, we examined the effect of perturbations caused by the invading pathogen on the extent and regulation of rhizosphere microbial diversity and its impact on biogeochemical cycles that regulate soil fertility and ecosystem function in citrus groves. The results of this study were in accordance with our initial hypothesis that pathogen infection causes fluctuations in the rhizosphere microbial community structure and influence bacterial guilds known to perform important ecological processes.
The bacterial community composition and function in the rhizosphere is likely to be determined by many selection factors that influence the growth and size of different bacterial populations. The results of this study showed that ‘Ca. L. asiaticus’, an obligate endophytic bacterium can drastically influence the composition of rhizobacterial community. This is an interesting observation as it indicates that the pathogen exposure can restructure the native microbial community even when the direct competition effect is lacking. Matilla et al. (2007) have stated that the composition of microbial community in the rhizosphere is related to the diversity of the nutrients and plant secondary metabolites present in the root exudates. Introduction of phytopathogens and nonnative bacteria have been shown to change the amount and composition of organic acids, sugars and other essential nutrients in the root exudates (Phillips et al., 2004; Kamilova et al., 2006; Neumann and Römheld, 2007). Our model disease, HLB is known to cause carbohydrate partitioning imbalances throughout the tree (Etxeberria et al., 2009). HLB-infected trees accumulate a much higher level of starch in the leaves, whereas the roots are depleted of any starch. Blockage of the photoassimilate transport causes the roots to consume the stored starch to sustain their metabolic activities resulting in root death and eventual tree decline (Etxeberria et al., 2009; Kim et al., 2009). We assume that alteration in plant physiology leading to quantitative and qualitative changes in partitioning the photoassimilates is primarily responsible for the shift in microbial diversity in the rhizosphere of HLB-diseased and healthy trees.
In this study, clone library and qPCR analyses revealed distinct abundance and phylogenetic differences between ‘Ca. L. asiaticus’-infected and healthy samples from citrus rhizosphere. ‘Ca. L. asiaticus’ caused a significant reduction in the diversity and abundance of bacterial groups belonging to Proteobacteria. Previous studies have identified various members of Proteobacteria such as Pseudomonas spp. and Burkholderia spp. as highly rhizocompetent genera (Vancanneyt et al., 1996; Lugtenberg et al., 2001; Berg et al., 2005). Proteobacteria has been described as the dominant bacterial group in the rhizosphere of various plant species, presumably because of their relatively rapid growth rates (Cardon and Whitbeck, 2007; Fierer et al., 2007; DeAngelis et al., 2009). In contrast, ‘Ca. L. asiaticus’ infection promoted the diversity and abundances of bacteria belonging to Actimomycetes, Firmicutes (including Bacillus spp.) and Acidobacteria. These groups have been implicated to be typical bulk-soil inhabitants and represent stable components of the microbial ecosystems the within-group members of which remain unaffected by changes in the nutrients, or are less sensitive to such changes (Crawford, 1978; Fierer et al., 2007; Kielak et al., 2008; Yergeau et al., 2009). These stable patterns may be explained by the oligotrophic nature of these bacteria. These populations respond slowly to changes in the root environment including those brought about by root exudates (Duineveld et al., 1998). Our results provided evidence for the previously presumed levels of interactions of these groups with plants, with rhizosphere bacteria reacting more strongly to changes in plant physiology and exudation induced by phytopathogen infection.
As microbial ecology involves the study of both the structure and the function on an ecosystem, meaningful assessments of microbial community must consider not only the abundance and distribution of species but also the functional diversity present in microbial community (Konopka, 2009). To analyze the effects of phytopathogen infection on functional diversity, we determined the composition and abundance of various bacterial groups involved in different functional processes using qPCR and GeoChip 3.0. We found that phytopathogen infection has drastic effects on various functional guilds of bacteria. Some limitations in our ability to draw conclusions about the active function of microbes from this analysis can arise because of the fact that the presence of a functional gene does not always mean that the function is operating (Berthrong et al., 2009). More direct connections can be drawn by measuring the actual processes over time or by the analysis of mRNA. However, changes in functional gene abundances of various groups of bacteria have been correlated with the actual processes rates in the ecosystem (Nicol et al., 2008; Wakelin et al., 2009). Recent research on environmental samples using both mRNA and genomic DNA microarrays has also shown that the dominant species identified by mRNA arrays are also most abundant in terms of genomic DNA (Bulow et al., 2008). This suggests that reasonable connections can be drawn up between the genomic DNA and biogeochemical cycles by using qPCR and GeoChip 3.0 analyses.
Carbon cycle gene distribution in the rhizosphere of citrus was significantly affected by ‘Ca. L. asiaticus’ infection. Carbon degradation is a major biogeochemical process in terrestrial ecosystems. The rate of degradation depends on a number of factors, including availability and types of carbon substrates, as well as the microbial consortium present (Waldrop et al., 2000). In addition, previous reports have shown that the composition of the microbial community affects the degradation rates of soil carbon compounds independent of environment variables (Waldrop et al., 2000). Our results showed that a specialized community has developed in the rhizosphere of ‘Ca. L. asiaticus’-infected citrus that has shifted away from using more easily degraded carbon sources to more recalcitrant forms. This could be attributed to the limited availability of readily available carbon in the soil because of the disruption of root exudations patterns and rhizodeposition. Reduction in organic matter inputs may activate a specialized group of microorganisms with capabilities to produce enzymes which can degrade recalcitrant carbon sources (Waldrop et al., 2000). In fact, we observed an increase in the abundance bacteria belonging to Actinobacteria and Acidobacteria, both of which have been reported to efficiently use more oxidized forms of carbon (Crawford, 1978; Waldrop et al., 2006; Yergeau et al., 2009; Zinger et al., 2009). Characterization based on the ecological theory of the life strategies suggested that the rhizosphere of healthy citrus had greater fraction of known r-strategists, whereas ‘Ca. L. asiaticus’ infection increased the fraction of k-strategists in the rhizosphere. R-strategists preferentially consume labile soil organic carbon pools, have high nutritional requirements and can exhibit high growth rates when resource conditions are abundant. In contrast, k-strategists exhibit slower growth rates and are likely to outcompete r-strategists in conditions of low nutrient availability because of their high substrate affinities (Fierer et al., 2007). GeoChip 3.0 also detected a significantly low abundance of various genes involved in carbon fixation. From a perspective of mitigating greenhouse gas accumulation, soil carbon not only stores carbon dioxide (CO2) but also offers other benefits, such as acting as a chemical filter (through soil minerals) for cleaning water, reducing soil erosion, conserving water, providing microbial habitats and sources of long-term slow-release nutrients, as well as improving soil structure and productivity (IPCC, 2007). The shift in patterns of rhizodeposition and changes in the carbon utilization and fixation potential of the microbial community in response to ‘Ca. L. asiaticus’ infection can have long-term effects on carbon storage and sequestering.
Nitrogen is the most limiting nutrient to plant growth in temperate terrestrial ecosystems (Curl and Truelove, 1986), in part because plants cannot access macromolecular organic nitrogen. Perhaps nowhere are the links between microbial community composition and function more important than in nitrogen cycling, in which the majority of transformations are microbially mediated and many individual processes are carried out by specialized organisms (Zumft, 1997; de Boer and Kowalchuk, 2001). GeoChip 3.0 and qPCR analyses (Figure 6) revealed that ‘Ca. L. asiaticus’ infection leads to decreased abundance of various genes involved in nitrogen cycling, independently of the taxonomic origin. Enhanced root exudation and high amounts of nutrients in the rhizosphere are considered to be optimal conditions for nitrogen fixers (Bürgmann et al., 2005) and denitrifiers (Mayer et al., 2004; Hai et al., 2009). The effect of ‘Ca. L. asiaticus’ infection on the abundance of genes involved in nitrogen cycling was probably due to the differential inputs of readily available carbon through root exudates or changes in root turnover. We also observed significant decreases in the types and abundance of various bacterial groups possessing genes for nitrification and denitrification. For example, our data showed a reduction of 24.6% and 40.7% in the signal intensities of various genes involved in nitrification and denitrification, respectively. Changes in the community structure have been shown to have functional significance for processes mediated by a narrow spectrum of organisms, such as nitrogen fixation and nitrification (Hsu and Buckley, 2009). Shifts in the microbial community of these specialists’ bacteria can have strong impact on ecosystem functionality. Decrease in the abundance of the microbial communities involved in nitrogen turnover may affect overall nitrogen budget through decreased nitrogen fixation or denitrification in the citrus-producing agro-ecosystem, although the linkage between decreased abundances or system level process rates require further study.
Only few studies have examined the influence of phytopathogens on the microbial diversity of plant-associated bacterial community (McSpadden Gardener and Weller, 2001; Yang et al., 2001; Araujo et al., 2002; Reiter et al., 2002; Filion et al., 2004; Trivedi et al., 2010, 2011). To the best of our knowledge, ours is the first study to comprehensively report the changes in the functional diversity of plant-associated bacteria in response to pathogen infection. In accordance with previous studies, we also observed shifts in the structure of bacterial community in diseased plants, but there were large differences in the extent of these changes. Similar to the results of this study, Filion et al. (2004) have reported negative effects of root rot disease on the stability of the rhizosphere bacterial community of black spruce (Picea mariana) seedlings. In contrast, few studies have reported increase in bacterial diversity in pathogen-infested plants (Yang et al., 2001; Reiter et al., 2002). Interestingly, shifts in the structure of fungal community in response to pathogen infection have also been observed (Filion et al., 2004; Vujanovic et al., 2007). We postulate that the nature of fluctuations in the structure and function of plant-associated microbial community would be differentially determined by the virulence mechanism(s) of the pathogen and the response of host plant-to-pathogen infection. This study could serve as a model for other diseases which involve the blockage of vascular tissues and can cause carbohydrate partitioning imbalances throughout the host plant.
Although soil microbial communities are robust, relationships between diversity and stability need to be considered in developing a predictive understanding of microbial community responses to environment perturbations (Girvan et al., 2005). According to the insurance hypothesis (Yachi and Loreau, 1999), species richness has a positive effect on ecosystem productivity through a buffering effect against disturbances. As shown in the study, phytopathogen infection can drastically influence the structure and function of rhizosphere microbial communities through altered carbon partitioning to the roots leading to severe consequences in the stability and productivity of ecosystem. Although the microbial communities in the rhizosphere exhibit a high degree of functional redundancy (Yachi and Loreau, 1999; Allison and Martiny, 2008), shrinking genetic and functional diversity in response to phytopathogen infection, will compromise capacity of adaptive responses to further perturbation. The findings of this study support the view that decreased diversity could lead to decreased stability following perturbations caused by biotic or abiotic factors. Microbial community analysis reveals a complex relationship between genetic diversity, function and stability of rhizosphere microbial community in response to phytopathogen infection. Understanding how essential ecosystem functioning relates to soil biodiversity will therefore contribute to the maintenance of ecosystem sustainability.
Conclusions
The local feedback interactions between plants and soil microbes have been shown to strongly influence both plant and microbial community composition and ecosystem processes. Infection by a phytopathogen can change the productivity of plant communities, thereby altering the quality and quantity of organic matter entering into soil as litter and/or root exudates. Our study provides evidence that suggests that changes in plant physiology mediated by ‘Ca. L. asiaticus’ infection could elicit drastic shifts in the composition and functional potential of soil microbial communities. This is particularly important as it is widely acknowledged that microorganisms are vital to plant productivity and also act as key players in global biogeochemical cycles. We propose to look the plant disease not only in context of plant yields but also as potential agents causing disturbances in ecosystem equilibrium. However, further study is required on understanding the relative importance of perturbations caused by phytopathogens on the microbial biodiversity and activity and the effect of these shifts in local and global-scale ecological phenomenon.
References
Allison SD, Martiny JBH . (2008). Resistance, resilience, and redundancy in microbial communities. Proc Nat Acad Sci USA 105: 11512–11519.
Amann RI, Ludwig W, Schleifer KH . (1995). Phylogenetic identification and in situ detection of individual microbial cells without cultivation. Microbiol Rev 59: 143–169.
Araújo WL, Marcon J, Maccheroni W, van Elsas JD, van Vuurde JWL, Azevedo JL . (2002). Diversity of endophytic bacterial populations and their interaction with Xylella fastidiosa in citrus plants. Appl Environ Microbiol 68: 4906–4914.
Ashelford KE, Chuzhanova NA, Fry JC, Jones AJ, Weightman AJ . (2006). New screening software shows that most recent large 16S rRNA gene clone libraries contain chimeras. Appl Environ Microbiol 72: 5734–5741.
Aubert B . (1992). Citrus greening disease, a serious limiting factor for citriculture in Asia and Africa. Proc Intern Soc Citricult 1: 817–820.
Bais HP, Weir TL, Perry LG, Gilroy S, Vivanco JM . (2006). The role of root exudates in rhizosphere interactions with plants and other organisms. Ann Rev Plant Biol 57: 233–266.
Bardgett RD, Freeman C, Ostle NJ . (2008). Microbial contributions to climate change through carbon cycle feedbacks. ISME J 2: 805–814.
Berg G, Eberl L, Hartmann A . (2005). The rhizosphere as a reservoir for opportunistic human pathogenic bacteria. Environ Microbiol 7: 1673–1685.
Berg IA, Kockelkorn D, Buckel W, Fuchs G . (2007). A 3-hydroxypropionate/4-hydroxybutyrate autotrophic carbon dioxide assimilation pathway in Archaea. Science 318: 1782–1786.
Berthrong ST, Schadt CW, Piñeiro G, Jackson RB . (2009). Afforestation alters the composition of functional genes in soil and biogeochemical processes in South American grasslands. Appl Environ Microbiol 75: 6240–6248.
Bové J . (2006). Huanglongbing: a destructive, newly-emerging, century-old disease of citrus. Plant Mol Biol Pathol 88: 7–37.
Bulow SE, Francis CA, Jackson GA, Ward BB . (2008). Sediment denitrifier community composition and nirS gene expression investigated with functional gene microarrays. Environ Microbiol 10: 3057–3069.
Bürgmann H, Meier S, Bunge M, Widmer F, Zeyer J . (2005). Effects of model root exudates on structure and activity of a soil diazotroph community. Environ Microbiol 7: 1711–1724.
Cardon Z, Whitbeck J . (2007). The Rhizosphere: An Ecological Perspective, 1st edn. Academic Press: San Diego, USA, 232pp.
Cheatham MR, Rouse MN, Esker PD, Igancio S, Pradel W, Raymundo R et al. (2009). Beyond yields-plant disease in the context of ecological services. Phytopathology 99: 1228–1236.
Crawford DL . (1978). Lignocellulose decomposition by selected Streptomyces strains. Appl Environ Microbiol 6: 1041–1045.
Curl EA, Truelove B . (1986). The Rhizosphere. Advanced Series in Agricultural Sciences, Vol. 15. Springer-Verlag: New York, 298pp.
DeAngelis KM, Brodie EL, DeSantis TZ, Andersen GL, Lindow SE, Firestone MK . (2009). Selective progressive response of soil microbial community to wild oat roots. ISME J 3: 168–178.
de Boer W, Kowalchuk GA . (2001). Nitrification in acid soils: micro-organisms and mechanisms. Soil Biol Biochem 33: 853–866.
Dennis PG, Miller AJ, Hirsch PR . (2010). Are root exudates more important than other sources of rhizodeposits in structuring rhizosphere bacterial communities? FEMS Microbial Ecol 72: 313–327.
Duineveld BM, Rosado AS, van Elsas JD, van Veen JA . (1998). Analysis of the dynamics of bacterial communities in the rhizosphere of the chrysanthemum via denaturing gradient gel electrophoresis and substrate utilization patterns. Appl Environ Microbiol 64: 4950–4957.
Eisen M, Spellman P, Brown P, Botstein D . (1998). Cluster analysis and display of genome-wide expression patterns. Proc Nat Acd Sci USA 95: 14863–14868.
Etxeberria E, Gonzalez P, Achor D, Albrigo G . (2009). Anatomical distribution of abnormally high levels of starch in HLB-affected Valencia orange trees. Physiol Mol Plant P 74: 76–83.
Fierer N, Bradford MA, Jackson RB . (2007). Toward an ecological classification of soil bacteria. Ecology 88: 1354–1364.
Filion M, Hamelin RC, Bernier L, St-Arnaud M . (2004). Molecular profiling of rhizosphere microbial communities associated with healthy and diseased black spruce (Picea mariana) seedlings grown in a nursery. Appl Environ Microbiol 70: 3541–3551.
Fitter AH, Gilligan CA, Hollingworth K, Kleczkowski A, Twyman RM, Pitchford JW . (2005). Biodiversity and ecosystem function in soil. Funct Ecol 19: 369–377.
Girvan MS, Campbell CD, Killham K, Prosser JI, Glover LA . (2005). Bacterial diversity promotes community stability and functional resilience after perturbation. Environ Microbiol 7: 301–313.
Good IJ . (1953). The population frequencies of species and the estimation of population parameters. Biometrika 40: 237–264.
Gottwald TR . (2010). Current epidemiological understanding of citrus Huanglongbing. Annu Rev Phytopathol 48: 119–139.
Hai B, Diallo NH, Sall S, Haesler F, Schauss K, Bonzi M et al. (2009). Quantification of key genes steering the microbial nitrogen cycle in the rhizosphere of sorghum cultivars in tropical agroecosystems. Appl Environ Microbiol 75: 4993–5000.
Haichar FZ, Marol C, Berge O, Rangel-Castro JI, Prosser JI, Balesdent J et al. (2008). Plant host habitat and root exudates shape soil bacterial community structure. ISME J 2: 1221–1230.
Halbert S, Manjunath K . (2004). Asian citrus psyllids (Sternorrhyncha: Psyllidae) and greening disease of citrus: a literature review and assessment of risk in Florida. Fla Entomol 87: 330–353.
He Z, Deng Y, Van Nostrand JD, Tu Q, Xu M, Hemme CL et al. (2010a). GeoChip 3.0 as a high-throughput tool for analyzing microbial community composition, structure and functional activity. ISME J 4: 1167–1179.
He Z, Gentry TJ, Schadt CW, Wu L, Liebich J, Chong SC et al. (2007). GeoChip: a comprehensive microarray for investigating biogeochemical, ecological and environmental processes. ISME J 1: 67–77.
He Z, Xu M, Deng Y, Kang S, Kellogg L, Wu L et al. (2010b). Metagenomic analysis reveals a marked divergence in the structure of belowground microbial communities at elevated CO2 . Ecol Lett 13: 564–575.
He Z, Zhou J . (2008). Empirical evaluation of a new method for calculating signal-to-noise ratio for microarray data analysis. Appl Environ Microbiol 74: 2957–2966.
Hill MO, Gauch HG . (1980). Detrended correspondence analysis: an improved ordination technique. Vegetatio 42: 47–58.
Holland S . (1998). aRarefactWin. University of Georgia: Athens, GA, http://www.uga.edu/strata/software/anRareReadme.html.
Hsu SF, Buckley DH . (2009). Evidence for the functional significance of diazotroph community structure in soil. ISME J 3: 124–136.
IPCC (2007). Climate Change: Fourth Assessment Report of the Intergovernmental Panel on Climate Change. Cambridge University Press-Cambridge: United Kingdom and New York, NY, USA.
Kamilova F, Kravchenko LV, Shaposhnikov AI, Azarova T, Makarova N, Lugtenberg B . (2006). Organic acids, sugars, and l-tryptophan in exudates of vegetables growing on stonewool and their effects on activities of rhizosphere bacteria. Mol Plant–Microbe Interactions 19: 250–256.
Kent AD, Triplett EW . (2002). Microbial communities and their interactions in soil and rhizosphere ecosystems. Ann Rev Microbiol 56: 211–236.
Kielak A, Pijl AS, van Veen JA, Kowalchuk GA . (2008). Differences in vegetation composition and plant species identity lead to only minor changes in soil-borne microbial communities in a former arable field. FEMS Microbiol Ecol 63: 372–382.
Kim JS, Sagaram US, Burns JK, Li JL, Wang N . (2009). Response of sweet orange (Citrus sinensis) to ‘Candidatus Liberibacter asiaticus’ infection: microscopy and microarray analyses. Phytopathology 99: 50–57.
Konopka A . (2009). What is microbial community ecology? ISME J 3: 1223–1230.
Lozupone C, Knight R . (2005). UniFrac: a new phylogenetic method for comparing microbial communities. Appl Environ Microbiol 71: 8228–8235.
Lugtenberg B, Kamilova F . (2009). Plant-growth promoting rhizobacteria. Annu Rev Microbiol 63: 541–556.
Lugtenberg BJJ, Dekkers L, Bloemberg GV . (2001). Molecular determinant of rhizosphere colonization by Pseudomonas. Annu Rev Phytopathol 39: 461–490.
Luo J, Wu W, Fienen MN, Jardine PM, Mehlhorn TL, Watson DB et al. (2006). A nested-cell approach for in situ remediation. Ground Water 44: 266–274.
Matilla MA, Espinosa-Urgel M, Rodriguez-Herva JJ, Ramos JL, Ramos-Gonzales MI . (2007). Genomic analysis reveals the major driving forces of bacterial life in the rhizosphere. Genome Biol 8: R179.
Mayer J, Buegger F, Jensen E, Schloter M, Heß J . (2004). Turnover of grain legume N rhizodeposits and effect of rhizodeposition on the turnover of crop residues. Biol Fertil Soils 39: 153–164.
McCune B, Mefford MJ . (1999). Multivariate Analysis of Ecological Data, Version 4.25. MjM Software Design: Gleneden Beach, OR, USA.
McSpadden Gardener BB, Weller DM . (2001). Changes in population of rhizosphere bacteria associated with take-all disease of wheat. Appl Environ Microbiol 67: 4414–4425.
Montesinos E, Bonaterra A, Badosa E, Frances J, Alemany J, Llorente I et al. (2002). Plant-microbe interactions and the new biotechnological methods of plant disease control. Int Microbiol 5: 169–175.
Nannipieri P, Ascher J, Ceccherini MT, Landi L, Pietramellara G, Renella G . (2003). Microbial diversity and soil functions. Eur J Soil Sci 54: 655–670.
Neumann G, Römheld V . (2007). The release of root exudates as affected by the plant physiological status. In: Pinton R, Varanini Z, Nannipieri P (eds), The Rhizosphere Biochemistry and Organic Substances at the Soil–Plant Interface, 2nd edn. CRC Press/Taylor and Francis: New York, pp 23–72.
Nicol GW, Leininger S, Schleper C, Prosser JI . (2008). The influence of soil pH on the diversity, abundance and transcriptional activity of ammonia oxidizing archaea and bacteria. Environ Microbiol 10: 2966–2978.
Phillips DA, Fox TC, King MD, Bhuvaneswari TV, Teuber LR . (2004). Microbial products trigger amino acid exudation from plant roots. Plant Physiol 136: 2887–2894.
Reiter B, Pfeifer U, Schwab H, Sessitsch A . (2002). Response of endophytic bacterial communities in potato plants to infection with Erwinia carotovora subsp. atroseptica. Appl Environ Microbiol 68: 2261–2268.
Sagaram US, De Angelis KM, Trivedi P, Andersen GL, Lu SE, Wang N . (2009). Bacterial diversity analysis of Huanglongbing pathogen-infected citrus, using PhyloChip arrays and 16S rRNA gene clone library sequencing. Appl Environ Microbiol 75: 1566–1574.
Schloss PD, Handelsman J . (2005). Introducing DOTUR, a computer program for defining operational taxonomic units and estimating species richness. Appl Environ Microbiol 71: 1501–1506.
Schloss PD, Handelsman J . (2006). Introducing SONS, a tool for operational taxonomic unit-based comparisons of microbial community memberships and structures. Appl Environ Microbiol 72: 6773–6779.
Thompson JD, Higgins DG, Gibson TJ . (1994). CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22: 4673–4680.
Trivedi P, Duan Y, Wang N . (2010). Huanglongbing, a systemic disease, restructures the bacterial community associated with citrus roots. Appl Environ Microbiol 76: 3427–3436.
Trivedi P, Sagaram US, Kim JS, Brlansky RH, Rogers ME, Stelinski LL et al. (2009). Quantification of viable Candidatus Liberibacter asiaticus in hosts using quantitative PCR with the aid of ethidium monoazide (EMA). Eur J Plant Pathol 124: 553–563.
Trivedi P, Spann TM, Wang N . (2011). Isolation and characterization of beneficial bacteria associated with citrus roots in Florida. Microbial Ecol: e-pub ahead of print 28 February 2011.
Vancanneyt M, Witt S, Abraham WR, Kersters K, Fredrickson HL . (1996). Fatty acid content in whole-cell hydrolysates and phospholipid fractions of pseudomonads: a taxonomic evaluation. Syst Appl Microbiol 19: 528–540.
van der Putten WH, Klironomos JN, Wardle DA . (2007). Microbial ecology of biological invasions. ISME J 1: 28–37.
Vujanovic V, Hamelin RC, Bernier L, Vujanovic G, St-Arnaud M . (2007). Fungal diversity, dominance, and community structure in the rhizosphere of clonal Picea mariana plants throughout nursery production chronosequences. Microb Ecol 54: 672–684.
Waldrop MP, Balser TC, Firestone MK . (2000). Linking microbial community composition to function in a tropical soil. Soil Biol Biochem 32: 1837–1846.
Waldrop MP, Zak DR, Blackwood CB, Curtis CD, Tilman D . (2006). Resource availability controls fungal diversity across a plant diversity gradient. Ecol Lett 9: 1127–1135.
Wakelin SA, Gregg AL, Simpson RJ, Li GD, Riley IT, Mckay AC . (2009). Pasture management clearly affects soil microbial community structure and N-cycling bacteria. Pedobiologia 52: 237–251.
Wang Q, Garrity GM, Tiedje JM, Cole JR . (2007). Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73: 5261–5267.
Wu L, Liu X, Schadt CW, Zhou JZ . (2006). Microarray-based analysis of subnanogram quantities of microbial community DNAs by using whole-community genome amplification. Appl Environ Microbiol 72: 4931–4941.
Yachi S, Loreau M . (1999). Biodiversity and ecosystem productivity in a fluctuating environment: the insurance hypothesis. Proc Nat Acad Sci USA 96: 57–64.
Yang C, Crowley DE, Menge JA . (2001). 16S rDNA fingerprinting of rhizosphere bacterial communities associated with healthy and Phytophthora infected avocado roots. FEMS Microbiol Ecol 35: 129–136.
Yergeau E, Schoondermark-Stolk SA, Brodie EL, Dejean S, Desantis TZ, Goncalves O et al. (2009). Environmental microarray analyses of Antarctic soil microbial communities. ISME J 3: 340–351.
Zhou J . (2003). Microarrays for bacterial detection and microbial community analysis. Curr Opin Microbiol 6: 288–294.
Zhou J, He Z, Van Nostrand JD, Wu L, Deng Y . (2010). Applying GeoChip analysis to disparate microbial communities. Microbe 5: 60–65.
Zhou J, Xia B, Treves DS, Wu LY, Marsh TL, O’Neill RV et al. (2002). Spatial and resource factors influencing high microbial diversity in soil. Appl Environ Microbiol 68: 326–334.
Zinger L, Shanhavaz B, Baptist F, Geremia RA, Choler P . (2009). Microbial diversity in alpine tundra soils correlates with snow cover dynamics. ISME J 3: 850–859.
Zumft WG . (1997). Cell biology and molecular basis of denitrification. Microbiol Mol Biol Rev 61: 533–616.
Acknowledgements
This study was partially supported by the United States Department of Agriculture (Project 2007-35319-18305) through NSF-USDA Microbial Observatories Program.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Trivedi, P., He, Z., Van Nostrand, J. et al. Huanglongbing alters the structure and functional diversity of microbial communities associated with citrus rhizosphere. ISME J 6, 363–383 (2012). https://doi.org/10.1038/ismej.2011.100
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2011.100
Keywords
This article is cited by
-
Effects of above ground pathogen infection and fungicide application on the root-associated microbiota of apple saplings
Environmental Microbiome (2023)
-
Climate change impacts on plant pathogens, food security and paths forward
Nature Reviews Microbiology (2023)
-
Rhizosphere soil microbial communities under foxtail millet continuous and rotational cropping systems and their feedback effects on foxtail millet downy mildew suppression
Plant Growth Regulation (2023)
-
Fungal but not bacterial β-diversity decreased after 38-year-long grazing in a southern grassland
Plant and Soil (2023)
-
Spatio-temporal variation in the root-associated microbiota of orchard-grown apple trees
Environmental Microbiome (2022)