Abstract
Both genetic variations and diet-disrupted gut microbiota can predispose animals to metabolic syndromes (MS). This study assessed the relative contributions of host genetics and diet in shaping the gut microbiota and modulating MS-relevant phenotypes in mice. Together with its wild-type (Wt) counterpart, the Apoa-I knockout mouse, which has impaired glucose tolerance (IGT) and increased body fat, was fed a high-fat diet (HFD) or normal chow (NC) diet for 25 weeks. DNA fingerprinting and bar-coded pyrosequencing of 16S rRNA genes were used to profile gut microbiota structures and to identify the key population changes relevant to MS development by Partial Least Square Discriminate Analysis. Diet changes explained 57% of the total structural variation in gut microbiota, whereas genetic mutation accounted for no more than 12%. All three groups with IGT had significantly different gut microbiota relative to healthy Wt/NC-fed animals. In all, 65 species-level phylotypes were identified as key members with differential responses to changes in diet, genotype and MS phenotype. Most notably, gut barrier-protecting Bifidobacterium spp. were nearly absent in all animals on HFD, regardless of genotype. Sulphate-reducing, endotoxin-producing bacteria of the family, Desulfovibrionaceae, were enhanced in all animals with IGT, most significantly in the Wt/HFD group, which had the highest calorie intake and the most serious MS phenotypes. Thus, diet has a dominating role in shaping gut microbiota and changes of some key populations may transform the gut microbiota of Wt animals into a pathogen-like entity relevant to development of MS, despite a complete host genome.
Similar content being viewed by others
Introduction
Metabolic syndrome (MS) is a combination of medical disorders, including obesity and insulin resistance that increase the risk of developing diabetes and cardiovascular diseases, which have become a devastating epidemic worldwide. Many large-scale epidemiological studies have indicated that the change from a plant-based diet to the one that is animal-based may be the most important factor associated with the rapid increase in the prevalence of MS (Campbell and Campbell, 2005). Altered gut microbiota has recently been suggested to be critical in the development of obesity, diabetes and hypertension (Turnbaugh et al., 2006; Cani et al., 2007b; Holmes et al., 2008; Wen et al., 2008). Gut microbiota, which are closely associated with host nutrition, metabolism and immunity, acts as a ‘second genome’ for modulating the health phenotype of the superorganism host (Jia et al., 2008). Gut microbiota are likely indispensable for obesity development, as germ-free animals are resistant to high-fat diet (HFD)-induced obesity, indicating that high-calorie food alone is not sufficient to induce obesity and insulin resistance (Backhed et al., 2007). Low-grade, systemic and chronic inflammation has been identified as one primary pathological condition underlying the development of MS (Wellen and Hotamisligil, 2005). Recently, endotoxins produced by opportunistic pathogenic members of the gut microbiota have been identified as the primary mediator for triggering the low-grade inflammation responsible for MS development (Cani et al., 2007b). It has been shown that HFDs disrupt gut microbiota in two ways, diminishing levels of gut barrier-protecting bifidobacteria and promoting the growth of endotoxin producers (Cani et al., 2007a). These changes eventually result in higher levels of lipopolysaccharide (LPS) in the host blood, causing inflammation, and consequently, obesity and insulin resistance (Cani et al., 2007a, 2007b). Although specific endotoxin producers remain to be identified, these observations highlight the important mediating role of gut microbiota in diet-induced MS.
For many years, scientists have focused on finding some common variants in human genes that may lead to MS. For example, the variant in the fat mass and obesity-associated gene (FTO) is associated with body mass index in populations of European origin (Frayling et al., 2007), but not in the Chinese populations (Li et al., 2008a). In humans, one of the hallmarks of MS is a decreased plasma concentration of high-density lipoprotein (HDL) and its major component, apolipoprotein a-I (Apoa-I). It has long been postulated that Apoa-I and HDL levels are inversely associated with the incidence of MS, which are associated with an increased risk of type II diabetes and cardiovascular diseases (Dandona et al., 2005; Eckel et al., 2005). Recently, Han et al. (Han et al., 2007) showed that Apoa-I−/− mice have impaired glucose tolerance (IGT) and increased body fat because of the fact that Apoa-I activates AMP-activated protein kinase (AMPK) and thus affects the energy and glucose metabolism of the cell. Given the fact that variations in both host genetics and gut microbiota predispose animals to MS, it is compelling to investigate the relative contributions of diet-disrupted gut microbiota and host gene mutations to MS development.
In this study, Apoa-I−/− mice and their wild-type (Wt) counterparts were maintained on either a normal chow (NC) diet or HFD for a long period of time to reflect the chronic nature of MS development in humans. Faecal samples were collected from each of the animals at the end of the trial and subjected to multivariate statistical analysis of gut microbiota using two 16S rRNA gene fingerprinting methods (denaturing gradient gel electrophoresis (DGGE), and terminal restriction fragment length polymorphism (T-RFLP)), followed by bar-coded pyrosequencing. Our results indicate that diet-induced changes of gut microbiota relevant to MS phenotype development override host gene mutations in the mouse lines studied. This finding calls for further studies to assess the relative contributions of diet-disrupted gut microbiota and host gene mutations relevant to development of MS.
Materials and methods
Animal treatment
Apoa-I−/− and Wt C57BL/6J mice (male, at age 10–12 weeks) were purchased from the Jackson Laboratory (Bar Harbor, ME, USA) and raised in the same room with a regular 12-h dark/light cycle. After acclimatization, the Apoa-I−/− and the Wt animals were fed with both NC diet (containing 5.2% fat, 3.2–3.4 kcal g−1, from SLAC Inc., Shanghai, China) and HFD (containing 34.9% fat, 5.21 kcal g−1, from Research Diets, Inc., New Brunswick, NJ, USA); each group contained five animals. Each group had the first two animals placed in one cage and the other three in another, whereas all were reared in the same room with uniform conditions carefully maintained among treatments. The food intake and body weight of each animal was measured every 2 weeks. For the glucose tolerance test, mice at 25 weeks were fasted for 3 h, then injected intraperitoneally with glucose (2 g per kg body weight), and blood glucose levels were monitored before and at 30, 60 and 120 min after injection using a glucometer (FreeStyle; TheraSense, Alameda, CA, USA). The day before blood glucose testing, fresh faecal matter was collected from each of the mice and immediately stored at −20 °C for subsequent analysis. The protocol for animal use was approved by the Experimental Animal Committee of the Institute for Nutritional Sciences, Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences.
16S rRNA gene-based analysis
Faecal DNA was extracted using the bead-beating method, as previously described (Zoetendal et al., 2002). Isolated faecal DNA was then used as a template for the amplification of the V3 regions of the 16S rRNA gene using the universal primers P3 (5′-CGCCCGCCGCGCGCGGCGGGCGGGGCGGGGGCACGGGGGGCCTACGGGAGGCAGCAG-3′) and P2 (5′-ATTACCGCGGCTGCTGG-3′) and the hot-start touchdown protocol described by Muyzer et al. (Muyzer et al., 1993). DGGE was carried out with a Dcode System apparatus (Bio-Rad, Herts, UK) and a gradient from 27–52%. Phylogenetic identification of important bands was conducted, as described (Muyzer et al., 1993) (Supplementary Table 2).
The PCR products from the amplification of the 16S rRNA gene with primers 8F (5′-GAGAGTTTGATCCTGGCTCAG-3′), the 5′ end labelled D4 and 1492R (5′-GGC/TTACCTTGTTACGACTT-5′) (Hayashi et al., 2002) were used for T-RFLP analysis. Five restriction endonucleases, AluI, HaeIII, HhaI, MspI or Csp6I, were used for T-RFLP analysis with a CEQ 8000 genetic analysis system (Liu et al., 1997).
For bar-coded pyrosequencing, the primers P1 (5′-NNNNNNNNCCTACGGGAGGCAGCAG-3′) and P2 (5′-NNNNNNNNATTACCGCGGCTGCT-3′), marked at the 5′ end with a sample unique DNA bar code of eight nucleotide sequences, were used to amplify the V3 regions from each faecal sample. The products from different samples were mixed in equal ratios for pyrosequencing with the GS FLX platform (McKenna et al., 2008).
Statistical and bioinformatics analysis
Terminal restriction fragment length polymorphism fingerprints were digitalized with software from the CEQ 8000 system (Beckman Coulter, Fullerton, CA, USA), then analysed with the principal components analysis and multivariate analysis of variance (MANOVA) in a Matlab (ver. 7.1, The MathWorks, Inc., Natick, MA, USA) environment.
On the basis of several previous reports describing sources of errors in 454 sequencing runs (Margulies et al., 2005; Sogin et al., 2006; McKenna et al., 2008), the standards used for quality control are described in the Supplementary Information. The usable V3 unique sequences were aligned using NAST (DeSantis et al., 2006), and then imported into the ARB (Ludwig et al., 2004) to construct a neighbour-joining tree for online UniFrac analysis (Lozupone et al., 2006). Operational taxonomic unit (OTU) was classified with Distance-Based OTU and Richness (DOTUR) (Schloss and Handelsman, 2005). One sequence randomly selected from each OTU was BLAST searched against the Ribosomal Database Project (RDP, version 9.33) to identify the taxonomic group, and then inserted into pre-established phylogenetic trees of full-length 16S rRNA gene sequences in ARB. PLS-DA (Perez-Enciso and Tenenhaus, 2003) was used to discriminate groups by diets, host genotypes or healthy phenotypes. Martens’ uncertainty test (Westad and Martens, 2000) and one-way ANOVA (P<0.05) were used to select key OTUs contributing to the classification. The PLS-DA models were tested with leave-one-out cross-validation (Osten, 1988).
Quantitative real-time PCR of Bifidobacterium spp.
Real-time PCR amplification and detection were performed with the DNA Engine Opticon 2 system (MJ Research, Waltham, MA, USA). The primers were the Bifidobacterium-specific primers, Bif164-f and Bif662-r (Satokari et al., 2001). The details of the PCR program and analysis are in the Supplementary Information.
Results
Long-term effects of HFD intake and host gene mutation on health phenotypes
Apoa-I−/− mice and their Wt counterparts were fed on either NC or HFD (n=5 for each group) for 25 weeks. Phenotyping data indicate that the Wt/NC animals were the healthiest, with normal weight and glucose tolerance, whereas the Apoa-I−/−/NC animals had IGT, and the two genotypes on HFD showed both IGT and obesity (Figure 1). Intriguingly, Wt/HFD animals were the most obese with the highest insulin resistance. The much less severe MS phenotype of Apoa-I−/−/HFD animals showed reduced food intakes.
Phenotypes of mouse groups, with different genotypes fed on different diets. (a) Average food intake per day of the four groups of mice. (b) Growth curve established using average body weight of four groups every 2 weeks. (c) Glucose tolerance test (n=5 in each group). Mean values±s.d. are shown. Colour code for each treatment group in the figures: green, wild type (Wt)/normal chow (NC); black, Wt/high-fat diet (HFD); blue, Apoa-I−/−/NC; red, Apoa-I−/−/HFD. Student t-test was used to compare the groups with the same diet (*P<0.05 and **P<0.01). Please note that part of the data from the groups of mice fed with NC has been previously reported (Han et al., 2007). However, these data are included here for better comparison with the mice fed with HFD.
Overall structural responses of gut microbiota to long-term intake of HFD and a host gene mutation
Two DNA fingerprinting methods show that the most significant differences in the composition of gut microbiota were between groups of animals on different diets. Animals on an NC diet had dramatically different predominant DGGE bands (for example, b6 in Figure 2a) from those fed on an HFD (for example, b4). The principal components analysis scores plot based on the T-RFLP data shows that diet-related differences were mainly observed along PC1, which accounts for 57% of the total variations, whereas the two genotypes on an NC diet had much smaller differences along PC3 (12% of total variations) (Figure 2b). Interestingly, the gut microbiota differences between genotypes were significantly reduced on HFD, indicating that abnormal diet may diminish the differences in gut microbiota structures imposed by differing host genotypes. A multivariate analysis of variance (MANOVA) test also indicated that the four groups were first separated as two clades based on diet differences, with a much smaller within-the-clade distance between genotypes on an HFD compared with those on an NC diet (Figure 2c). The Wt/NC animals clustered in a different space from the other three groups with IGT, suggesting that the difference in gut microbiota structure might be commensurate with the host health phenotypes (Figure 2b). Another interesting phenomenon is that all the animals with IGT/obesity had much wider interindividual variations than the healthy Wt/NC animals, particularly between animals in different cages (Figures 2a and b).
Comparison of gut microbiota composition between the mouse groups with different genotypes on different diets. (a) Denaturing gradient gel electrophoresis (DGGE) fingerprinting of V3 region of 16S rRNA genes from faecal bacterial communities. M: DGGE marker. (b) The principal components analysis (PCA) scores plot of the terminal terminal restriction fragment length polymorphism (T-RFLP) data from faecal bacterial communities. Near full-length 16S rRNA genes were digested with five restriction enzymes to generate polymorphism patterns. Each sample had three replicates. (c) Clustering of gut microbiota based on distances between different groups calculated with multivariate analysis of variance test of the first nine PCs of T-RFLP data. The Mahalanobis distances between group means are shown. **P<0.01. Animal groups are colour coded as in Figure 1.
To confirm the findings from these DNA fingerprinting methods, a bar-coded pyrosequencing of the 16S rRNA gene V3 region was used for a deep molecular inventory of the gut microbiota samples. A total of 29 314 useable reads were obtained with 4156 unique sequences, of which, 3145 were detected only once in all the samples; a substantial ‘rare biosphere’. In all, 516 OTUs were delineated at a 97% homology cutoff (Supplementary Figure 1). Both the principal coordinate analysis scores plot and hierarchical clustering analysis with UniFrac metrics (Lozupone et al., 2006) show that the differences between the HFD and NC groups were more significant than those between the two genotypes on the same diet, confirming the above results (Supplementary Figure 2). All replicate samples from the same animal clustered together, showing satisfactory reproducibility of this bar-coded pyrosequencing method.
Identification of the key phylotypes responsible for differentiation between different groups
The combination of pyrosequencing with a multivariate statistical method, Partial Least Square Discriminate Analysis (PLS-DA) was used to reveal detailed structural changes in the gut microbiota in response to variations in host genetics, diet and health phenotypes. Scores plots based on the first two components show that groups with different genotypes, diets or IGT/obesity phenotypes can be well separated (Supplementary Figure 3). Leave-one-out cross-validation yielded high prediction rates for all the classification models (Osten 1988). The Martens’ uncertainty test (Westad and Martens, 2000) and a one-way ANOVA test then selected a total of 65 phylotypes as key variables for separating gut microbiota under different genotypes, diets or IGT/obesity phenotypes (Figure 3, Supplementary Figure 4 and Supplementary Table 1). The results showed that 48 out of the 65 phylotypes responded mainly to diet, supporting the DNA fingerprinting results.
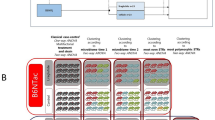
Abundance distribution of the 65 phylotypes identified as key variables for discrimination among the groups of treated mice. These operational taxonomic units (OTUs) from pyrosequencing (97% identify threshold), selected using Partial Least Square Discriminate Analysis (PLS-DA) and Martens’ uncertainty test, include variables with significant differences between the mice given different diets, having different genotypes, or in different health states. To show the distribution of the OTUs with lower abundance, the coloured squares of each column have been scaled to indicate the relative ratios of the OTU among 20 mice. These identified phylotypes are distributed across all the major phyla: 47 in Firmicutes, 11 in Bacteroidetes, 1 in Proteobacteria and 6 in Actinobacteria.
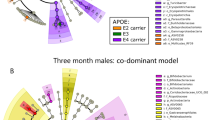
We found that four different lineages (M1–M4) within the Erysipelotrichaceae family responded differentially to diet or host health phenotypes (Figure 4a). In most animals, Erysipelotrichaceae is the most predominant clade (Supplementary Figure 5). Total phylotypes in M1 were abundant (about 20% of total microbiota populations) in healthy Wt/NC animals, but significantly reduced (<2% of total populations) in the other three groups with IGT. Groups M2–M4 responded only to diet changes. M2 and M4 were predominant in HFD groups, whereas M3 was more prevalent in NC animals (Figure 4b).
Phylogeny and differential abundance and distribution of phylotypes in the family Erysipelotrichaceae among treatment groups. (a) Phylogeny of phylotypes in the family Erysipelotrichaceae. One sequence was randomly selected from each operational taxonomic unit (OTU) (97% identify threshold) and was inserted into pre-established phylogenetic trees of full-length 16S rRNA gene sequences in ARB. The tree shown here only includes some reference species of Firmicutes and the sequences from this study in the family Erysipelotrichaceae. These phylotypes fall into four lineages, M1–M4. The DGGE bands in M1–M4 are also shown. (b) Abundance distribution of M1, M2, M3 and M4 in the four groups of animals.
Four members in the family Bifidobacteriaceae were present in most animals on an NC diet, but completely disappeared in those on an HFD, regardless of genotype, as confirmed by bifidobacteria-specific real-time PCR (Supplementary Figure 7). Furthermore, we found that one phylotype in the family Desulfovibrionaceae was more prevalent in each of the IGT/obese groups, most notably in the Wt/HFD group, which had the most serious obesity and IGT phenotypes (Figures 1 and 3). Specifically, there was a nearly sevenfold increase of this phylotype in Wt/HFD relative to that observed in the Wt/NC group (4.67 vs 0.7%, one-way ANOVA test, P=0.0075, Supplementary Table 1).
Discussion
Animals can be regarded as walking bioreactors, maintaining a highly diverse mixed chemostat culture that utilizes their diet as a continuous feeding medium (Sonnenburg et al., 2004). It is not surprising that significant and long-term changes in diet composition will lead to profound alterations in gut microbiota structure (Finegold et al., 1974; Tajima et al., 2001). Significant differences in the relative abundance of Firmicutes and Bacteroidetes in gut microbiota have been observed between host groups with obesity and lean phenotypes (Turnbaugh et al., 2006, 2008). Phylum-wide changes in gut microbiota composition were not observed in our diet-induced obese animals, regardless of genotypes. However, we revealed the changes at much smaller phylogenetic lineages than phyla. Blooming of Mollicutes (equivalent to the class Erysipelotrichi in this study) was previously observed in diet-induced obese animals (Turnbaugh et al., 2008), and we found that four different lineages (M1–M4) within the family Erysipelotrichaceae of this class responded differentially to diet or host health phenotypes. The differential or even contrasting behaviours of lineages within the same family emphasizes the importance of phylotype-specific analysis for understanding the role of gut microbiota composition in determining host health or disease development. The lineages (M1–M4) identified here are major discrete phylogenetic clusters with a ‘fan-like’ structure in the phylogenetic tree (Acinas et al., 2004). This suggests that each lineage consists of closely related phylotypes with no competitive pressure to purge its diversity from within, although different lineages may compete with each other under selective pressures imposed by changing diet, host genotype or health phenotype. Furthermore, most rare phylotypes in these lineages showed behaviour similar to the identified predominant ones (Supplementary Figure 6). This suggests that phylotypes in the same phylogenetic cluster may share similar functions relevant to host phenotypes, and that the ‘rare biosphere’ is not negligible (Sogin et al., 2006).
Endotoxins produced by the HFD-altered gut microbiota have been identified as important mediators in triggering inflammation, a key underlying pathological condition in MS development (Cani et al., 2007b). Cani et al. showed that continuous subcutaneous infusion of purified LPS induces inflammation and then obesity and insulin resistance in mice fed on NC diet. Knockout of CD14 abolishes these triggering effects of LPS. Inclusion of oligofructose in the HFD maintains the integrity of the bifidobacteria population and the normal gut barrier, helping the animals ward off LPS from gut microbiota and maintain normal weight despite a high-calorie diet. All of this evidence supports a key mediating role of LPS in diet-induced MS by way of triggering inflammation. Our study is in agreement with their report on the association between reduced population levels of bifidobacteria in the gut and increased inflammation of the host, potentially due to the increased gut barrier permeability to endotoxins from loss of these gut barrier-protecting bacteria (Cani et al., 2007b). In our study, this group of bacteria was actually ‘eliminated’ from guts fed with HFD, regardless of genotype differences, possibly due to the much longer feeding time. Cani et al. did not identify specific endotoxin producers with the fluorescent in situ hybridization (FISH) technology they used. This may be due to the lack of appropriate probes in their experiment, as FISH can only detect what has already been characterized. Here, through the use of a combination of bar-coded pyrosequencing and multivariate statistics, we were able to identify sulphate-reducing bacteria in family Desulfovibrionaceae as the potentially important endotoxin producers whose abundance changes were associated with the development of MS in our animal models. Members of this family are Gram-negative, opportunistic pathogen, endotoxins producers (Loubinoux et al., 2000; Weglarz et al., 2003) and are also capable of reducing sulphate to H2S, damaging the gut barrier (Beerens and Romond, 1977). These findings are in agreement with the metabolic endotoxemia hypothesis on MS onset and development.
We were able to identify specific phylotypes whose population changes were more relevant to MS development through the utilization of the Apoa-I−/− mouse model because of the careful manipulation of rearing conditions to minimize stochastic interindividual variations of gut microbiota. In addition, our use of long-term feeding experiments with different diets ensured stabilization of physiological integration between gut microbiota and hosts, to reflect the chronic nature of MS in humans (Zoetendal et al., 1998; Wei et al., 2004).
The utilization of a combination of DNA pyrosequencing with appropriate multivariate statistics facilitated the pattern discovery process. Although standard statistical methodologies do not work well when the number of samples are much less than that of the variables (thousands of OTUs as variables from dozens of samples) and the abundance distributions of variables are extremely uneven, PLS-DA is a useful multivariate analysis tool to deal with this type of data (Nguyen and Rocke, 2002; Perez-Enciso and Tenenhaus, 2003; Wang et al., 2004; Zhang et al., 2009). The application of PLS-DA to bar-coded pyrosequencing data was robust and also sensitive enough to identify specific members of gut microbiota with health-relevant responses to diet, genotype or health phenotypes, including both minor species (at 1–5%) and major species (>10% of total gut populations). This new methodology also confirms and expands results from the two classical DNA fingerprinting methods, PCR–DGGE and T-RFLP. Therefore, this strategy has provided both a new methodology and helpful insights for metagenomic studies on the overwhelmingly complex microbiota for understanding MS development in humans. As the key phylotypes to MS development are intermingled with those that behave stochastically, the current metagenomic strategy with a one time point ‘snap-shot’ and low-coverage sequencing of total community DNAs can be problematic, as the genes in key functional members can be significantly ‘diluted’ by genes from irrelevant species, making functional attribution difficult or perhaps misleading. This is especially true if the functionally important members, such as Bifidobacteriaceae or Desulfovibrionaceae, are relatively minor within the community compared with those that respond mainly to diet. Therefore, new strategies will need to be developed for assigning sequencing reads to their corresponding phylogenetic bins with relevance to host health phenotypes. Covariation analysis, such as that used in finding associations between specific phylotypes and host metabotypes through dynamic monitoring of a cohort, may help identify linkages between 16S rRNA gene markers and related functional genes from metagenomic data (Li et al., 2008b).
Most intriguingly, in this study, the Wt/HFD animal group developed the worst MS phenotype and contained the highest increase of sulphate-reducing bacteria. Apoa-I−/− mice on an HFD showed less severe MS phenotypes than did Wt/HFD mice, possibly due to significantly lower food intake and much smaller alterations in gut microbiota, particularly a much lower population level of the sulphate-reducing bacteria. Taken together, these results indicate a possibly dominating role of gut microbiota disrupted by long-term, unlimited feeding of HFD in MS development over variations of host genetic predisposition to the disease. This is supported by the large amount of epidemiological data that diet changes are the most important contributors for the increasing epidemic of metabolic diseases among human populations and the fact that the incidence of metabolic diseases has increased dramatically in a relatively short period of time during which human genetic diversity hardly changed. However, this conclusion still only holds true for the particular gene mutation we used in our study. With similar strategies used in this study, more genetic mutations should be tested for assessing the relative contributions of genotypes and gut microbiota for predisposing hosts to MS development. To further establish whether there is a causal link between specific changes of gut microbiota and MS development, the animals should be sampled more frequently in future studies.
The results of this study indicate that the composition of gut microbiota is tightly interwoven with long-term diet patterns and health phenotypes of the host, with changes of some specific phylotypes most relevant to MS development. With diets having a possibly dominating role in transforming gut microbiota into a ‘pathogen-like entity’ for metabolic diseases, it implies a vast possibility of combating these diseases by modulating gut microbiota with designed diet interventions, targeting the promotion of gut barrier protectors and the suppression of endotoxin producers.
Accession numbers
The unique sequences obtained from pyrosequencing are available in the GenBank database under accession numbers FJ032696–FJ036849. The sequences from key DGGE bands obtained in this study are available in the GenBank database under accession numbers EU584214–EU584231.
Accession codes
References
Acinas SG, Klepac-Ceraj V, Hunt DE, Pharino C, Ceraj I, Distel DL et al. (2004). Fine-scale phylogenetic architecture of a complex bacterial community. Nature 430: 551–554.
Backhed F, Manchester JK, Semenkovich CF, Gordon JI . (2007). Mechanisms underlying the resistance to diet-induced obesity in germ-free mice. Proc Natl Acad Sci USA 104: 979–984.
Beerens H, Romond C . (1977). Sulfate-reducing anaerobic bacteria in human feces. Am J Clin Nutr 30: 1770–1776.
Campbell TC, Campbell TM . (2005). The China Study: The Most Comprehensive Study of Nutrition Ever Conducted and the Startling Implications for Diet, Weight Loss and Long-Term Health. Benbella Books: Texas, USA.
Cani PD, Neyrinck AM, Fava F, Knauf C, Burcelin RG, Tuohy KM et al. (2007a). Selective increases of bifidobacteria in gut microflora improve high-fat-diet-induced diabetes in mice through a mechanism associated with endotoxaemia. Diabetologia 50: 2374–2383.
Cani PD, Amar J, Iglesias MA, Poggi M, Knauf C, Bastelica D et al. (2007b). Metabolic endotoxemia initiates obesity and insulin resistance. Diabetes 56: 1761–1772.
Dandona P, Aljada A, Chaudhuri A, Mohanty P, Garg R . (2005). Metabolic syndrome: a comprehensive perspective based on interactions between obesity, diabetes, and inflammation. Circulation 111: 1448–1454.
DeSantis Jr TZ, Hugenholtz P, Keller K, Brodie EL, Larsen N, Piceno YM et al. (2006). NAST: a multiple sequence alignment server for comparative analysis of 16S rRNA genes. Nucleic Acids Res 34: W394–W399.
Eckel RH, Grundy SM, Zimmet PZ . (2005). The metabolic syndrome. Lancet 365: 1415–1428.
Finegold SM, Attebery HR, Sutter VL . (1974). Effect of diet on human faecal flora: comparison of Japanese and American diets. Am J Clin Nutr 27: 1456–1469.
Frayling TM, Timpson NJ, Weedon MN, Zeggini E, Freathy RM, Lindgren CM et al. (2007). A common variant in the FTO gene is associated with body mass index and predisposes to childhood and adult obesity. Science 316: 889–894.
Han R, Lai R, Ding Q, Wang Z, Luo X, Zhang Y et al. (2007). Apolipoprotein A-I stimulates AMP-activated protein kinase and improves glucose metabolism. Diabetologia 50: 1960–1968.
Hayashi H, Sakamoto M, Benno Y . (2002). Phylogenetic analysis of the human gut microbiota using 16S rDNA clone libraries and strictly anaerobic culture-based methods. Microbiol Immunol 46: 535–548.
Holmes E, Loo RL, Stamler J, Bictash M, Yap IK, Chan Q et al. (2008). Human metabolic phenotype diversity and its association with diet and blood pressure. Nature 453: 396–400.
Jia W, Li H, Zhao L, Nicholson JK . (2008). Gut microbiota: a potential new territory for drug targeting. Nat Rev Drug Discov 7: 123–129.
Li H, Wu Y, Loos RJ, Hu FB, Liu Y, Wang J et al. (2008a). Variants in the fat mass- and obesity-associated (FTO) gene are not associated with obesity in a Chinese Han population. Diabetes 57: 264–268.
Li M, Wang B, Zhang M, Rantalainen M, Wang S, Zhou H et al. (2008b). Symbiotic gut microbes modulate human metabolic phenotypes. Proc Natl Acad Sci USA 105: 2117–2122.
Liu WT, Marsh TL, Cheng H, Forney LJ . (1997). Characterization of microbial diversity by determining terminal restriction fragment length polymorphisms of genes encoding 16S rRNA. Appl Environ Microbiol 63: 4516–4522.
Loubinoux J, Mory F, Pereira IA, Le Faou AE . (2000). Bacteremia caused by a strain of Desulfovibrio related to the provisionally named Desulfovibrio fairfieldensis. J Clin Microbiol 38: 931–934.
Lozupone C, Hamady M, Knight R . (2006). UniFrac—an online tool for comparing microbial community diversity in a phylogenetic context. BMC Bioinformatics 7: 371.
Ludwig W, Strunk O, Westram R, Richter L, Meier H, Yadhukumar et al. (2004). ARB: a software environment for sequence data. Nucleic Acids Res 32: 1363–1371.
Margulies M, Egholm M, Altman WE, Attiya S, Bader JS, Bemben LA et al. (2005). Genome sequencing in microfabricated high-density picolitre reactors. Nature 437: 376–380.
McKenna P, Hoffmann C, Minkah N, Aye PP, Lackner A, Liu Z et al. (2008). The macaque gut microbiome in health, lentiviral infection, and chronic enterocolitis. PLoS Pathog 4: e20.
Muyzer G, de Waal EC, Uitterlinden AG . (1993). Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl Environ Microbiol 59: 695–700.
Nguyen DV, Rocke DM . (2002). Tumor classification by partial least squares using microarray gene expression data. Bioinformatics 18: 39–50.
Osten DW . (1988). Selection of optimal regression models via cross-validation. J Chemom 2: 39–48.
Perez-Enciso M, Tenenhaus M . (2003). Prediction of clinical outcome with microarray data: a partial least squares discriminant analysis (PLS-DA) approach. Hum Genet 112: 581–592.
Satokari RM, Vaughan EE, Akkermans AD, Saarela M, de Vos WM . (2001). Bifidobacterial diversity in human feces detected by genus-specific PCR and denaturing gradient gel electrophoresis. Appl Environ Microbiol 67: 504–513.
Schloss PD, Handelsman J . (2005). Introducing DOTUR, a computer program for defining operational taxonomic units and estimating species richness. Appl Environ Microbiol 71: 1501–1506.
Sogin ML, Morrison HG, Huber JA, Mark Welch D, Huse SM, Neal PR et al. (2006). Microbial diversity in the deep sea and the underexplored ‘rare biosphere’. Proc Natl Acad Sci USA 103: 12115–12120.
Sonnenburg JL, Angenent LT, Gordon JI . (2004). Getting a grip on things: how do communities of bacterial symbionts become established in our intestine? Nat Immunol 5: 569–573.
Tajima K, Aminov RI, Nagamine T, Matsui H, Nakamura M, Benno Y . (2001). Diet-dependent shifts in the bacterial population of the rumen revealed with real-time PCR. Appl Environ Microbiol 67: 2766–2774.
Turnbaugh PJ, Backhed F, Fulton L, Gordon JI . (2008). Diet-induced obesity is linked to marked but reversible alterations in the mouse distal gut microbiome. Cell Host Microbe 3: 213–223.
Turnbaugh PJ, Ley RE, Mahowald MA, Magrini V, Mardis ER, Gordon JI . (2006). An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444: 1027–1031.
Wang Y, Holmes E, Nicholson JK, Cloarec O, Chollet J, Tanner M et al. (2004). Metabonomic investigations in mice infected with Schistosoma mansoni: an approach for biomarker identification. Proc Natl Acad Sci USA 101: 12676–12681.
Weglarz L, Dzierzewicz Z, Skop B, Orchel A, Parfiniewicz B, Wisniowska B et al. (2003). Desulfovibrio desulfuricans lipopolysaccharides induce endothelial cell IL-6 and IL-8 secretion and E-selectin and VCAM-1 expression. Cell Mol Biol Lett 8: 991–1003.
Wei G, Pan L, Du H, Chen J, Zhao L . (2004). ERIC-PCR fingerprinting-based community DNA hybridization to pinpoint genome-specific fragments as molecular markers to identify and track populations common to healthy human guts. J Microbiol Methods 59: 91–108.
Wellen KE, Hotamisligil GS . (2005). Inflammation, stress, and diabetes. J Clin Invest 115: 1111–1119.
Wen L, Ley RE, Volchkov PY, Stranges PB, Avanesyan L, Stonebraker AC et al. (2008). Innate immunity and intestinal microbiota in the development of type 1 diabetes. Nature 455: 1109–1113.
Westad F, Martens H . (2000). Variable selection in near infrared spectroscopy based on significance testing in partial least squares regression. Journal of Near Inffrared Spectroscopy 8: 117–124.
Zhang M, Zhang M, Zhang C, Du H, Wei G, Pang X et al. (2009). Pattern extraction of structural responses of gut microbiota to rotavirus infection via multivariate statistical analysis of clone library data. FEMS Microbiol Ecol 70: 21–29.
Zoetendal EG, Akkermans AD, De Vos WM . (1998). Temperature gradient gel electrophoresis analysis of 16S rRNA from human faecal samples reveals stable and host-specific communities of active bacteria. Appl Environ Microbiol 64: 3854–3859.
Zoetendal EG, von Wright A, Vilpponen-Salmela T, Ben-Amor K, Akkermans AD, de Vos WM . (2002). Mucosa-associated bacteria in the human gastrointestinal tract are uniformly distributed along the colon and differ from the community recovered from feces. Appl Environ Microbiol 68: 3401–3407.
Acknowledgements
This work was supported by the National Natural Science Foundation of China Program Grants 30730005, 30821005 and 20875061, 973 Program Grants 2007CB513002 and 2004CB518600, 863 Program Grant 2008AA02Z315, the International Cooperation Program Grants 2007DFC30450 and 075407001, and the Chinese Academy of Sciences (Knowledge Innovation Program KSCX1-YW-02).
Author information
Authors and Affiliations
Corresponding authors
Additional information
Supplementary Information accompanies the paper on The ISME Journal website (http://www.nature.com/ismej)
Supplementary information
Rights and permissions
About this article
Cite this article
Zhang, C., Zhang, M., Wang, S. et al. Interactions between gut microbiota, host genetics and diet relevant to development of metabolic syndromes in mice. ISME J 4, 232–241 (2010). https://doi.org/10.1038/ismej.2009.112
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2009.112
Keywords
This article is cited by
-
Correlation of Differentially Expressed lncRNAs with Intestinal Flora Imbalance, Small Intestinal Permeability, and Glucose Uptake in T2DM Mice
Applied Biochemistry and Biotechnology (2024)
-
Analysis of strain, sex, and diet-dependent modulation of gut microbiota reveals candidate keystone organisms driving microbial diversity in response to American and ketogenic diets
Microbiome (2023)
-
Gut microbiota in a mouse model of obesity and peripheral neuropathy associated with plasma and nerve lipidomics and nerve transcriptomics
Microbiome (2023)
-
The oral microbiome in autoimmune diseases: friend or foe?
Journal of Translational Medicine (2023)
-
Individual and combined effects of fluoride and arsenic on gut bacteria: a recent update
The Nucleus (2023)