Abstract
In nature, the complexity and structure of microbial communities varies widely, ranging from a few species to thousands of species, and from highly structured to highly unstructured communities. Here, we describe the identity and functional capacity of microbial populations within distinct layers of a pristine, marine-derived, meromictic (stratified) lake (Ace Lake) in Antarctica. Nine million open reading frames were analyzed, representing microbial samples taken from six depths of the lake size fractionated on sequential 3.0, 0.8 and 0.1 μm filters, and including metaproteome data from matching 0.1 μm filters. We determine how the interactions of members of this highly structured and moderately complex community define the biogeochemical fluxes throughout the entire lake. Our view is that the health of this delicate ecosystem is dictated by the effects of the polar light cycle on the dominant role of green sulfur bacteria in primary production and nutrient cycling, and the influence of viruses/phage and phage resistance on the cooperation between members of the microbial community right throughout the lake. To test our assertions, and develop a framework applicable to other microbially driven ecosystems, we developed a mathematical model that describes how cooperation within a microbial system is impacted by periodic fluctuations in environmental parameters on key populations of microorganisms. Our study reveals a mutualistic structure within the microbial community throughout the lake that has arisen as the result of mechanistic interactions between the physico-chemical parameters and the selection of individual members of the community. By exhaustively describing and modelling interactions in Ace Lake, we have developed an approach that may be applicable to learning how environmental perturbations affect the microbial dynamics in more complex aquatic systems.
Similar content being viewed by others
Introduction
Microorganisms drive biogeochemical processes that are critical for maintaining the planet in a habitable state (Falkowski et al., 2008). They achieve primary production and global cycling of nutrients through the action of individuals with specific functional traits, performing often diverse roles within a significantly larger community. The make-up and functional effectiveness of a whole community is influenced by numerous biotic and abiotic factors, including the energy and nutrient fluxes of the system and the evolutionary history and trajectory of the microorganisms colonizing the system.
Although the important abiotic characteristics of specific ecosystems can and have been described, it is only recently that both phylogenetic and biological functional inventories have begun to be assembled in ways which can enable the main biological processes within a whole ecosystem to be inferred. Through the use of metagenomics (shotgun sequencing of an environmental sample) and associated functional studies, whole ecosystems have begun to be described. In an acid mine drainage system dominated by only a few species, almost complete composite genomes and metabolic reconstruction of pathways has been achieved (Tyson et al., 2004; Ram et al., 2005). In contrast, the Global Ocean Survey has uncovered an enormous extent of microbial diversity and functional potential of oceanic microbial communities (Rusch et al., 2007).
What has not yet been achieved is to determine if a structured ecosystem with distinct microbial communities can be described in its entirety. Our aim in this study was to not only define the identity and functional capacity of microorganisms, but to establish the interactions between populations that fulfill nutrient cycling and that shape the evolution of the microbial communities. Ultimately, we wished to derive a model that could be used to test the effects of environmental perturbations on the system.
To achieve this goal, we chose to study a pristine, meromictic (permanently stratified) lake, Ace Lake (68.473°S, 78.189°E), located in the Vestfold Hills of East Antarctica (Supplementary Figure S1). Ace Lake was formed at the end of the Quaternary ∼12 000 years BP, and was initially fresh (Gibson, 1999; Rankin et al., 1999; Coolen et al., 2004, 2006; Cromer et al., 2005). Seawater invaded the lake basin during an early Holocene sea level highstand (∼7000 years BP), and the lake was reformed after subsequent sea-level fall after ∼5000 years BP. The ice cover (∼11 months of the year) and salinity gradient (increasing to 4.3% at the bottom) sustain an upper oxic mixolimnion separated by a distinct oxycline to a very stable anoxic monimolimnion in which a methane cycle has been established for ∼3000 years (Figure 1). Light penetration in the lake distinguishes it from other temperate or Alpine meromictic lakes as primary productivity beyond the top few meters can only occur during the summer and the system is largely unaffected by diel cycling.
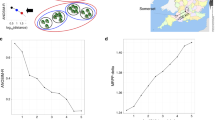
Physicochemical and biological structure of Ace Lake. The habitat of Ace Lake can be divided into three main zones: an upper oxic mixolimnion that extends to 11.5 m, a transition zone corresponding to the halo- and oxycline centered at approximately 12.7 m depth and a lower anoxic monimolimnion (Rankin et al., 1999). Owing to light intensity and snow/ice cover, photosynthetically active radiation (PAR) penetrates ∼2 m in winter and ∼12 m in summer (Rankin et al., 1999). In the panel describing nutrient cycling, both biotic (red arrows) and abiotic (blue arrows) interactions are shown. In the panel describing the biochemical pathways, the width of each arrow is approximately proportional to the significance of that pathway at the depth shown (see Figure 4).
Scientists first visited Ace Lake in 1974, and continuing studies have resulted in it being the most well-characterized meromictic lake in Antarctica (Rankin et al., 1999). The lake is a structured, relatively closed system with marine-derived biota making it ideal for studying the short-term evolution of species and ecosystem function. A range of studies have described archaea, bacteria, eucarya and viruses in the lake, including several unusual bacteria, and the psychrophilic methanogens Methanogenium frigidum and Methanococcoides burtonii (Gibson, 1999; Rankin et al., 1999; Bowman et al., 2000; Coolen et al., 2004, 2006; Cromer et al., 2005; Laybourn-Parry et al., 2005; Madan et al., 2005; Powell et al., 2005; Cavicchioli, 2006; Allen et al., 2009). The lake is sensitive to climatic influences and is a sentinel for monitoring climate change.
Our analyses of this highly stratified ecosystem describe: (a) how the structure of the community and the interactions of microbial populations define biogeochemical fluxes; (b) the importance of size fractionation for generating a greatly improved understanding of microbial ecosystems in the context of resource partitioning; (c) the community response to resource limitation (especially nitrogen) by short circuiting of biogeochemical cycles; and (d) the development of a mathematical model that shows how the apparent cooperation within the microbial ecosystem can be explained by the effects of periodic fluctuations in environmental parameters on key populations of microorganisms.
Materials and methods
Ace Lake samples
Water samples were collected from Ace Lake (68° 24′ S, 78° 11′E), Vestfold Hills, Antarctica on 21 and 22 December 2006. A 2 m hole positioned above the deepest point (25 m depth) of the lake was drilled through the ice cover of Ace Lake to reach the lake surface. A volume of 1–10 l was collected by sequential size fractionation through a 20 μm pre-filter directly onto filters 3.0, 0.8 and 0.1 μm pore-sized, 293 mm polyethersulfone membrane filters (Rusch et al., 2007), along the depth profile as described previously (Ng et al., 2010). Samples were taken in the order, 23, 18, 14, 12.7, 5 and finally, 11.5 m. After samples from each depth were collected, the sample racks were sequentially washed with 2 × 25 l 0.1 N NaOH, 2 × 25 l 0.053% NaOCl, and 2 × 25 l fresh water. The sample hose was flushed with water from each depth before being applied to the filters. A Chlorobium signature was identified at 5 m, but not immediately above the green sulfur bacteria (GSB) layer at 11.5 m. As the next sample taken after sampling at 12.7 m was at 5 m, and then 11.5 m, despite all equipment being thoroughly washed with bleach, sodium hydroxide and water, the simplest explanation for the GSB signature at 5 m is carry-over from sampling of the dense biomass at 12.7 m. A sonde probe (YSI model 6600, YSI Inc., Yellow Springs, OH, USA) was used to record depth, dissolved oxygen content, pH, salinity, temperature and turbidity throughout the water column of the lake. Total organic carbon was determined using a total organic carbon analyzer, TOC-5000A (Shimadzu, Kyoto, Japan) equipped with a ASI-5000A auto sampler (Shimadzu), and particulate organic carbon by standard protocols (http://www.epa.gov/glnpo/lmmb/methods/about.html) at the Centre for Water and Waste Technology, UNSW.
DNA sequencing and data cleanup
DNA extraction and Sanger sequencing was performed on 3730xl capillary sequencers (Applied Biosystems, Carlsbad, CA, USA) and pyrosequencing on GS20 FLX Titanium (Roche, Branford, CT, USA) at the J Craig Venter Institute in Rockville, MD, USA (Rusch et al., 2007). The scaffolds and annotations will be available via Community Cyberinfrastructure for Advanced Microbial Ecology Research & Analysis and public sequence repositories such as National Center for Biotechnology Information and the reads will be available via the NCBI Trace Archive. Sanger reads were trimmed according to quality clear ranges. The quality of pyrosequencing reads was assessed as follows: a Blast nucleotide database was created from the Sanger reads of the 0.1 μm fraction of samples GS230, GS231 and GS232. After blasting the corresponding pyrosequencing reads against each database with a minimum bitscore of 80 and maximum e-value of 0.1, reads were binned according to length. The percentage of reads for each bin lacking a match to the Sanger read database was recorded. The percentage reads at least 25% repetitive after MDUST (Morgulis et al., 2006) analysis at default settings, and the percentage of reads containing N's, were assessed. In contrast to earlier pyrosequencers (Huse et al., 2007), no length-dependent bias in reads containing N's was observed. However, short reads had a disproportionately high number of repeats. Moreover, based on the proportion of reads with no match to the Sanger data set, both very short and very long reads had a disproportionately high number of errors; an observation that was previously reported (Huse et al., 2007). On the basis of this analysis, a three step filtering process was applied to each sample: reads were initially run through the Celera sffToCA (Miller et al., 2008) pre-processor followed by Lucy (Chou and Holmes, 2001) and finally, excluding the bottom 8% and top 3% of reads determined from the read-length distribution. As the sffToCA (v5.3) pre-processor removes all reads with a perfect prefix of any other read it overcomes the ‘perfect duplicates’ problem (Gomez-Alvarez et al., 2009). After this process, <5% of the reads belonged to clusters of duplicates with three or more reads, and Clusters of Orthologous Groups of proteins (COG) classification of these reads showed an over-representation of category L (replication, recombination and repair) that includes mobile genetic elements, which are often duplicated, suggesting a potential biological significance for the duplicated reads. It is possible these residual duplications are a result of high gene copy number or localized fragility of DNA sequences that might be biasing the shear points.
Phylogenetic analyses
Sequence data were compared with the ribosomal database project release 10, update 19 (Cole et al., 2009) using Blastn (e-5) to identify sequences containing bacterial and archaeal 16S ribosomal RNA (rRNA) gene fragments. Only sequences displaying a >100 bp overlap with high scoring pairs were used for further analysis. The taxonomic identification of high scoring pairs was retrieved using the ribosomal database project online tools ‘Seq Cart’ and ‘Classifier’. The percent identity of sequences to ribosomal database project high scoring pairs was taken as a measure of the phylogenetic novelty of each sequence. Sequence data were further compared (Blastn, e-5) with the SILVA100 SSURef database (http://www.arb-silva.de), which contains 47 448 eucaryal 18S rRNA gene sequences. Sequences displaying a >100 bp overlap with high scoring pairs were taxonomically identified by alignment and phylogenetic tree analysis using ARB (www.arb-home.de). The percentage composition of each major clade within each sample and size fraction was displayed using the interactive tree of life (Letunic and Bork, 2006). Sequence data were compared with a manually curated database contain 618 internal transcribed spacer regions from the SAR11 clade (Brown and Fuhrman, 2005). Sequences containing a SAR11-like internal transcribed spacer fragment were manually aligned and inserted into the internal transcribed spacer tree topology using parsimony methods in ARB. Comparison of samples based on ‘order’ level phylogenetic composition was carried out using square-root transformed Bray–Curtis similarities obtained from the an ‘order’ level relative abundance matrix and analyzed by UPGMA clustering and non-metric multidimensional scaling using the Primer v6 package (Clarke and Warwick, 2001).
Read-based analyses
The set of protein-encoding open reading frames (ORFs) for each sample was predicted with MetaGene (Noguchi et al., 2006). Each ORF was translated, all the predicted proteins shorter than 30 amino acids were discarded and the remainder were searched with Blastp against the COG protein database (Tatusov et al., 2001) with a cutoff of 30% minimum identity and e-value of 1e-5 and against the Kyoto Encyclopedia of Genes and Genomes (KEGG) protein database with a maximum e-value of 1e-5. The Blast results were used to assign predicted ORFs to COGs using custom Perl scripts and to KEGGs using KOBAS (Wu et al., 2006). Statistical over-representation of pathways and COG categories was determined by pairwise comparisons of each metagenomic sample using Fisher's exact test, with confidence intervals at 99% significance calculated by the Newcombe–Wilson method and Holm–Bonferroni correction (P-value cutoff of 1e-5) as implemented in STAMP (Parks and Beiko, 2010). Hierachical clustering and heat-plots were generated with R (R Development Core Team, 2009) using the library ‘seriation’. Only COG categories and KEGG pathways with at least 5000 protein assignments throughout the lake were considered. The genetic potential to perform specific steps in biogeochemical cycles of the lake was assessed using a combination of marker genes. Each gene combination was averaged (multiple enzymes/subunits in the same conversion step) or summed (multiple pathways performing the same conversion step) and normalized to 100 000 proteins. As the average genome size for the microbial community at each fraction and depth was similar (2–2.5 Mb), no further normalization was applied (Beszteri et al., 2010). The phylogenetic placement of each read was inferred searching the RefSeq database (Pruitt et al., 2007) with tBlastx and parsing the results with GAAS (Angly et al., 2009) with library hits normalized by database sequence/taxon length using the following cutoffs: minimum percentage identity 60% over a minimum of 60% of the query length and maximum e-value of 1e-03. A Simpson index of diversity was calculated for each sample from the phylogenetic composition using the formula: Ď=1−Σ (ni/N)2, where ni/N is the normalized fraction of the community represented by the i-th species.
Analysis of trophic strategy
Each metagenomic sample was considered in the context of the trophic strategy of the dominant organism(s) as described previously (Lauro et al., 2009) with modifications. As the shorter read length of pyrosequencing data affected the detection of protein localization, a comparison of each 0.1 μm 454 data set with the corresponding Sanger data set was performed. Using the assumption that percentage localizations calculated using PSORT (Gardy et al., 2005) should be the same between the two data sets, a correction factor was computed and applied. Genome size was inferred with GAAS (Angly et al., 2009) and the average number of rRNA operons was estimated by dividing the number of 16S rRNA genes detected in each sample by the average number of proteins assigned to COGs of 32 single-copy genes (COG0012, COG0016, COG0048, COG0049, COG0052, COG0080, COG0081, COG0087, COG0088, COG0090, COG0091, COG0092, COG0093, COG0094, COG0096, COG0097, COG0098, COG0099, COG0100, COG0102, COG0103, COG0124, COG0184, COG0185, COG0186, COG0197, COG0200, COG0201, COG0256, COG0522, COG0533 and COG0541).
Metaproteomic analysis
Proteins were extracted from membrane filters from all 0.1 μm fractions from the six depths (5, 11.5, 12.7, 14, 18 and 23 m), and one-dimensional sodium dodecyl sulfate–polyacrylamide gel electrophoresis and in gel trypsin digestion, liquid chromatography and mass spectrometry (MS), and MS/MS data analysis and validation of protein identifications performed as previously described (Ng et al., 2010), with minor modifications. The spectra generated were searched against the protein sequence database corresponding to that depth constructed from the 0.1 μm mosaic assemblies. Mosaic assemblies were generated for each sample fraction using Celera WGS Assembler v5.3 (Myers et al., 2000). For each assembly, the runtime parameters used were as outlined for 454 sequencing data in the published Standard Operating Procedure (http://sourceforge.net/apps/mediawiki/wgs-assembler/index.php?title=SFF_SOP). As none of the samples can be considered clonal, these are regarded as stringent assemblies (Rusch et al., 2007). Each 0.1 μm fraction assembly was a hybrid of Sanger and 454 read data, wherein the estimated genome size was manually set to minimize the number of unitigs from abundant organisms being falsely classified as degenerate (Rusch et al., 2007). Annotation of each sample fraction assembly was carried out using an in-house pipeline, wherein the pipeline stages consisted of genomic feature detection and subsequent annotation. Detected features consisted of ORFs, transfer RNA and rRNA. Each detected ORF was further annotated by Blast comparison against NR, Swissprot and KEGG-peptide sequence databases and by HMMER comparison against TIGRFAM (Haft et al., 2001), COG (Tatusov et al, 1997; Tatusov et al., 2003) and known marker genes (von Mering et al., 2007). In all cases the cutoff e-value was a maximum of 1e-5. The number of protein sequences in each database were as follows: 5 m, 138 208; 11.5 m, 133 948; 12.7 m, 27 142; 14 m, 62 436; 18 m, 71 512; and 23 m, 128 878. Scaffold (version Scaffold_2_05_01, Proteome Software Inc., Portland, OR, USA) was used to validate MS/MS-based peptide and protein identifications. Peptide and protein identifications were accepted if they could be established at >95% and 99% probability, respectively, as specified by the Peptide Prophet algorithm (Keller et al., 2002). Protein identifications required the identification of at least two peptides. Proteins that contained similar peptides and could not be differentiated based on MS/MS analysis alone were grouped to satisfy the principles of parsimony and are referred to as a protein group. Spectral counting was used to semi-quantitively estimate protein abundance. The total assigned spectra that matched to each identified protein were exported from Scaffold 2.0. For similar proteins that have shared peptides (a protein ambiguity group), spectra were assigned to the protein with the most unique spectra. To normalize for variation in total spectra acquired between sample replicates, the number of spectra of each protein was multiplied by the average total spectra divided by the total spectra of the individual replicate. The spectral count of each protein was averaged across the replicates. As longer proteins are more likely to be detected, the average spectral counts were divided by the length of the protein. This value is equivalent to the normalized spectral abundance factor (Florens et al., 2006; Zybailov et al., 2006). In order to compare the relative abundance of proteins between depths, the normalized spectral abundance factor was divided by the average read depth of the contig (scaffold or degenerate) to which the protein mapped. If >90% of a scaffold's length consisted of surrogate (highly degenerate unitig) sequence, the average read depth of the surrogate was used. For identified proteins that were part of a protein group the longest protein length and largest read depth value in the group was used. Pairwise comparisons of each zone were conducted on COG assigned proteins. The normalized spectral counts from each protein was aggregated based on their COG annotation. All proteins that were part of an ambiguity group were confirmed to share the same COG annotation to ensure counts were not biased because of the common spectra. The summed spectral counts from 5 and 11.5 m (mixolimnion), and 14, 18 and 23 m (monimolimnion) were pooled. Statistical significance of differences between each zone was assessed using Fisher's exact test, with confidence intervals at 99% significance calculated by the Newcombe–Wilson method and Holm–Bonferroni correction (P-value cutoff of 1e-5) in STAMP (Parks and Beiko, 2010). All proteins identified, including their gene identifier, normalized spectral abundance, COG and KEGG orthology identifiers, KEGG locus tag and matching COG or KEGG description are provided in Supplementary Table S1.
Epifluorescence microscopy
Samples of unfiltered lake water and the flow-through from 3.0 and 0.8 μm filters from all depths were collected on November 2008 and fixed on site in formalin 1% (v/v). The samples were stored at −80 °C for subsequent direct counts of cells and viral-like particles (VLPs). Enumeration was performed according to the method of Patel et al. (2007) with the following modifications. Lake water samples were filtered onto 0.01 μm pore-size polycarbonate filters (25 mm Poretics, GE Osmonics, Minnetonka, MN, USA). Filters were air dried, then placed with the back of the filter on top of a 30 μl aliquot of 0.1% (w/v) molten low-gelling-point agarose and allowed to dry at 30 °C. Samples were stained by the addition of 1 μl working solution (1/400 dilution in 0.02 μm filtered sterile Milli-Q) of SYBR Gold (Molecular Probes, Eugene, OR, USA) to 25 μl of mounting medium (VECTASHIELD HardSet, Vector Laboratories, Burlingame, CA, USA). Stained samples were counted immediately, or stored at −20 °C for up to a week before counting. Samples were visualized under wide-blue filter set (excitation 460–495 nm, emission 510–550 nm) with an epifluorescence microscope (Olympus BX61, Hamburg, Germany).
Development and application of the predator–prey model for phototrophic microorganisms
To establish the populations dynamics of the phytoplankton of Ace Lake, we developed a modification of the Lotka–Volterra predator–prey model with a phage predator and a bacterial prey. The original model is described by the following two differential equations:


where P is the number of phage (predator) and A the number of bacteria (prey) and α, ɛ, θ, μ are four parameters, which define the interactions between the phage and the bacterial prey.
We expanded the model assuming that the growth of the population of prey (A) was linearly dependent on light availability (Parkin and Brock, 1980). The light–dark cycles are simulated by a discontinuous parametric sinusoidal function (described in Equation (3)). The prey population can exist in two different states: actively growing (A) and inactive (I). The sum of the active and inactive is the total population (G; Equation (7)). When light is available only the active population can grow. The growth of A is also limited by the carrying capacity (C) of the system. Together, these assumptions make up the first terms of Equation (4). The second term describes the death of the active population in the absence of predation, the third term describes the conversion of inactive cells into active cells (light dependent), the fourth term describes the conversion of active into inactive cells (light independent) and the fifth term describes the effect of predator-caused microbial death, which can only act on the active population. Similarly, Equation (5) describes the changes in the population of inactive cells. This is affected by the flux of cells entering the active state (first term), the flux of cells becoming inactive and the death rate of the inactive cells. It is important to note that the flux of cells becoming active has to be greater than for cells becoming inactive (γ>δ) or the system would spontaneously die out. Also, although not strictly necessary, the model assumes a higher death rate for the inactive population than the active population. This simulates, at least in part, bacterial senescence. Equation (6) is unchanged from the original model of Lotka–Volterra (Equation (2)) and describes the prey-dependent growth of the phage and its death.
The variations of the populations over time are a result of the integration over time of the following system of differential equations:





The integration was performed with the COmplex PAthway SImulator (COPASI) software (Hoops et al., 2006) using the deterministic Livermore solver for ordinary differential equations (LSODA) method. All the parameters were estimated from the available data. The shape of the seasonal light profile was fitted to the seasonal variations in the photosynthetically active radiation available in Ace Lake (Powell et al., 2005). The maximum growth rate was equal, for the Synechococcus strains, to the in situ rates measured in Ace Lake (Powell et al., 2005), and for the GSB strains, to the growth rate at a similar growth temperature of a sister taxon (Overmann and Pfennig, 1989). This rate is much higher than the in situ rates measured in meromictic lakes of temperate waters and is therefore an upper limit. Our model holds true for all GSB slower growth rates. The rates of transition between the active and inactive cell population were chosen to give a growth lag period of approximately 44 h (under constant illumination). This was chosen as a conservative estimate of lag based on a lag of 48 h for sister taxa of GSB when subjected to a shift in nutrients during optimal growth conditions (Overmann and Pfennig, 1989). The graze-free death rates are comparable to those available for natural microbial populations (Pace, 1988) and the model is largely unaffected (within reason) by variations in these parameters. The carrying capacity for each population was set at 1 × 109. The recovery times for the phototrophic populations following an extinction event were simulated by assuming a seed population of a 1000 cells ml−1 in the inactive state. The COPASI implementations of the Ace Lake solution are available on request.
Results and discussion
Phylogenetic stratification of the microbial community
To baseline and monitor microbial community composition and functional microbial processes throughout the lake, a combined metaproteogenomic analysis of Ace Lake was performed (Supplementary Figure S1). Biomass was captured directly through a 20 μm prefilter sequentially onto 3.0, 0.8 and 0.1 μm filters from the aerobic mixolimnion (5 and 11.5 m), the oxycline (12.7 m) and the anoxic monimolimnion (14, 18 and 23 m) (Figure 1 and Supplementary Figure S1). Shotgun sequencing was performed on biomass from all size fractions and depths yielding 741 Mb of Sanger sequencing data and 3.0 Gb of Titanium pyrosequencing data. The Sanger data were trimmed according to quality scores and a novel set of filtering parameters was applied to the 454 data. A total of 8 926 759 ORFs were identified, of which ∼28% were assigned matches to a COG category, and ∼26% to a KEGG pathway (Supplementary Figure S1D). Metaproteomics was performed on matched samples from 0.1 μm filters (Ng et al., 2010), generating 490 612 mass spectra and a total of 1824 protein identifications.
The phylogenetic diversity in the lake was determined by two approaches: analyzing a total of 5054 16S and 218 18S rRNA gene sequences retrieved from the metagenomic data set, and by read-based taxonomic binning of a total of 8 103 379 reads with matches to RefSeq. Read-based taxonomic binning detected organisms lacking rRNA (that is, viruses) and was normalized to genome size, although clades lacking sequenced representatives in RefSeq (that is, candidate divisions) were not able to be detected. Both approaches identified the same major types of cellular phyla within the microbial community throughout the lake (Figure 2 and Supplementary Figures S2 and S3). The Simpson indices of diversity for the read-based analysis (Figure 2) were high in the larger fractions (0.8 and 3.0 μm) in both the mixolimnion and monimolimnion and low in these fractions at the oxycline. In contrast, the 0.1 μm fraction showed a relatively low diversity in the mixolimnion with communities dominated by large algal viruses. The diversity dropped to an even lower level at the oxycline, then increased with depth in the monimolimnion to very high values. Overall, the phylogenetic analyses show that the cellular microorganisms dominating the lake are bacteria, with relatively few eucarya present (mainly in the mixolimnion), and few archaea in the monimolimnion.
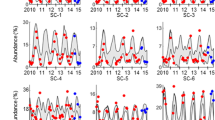
Read-based taxonomic binning of the metagenomic data of Ace Lake. At each depth, the area of each circle is proportional to the percent composition of that clade for the 0.1 μm (red), 0.8 μm (green) and 3.0 μm (blue) fractions. The corresponding Simpson index of diversity ranging from 0 to 1 (right hand y axis) is superimposed as a line graph (calculated as described in Materials and methods) and is shown using the same color scheme for the three size fractions. Within the Proteobacteria, vertical bars represent the fraction of Alphaproteobacteria (purple), Betaproteobacteria (pink), Gammaproteobacteria (yellow), Deltaproteobacteria (green) and Epsilonproteobacteria (blue). The SAR11 clade is the major contributor to the Alphaproteobacteria. Within the viruses, vertical bars represent the fraction of Phycodnaviridae (purple), Siphoviridae (pink), Myoviridae (yellow), Podoviridae (green) and other (blue) viral families.
The read-based phylogenetic assignments showed that all size fractions in the mixolimnion were dominated by large algal viruses (Phycodnaviridae), and was consistent with total bacterial (∼9 × 105 cells ml−1) and viral (∼5 × 106 VLP ml−1) counts (Supplementary Figure S4). The algal virus was also detected in the monimolimnion, although no eucaryal sequences (belonging to an algal host) were detected in the monimolimnion indicating that viruses are likely to be associated with sinking particulate organic matter. A large proportion (22%) of the 18S rRNA gene sequences detected in the mixolimnion were related to Mantoniella (Supplementary Figure S3A), a picoeucaryal member of the Chlorophyta, which is likely to be the host for the Phycodnavirus. The abundance of photosynthetic nanoflagellates has been reported to peak early in summer and decline through December (Madan et al., 2005). However, because of a lack of or negative correlation between protists and VLPs in the mixolimnion, it was reported that protists were not the viral hosts and that the VLPs were bacteriophages (Madan et al., 2005). Instead, our data indicate that by late December the viral population is dominated by Phycodnaviruses that may control the abundance of Mantoniella and thereby make an important contribution to the seasonal algal population dynamics, similar to their role in some freshwater Antarctic lakes (López-Bueno et al., 2009).
Consistent with Ace Lake's marine origin, the ubiquitous marine SAR11 clade represented ∼20% of the 16S rRNA genes and >40% of the read-based phylogenetic assignments normalized to genome size as a proportion of the bacteria in the 0.1 μm fraction of the mixolimnion (Figure 2 and Supplementary Figure S2). Analysis of SAR11-related 16S–23S rRNA internal transcribed spacer region sequences revealed representatives from three clusters generally found at different depths and habitat types (Supplementary Figure S3B). The SAR11 surface 1 cluster sequences were predominant, and closely related to Pelagibacter ubique HTCC1062 and HTCC1002 strains. Representatives from this cluster are found in cold-temperate or polar regions, including Antarctica (Brown and Fuhrman, 2005). Two SAR11 surface 3 cluster sequences were associated with a cluster that only contained other sequences derived from Antarctic samples (Garcia-Martinez and Rodriguez-Valera, 2000). The Actinobacteria sequences in the mixolimnion were associated with a diverse phylogenetic cluster (Luna cluster) mainly contributed by freshwater ultramicrobacteria (Hahn et al., 2003). Several Luna cluster isolates contain rhodopsin genes (Sharma et al., 2008) and similar genes were present in the Ace Lake mixolimnion and from metaproteomic data found to be expressed (167820670 and 163154474). However, the major 16S rRNA gene assembly was most closely related to sequences from Lake Bonney, a saline lake in the McMurdo Dry Valley region. The other major bacterial component of the mixolimnion was cyanobacteria of the genus Synechococcus, which dominated the 0.8 and, to a lesser extent, 3.0 μm fractions. Synechococcus, representing a unique and highly adapted clade, have previously been identified in Ace Lake with numbers peaking in summer (up to 8 × 106 cells ml−1) at depth (10–11 m) where they represented the dominant type of phytoplankton (Powell et al., 2005).
Overall the community composition of the mixolimnion is sufficiently similar to marine surface waters to indicate that important marine species are capable of adapting to the lacustrine conditions prevailing in Ace Lake. However, the species richness in the mixolimnion is approximately one order of magnitude lower than the open ocean. Moreover, some clades (for example, Actinobacteria) are over-represented in the mixolimnion of Ace Lake. Collectively, these observations suggest that re-seeding is a relatively rare phenomenon and that a strong selection against immigrant species is now in place in the mixolimnion.
Proteobacteria were significant members of the community at all depths but both the 16S rRNA gene and the read-based analyses detected a shift from Alphaproteobacteria and Deltaproteobacteria in the mixolimnion to Deltaproteobacteria, Epsilonproteobacteria and Gammaproteobacteria at depth (Figure 2 and Supplementary Figure S2). The proportion of Gammaproteobacteria and Deltaproteobacteria was higher in the larger size fractions, indicative of large-sized cells, cell aggregates or cells attached to particles. The size fractionation not only reflected cell size differences (Supplementary Figure S4), but also trophic distinction (Lauro et al., 2009), with the cells captured on the 0.8 and 3.0 μm fractions being overall less oligotrophic than the 0.1 μm fraction (Supplementary Figure S5).
At 12.7 m depth, the light levels and the sharp transition in oxygen content and salinity (Supplementary Figure S1) favor the dominance of a very high-density (2.2 × 108 cells ml−1) of a single type of GSB of the genus Chlorobium, referred to as C-Ace (Ng et al., 2010). Viral signatures were essentially absent from this zone. This is consistent with the identification of clustered regularly interspaced short palindromic repeats (CRISPR)-associated (CAS) proteins Cse2, Cse3 and Cse4 (165526330, 165526332 and 165526334, respectively) in the 12.7 m metaproteome. The CAS gene locus (cas3, cse1, cse2, cse3, cse4, cas5 and cas1b), to which the proteins map, shares its organization with CAS loci of sequenced GSB, and groups with the Escherichia coli subtype/variant 2. The CRISPR/CAS system is likely to confer phage resistance to C-Ace, akin to its role in other organisms (Horvath and Barrangou, 2010). This is further discussed in ‘community dynamics in response to periodic fluctuations in light’ below.
Below the GSB zone, genomic signatures were for a highly diverse and novel community of anaerobic bacteria and archaea that would be involved in remineralizing organic matter. Candidate divisions OD1 and OP11 were detected in the 16S rRNA gene analysis and because of the lack of reference genomes in RefSeq, reads were probably assigned to sister phyla Chloroflexi and Spirochaetes, respectively (Supplementary Figure S2). Microbial counts in the monimolimnion were as high as 1.6 × 107 cells ml−1 (Supplementary Figure S4). The larger size fractions (0.8 and 3.0 μm) contained a high proportion of GSB and cyanobacteria (Figure 2), and functionally clustered with the 12.7 m/3.0 μm fraction (Figure 3 see ‘stratification of microbial processes and nutrient cycles’ below). The metaproteomic signatures for the 0.1 μm fraction were consistent with reduced metabolic activity of GSB in the monimolimnion (for example, reduced detection, and hence abundance of GSB ribosomal proteins and key anaerobic carbon fixation and fermentation enzymes; Supplementary Figure S6). These data are consistent with the monimolimnion receiving microbial aggregates sinking from shallower depths of the water column (Figure 1).
Heatmap plot and functional clustering of the predicted ORFs from the metagenomic reads based on COG categories. Supplementary Figure S7 shows a similar clustering based on KEGG pathways. Plots were generated as described in Materials and methods. M, Cell wall/membrane/envelope biogenesis; J, Translation, ribosomal structure and biogenesis; L, Replication, recombination and repair (including transposases); R, General function prediction only; E, Amino acid transport and metabolism; C, Energy production and conversion; G, Carbohydrate transport and metabolism; K, Transcription; O, Posttranslational modification, protein turnover, chaperones; S, Function unknown; F, Nucleotide transport and metabolism; H, Coenzyme transport and metabolism; P, Inorganic ion transport and metabolism; I, Lipid transport and metabolism; T, Signal transduction mechanisms; V, Defense mechanism; U, Intracellular trafficking, secretion and vesicular transport; Q, Secondary metabolites biosynthesis, transport and catabolism; D, Cell cycle control, cell division and chromosome partitioning; N, Cell motility. The heat scale is the percentage of ORFs assigned to each individual COG category. Among the COG categories of genes that defined the uniqueness of the 0.1 μm fractions of the monimolimnion were those for replication (L) and cell envelope (M); categories, which have previously been associated with temperature adaptation in M. burtonii (Allen et al., 2009). Eucaryotic-specific COG categories (B, Z, W, Y) were present in numbers below the cutoff (<5000 protein assignments throughout the lake) and are therefore not shown.
Viral signatures re-appeared in the monimolimnion dominated by bacteriophages (Siphoviridae, Myoviridae and Podoviridae). Owing to the size of the bacteriophages, extracellular forms are likely to pass through the 0.1 μm filter. The presence of a significant number of bacteriophage sequences is therefore indicative of predominantly lysogenic forms or cell-associated virions. The ratio of bacteriophage to total viral population increased proportionally in the larger size fractions consistent with trophic analyses that indicate that the larger size fractions are mostly copiotrophic (Supplementary Figure S5) particle-attached bacteria and therefore likely to be sensitive to lysogenic phage infection (Lauro et al., 2009). The 23 m unfiltered lake water contained very high levels (1.3 × 108 VLPs ml−1) of VLPs. The high diversity of bacteria and archaea in all size fractions of the monimolimnion (Figure 2) is consistent with the presence of a high viral population (Rodriguez-Valera et al., 2009).
Stratification of microbial processes and nutrient cycles
COG and KEGG functional gene assignments were compared across all size fractions and depths (Figure 3 and Supplementary Figure S7). The 0.1 μm fractions of the monimolimnion were found to be most functionally similar to each other while also being characterized by extremely high diversity (Figure 2 Simpson index >0.9). Although a large number of predicted genes (11–35%) were able to be assigned to KEGG and COG categories, the percentage of reads that could be assigned to a phylum decreased with depth, dropping to <3% at 23 m (Supplementary Figure S1D). This implies a high degree of phylogenetic novelty in the anoxic zone, but also a high degree of functional similarity. In parallel with taxonomic diversity increasing with depth (with the exception of the GSB layer), the rate of metaproteomic identifications decreased with depth; for example, at 23 m, 67% were hypothetical proteins that tended to lack orthologs in well-characterized organisms.
Across all zones in the lake, genes with transport functions were represented in the metagenome data. The diversity and abundance of ABC-transporters was lowest in the 0.1 μm fractions at 23 m (Figure 3 and Supplementary Figure S7), and a correspondingly low number were detected in the metaproteome. In contrast, numerous transporters, predominantly ABC type, were identified in the metaproteome of the mixolimnion samples, with a high COG representation of transporters for carbohydrates (∼34% of normalized spectra), amino acids (∼32%) and inorganic ions (∼9%) (Supplementary Figure S6). All transporters in the metaproteome were of bacterial origin and conservative phylum-level assignments of the normalized spectra showed the majority to originate from Proteobacteria (69%), of which SAR11 comprised 46% and Actinobacteria 19%. From metaproteome analyses, a high proportion of expressed genes with transport functions have also been reported for SAR11 from coastal (Poretsky et al., 2010) and open ocean waters (Sowell et al., 2009; Morris et al., 2010). Oligotrophs, such as SAR11 not only possess a low diversity of high-affinity transporters (Lauro et al., 2009) but regulate the relative abundance of transporters expressed in response to dissolved organic carbon (DOC) availability (Poretsky et al., 2010). The prevalence of amino acid and simple sugar transporters, and the low DOC concentration in the Ace Lake mixolimnion (Figure 1) are likely to reflect efficient utilization of these substrates from the DOC pool.
Two SAR11 transport proteins that were detected in the Ace Lake metaproteome were not detected in the Sargasso Sea (Sowell et al., 2009): an ectoine/hydroxyectoine (167807477 and 167892279) and a zinc ABC transporter (167933120). The zinc ABC transporter is likely to support zinc efflux in response to zinc concentrations, which are ∼70-fold higher in the mixolimnion of Ace Lake compared with seawater (Rankin et al., 1999). Conversely, phosphate transporters were a major class detected from the Sargasso Sea (Sowell et al., 2009) but were absent from the Ace Lake metaproteome, consistent with lower phosphate levels in the Sargasso Sea (<5 nM) compared with Ace Lake (1–12 μM). The differences in transporter expression identified from metaproteome data between Ace Lake and oceanic SAR11 are likely to signify adaptive growth strategies, which have evolved in the Ace Lake SAR11 community.
Gene function (Figure 3 and Supplementary Figure S7), expression (Supplementary Figure S6) and taxonomy (Figure 2 and Supplementary Figure S2 and S3) were used in combination with physical data (Supplementary Figure S1) to infer the ways in which carbon, nitrogen and sulfur are cycled (Figure 4) by the microbial communities throughout the lake (Figure 1). The data point to carbon cycling being strongly influenced by anaerobic carbon fixation at the oxycline by GSB in all fractions and sulfate-reducing bacteria (SRB) in the 0.8 and 3.0 μm fractions, and fermentation in the monimolimnion by communities in the 0.8 and 3.0 μm fractions (Figure 4a). Aerobic carbon fixation would occur in the mixolimnion mediated by cyanobacteria present in the 0.8 μm fraction (Figure 4a).
The carbon, nitrogen and sulfur cycles in Ace Lake. The genetic potential for each conversion step was estimated in each of the three size fractions (red, 0.1 μm; green, 0.8 μm; and blue, 3.0 μm) using a combination of normalized marker gene ratios as summarized below. The size of each arrow is proportional to the total potential flux (sum of all the genetic potential for that step in the lake). The phylogeny of sequences with KEGG hits were examined in order to infer which types of microorganisms were likely to be capable of specific functional processes. (a) Carbon cycle. Anaerobic carbon fixation=K00174+K00175+K00244+K01648)/4+(K00194+K00197)/2; aerobic carbon fixation=(K00855+K01602)/2; fermentation=K00016+(K00169+K00170)/2; aerobic respiration=(K02256+K02262)/2+(K02274+K02276)/2; methanogenesis=(K00400+K00401)/2; aerobic methane oxidation=K08684; CO oxidation=(K03518+K03519+K03520)/3. The KO (KEGG Orthology) numbers correspond to the following marker genes: adenosine triphosphate (ATP) citrate lyase (K01648); 2-oxoglutarate:ferredoxin oxidoreductase subunit alpha (K00174); 2-oxoglutarate:ferredoxin oxidoreductase subunit beta (K00175); fumarate reductase flavoprotein subunit (K00244); CO dehydrogenase subunit delta (K00194); CO dehydrogenase subunit gamma (K00197); phosphoribulokinase (K00855); RuBisCO small chain (K01602); L-lactate dehydrogenase (K00016); pyruvate:ferredoxin oxidoreductase alpha subunit (K00169); pyruvate:ferredoxin oxidoreductase beta subunit (K00170); cytochrome c oxidase subunit I (coxI) (K02256); cytochrome c oxidase subunit III (coxIII) (K02262); cytochrome c oxidase subunit I (coxA) (K02274); cytochrome c oxidase subunit III (coxC) (K02276); coenzyme M methyl reductase beta subunit (mcrB) (K00401); methyl coenzyme M reductase system, component A2 (K00400); methane monooxygenase (K08684); CO dehydrogenase small subunit (coxS) (K03518); CO dehydrogenase medium subunit (coxM) (K03519); CO dehydrogenase large subunit (coxL) (K03520). (b) Nitrogen cycle. Nitrogen assimilation=(K00360+K00367+K01915+K00265+K00284)/3; denitrification=(K02305+K04561+K00376)/3; nitrogen fixation=(K00531+K02586+K02588+K02591)/4; ammonification=K05904+K03385; mineralization=K00260+K00261+K00262; anammox=K10535; Nitrification=no genes for the oxidation of ammonia (K10944, K10945, K10946) were detected. The KO (KEGG Orthology) numbers correspond to the following marker genes: assimilatory nitrate reductase (K00360); assimilatory nitrate reductase (K00367); glutamine synthetase (glnA) (K01915); glutamate synthase (NADPH/NADH) large chain (gltB) (K00265); glutamate synthase (ferredoxin-dependent) (gltS) (K00284); nitric-oxide reductase (norC) (K02305); nitric-oxide reductase (norB) (K04561); nitrous oxide reductase (nosZ) (K00376); nitrogenase (K00531); nitrogenase molybdenum-iron protein alpha chain (nifD) (K02586); nitrogenase iron protein (nifH) (K02588); nitrogenase molybdenum-iron protein beta chain (nifK) (K02591); cytochrome c nitrite reductase (nrfA) (K05904); formate-dependent nitrite reductase periplasmic cytochrome c552 (nrfA) (K03385); glutamate dehydrogenase (K00260, K00261, K00262); hydroxylamine oxidoreductase/hydrazine oxidoreducatse (hao/hzo) (K10535); ammonia monooxygenase subunit A (amoA) (K10944); ammonia monooxygenase subunit B (amoB) (K10945); ammonia monooxygenase subunit C (amoC) (K10946). (c) Sulfur cycle. Assimilatory sulfate reduction=(K00860+K00956+K00957)/3; mineralization=K00456+K01011; dissimilatory sulfate reduction*=(K00394+K00395+K00396)/3; sulfide oxidation*=(K00394+K00395+K00396)/3. *As marker genes K00394, K00395, K00396 can operate in both an oxidative and a reductive way, they were assigned to the sulfate reduction step if they had a best match within KEGG to an ortholog from a sulfate-reducing clade. Similarly, they were assigned to sulfide oxidation if the best match was to an ortholog from a sulfur-oxidizing clade. The KO (KEGG Orthology) numbers correspond to the following marker genes: adenylylsulfate kinase (cysC) (K00860); sulfate adenylyltransferase subunit 1 (cysN) (K00956); sulfate adenylyltransferase subunit 2 (cysD) (K00957); adenylylsulfate reductase subunit A (aprA) (K00394); adenylylsulfate reductase subunit B (aprB) (K00395); sulfite reductase (dsrA) (K00396); cysteine dioxygenase (K00456); 3-mercaptopyruvate sulfurtransferase (K01011).
In the mixolimnion, the bacterial members responsible for remineralization of particulate organic carbon to DOC include members of the Flavobacteria and Gammaproteobacteria, which are likely to be particle-associated copiotrophs (Supplementary Figure S5). Heterotrophic conversion would be further performed by free-living, oligotrophic Actinobacteria and members of the SAR11 clade. In the monimolimnion, the combination of fermentative, sulfate-reducing and methanogenic microorganisms would decompose particulate organic carbon into smaller molecules, ultimately to CO2 and CH4. The incomplete biotic oxidation of these organic compounds would lead to the production of carbon monoxide (CO). Consistent with this, CO dehydrogenase genes are found at significant levels throughout the lake indicating that CO oxidation might be an important pathway for energy generation. CO oxidation could explain why the lake is supersaturated in inorganic carbon with a net export of carbon from the monimolimnion to the atmosphere (during periods that are ice free) (Rankin et al., 1999). CO dehydrogenase activity from anaerobic carboxydotrophs (for example, SRB, archaea) would generate other energy-yielding substrates (for example, H2 and acetate) while maintaining non-inhibitory CO concentrations (Oelgeschläger and Rother, 2008). Consistent with this hypothesis the phylogenetic affiliation of CO dehydrogenase was primarily to Alphaproteobacteria and Betaproteobacteria in the mixolimnion and to Archaea in the monimolimnion.
High levels of methane present at depth can be linked to slow rates of accumulation by a relatively low abundance of Euryarchaeota (∼2–5% of the total community), coupled to the very low potential for aerobic methane oxidation and the absence of 16S rRNA signatures for ANME Euryarchaeota. One of the two methanogenic archaea isolated from the lake, M. burtonii, possesses several genes most closely related to Epsilonproteobacteria and Deltaproteobacteria (Allen et al., 2009). The relative abundance of Epsilonproteobacteria and Deltaproteobacteria (Figure 2), and an overrepresentation of transposable elements (Figure 3 and Supplementary Figure S6) is consistent with gene exchange occurring within the microbial community in this zone, particularly between the Epsilonproteobacteria and Deltaproteobacteria and M. burtonii; the normalized number of transposases throughout the Ace Lake metagenome is two orders of magnitude higher than in marine metagenomes (for example, Global Ocean Survey). The high numbers of bacteriophages in the monimolimnion (detected by microscopy, Supplementary Figure S4; metaproteomics, Supplementary Figure S6; metagenomics, Figure 2), and increase in DOC observed at depth (Figure 1), also indicates that carbon turnover in the monimolimnion is likely to be tightly coupled to the carbon flux going through a viral shunt, as proposed for open ocean systems (Suttle, 2005). The bacteriophages are also likely vehicles for mediating gene exchange.
Nitrogen fixation in the lake (Figure 4b) is likely to be driven by the dense population of GSB. A previous study reported that Nif proteins were not detected in the metaproteome and that the GSB were likely to be assimilating ammonia (Ng et al., 2010). Although it is likely that at the time of sampling the levels of ammonia present at the oxycline were sufficient to inhibit the expression of nitrogen fixation genes, when ammonia levels drop diazotrophy would need to occur. The concentration of reduced nitrogen (ammonium plus amino acids) measured in the lake is low in the mixolimnion, rising through the GSB zone to reach maximum levels in the monimolimnion of ∼55 μM (Rankin et al., 1999). A similar increase in total nitrogen occurs through the monimolimnion, reaching highest levels at the bottom of the lake (Rankin et al., 1999). Diffusion of reduced nitrogen into the mixolimnion would provide an important source of nitrogen for phototrophs. Low rates of diazotrophy are also likely to occur in the mixolimnion (for example, by cyanobacteria) and monimolimnion (Figure 4b). In the monimolimnion, methanogens that encode nif genes, such as M. burtonii (Allen et al., 2009), could benefit from nitrogen fixation. Symbiotic relationships between diazotrophic archaea and SRB that have been reported to occur in marine environments (Dekas et al., 2009) may also occur in Ace Lake.
Most of the genetic potential to cycle the nitrogen pool appears to be limited to nitrogen assimilation throughout the lake and remineralization in the monimolimnion (Figure 4b). The detection of glutamine synthetase (GlnA) and glutamate synthases (GltBS) in the metaproteome are supportive of active nitrogen assimilation. In the mixolimnion, GlnA was linked to SAR11 and Actinobacteria, and they are likely to be responsible for nitrogen absorption in the oxic zone. At the oxycline, GlnA and GltB from GSB were abundant, indicating an important role for nitrogen assimilation at this zone in the lake. In contrast, the microbial community in Ace Lake is characterized by the absence of genes for ammonia oxidation (amoA, amoB and amoC), and a very low number of 16S rRNA genes for typical nitrifying bacteria and archaea. Nitrate levels in the lake remain low throughout the year (Rankin et al., 1999), although the limited input from snow melt may select for the retention of the small genetic capacity for ammonification (Figure 4b). The persistent lack of bioavailable nitrogen (nitrate, nitrite) may not only lead to a lack of nitrification, but could cause the loss of species that can perform nitrification. The absence of nitrification could also be a mechanism to conserve bioavailable nitrogen.
The only potential biotic source of nitrogen loss appears to be from anaerobic ammonia oxidation (anammox) in the monimolimnion mediated by putative HAO/HZO (K10535) proteins. If this occurs, it may be carried out by Planctomycetes, which are known to perform anammox and whose molecular signatures increase with depth (Supplementary Figure S2). Overall, the higher capacity to produce and consume ammonia in the anoxic zone of the lake could lead to a periodic accumulation of ammonia and hence regulation of the input flux of atmospheric nitrogen. The absence of nitrification and low levels of denitrification are fundamental to the survival of the community because of their dependence on the seasonal nitrogen input from the GSB layer. The pseudo-cooperative behavior can be explained by the nitrifiers being outcompeted by heterotrophs assimilating nitrogen during periods of extreme nitrogen limitation (Taylor and Townsend, 2010).
The main turnover of sulfur compounds occurs at the oxycline (Figure 4c). The GSB that dominate the 0.1 μm fraction (predominantly short rods; Supplementary Figure S4B), consume sulfide and appear to be oligotrophic (Supplementary Figure S5). In turn, the SRB that dominate the 3.0 μm fraction (largely filamentous cells; Supplementary Figure S4B) reduce sulfate copiotrophically (Supplementary Figure S5). Genes for assimilatory sulfate reduction were present in metagenome data of all three fractions at all depths, although they were lowest at the oxycline. However, there was no evidence for expression of the genes as assimilatory sulfate reduction proteins were not detected by metaproteomics. In contrast, multiple subunits of the GSB dissimilatory sulfide reductase complex were identified (Ng et al., 2010) indicating functionality of this pathway at the oxycline. GSB likely utilize the dissimilatory sulfide reductase system to convert sulfur to sulfite and the polysulfide-reductase-like complex 3 to oxidize sulfite to sulfate. SRB may then reform sulfide completing the sulfur cycle between the GSB and SRB (Ng et al., 2010). Although SRB were detected at the three depths of the monimolimnion, sulfate is depleted in the water column and sediment at the bottom of the lake limiting their dissimilatory capacity (Rankin et al., 1999). Finally, sulfate in the mixolimnion can be linked to the activity of reverse adenosine-5′-phosphosulfate reductase that has been observed in SAR11 genomes (Meyer and Kuever, 2007) and a concomitant incapacity to perform sulfur reduction.
Community dynamics in response to periodic fluctuations in light
In an Alpine freshwater meromictic lake (Lake Cadagno), a Chlorobium species was reported to have become dominant (by 2006), after replacing purple sulfur bacteria and other strains of GSB that had previously been abundant (before 2000) (Halm et al., 2009; Gregersen et al., 2009). However, the available data (Coolen et al., 2006) suggest that the same species has been dominant, if not unique, in the Ace Lake oxycline since its inception. In fact, the GSB in Ace Lake is essentially monophyletic (Ng et al., 2010), and we observe high cell counts, low VLP counts (Supplementary Figure S4) and no viral sequences in the GSB zone. As a result, we hypothesize that the GSB population in Ace Lake represents an exception to the viral/bacterial population dynamics that has been generally ascribed to aquatic ecosystems (Rodriguez-Brito et al., 2010) and suggest that this may be due to synchronicity in the GSB population caused by the annual, polar dark–light cycles.
To test this hypothesis, we developed a modified Lotka–Volterra model in which the growth of a GSB population was dependent on light intensity (Figure 5). The model is based on the assumption that the population can exist either in an active state in which growth and predation can occur, or an inactive state (Figure 5a). A constant proportion of the bacterial population enters the inactive state while reversion to the active state happens at much higher rates and only in the presence of light. The yearly photosynthetically active radiation profile at the oxycline in Ace Lake was included (Figure 5c) based on published data (Powell et al., 2005). The model predicts that the ability of the GSB population to persist in Ace Lake is dependent on the absence of phage predators; in the presence of phage, the GSB population would be almost eradicated (Figure 5h) and not recover for many years (>5 years) (Figure 5d). As a result of the lake's dependence on the cycling of carbon, nitrogen and sulfur by the GSB, phage-mediated destruction of the GSB would cause mass extinction events throughout the whole lake community. We do not know if the lack of phage predation is a result of decreased virulence (Wild et al., 2009) or an inherent property of the strain of Chlorobium in Ace Lake. We do note that the presence of a highly virulent phage could have led to the extinction of the community in the past, but this appears unlikely as it would imply that the only Chlorobium strain capable of occupying the oxycline in Ace Lake is the current one. It is also possible that the creation of a phage-free niche for the GSB stems from the incapacity of the viral population to adapt to the long cycles of inactivity of hosts leading to their disappearance early in the development of the community.
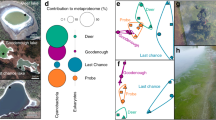
Effects of phage predation on phototrophs and time of population recovery in Ace Lake assessed with a modified Lotka–Volterra model. Schematic diagram describing the model including parameters used for the model (a). The cyclic abundance of the populations of GSB (b) and cyanobacteria (e) in Ace Lake was dependent on the seasonal light–dark cycles at the oxycline (c) and mixolimnion (f), respectively. Similarly, the establishment in available niche-space of a new population of GSB (d) and cyanobacteria (g) is simulated over time. The introduction of phage (purple) predation in Ace Lake during the summer (note the different time scale) leads to either the extinction of the GSB (h) or seasonal oscillations in the cyanobacterial (i) populations. The effects are shown of the introduction of phage (purple) predation in a slow-growing phototrophic population (for example, GSB) (j) or fast-growing phototrophic population (for example, cyanobacteria) (k) and the recovery of the slow- (l) and fast-growing populations (m) in a meromictic lake with diel cycling. Number of GSB active cells (blue); number of GSB inactive cells (green); total number of GSB cells (cyan); number of phage particles (purple); light (red).
The model also predicts that the GSB population of Ace Lake undergoes approximately a 10-fold fluctuation in number between summer and winter, and a major transition between a population that is almost completely active in summer and essentially inactive at the end of the winter (Figure 5b). The population fluctuation in the absence of predation is shown (Figure 5e). In addition to GSB, the model predicts that other phototrophs of Ace Lake with higher maximum growth rates than GSB that occupy niches in shallower waters (with different light profiles) would experience strain cycling (Figures 5g and i). This is compatible with the metagenomic signatures of multiple strains of Synechococcus in the mixolimnion and knowledge that Synechococcus bloom and wane (Powell et al., 2005). This prediction is also consistent with our data pointing to the end of a eucaryal algal bloom having just occurred in the lake (see above).
Finally, our model offers a view about the cycling and coexistence of multiple strains of Chlorobium with niche overlap in meromictic lakes with higher periodicity, such as Lake Cadagno (Figure 5j). Even in a more rapidly oscillating environment, the persistence of very high numbers of GSB could be explained by a significant fraction of the population being inactive and therefore resistant to phage predation. The model predicts that, when present, a phage predator would introduce persistent oscillations in the population of each strain (Figure 5j) while the niches left available could be occupied by further strains (or different species of a similar ecotype) in <2 years (Figure 5l). In the case of faster growing phototrophs, the increased frequency of the oscillations would maintain the populations under stricter control resulting in bacterial numbers that peak one order of magnitude lower than those of slow growing GSB (Figure 5k) and would allow for the rapid niche colonization by new strains (Figure 5m). These predictions are consistent with available data for GSB and higher water phototrophs from Lake Cadagno (Musat et al., 2008).
The loss of pathways (for example, nitrification), the expansion of gene families (for example, mobile elements) and the selection of genomes (for example, GSB, viruses), which are key elements for the functioning of the Ace Lake ecosystem are all necessary steps for the transition from a marine to a meromictic lake ecosystem. We posit that this occurred through the periodic extinction of populations and/or communities with overall low inclusive fitness during the early annual lake cycles until a stable combination was obtained. In this framework, the lake community would be at present very delicate because of its low diversity (McCann, 2000) and its reliance on key components, which would make the community overall extremely fit but also very specialized. In particular, as a result of the extreme lack of diversity of the GSB and their critical role in the lake's carbon, nitrogen and sulfur cycles, the GSB may be easily lost as a result of environmental perturbation, including climate change. The long (estimated in many years) recovery of the community would undoubtedly alter the lake biogeochemistry forever.
References
Allen M, Lauro FM, Williams TJ, Burg D, Siddiqui KS, De Francisci D et al. (2009). The genome sequence of the psychrophilic archaeon, Methanococcoides burtonii: the role of genome evolution in cold-adaptation. ISME J 3: 1012–1035.
Angly FE, Willner D, Prieto-Davó A, Edwards RA, Schmieder R, Vega-Thurber R et al. (2009). The GAAS metagenomic tool and its estimations of viral and microbial average genome size in four major biomes. PLoS Comput Biol 5: e1000593.
Beszteri B, Temperton B, Frickenhaus S, Giovannoni SJ . (2010). Average genome size: a potential source of bias in comparative metagenomics. ISME J 4: 1075–1077.
Bowman JP, McCammon SA, Rea SM, McMeekin TA . (2000). The microbial composition of three limnologically disparate hypersaline Antarctic lakes. FEMS Microbiol Lett 183: 81–88.
Brown MV, Fuhrman JA . (2005). Marine bacterial microdiversity as revealed by internal transcribed spacer analysis. Aquat Microb Ecol 41: 15–23.
Cavicchioli R . (2006). Cold adapted archaea. Nature Rev Microbiol 4: 331–343.
Chou H-H, Holmes MH . (2001). DNA sequence quality trimming and vector removal. Bioinformatics 17: 1093–1104.
Clarke KR, Warwick RM . (2001). Change in marine communities: an approach to statistical analysis and interpretation, 2nd edn. PRIMER-E: Plymouth.
Cole JR, Wang Q, Cardenas E, Fish J, Chai B, Farris RJ et al. (2009). The ribosomal database project: improved alignments and new tools for rRNA analysis. Nucleic Acids Res 37: D141–D145.
Coolen MJL, Hopmans EC, Rijpstraa WIC, Muyzera G, Schoutena S, Volkman JK et al. (2004). Evolution of the methane cycle in Ace Lake (Antarctica) during the Holocene: response of methanogens and methanotrophs to environmental changes. Org Geochem 35: 1151–1167.
Coolen MJL, Muyzer G, Schouten S, Volkman JK, Sinninghe Damste JS . (2006). Sulfur and methane cycling during the Holocene in Ace Lake (Antarctica) revealed by lipid and DNA stratigraphy. In: Neretin LN (ed). Past and Present Marine Water Column Anoxia (NATO Science Series: IV-Earth and Environmental Sciences). Springer: Dordrecht, The Netherlands, pp 41–65.
Cromer L, Gibson JAE, Swadling KM, Ritz DA . (2005). Faunal microfossils: indicators for ecological change in an Antarctic saline lake. Palaeogeogr Palaeoclimatol Palaeoecol 221: 83–97.
Dekas AE, Poretsky RS, Orphan VJ . (2009). Deep-Sea archaea fix and share nitrogen in methane-consuming microbial consortia. Science 326: 422–426.
Falkowski PG, Fenchel T, Delong EF . (2008). The microbial engines that drive Earth's biogeochemical cycles. Science 320: 1034–1039.
Florens L, Carozza MJ, Swanson SK, Fournier M, Coleman MK, Workman JL et al. (2006). Analyzing chromatin remodeling complexes using shotgun proteomics and normalized spectral abundance factors. Methods 40: 303–311.
Garcia-Martinez J, Rodriguez-Valera F . (2000). Microdiversity of uncultured marine prokaryotes: the SAR11 cluster and the marine archaea of group I. Mol Ecol 9: 935–948.
Gardy JL, Laird MR, Chen F, Rey S, Walsh CJ, Ester M et al. (2005). ‘PSORT-B v.2.0: expanded prediction of bacterial protein subcellular localization and insights gained from comparative proteome analysis’. Bioinformatics 21: 617–623.
Gibson JAE . (1999). The meromictic lakes and stratified marine basins of the Vestfold Hills, East Antarctica. Antarctic Sci 11: 175–192.
Gomez-Alvarez V, Teal TK, Schmidt TM . (2009). Systematic artifacts in metagenomes from complex microbial communities. ISME J 3: 1314–1317.
Gregersen LH, Habicht KS, Peduzzi S, Tonolla M, Canfield DE, Miller M et al. (2009). Dominance of a clonal green sulfur bacterial population in a stratified lake. FEMS Microbiol Ecol 70: 30–41.
Haft DH, Loftus BJ, Richardson DL, Yang F, Eisen JA, Paulsen IT et al. (2001). TIGRFAMs: a protein family resource for the functional identification of proteins. Nucleic Acids Res 29: 41–43.
Hahn MW, Lünsdorf H, Wu Q, Schauer M, Höfle MG, Boenigk J et al. (2003). Isolation of novel ultramicrobacteria classified as actinobacteria from five freshwater habitats in Europe and Asia. Appl Environ Microbiol 69: 1442–1451.
Halm H, Musat N, Lam P, Langlois R, Musat F, Peduzzi S et al. (2009). Co-occurence of denitrification and nitrogen fixation in a meromictic lake, Lake Cadagno (Switzerland). Environ Microbiol 11: 1945–1958.
Hoops S, Sahle S, Gauges R, Lee C, Pahle J, Simus N et al. (2006). COPASI—a COmplex PAthway SImulator. Bioinformatics 22: 3067–3074.
Horvath P, Barrangou R . (2010). CRISPR/Cas, the immune system of bacteria and archaea. Science 327: 167–170.
Huse SM, Huber JA, Morrison HG, Sogin ML, Welch DM . (2007). Accuracy and quality of massively parallel DNA pyrosequencing. Genome Biol 8: R143.
Keller A, Nesvizhskii AI, Kolker E, Aebersold R . (2002). Empirical statistical model to estimate the accuracy of peptide identifications made by MS/MS and database search. Anal Chem 74: 5383–5392.
Lauro FM, McDougald D, Thomas T, Williams TJ, Egan S, Rice S et al. (2009). The genomic basis of trophic strategy in marine bacteria. Proc Natl Acad Sci USA 106: 15527–15533.
Laybourn-Parry J, Marshall WA, Marchant HJ . (2005). Flagellate nutritional versatility as a key to survival in two contrasting Antarctic saline lakes. Freshw Biol 50: 830–838.
Letunic I, Bork P . (2006). Interactive tree of life (iTOL): an online tool for phylogenetic tree display and annotation. Bioinformatics 23: 127–128.
López-Bueno A, Tamames J, Velázquez D, Moya A, Quesada A, Alcamí A . (2009). High diversity of the viral community from an Antarctic lake. Science 326: 858–861.
Madan JA, Marshall WA, Laybourn-Parry J . (2005). Virus and microbial loop dynamics over an annual cycle. Freshw Biol 50: 1291–1300.
McCann KS . (2000). The diversity—stability debate. Nature 405: 228–233.
Meyer B, Kuever J . (2007). Molecular analysis of the distribution and phylogeny of dissimilatory adenosine-5′-phosphosulfate reductase-encoding genes (aprBA) among sulfur-oxidizing prokaryotes. Microbiology 153: 3478–3498.
Miller JR, Delcher AL, Koren S, Venter E, Walenz BP, Brownley A et al. (2008). Aggressive assembly of pyrosequencing reads with mates. Bioinformatics 24: 2818–2824.
Morgulis A, Gertz EM, Schäffer AA, Agarwala R . (2006). WindowMasker: window-based masker for sequenced genomes. Bioinformatics 22: 134–141.
Morris RM, Nunn BL, Frazar C, Goodlett DR, Ting YS, Rocap G . (2010). Comparative metaproteomics reveals ocean-scale shifts in microbial nutrient utilization and energy transduction. ISME J 4: 673–685.
Musat N, Halm H, Winterholler B, Hoppe P, Peduzzi S, Hillion F et al. (2008). A single-cell view on the ecophysiology of anaerobic phototrophic bacteria. Proc Natl Acad Sci USA 105: 17861–17866.
Myers EW, Sutton GG, Delcher AL, Dew IM, Fasulo DP, Flanigan MJ et al. (2000). A whole-genome assembly of Drosophila. Science 287: 2196–2204.
Ng C, DeMaere MZ, Williams TJ, Lauro FM, Raftery M, Gibson JA et al. (2010). Metaproteogenomic analysis of a dominant green sulfur bacterium from Ace Lake, Antarctica. ISME J 4: 1002–1019.
Noguchi H, Park J, Takagi T . (2006). MetaGene: prokaryotic gene finding from environmental genome shotgun sequences. Nucleic Acids Res 34: 5623–5630.
Oelgeschläger E, Rother M . (2008). Carbon monoxide-dependent energy metabolism in anaerobic bacteria and archaea. Arch Microbiol 190: 257–269.
Overmann J, Pfennig N . (1989). Pelodictyon phaeoclathratiforme sp. nov., a new brown-colored member of the Chlorobiaceae forming net-like colonies. Arch Microbiol 152: 401–406.
Pace ML . (1988). Bacterial mortality and the fate of bacterial production. Hydrobiologia 159: 41–49.
Parkin TB, Brock TD . (1980). Photosynthetic bacterial production in lakes: the effects of light intensity. Limnol Oceanogr 25: 711–718.
Parks DH, Beiko RG . (2010). Identifying biologically relevant differences between metagenomic communities. Bioinformatics 26: 715–721.
Patel A, Noble RT, Steele JA, Schwalbach MS, Hewson I, Fuhrman JA . (2007). Virus and prokaryote enumeration from planktonic aquatic environments by epifluorescence microscopy with SYBR Green I. Nat Protocols 2: 269–276.
Poretsky RS, Sun S, Mou X, Moran MA . (2010). Transporter genes expressed by coastal bacterioplankton in response to dissolved organic carbon. Environ Microbiol 12: 616–627.
Powell LM, Bowman JP, Skerratt JH, Franzmann PD, Burton HR . (2005). Ecology of a novel Synechococcus clade occurring in dense populations in saline Antarctic lakes. Mar Ecol Prog Ser 291: 65–80.
Pruitt KD, Tatusova T, Maglott DR . (2007). NCBI reference sequences (RefSeq): a curated non-redundant sequence database of genomes, transcripts and proteins. Nucleic Acids Res 35: D61–D65.
R Development Core Team (2009). R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing: Vienna, Australia. ISBN 3-900051-07-0.
Ram RJ, Verberkmoes NC, Thelen MP, Tyson GW, Baker BJ, Blake II RC et al. (2005). Community proteomics of a natural microbial biofilm. Science 308: 1915–1920.
Rankin LM, Gibson JAE, Franzmann PD, Burton HR . (1999). The chemical stratification and microbial communities of Ace Lake, Antarctica: a review of the characteristics of a marine-derived meromictic lake. Polarforschung 66: 35–52.
Rodriguez-Brito B, Li L, Wegley L, Furlan M, Angly F, Breitbart M et al. (2010). Viral and microbial community dynamics in four aquatic environments. ISME J 4: 739–751.
Rodriguez-Valera F, Martin-Cuadrado AB, Rodriguez-Brito B, Pasić L, Thingstad TF, Rohwer F et al. (2009). Explaining microbial population genomics through phage predation. Nature Rev Microbiol 7: 828–836.
Rusch DB, Halpern AL, Sutton G, Heidelberg KB, Williamson S, Yooseph S et al. (2007). The sorcerer II global ocean sampling expedition: Northwest Atlantic through eastern tropical Pacific. PLoS Biol 5: e77.
Sharma AK, Zhaxybayeva O, Papke RT, Doolittle WF . (2008). Actinorhodopsins: proteorhodopsin-like gene sequences found predominantly in non-marine environments. Environ Microbiol 10: 1039–1056.
Sowell SM, Wilhelm LJ, Norbeck AD, Lipton MS, Nicora CD, Barofsky DF et al. (2009). Transport functions dominate the SAR11 metaproteome at low-nutrient extremes in the Sargasso Sea. ISME J 3: 93–105.
Suttle CA . (2005). Viruses in the sea. Nature 437: 356–361.
Tatusov RL, Fedorova ND, Jackson JD, Jacobs AR, Kiryutin B, Koonin EV et al. (2003). The COG database: an updated version includes eukaryotes. BMC Bioinform 4: 41.
Tatusov RL, Koonin EV, Lipman DJ . (1997). A genomic perspective on protein families. Science 278: 631–637.
Tatusov RL, Natale DA, Garkavtsev IV, Tatusova TA, Shankavaram UT, Rao BS et al. (2001). The COG database: new developments in phylogenetic classification of proteins from complete genomes. Nucleic Acids Res 29: 22–28.
Taylor PG, Townsend AR . (2010). Stoichiometric control of organic carbon—nitrate relationships from soils to the sea. Nature 464: 1178–1181.
Tyson GW, Chapman J, Hugenholtz P, Allen EE, Ram RJ, Richardson PM et al. (2004). Community structure and metabolism through reconstruction of microbial genomes from the environment. Nature 428: 37–43.
von Mering C, Hugenholtz P, Raes J, Tringe SG, Doerks T, Jensen LJ et al. (2007). Quantitative phylogenetic assessment of microbial communities in diverse environments. Science 315: 1126–1130.
Wild G, Gardner A, West SA . (2009). Adaptation and the evolution of parasite virulence in a connected world. Nature 459: 983–986.
Wu J, Mao X, Cai T, Luo J, Wei L . (2006). KOBAS server: a web-based platform for automated annotation and pathway identification. Nucleic Acids Res 34: W720–W724.
Zybailov B, Mosley AL, Sardiu ME, Coleman MK, Florens L, Washburn MP . (2006). Statistical analysis of membrane proteome expression changes in Saccharomyces cerevisiae. J Proteome Res 5: 2339–2347.
Acknowledgements
We thank J Craig Venter, Karla Heidelberg, John Bowman, Louise (Cromer) Newman, Anthony Hull, John Rich, Martin Riddle, Jing Guan, Joachim Mai and Tassia Kolesnikow for assisting our Antarctic program. We thank the reviewers who provided very constructive critiques during the review process. The work of the Australian contingent was supported by the Australian Research Council, the Australian Antarctic Division, the University of New South Wales and NSW Government. The work of JCVI members was supported by funding from the Gordon and Betty Moore Foundation.
Author information
Authors and Affiliations
Corresponding author
Additional information
Data deposition
All metagenomic and metaproteomic data are available via the CAMERA portal.
Supplementary Information accompanies the paper on The ISME Journal website
Supplementary information
Rights and permissions
About this article
Cite this article
Lauro, F., DeMaere, M., Yau, S. et al. An integrative study of a meromictic lake ecosystem in Antarctica. ISME J 5, 879–895 (2011). https://doi.org/10.1038/ismej.2010.185
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2010.185
Keywords
This article is cited by
-
Genomic insights into cryptic cycles of microbial hydrocarbon production and degradation in contiguous freshwater and marine microbiomes
Microbiome (2023)
-
Evaluation of industrial ecology in the π-shaped curve area of China’s Yellow River based on the grey Lotka–Volterra model
Scientific Reports (2023)
-
Trajectories of freshwater microbial genomics and greenhouse gas saturation upon glacial retreat
Nature Communications (2023)
-
Freeze-thaw cycles alter the growth sprouting strategy of wetland plants by promoting denitrification
Communications Earth & Environment (2023)
-
Metagenomics Unveils Microbial Diversity and Their Biogeochemical Roles in Water and Sediment of Thermokarst Lakes in the Yellow River Source Area
Microbial Ecology (2023)