Abstract
In this review, our current understanding of the species Escherichia coli and its persistence in the open environment is examined. E. coli consists of six different subgroups, which are separable by genomic analyses. Strains within each subgroup occupy various ecological niches, and can be broadly characterized by either commensalistic or different pathogenic behaviour. In relevant cases, genomic islands can be pinpointed that underpin the behaviour. Thus, genomic islands of, on the one hand, broad environmental significance, and, on the other hand, virulence, are highlighted in the context of E. coli survival in its niches. A focus is further placed on experimental studies on the survival of the different types of E. coli in soil, manure and water. Overall, the data suggest that E. coli can persist, for varying periods of time, in such terrestrial and aquatic habitats. In particular, the considerable persistence of the pathogenic E. coli O157:H7 is of importance, as its acid tolerance may be expected to confer a fitness asset in the more acidic environments. In this context, the extent to which E. coli interacts with its human/animal host and the organism's survivability in natural environments are compared. In addition, the effect of the diversity and community structure of the indigenous microbiota on the fate of invading E. coli populations in the open environment is discussed. Such a relationship is of importance to our knowledge of both public and environmental health.
Similar content being viewed by others
Introduction
Escherichia coli occurs in diverse forms in nature, ranging from commensal strains to those pathogenic on human or animal hosts. On the basis of genomic information, the species has been divided into six (five major) different phylogenetic groups, denoted as A, B1, B2, C, D and E (Touchon et al., 2009). These subgroups encompass saprophytic (A) and pathogenic (in particular B2, D) types and are often considered to be the result of long (>1 Myr) evolution time. Although the physiology, genetics and biochemistry of E. coli have been intensively studied, it is not known in detail how the bacterium behaves in its natural habitats. Such habitats have been divided in primary, that is, animal/human host-associated (MacFarlane and MacFarlane, 1997), and secondary (that is, open or non-host-associated) habitats (Savageau, 1983). The versatile behaviour exhibited by E. coli in these habitats is reflected in the immense diversity within the species (Bergthorsson and Ochman 1998). In fact, the various commensal and pathogenic forms of E. coli are known to possess genomes that may differ by up to 20% (Ochman and Jones, 2000). The phenotypes of the different E. coli forms are related to such genomic differences and the ensuing patterns of gene expression. The occurrence of key genomic islands in E. coli that define the different behavioural types is clearly an underpinning factor (Dobrindt et al., 2004; Touchon et al., 2009).
The commensal form of E. coli, exemplified by strain MG1655 (Touchon et al., 2009), is traditionally considered as a harmless bacterium that lives in the intestinal system of mammals and assists its host in the breakdown of particular carbon compounds. On the other hand, the pathogenic forms of E. coli, such as the verotoxigenic (VTEC) (Taylor, 2008), enterohaemorrhagic (EHEC, a subclass of the VTEC class), enteroinvasive (EIEC) and uropathogenic/extraintestinal pathogenic (UPEC/ExPEC) classes, all possess capacities that are harmful to their hosts. The well-known E. coli O157:H7 is an example of a harmful VTEC, which has already caused mortality worldwide. VTEC strains are capable of producing verotoxins (genes denoted as stx) (Taylor, 2008) causing mild to bloody diarrhoea, which eventually culminates in the haemolytic uraemic syndrome. More than 150 serotypes of verotoxin-producing E. coli have been found, but the majority of outbreaks are related to serotype O157. E. coli O157:H7 is dangerous because of its resistance to low pH (∼2.5), which allows passage through the stomach, its low infective dose (as few as 10 cells) and its high pathogenicity (Tilden et al., 1996). Following tissue invasion, it may even cause death (Ritchie et al., 2003). Moreover, the stx genes were found to be transferable to non-pathogenic E. coli strains, allowing these to enhance their virulence (Herold et al., 2004).
In the light of current questions concerning the genomic makeup, origin and environmental circulation of—in particular—pathogenic E. coli, this review will examine our current knowledge of the fate of different E. coli forms in open environments, with a focus on E. coli O157:H7. We will emphasize, whenever possible, the impact of the genomic makeup of the organism.
Importance of genomic islands
All pathogenic strains of E. coli contain genomic regions (islands) loaded with a suite of virulence genes that encode key traits for adherence/colonization, invasion, secretion of toxic compounds and transport functions, as well as siderophore production (Touchon et al., 2009). Some of these capacities are also present in commensal E. coli strains and may enhance environmental fitness, although this is often not recognized. A full complement of island-borne traits, which together define behaviour, is often required for pathogenicity (Touchon et al., 2009). With a basis in recent genome-sequencing projects, the diversity across E. coli genomes is now better understood than ever before (Kudva et al., 2002; Touchon et al., 2009). It is clear that, in particular, horizontal gene transfers have been involved in the spread of the virulence-related genomic islands across different E. coli strains (Ochman and Jones, 2000; Touchon et al., 2009). Thus, the picture of an about 2000-gene core genome characteristic for the species, in addition to an open pan genome (currently about 18 000 genes), has recently emerged (Touchon et al., 2009), illustrating the great impact of horizontal gene transfer for genomic plasticity of the species (Figure 1). An analysis of the sites where insertions or deletions had preferentially occurred across all E. coli genomes recently identified 133 such hotspots (Touchon et al., 2009). Figure 2a illustrates a hypothetical display of such sites across the genomes of commensal, EHEC and UPEC strains. One concrete example, shown in Figure 2b, is offered by the pheV tRNA hotspot, displaying the different inserts across strains K-12 MG1655, O157:H7 (Sakai) and CTF073 (Touchon et al., 2009). Strikingly, all the four typical UPEC/ExPEC strains examined consistently contained a similar insert at this site, which is denoted as the pap operon. This operon was found to have a role in E. coli fitness in urinary tract invasion. Interestingly, some gene modules were common between the pheV islands of the UPEC strain CFT073 and strain K12 (Figure 2b). In contrast, the pheV insertion hotspot in the O157:H7 strain (Sakai) was composed of 32 strain-specific coding regions encompassing genes for hypothetical proteins next to those for putative enterotoxin and cytotoxin.
The average E. coli genome is shaped by a multitude of evolutionary forces derived from its primary (host) and secondary habitats, in which both biotic (predators, competitors, cheaters, host defense mechanisms) and abiotic (pH, temperature, UV, mineral depletion and so on) pressures are present. E. coli strains possess a core of about 2000 genes, which equip them with a versatile metabolism. The E. coli pan genome consists of about 18 000 genes, of which 11% belong to the core (dark blue), a large portion (62%, blue) is composed of so-called ‘persistent’ genes, and 26% can be considered as ‘volatile’ genes (pale blue) (Touchon et al., 2009). Events of gene acquisition and loss are consistently linked to insertion/deletion hotspots (red), and cooperatively shape the E. coli genome with selectively maintained core/persistent genes. These events may result in the evolution of gene clusters defining specific E. coli phenotypes, such as the pap operon, which is involved in the pathogenesis of urinary tract infection caused by UPEC strains (see Figure 2). The arrow represents a hypothetical gradient in which E. coli genomic features that are most likely associated with volatile, persistent and core genes are shown.
When complete genomes of E. coli strains are aligned, typical insertion/deletion hotspots can be identified at corresponding locations, as hypothetically displayed in (a). Both the size and gene composition of these regions may differ widely between strains, as exemplified in (b) for the pheV tRNA insertion hotspot. Here, the pheV hotspots of commensal (K-12 MG1655), enterohaemorrhagic (EHEC, O157:H7 Sakai) and uropathogenic (UPEC, CTF073) strains are compared after the data of Touchon et al. (2009). Sections of the insertion hotspots highlighted in different patterns/colours correspond to distinct gene modules (see Touchon et al., 2009 for details). The pap operon (red) has a role in urinary tract infection, is a genomic signature of UPEC/ExPEC strains and can be found in the pheV insertion hotspots of strains CFT073, APEC O1, S88, UMN026 and IAI39. Black sections represent genes/gene modules, which are specific to the corresponding strains. The pheV insertion hotspot in the O157:H7 strain Sakai is composed of 32 strain-specific CDs encoding, among others, 22 hypothetical proteins, 4 transposases, 1 putative enterotoxin and 1 putative cytotoxin. Gene modules highlighted in green are common to the pheV islands of strains K-12 and CFT073. Genes in these modules encode, among others, glycolate oxidases, a glycolate transporter and putative nucleoside triphosphate hydrolases. (A full colour version of this figure is available at The ISME Journal online).
In spite of the fact that we currently possess such detailed information about the often substantial within-E. coli genomic variation, we lack a general understanding of how the genomic makeup translates into specific behaviour/survival in a complex open environment. One real possibility is the fact that we often overlook the environmental relevance of particular island traits, which are thought of as being important in the pathogenic process. An example of this is the mobile iron uptake operon system ybt, originally denoted as a so-called high-pathogenicity island that is often found in E. coli at asn tRNA insertion sites (Schubert et al., 2004). An investigation of online genomic data taught us that this operon was actually spread across the different E. coli subgroups in both pathogenic and commensal forms (Figure 3). The operon may be considered to represent an environmentally relevant trait that is transferable by horizontal gene transfer given its occurrence in many species and genera in the family Enterobacteriaceae (Schubert et al., 2009). However, we do not know what it adds to the already substantial iron-scavenging capabilities of E. coli.
Occurrence of the mobile ybt operon encoding the yersiniabactin (Ybt) iron-acquisition system across E. coli strains of diverse phylogenetic groups and lifestyles. All strains shown, with the exception of ECOR 02 and O6:K5:H1 DSM6601, have their complete genomes sequenced. The island is heavily present in B2 and D phylotypes. Its occurrence in commensal representatives of the B2 group suggests ecological relevance and, possibly, acquisition by ancestral B2 types. A role of the ybt operon as a possible ‘saprophytic island’ can be postulated because of its wide occurrence in bacteria of divergent lifestyles, including E. coli strains of phylogenetic groups A, B1, B2, D and E (Hacker and Carniel, 2001). A general representation of the functional and ‘mobile’ sections of the island is adapted from Schubert et al. (2004) and shown in the upper panel. asn, asn tRNA gene; int, integrase-encoding gene. Genes in the functional operon code for irp, iron-regulated proteins, including irp2 (Ybt peptide synthetase) and irp1 (Ybt peptide/polyketide synthetase); ybtA, AraC-type transcriptional activator; and fyuA, outer membrane protein. Other genes/gene clusters frequently present in genomic islands that may enable E. coli strains to thrive in multiple niches or act as saprophytes, commensals or pathogens are those involved in multidrug/antibiotic resistance and in the production of adherence factors (reviewed in Hacker and Carniel, 2001).
The lifestyle of E. coli is biphasic
Over evolutionary time, the different E. coli subgroups may have largely experienced biphasic lifestyles, consisting of host-independent and host-associated phases. The interactions with the host may differ in severity in accordance with the E. coli and host type, and—given the complexity and interplay of current pathogenicity determinants on the genomic islands—may have evolved gradually. That is, complexity may have been progressively added to the gene repertoire involved in the host-interactive process. For instance, cells of E. coli that enter the human/animal stomach with ingested food are confronted with severe stress due to the often low pH in the intestinal tract. Following stomach passage, they may then encounter conditions that are amenable to growth and survival in the intestinal system. In contrast, conditions in the open environment are very different. Thus, fluctuating but generally low concentrations of available carbon/energy sources, temperatures, oxic versus anoxic conditions and variable (low to high) osmolarity will be present. In the light of these challenging conditions, most forms of E. coli seem to have conserved particular key evolutionary adaptations in their core genomes, as indicated by the presence in most strains of import systems, such as siderophore-mediated iron uptake systems (Schubert et al, 2004) or ABC transporters for uptake of amino acids and sugars. This suggests a high ability in E. coli to obtain diverse nutrients, aiding in its survival in open environments (Ihssen et al., 2007).
It was recently suggested that ‘the more particular types adapt to hosts in which they find suitable niches, the more they will lose the ability to grow in other, contrasting, environments’ (Franz and van Bruggen, 2008). However, this seems contentious, as the capability of E. coli to evolve and survive in a formerly occupied or new habitat may not be easily lost. Growth efficiency and competitiveness under conditions of low available nutrient levels likely represent the most important physiological factors leading to the successful persistence of E. coli in nutrient-limited open environments (Ihssen and Egli, 2005). Therefore, it is important to understand the mechanism(s) of adaptation to nutrient-limited environments, which affect the survival of E. coli in such conditions.
E. coli persistence in open environments—importance and threat
The extent to which E. coli, in particular pathogenic strains, can survive in open environments and which factors affect this survival rate are crucial issues from a fundamental point of view. Next to their capabilities to acquire nutrients, some E. coli strains produce filamentous structures that extend from the cell surface and help cells to attach to surfaces (for example, plant surfaces). Owing to this ability, E. coli coming from soil, manure, irrigation water (Solomon et al., 2002) or contaminated seeds (Itoh et al., 1998) may colonize (and thus find a refuge niche in) plants such as radish (Itoh et al., 1998) and lettuce (Solomon et al., 2002). Specifically, E. coli may access roots or leaf surfaces by splashing during rainfall or irrigation (Natvig et al., 2002). Internal plant compartments can also be colonized (Solomon et al., 2002), so that the E. coli cells cannot be easily washed from plant parts or be killed and removed by disinfectants and washing. Thus, from infested consumable products, the organism can further spread to uncontaminated products during food processing and packaging, resulting in dissemination in the food production chain. Pathogenic E. coli strains such as E. coli O157:H7 thus pose a threat to the food chain and represent a still underestimated environmental risk. It has been estimated that food-borne diseases cause ∼76 million illnesses, 325 000 hospitalizations and 5000 deaths each year in the United States (Mead et al., 2000), and a significant part of the outbreaks has been attributed to E. coli O157:H7. In Europe, such infections are also on the rise (Fisher and Meakins, 2006). Recently, a large multistate outbreak of E. coli O157:H7 from contaminated fresh spinach in the United States resulted in 187 cases of illness (including 97 hospitalizations and three deaths).
E. coli fate in natural habitats
Provided resource availability and key abiotic conditions are propitious, E. coli populations can survive and even grow in open environments. However, under fluctuating environmental conditions, such as those present in many soils and aquatic environments, growth may be differential and gross bacterial death may ensue if the death rate exceeds the growth rate. Both growth and death rates are determined by the environmental conditions at the local scale and by how the microorganism is able to cope with these local conditions by regulating its gene expression patterns.
For instance, an elegant study (Tao et al., 1999) used functional genomics to determine gene expression patterns in E. coli cells grown on a nutrient-rich versus a minimal medium. All 4290 protein-encoding genes were analyzed through microarray hybridization (RNA extraction followed by construction of hybridization probes through cDNA synthesis and hybridization to genomic DNA microarrays). The study showed that rapidly dividing cells in the rich growth medium had elevated expression of genes involved in translation and did not require expression of amino-acid biosynthesis genes in comparison to cells in the minimal medium. Hence, the environmental conditions clearly directed the expression of suites of genes geared toward optimal growth. Such global gene expression analysis can be further used to study whole-cell physiology under diverse conditions, mimicking those deemed relevant for open environments. In particular, the dynamics of gene expression in a habitat such as soil has been underexplored (Saleh-Lakha et al., 2008). For instance, the gene expression patterns of different E. coli commensal and pathogenic forms could be assessed to determine whether particular genes, including the iron acquisition and other virulence-related ones, are expressed in this (and other) environment(s) and to what extent this expression affects bacterial survival. In this context, it was found that over 300 strain O157:H7-specific genes responded to a growth transition in minimal glucose medium (exponential to stationary), with significant changes in gene expression including multiple genes in pathogenicity islands or toxin-converting bacteriophages (Bergholz et al., 2007).
Growth and survival of E. coli in open environments is often restricted by the availability of nutrients and energy sources. In a growing culture, starvation will ensue at a given moment due to a limitation of particular carbon or other substrates. Under such starvation conditions, the cells progressively metabolize their cellular carbohydrates, followed by proteins and RNA, while initially protecting the DNA. In most open environments, E. coli may behave similarly when nutrients exhaust. However, it will first respond to the different—often complex—energy sources and organic compounds, which are usually present in low concentrations. Under such low-nutrient conditions, different alternative catabolic functions and binding proteins will become derepressed, as found in cultures under glucose or arabinose limitation (Ihssen and Egli, 2005). Similarly, iron may become limiting and thus iron-acquisition operons might become derepressed. It would thus be interesting to test the environmental fitness and gene expression of the iron-acquisition-island-containing strains (Figure 3). Furthermore, E. coli was also found to exhibit a high degree of catabolic flexibility, which conferred a clear fitness advantage in its secondary habitats such as soil and water (Ihssen and Egli, 2005). E. coli O157:H7 may survive and even grow in sterile freshwater at low carbon concentrations, which stands in contrast to the common conception that the organism will die out over time in such strongly carbon-limited environments (Vital et al., 2008). Moreover, because of the production of RpoS (subunit of RNA polymerase, represents a major factor involved in starvation survival), E. coli is able to rapidly adapt to, and tolerate, diverse stress conditions (Lange and Hengge-Aronis, 1991). It was thus shown that high osmolarity (Muffler et al., 1996), extremes and fluctuations of temperature (Muffler et al., 1997), low pH (Bearson et al., 1996) and low growth rate (Ihssen and Egli, 2004) induce rpoS in E. coli cells. Supporting the importance of rpoS, an rpoS− E. coli mutant showed decreased survival (colony-forming unit counts) in stationary phase in seawater (Rozen and Belkin, 2001).
Finally, E. coli can enter a ‘dormant’ (previously denoted as viable but non-culturable) state. In this state, cells cannot be easily recovered on standard laboratory media, but are still present as viable cells. For instance, in an experiment with E. coli O157:H7 in manure, significantly higher numbers of the organism were found by direct microscopic counts than by plating on a selective medium (Semenov et al., 2007). This indicated the prevalence of ‘dormant’ cells in the total E. coli O157:H7 population. The state can be triggered by stress conditions that are imposed, for instance, by low temperature (for example, 4 °C) or toxic metals (for example, copper, lead, mercury and cadmium) (Klein and Alexander, 1986). Although the resistance to starvation of E. coli leads to its persistence in open environments, we are unaware of the impact of cells entering this state, that is, whether it represents a state of enhanced environmental injury or a true survival mode.
Factors affecting E. coli fate in open environments
General observations on E. coli survival
E. coli can, to varying extents, survive in different open environments such as soil, manure and water (Kudva et al., 1998; Jiang et al., 2002; Vital et al., 2008). There are also possibilities for migration between these habitats. For instance, E. coli may reach the groundwater from top soil layers, as revealed in several studies (Mankin et al., 2007). Soil factors such as porosity, surface area, bulk density and macropore structure have important roles in the leaching of invading bacteria by their influence on adsorption and gravitational movement with water (van Elsas et al., 1991). However, for organic substrates such as manure and slurry, the adsorption and desorption behaviour of bacteria is not only linked to differences in physical characteristics of the substrate, but also to biophysical properties of the organic matter (Mankin et al., 2007; Semenov et al., 2009).
As a general observation, the population sizes of E. coli in soil and soil-related (manure) habitats have shown progressive declines in all habitats studied. On the other hand, the fate of E. coli populations under complex natural conditions is often not accurately predictable (Semenov et al., 2008). Although the conditions for survival of E. coli in soil, manure and water are considered to be less favourable than in the intestinal system (Tauxe et al., 1997), the organism has been observed to survive for days (at physiological (>30 °C) temperature, aerobic and under nutrient-limiting conditions) to almost a year in the former habitats (Kudva et al., 1998; Fukushima et al., 1999; Jiang et al., 2002).
It is important to understand the environmental controls that affect the survival of E. coli in its secondary habitats (Table 1) (Habteselassie et al., 2008), as in such habitats, the bacterium will be faced with conditions such as low or fluctuating levels of energy sources, high to low levels of oxygen, fluctuating and often extreme temperatures, low pH and/or high osmolarity.
Availability of resources
The availability of resources such as carbon substrates probably is the main critical factor that affects the persistence of E. coli in open environments such as soil and water. The resource availability will relate to the local conditions that typify the habitat. For a chemoheterotroph such as E. coli, obtaining sufficient carbon compounds will clearly impact its chances of survival. Thus, during adaptation to glucose-limited conditions, E. coli cells were shown to be ‘primed’ for the efficient uptake of various carbon sources due to the upregulation of a large number of genes encoding periplasmic binding proteins (Franchini and Egli, 2006). Such priming would allow these cells to capture a range of resources that occur in low concentrations. However, a stronger limitation of resources in environments inhabited or invaded by E. coli will govern its gene expression in another way, affecting survival. Nutrient scarcity in water systems as well as ‘oligotrophication’ of farm environments (when easily available nutrients are in immobilized form) have been suggested as factors or strategies that reduce the chances of survival and thus prevalence of E. coli (for example, strain O157:H7) (Franz and van Bruggen, 2008). In addition, cells may also enter a dormant state if particular stressors are present in the resource-restricted environment (Semenov et al., 2007).
Temperature
Temperature and temperature regime are other important factors that influence E. coli survival and growth. In an animal host body, temperature is often stable, whereas it may be strongly fluctuating in a non-host environment such as soil or water. Until recently, the effect of fluctuating temperature on E. coli survival and adaptation (compared with stable temperatures) in soil was poorly understood, as previous survival experiments had all been carried out under temperature-stable conditions (Kudva et al., 1998; Himathongkham et al., 1999; Franz et al., 2005). Recent results obtained with E. coli O157:H7 showed that survival in manure under fluctuating temperatures was generally lower than that under constant temperature (Semenov et al., 2007). Moreover, the reduction in survival of the organism was more pronounced when the amplitude in the temperature oscillations was larger (7 °C) than at smaller amplitudes (4 °C). Temperature increase might constitute greater stress and energy expenditure for the organism than decrease in temperature (Semenov et al., 2007). Moreover, gene expression patterns are probably constantly altered under temperature fluctuations. It was recently shown that the histone-like nucleoid structuring (H-NS) protein in E. coli controls a majority of thermoregulated genes at 37 °C and 23 °C (White-Ziegler and Davis, 2009). Hence, differential gene regulation in E. coli may occur across a broad temperature range, next to the regulations occurring at single temperatures (for example, the optimal growth temperature). The H-NS protein may provide a very efficient mechanism of gene expression control under fluctuating temperatures, but we need to increase our understanding of the relationships, if any, to heat and cold shock proteins.
pH
In several experiments, the survival of E. coli in soil was shown to be related to the local pH, and in particular soil acidity was detrimental. Expression, under such conditions, of the alternate sigma factor rpoS in E. coli was related to the induction of acid resistance. In particular, E. coli O157:H7 possesses systems for survival at low pH and therefore can be considered to be an intrinsically acid-resistant bacterium (Foster and Spector, 1995; Lin et al., 1996; Sang et al., 2000). Although E. coli O157:H7 strains still showed different capacities to survive in acidic environments (Lin et al., 1996), they all were superior in their survival over non-O157 EHEC (Bergholz and Whittam, 2007). Variation in rpoS induction levels might explain the variability in acid resistance for the different E. coli O157:H7 strains. Although three known acid resistance mechanisms, that is, two amino acid decarboxylase-dependent systems (glutamate and arginine) and a glucose catabolite-repressed system, have been tested (Large et al., 2005), none of these incited acid resistance in E. coli O157:H7. This suggested that the organism possibly uses combined mechanisms or even as-yet-unidentified alternative mechanisms. Strikingly, particular E. coli O157:H7 strains almost appeared as acidiphiles, as their survival was higher at low pH than at relatively high pH (Franz et al., 2005). However, a significant variation in the ability to survive in low-pH environments was found among isolates within a single serotype (Buchanan and Edelson, 1999).
Availability of water
The availability of water is another key determinant of E. coli survival and growth. Severe lowering of the water content (for example, in soil) incites increasing water stress around the cells, whereas water saturation will rapidly surround cells with water and induce anoxic conditions. Both extremes have severe consequences for E. coli physiology and survival in the system, given that extreme drought might result in massive cell death, whereas flooding will shift cellular metabolism to anaerobic processes. However, the behaviour of E. coli in open environments, in particular how the organism copes with fluctuations in water availability, is still unclear.
Presence and diversity of an indigenous microflora
In all natural habitats, E. coli populations will interact, in a loose or intricate way, with the local biota, including the microbial communities. Thus, different types of interactions can be expected in the intestinal tract of humans/animals, in manure, soil, irrigation water and on/in plants. One major microbial factor is the presence of protozoa, which can have a negative effect on E. coli survival as a result of predation, but sometimes even enhances survival. The latter was shown to occur for E. coli O157 in the environmental protozoan Acanthamoeba polyphaga (Barker et al., 1999). In contrast, the cumulative effect of the total indigenous microflora on E. coli survival is often negative as a result of predation, substrate competition and antagonism (Jiang et al., 2002; Unc et al., 2006; Semenov et al., 2007).
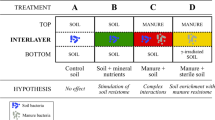
Contrary to the declining populations that are often seen in natural habitats, populations of E. coli can increase in such substrates under sterile conditions, that is, without predatory, antagonistic or competing organisms. This indicates that the natural microbiota in such cases has an overriding effect on survival (Jiang et al., 2002; Unc et al., 2006; Semenov et al., 2007). The diversity of the indigenous microbial communities has been brought up as an important factor that regulates the population dynamics of invading E. coli (van Elsas et al., 2007). According to this view, ecosystems with a higher level of biodiversity (Trevors, 1998) are more resistant to perturbances than those with a lower diversity (Tilman, 1997). Consequently, the former habitats would be less susceptible to invasion by E. coli than the latter (Girvan et al., 2005; Semenov et al., 2008). It is an old paradigm that most ecosystems are microbiostatic, that is, they have filled ecological niches and are difficult to invade. Although the exact influence of autochthonous microbial diversity and community structure on E. coli survival is still unclear, these two aspects of the microbiota are considered to be important. Indeed, the survival in soil of an introduced E. coli O157:H7 derivative was inversely proportional to the diversity of the microbial community present, established through differential fumigation and regrowth (van Elsas et al., 2007). The progressively changed microbial diversities and community compositions coincided with an enhancement of the survival rate of the invading pathogen. However, the impact of the indigenous microflora on E. coli might have been exacerbated, as the nutritional conditions are remote from those in the natural reservoir of the organism. To explain the effect, we hypothesized that lowering of the complexity of the soil microbiota probably resulted in a reduction of functional redundancy, which enhances the chances for the introduced organism to occupy a niche in the system and persist as a member of the community. However, we did not examine to what extent different functional groups in the indigenous microflora affected survival of the invading E. coli. Some organisms could be direct competitors by occupying the same niche, whereas others might have antagonistic or predatory activities. Isolation and identification of microorganisms that promote the decline of the invader may pinpoint inhibitory species, which might be applied as probiotics to reduce the survivability of pathogenic E. coli (Tabe et al., 2008). Conversely, the presence of cellulose- and lignin-degrading organisms might provide E. coli with easily available carbon sources.
Is there a relationship between E. coli genotype and survival capability in the open environment?
E. coli is closely related to Salmonella, with both species belonging to the same family, the Enterobacteriaceae. The two species resemble each other in many ways; however, they differ in essential details. Extensive data indicate that the overall genome complexity in Salmonella is generally 10–20% greater than that in E. coli. In addition, S. typhimurium is 1–4% higher in guanine-plus-cytosine content than most E. coli strains (Ingraham and Neidhardt, 1987). In the light of these differences, the survival of the two species in open environments has been shown to be different. Thus Salmonella strains survived significantly longer in terrestrial habitats in the majority of cases in comparison to E. coli strains (Himathongkham et al., 1999; Franz et al., 2005; Semenov et al., 2007). Such differential survival might relate to genome size or content or gene expression patterns between the two species. We surmised that also the within-E. coli genomic differences might correlate with divergent survival rates. For instance, two E. coli strains, C278 and C279 (different by enterobacterial repetitive intergenic consensus sequence (ERIC)-PCR-based genomic fingerprinting), survived differently in soil amended with swine manure (Topp et al., 2003). Moreover, basal production of toxin (Stx) by haemolytic uraemic syndrome-associated E. coli was higher than that by bovine-associated E. coli strains (Ritchie et al., 2003). In addition, a reconsideration of the work of Kudva et al. (1998) showed that at 23 °C a toxin-negative strain of E. coli O157:H7 grew and survived better than its toxin-positive counterpart, whereas no such differences were found at lower temperatures. This differential survival was found between days 15 and 55 of the experiment (Kudva et al., 1998), confirming an influence of genetic makeup (here, the production of a toxin) on survival characteristics. Shiga toxin-producing E. coli O157:H7, O11:H- and O26:H11 survived for comparable periods of time (up to 8 weeks) in bovine faeces, with rather similar average decay rates at 25 °C (Fukushima et al., 1999), although at 5 °C E. coli O157:H7 was superior in survival after the first 4 weeks over the other two strains. Similar results were shown for seawater, where different specific mutations (rpoS, otsA, relA, spoT, ompC and ompF) significantly influenced E. coli survival (Rozen and Belkin, 2001). Hence, particular characteristics encoded by the genome present in some E. coli strains, but not in others (like, for instance, the capacity to survive low temperature under cow dung conditions), may have been the cause of this differential survival, as the production of toxins requires energy and therefore imposes a fitness cost.
Fitness (growth rate) tests performed in rich Tryptic Soy Broth (TSB), poor and rumen medium under changing (aerobic versus anaerobic) conditions showed no difference in growth rates between E. coli O157:H7 and commensalistic E. coli strains. However, both types of strains had different carbon substrate utilization patterns (Durso et al., 2004): of 95 carbon sources, 27 were oxidized by the commensal E. coli but not by E. coli O157:H7 (Durso et al., 2004). This difference did not affect their growth in common media and also could not be linked to any differential survival in the cow stomach or dung. Finally, the variability across 57 commensal and pathogenic E. coli strains in utilization of carbon and energy sources as well as catabolic and stress protection genes was low, but such variation was clearly found across the different E. coli strains (Ihssen et al., 2007). These results suggest that functions affecting the microorganism's survival and growth, such as those of the central metabolism that determine growth rates, are actually quite broadly shared across the E. coli strains, potentially yielding grossly comparable survival rates in complex natural systems. On top of that, particular E. coli strains may have acquired or evolved specific systems—such as particular iron-acquisition operons (Figure 3), and/or capacities to withstand acidity and/or low temperature—that allow them to be fitter in some secondary habitats. However, these latter contentions need experimental validation.
Given the finding of a 2000-gene common E. coli core genome (Figure 1), which might encompass all genes that are of importance for environmental hardiness and survival (but does not encompass identifiable extraintestinal virulence-specific genes; Touchon et al., 2009), we postulate that E. coli genomes, next to providing overall cores that largely determine metabolic flux on the one, and stress tolerance on the other hand, incidentally (per strain or group of strains) provide specific traits that enhance their survival in open habitats.
On another matter, genes involved in virulence are variably present across E. coli strains, and can even be found on plasmids (Tóth et al., 2009), explaining their ready and rapid spread. However, only 131 E. coli O157:H7-specific proteins out of 1632 were associated with virulence, indicating that there is a limit to what genomes comprise. If E. coli O157:H7 would be transmitted directly from human to human, then the mere presence of the virulence genes—next to the core—might have been sufficient for rapid pathogenesis, resulting in O157:H7 outcompeting commensal E. coli. However, outbreaks of E. coli O157:H7 usually come from infested primary food, suggesting that pathogenic and commensal E. coli differ in other traits that affect their survival, allowing E. coli O157:H7 to be (overall) successful (Durso et al., 2004).
Conclusions and prospects for further research
It is known that the combination of abiotic (availability of energy and nutrient sources, pH, moisture and temperature) as well as biotic (indigenous microflora, including protozoa) factors sets the conditions under which E. coli needs to survive. Extreme or fluctuating values of each parameter pose varying levels of stress to the cells, leading to different survival times. Hence, these factors per se will determine E. coli survivability in natural systems by their direct effects on the exposed cells. Given the complexity and heterogeneity of most natural environments, it is intrinsically difficult to predict the fate of E. coli populations facing the combined environmental effects. Recent studies on the effect of microbial diversity and community structure on the persistence of introduced E. coli O157:H7 in soil have nevertheless shown the emergence of relationships between the microbial diversity of the system, abiotic factors and the pathogen's invasibility (van Elsas et al., 2007; Semenov et al., 2008).
In addition, the relationship between E. coli genotype and its phenotype, in terms of its environmental persistence, is not at all clear. As outlined in the foregoing, the genotypes within the species share a core set of genes (Figure 1; Touchon et al., 2009) that collectively broadly determines the organism's growth and metabolic characteristics as well as its stress resistance in natural environments. Traits that are additive to the ones encoded by the core set of genes, for instance the extra capacity to acquire iron in many strains (Figure 3) or the acid resistance in E. coli O157:H7, can be present in particular E. coli genotypes, but we lack extensive knowledge about their occurrence, functioning and fitness-enhancing effects. Future research should address the intriguing hypothesis that some of the E. coli strains, next to their evolution to fitness enhancement in their primary environment (in association with their host), also have evolved towards an enhanced fitness in open environments.
As a conclusion, we have to reexamine our knowledge about the survival of E. coli in open systems with respect to the effect of the environmental factors and the organism's genotype. It has been traditionally assumed that E. coli shows natural declines in open environments. However, forming an environmental reservoir of a given size in different conditions, the bacterium can cause risks to human health. The factors that determine the rate of survival of particular E. coli strains, and thus the risks, are not (yet) easily predictable, and our capability to understand the effects of the secondary habitat (especially soil and water resources) on E. coli behaviour will be paramount to our abilities to manage the organism from both environmental and public health perspectives.
References
Artz RRE, Townend J, Brown K, Towers W, Killham K . (2005). Soil macropores and compaction control the leaching potential of Escherichia coli O157:H7. Environ Microbiol 7: 241–248.
Bach SJ, Stanford K, McAllister TA . (2005). Survival of Escherichia coli O157:H7 in feces from corn- and barley-fed steers. FEMS Microbiol Lett 252: 25–33.
Barker J, Humphrey TJ, Brown MWR . (1999). Survival of Escherichia coli O157 in a soil protozoan: implications for disease. FEMS Microbiol Lett 173: 291–295.
Bearson SM, Benjamin Jr WH, Swords WE, Foster JW . (1996). Acid shock induction of RpoS is mediated by the mouse virulence gene mviA of Salmonella typhimurium. J Bacteriol 178: 2572–2579.
Bergholz TM, Whittam TS . (2007). Variation in acid resistance among enterohaemorrhagic Escherichia coli in a simulated gastric environment. J of Appl Microbiol 102: 352–362.
Bergholz TM, Wick LM, Qi W, Riordan JT, Ouellette LM, Whittam TS . (2007). Global transcriptional response of Escherichia coli O157:H7 to growth transitions in glucose minimal medium. BMC Microbiol 7: 97.
Bergthorsson U, Ochman H . (1998). Distribution of chromosome length variation in natural isolates of Escherichia coli. Mol Biol Evol 15: 6–16.
Buchanan RL, Bagi LK, Goins RV, Phillips JG . (1993). Response surface models for the growth kinetics of Escherichia coli O157:H7. Food Microbiol 10: 303–315.
Buchanan RL, Edelson SG . (1999). Effect of pH-dependent, stationary phase acid resistance on the thermal tolerance of Escherichia coli O157:H7. Food Microbiol 16: 447–458.
Dobrindt U, Hochhut B, Hentschel U, Hacker J . (2004). Genomic islands in pathogenic and environmental microorganisms. Nature Rev Microbiol 2: 414–424.
Durso LM, Smith D, Hutkins RW . (2004). Measurements of fitness and competition in commensal Escherichia coli and E. coli O 157:H7 strains. Appl Environ Microbiol 70: 6466–6472.
England LS, Lee H, Trevors JT . (1993). Bacterial survival in soil: Effect of clays and protozoa. Soil Biol Biochem 25: 525–531.
Fisher IS, Meakins S . (2006). Surveillance of enteric pathogens in Europe and beyond: Enter-net annual report for 2004. Euro Surveillance: European Communicable Disease Bulletin 11.
Foster JW, Spector MP . (1995). How Salmonella survive against the odds. Annual Rev of Microbiol 49: 145–174.
Franchini AG, Egli T . (2006). Global gene expression in Escherichia coli K-12 during short-term and long-term adaptation to glucose limited continuous culture conditions. Microbiology 152: 2111–2127.
Franz E, Semenov AV, Termorshuizen AJ, de Vos OJ, Bokhorst JG, van Bruggen AHC . (2008). Manure-amended soil characteristics affecting the survival of E. coli O157:H7 in 36 dutch soils. Environ Microbiol 10: 313–327.
Franz E, van Bruggen AHC . (2008). Ecology of E. coli O157:H7 and Salmonella enterica in the primary vegetable production chain. Crit Rev Microbiol 34: 143–161.
Franz E, van Diepeningen AD, de Vos OJ, van Bruggen AHC . (2005). Effects of cattle feeding regimen and soil management type on the fate of Escherichia coli O157:H7 and Salmonella enterica serovar typhimurium in manure, manure-amended soil, and lettuce. Appl Environ Microbiol 71: 6165–6174.
Fremaux B, Delignette-Muller ML, Prigent-Combaret C, Gleizal A, Vernozy-Rozand C . (2007). Growth and survival of non-O157:H7 Shiga-toxin-producing Escherichia coli in cow manure. J Appl Microbiol 102: 89–99.
Fukushima H, Hoshina K, Gomyoda M . (1999). Long-term survival of Shiga toxin-producing Escherichia coli O26, O111, and O157 in bovine feces. Applied and Environ Microbiol 65: 5177–5181.
Gagliardi JV, Karns JS . (2000). Leaching of Escherichia coli O157:H7 in diverse soils under various agricultural management practices. Appl Environ Microbiol 66: 877–883.
Girvan MS, Campbell CD, Killham K, Prosser JI, Glover LA . (2005). Bacterial diversity promotes community stability and functional resilience after perturbation. Environ Microbiol 7: 301–313.
Habteselassie M, Bischoff M, Blume E, Applegate B, Reuhs B, Brouder S . (2008). Environmental controls on the fate of Escherichia coli in soil. Water Air Soil Pollut 190: 143–155.
Hacker J, Carniel E . (2001). Ecological fitness, genomic islands and bacterial pathogenicity. EMBO Rep 5: 376–381.
Herold S, Karch H, Schmidt H . (2004). Shiga toxin-encoding bacteriophages—genomes in motion. Int J Med Microbiol 294: 115–121.
Himathongkham S, Bahari S, Riemann H, Cliver D . (1999). Survival of Escherichia coli O157:H7 and Salmonella typhimurium in cow manure and cow manure slurry. FEMS Microbiol Lett 178: 251–257.
Ihssen J, Egli T . (2004). Specific growth rate and not cell density controls the general stress response in Escherichia coli. Microbiol 150: 1637–1648.
Ihssen J, Egli T . (2005). Global physiological analysis of carbon- and energy-limited growing Escherichia coli confirms a high degree of catabolic flexibility and preparedness for mixed substrate utilization. Environ Microbiol 7: 1568–1581.
Ihssen J, Grasselli E, Bassin C, Francois P, Piffaretti JC, Koster W et al. (2007). Comparative genomic hybridization and physiological characterization of environmental isolates indicate that significant (eco-) physiological properties are highly conserved in the species Escherichia coli. Microbiol 153: 2052–2066.
Ingraham JL, Neidhardt FC . (1987). Escherichia coli and Salmonella typhimurium: Cellular and Molecular Biology. American Society for Microbiology: Washington, USA.
Itoh Y, Sugita-Konishi Y, Kasuga F, Iwaki M, Hara-Kudo Y, Saito N et al. (1998). Enterohemorrhagic Escherichia coli O157:H7 present in radish sprouts. Appl Environ Microbiol 64: 1532–1535.
Jiang X, Morgan J, Doyle MP . (2002). Fate of Escherichia coli O157:H7 in manure-amended soil. Appl Environ Microbiol 68: 2605–2609.
Klein TM, Alexander M . (1986). Bacterial inhibitors in lake water. Appl Environ Microbiol 52: 114–118.
Kudva IT, Blanch K, Hovde CJ . (1998). Analysis of Escherichia coli O157:H7 survival in ovine or bovine manure and manure slurry. Appl Environ Microbiol 64: 3166–3174.
Kudva IT, Evans PS, Perna NT, Barrett TJ, Ausubel FM, Blattner FR . (2002). Strains of Escherichia coli O157:H7 differ primarily by insertions or deletions, not single-nucleotide polymorphisms. J Bacteriol 184: 1873–1879.
Lange R, Hengge-Aronis R . (1991). Identification of a central regulator of stationary-phase gene expression in Escherichia coli. Mol Microbiol 5: 49–59.
Large TM, Walk ST, Whittam TS . (2005). Variation in acid resistance among shiga toxin-producing clones of pathogenic Escherichia coli. Appl Environ Microbiol 71: 2493–2500.
Lin J, Smith MP, Chapin KC, Baik HS, Bennett GN, Foster JW . (1996). Mechanisms of acid resistance in enterohemorrhagic Escherichia coli. Appl Environ Microbiol 62: 3094–3100.
Macfarlane GT, Macfarlane S . (1997). Human colonic microbiota: ecology, physiology and metabolic potential of intestinal bacteria. Scand J Gastroenterol Suppl 222: 3–9.
Mankin KR, Wang L, Hutchinson SL, Marchin GL . (2007). Escherichia coli sorption to sand and silt loam soil. Transactions of the ASABE 50: 1159–1165.
Mead PS, Slutsker L, Dietz V, McCaig LF, Bresee JS, Shapiro C et al. (2000). Food-related illness and death in the United States. J Environ Health 62: 9–18.
Muffler A, Barth M, Marschall C, Hengge-Aronis R . (1997). Heat shock regulation of sS turnover: a role for DnaK and relationship between stress responses mediated by sS and s32 in Escherichia coli. J Bacteriol 179: 445–452.
Muffler A, Traulsen DD, Lange R, Hengge-Aronis R . (1996). Post-transcriptional regulation of the sigma S subunit of RNA polymerase in Escherichia coli. J Bacteriol 178: 1607–1613.
Natvig EE, Ingham SC, Ingham BH, Cooperband LR, Roper TR . (2002). Salmonella enterica serovar Typhimurium and Escherichia coli contamination of root and leaf vegetables grown in soils with incorporated bovine manure. Appl Environ Microbiol 68: 2737–2744.
Nicholson FA, Groves SJ, Chambers BJ . (2005). Pathogen survival during livestock manure storage and following land application. Bioresour Technol 96: 135–143.
Ochman H, Jones IB . (2000). Evolutionary dynamics of full genome content in Escherichia coli. EMBO J 19: 6637–6643.
Oliver DM, Clegg CD, Heathwaite AL, Haygarth PM . (2007). Preferential attachment of Escherichia coli to different particle size fractions of an agricultural grassland soil. Water Air Soil Pollut 185: 369–375.
Ritchie JM, Campbell GR, Shepherd J, Beaton Y, Jones D, Killham K et al. (2003). A stable bioluminescent construct of Escherichia coli O157:H7 for hazard assessments of long-term survival in the environment. Appl Environ Microbiol 69: 3359–3367.
Rozen Y, Belkin S . (2001). Survival of enteric bacteria in seawater. FEMS Microbiol Rev 25: 513–529.
Saleh-Lakha S, Shannon KE, Goyer C, Trevors JT, Zebarth BJ, Burton DL . (2008). Nitric oxide reductase gene expression and nitrous oxide production in nitrate-grown Pseudomonas mandelii. Appl Environ Microbiol 74: 6876–6879.
Sang Ho C, Baumler DJ, Kaspar CW . (2000). Contribution of dps to acid stress tolerance and oxidative stress tolerance in Escherichia coli O157:H7. Appl Environ Microbiol 66: 3911–3916.
Savageau MA . (1983). Escherichia coli habitats, cell types and molecular mechanisms of gene control. Am Nat 122: 732–744.
Schubert S, Darlu P, Clermont O, Wieser A, Magistro G, Hoffmann C et al. (2009). Role of intraspecies recombination in the spread of pathogenicity islands within the Escherichia coli species. PLoS Pathogens 5: e1000257.
Schubert S, Dufke S, Sorsa J, Heesemann J . (2004). A novel integrative and conjugative element (ICE) of Escherichia coli: The putative progenitor of the Yersinia high-pathogenicity island. Molecular Microbiol 51: 837–848.
Semenov AV, Franz E, van Overbeek L, Termorshuizen AJ, van Bruggen AHC . (2008). Estimating the stability of Escherichia coli O157:H7 survival in manure amended soils with different management histories. Environ Microbiol 10: 1450–1459.
Semenov AV, van Bruggen AHC, van Overbeek L, Termorshuizen AJ, Semenov AM . (2007). Influence of temperature fluctuations on Escherichia coli O157:H7 and Salmonella enterica serovar Typhimurium in cow manure. FEMS Microbiol Ecol 60: 419–428.
Semenov AV, van Overbeek L, van Bruggen AHC . (2009). Percolation and survival of E. coli O157:H7 and Salmonella enterica serovar Typhimurium in soil amended with contaminated dairy manure or slurry. Appl Environ Microbiol 75: 3206–3215.
Sjogren RE . (1994). Prolonged survival of an environmental Escherichia coli in laboratory soil microcosms. Water Air Soil Pollut 75: 389–403.
Sjogren RE . (1995). Thirteen-year survival study of an environmental Escherichia coli in field mini-plots. Water Air Soil Pollut 81: 315–335.
Solomon EB, Yaron S, Matthews KR . (2002). Transmission of Escherichia coli O157:H7 from contaminated manure and irrigation water to lettuce plant tissue and its subsequent internalization. Appl Environ Microbiol 68: 397–400.
Tabe ES, Oloya J, Doetkott DK, Bauer ML, Gibbs PS, Khaitsa ML . (2008). Comparative effect of direct-fed microbials on fecal shedding of Escherichia coli O157:H7 and Salmonella in naturally infected feedlot cattle. J Food Protect 71: 539–544.
Tao H, Bausch C, Richmond C, Blattner FR, Conway T . (1999). Functional genomics: Expression analysis of Escherichia coli growing on minimal and rich media. J Bacteriol 181: 6425–6440.
Tauxe R, Kruse H, Hedberg C, Potter M, Madden J, Wachsmuth K . (1997). Microbial hazards and emerging issues associated with produce: a preliminary report to the National Advisory Committee on Microbiologic Criteria for Foods. J Food Protect 60: 1400–1408.
Taylor CM . (2008). Enterohaemorrhagic Escherichia coli and Shigella dysenteriae type 1-induced haemolytic uraemic syndrome. Pediatr Nephrol 23: 1425–1431.
Tilden Jr J, Majkowski J, Hollingsworth J, Morris Jr JG, Young W, Boesel B et al. (1996). A new route of transmission for Escherichia coli: infection from dry fermented salami. Am J Public Health 86: 1142–1145.
Tilman D . (1997). Community invasibility, recruitment limitation, and grassland biodiversity. Ecology 78: 81–92.
Topp E, Welsh M, Tien YC, Dang A, Lazarovits G, Conn K et al. (2003). Strain-dependent variability in growth and survival of Escherichia coli in agricultural soil. FEMS Microbiol Ecol 44: 303–308.
Tóth I, Schmidt H, Kardos G, Lancz Z, Creuzburg K, Damjanova I et al. (2009). Virulence genes and molecular typing of different groups of Escherichia coli O157 strains in cattle. Appl Environ Microbiol 75: 6282–6291.
Touchon M, Hoede C, Tenaillon O, Barbe V, Baeriswyl S, Bidet P et al. (2009). Organised genome dynamics in the Escherichia coli species results in highly diverse adaptive paths. PLoS Genet 5: e1000344.
Trevors JT . (1998). Bacterial biodiversity in soil with an emphasis on chemically-contaminated soils. Water Air Soil Pollut 101: 45–67.
Unc A, Gardner J, Springthorpe S . (2006). Recovery of Escherichia coli from soil after addition of organic wastes. Appl Environ Microbiol 72: 2287–2289.
Van Elsas JD, Jansson JK, Trevors JT . (2007). Modern Soil Microbiology, 2 edn. CRC Press: USA.
Van Elsas JD, Trevors JT, van Overbeek LS . (1991). Influence of soil properties on the vertical movement of genetically-marked Pseudomonas fluorescens through large soil microcosms. Biol Fertil Soils 10: 249–255.
Vital M, Hammes F, Egli T . (2008). Escherichia coli O157 can grow in natural freshwater at low carbon concentrations. Environ Microbiol 10: 2387–2396.
White-Ziegler CA, Davis TR . (2009). Genome-wide identification of H-NS-controlled, temperature-regulated genes in Escherichia coli K-12. J Bacteriol 191: 1106–1110.
Acknowledgements
This study was funded by the NATO Programme Security through Science ESP.EAP.CLG 981785. The Soil Biotechnology Foundation also provided support to R Costa.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
van Elsas, J., Semenov, A., Costa, R. et al. Survival of Escherichia coli in the environment: fundamental and public health aspects. ISME J 5, 173–183 (2011). https://doi.org/10.1038/ismej.2010.80
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2010.80
Keywords
This article is cited by
-
Analyzing the presence of microbial contaminants in water sourced from shallow wells within Dodoma city, Tanzania
International Journal of Energy and Water Resources (2024)
-
About the dark corners in the gene function space of Escherichia coli remaining without illumination by scientific literature
Biology Direct (2023)
-
A study on transport of Escherichia coli through saturated porous medium
International Journal of Environmental Science and Technology (2023)
-
Estimating the population-level prevalence of antimicrobial-resistant enteric bacteria from latrine samples
Antimicrobial Resistance & Infection Control (2022)
-
Synergistic cellulose-based nanocomposite packaging and cold plasma decontamination for extended saffron preservation
Scientific Reports (2022)