Abstract
Contemporary environmental disturbances and historical contingencies are considered to be major factors driving current differences in microbial diversity. However, little was known about their relative importance. This study combines culture-independent molecular techniques and advanced statistical analyses to examine quantitatively the relative importance of contemporary disturbances and historical contingencies in influencing large-scale soil bacterial diversity using a large set of manipulated field-based molecular data (212 samples). Contemporary disturbances were represented by applications of different fertilizers N, P, K and organic manure (OM) and historical contingencies by distinct geographic sampling locations and soil profiles. Multivariate regression tree (MRT) analysis showed that diversity estimates were mainly distinguished by sampling locations, which explained 40.8% of the variation in bacterial diversity, followed by soil profiles (19.5%), sampling time (13.1%), OM (3.7%) and P (1.8%). Aggregated boosted tree (ABT) analysis showed that the relative importance of different categorical factors on soil bacterial diversity variation was ranked as sampling locations, soil profiles, sampling time, OM and P. Both MRT and ABT analyses showed that historical contingencies were the dominant factor driving variation in bacterial diversity across a regional scale (about 1000 km), whereas some contemporary disturbances also caused variation in bacterial diversity at a local scale. This study demonstrated that past events and contemporary disturbances had similar influence on soil bacterial diversity to that documented for macroorganisms, indicating that there might be some common aspects of biogeography to all organisms.
Similar content being viewed by others
Introduction
For centuries, biologists have studied patterns of plant and animal diversity at different spatial scales. However similar studies were impossible for microorganisms for a long time because of the limitation of techniques that the spatial variation of soil microbial diversity has long been considered as ‘noise’ in microbiological studies (Ettema and Wardle, 2002; Finlay, 2002), although the composition and diversity of microbial communities are thought to have a direct influence on a wide range of ecosystem processes (Naeem and Li, 1997; Bell et al., 2005b; Madsen, 2005; Balvanera et al., 2006). Culture-independent molecular techniques now make it possible to explore microbial diversity more deeply and widely than ever before. As a result, a large body of studies has revealed the vast diversity of microorganisms missed by past culture-dependent studies (Torsvik et al., 1990; Gans et al., 2005) and distinguished a wide range of biotic and abiotic factors influencing microbial diversity and community such as vegetation (McArthur et al., 1988; He et al., 2005, 2006), trophic status (Lefranc et al., 2005), spatial distance (Cho and Tiedje, 2000), salinity (Crump et al., 2004; Lozupone and Knight, 2007), profile depth (Øvreås et al., 1997; Fierer et al., 2003), soil pH (Fierer and Jackson, 2006; He et al., 2007) and heavy metals (Sandaa et al., 1999; Gans et al., 2005). However, almost all of such studies just documented the variation and potential patterns of microbial diversity alone and certain ecological variation or gradient cannot be interpreted within a theory framework to explore the mechanisms that generate the patterns, as such studies failed to simultaneously consider two types of factors—contemporary environmental variations and historical contingencies—that were thought to be the major factors driving the current diversity variation by the current theory of prokaryotic biogeography and diversification (Martiny et al., 2006).
The relative contribution of contemporary environmental factors and the legacies of historical events on the present community composition and diversity is a long-standing theme of traditional biogeography (Willis and Whittaker, 2002; Ricklefs, 2004; Rajaniemi et al., 2006; Qian et al., 2007). The simultaneous consideration of contemporary disturbances and historical contingencies could structure a meaningful theory framework for exploring the four alternative hypotheses of the biogeography of microorganisms (Martiny et al., 2006). The first hypothesis is that microbial composition and diversity are randomly distributed over space. In this case, microbial composition and diversity cannot be distinguished by any of the two factors. The second hypothesis is that the differences in microbial composition and diversity merely reflect the influence of contemporary environmental variation. This would imply that different contemporary environments maintain distinctive microbial assemblages and that the effects of past evolutionary and ecological events can be rapidly erased because of the enormous dispersal capabilities of microorganisms. The third hypothesis is that the differences in microbial composition and diversity are merely due to the lingering effects of past evolutionary and ecological events (for example, distance isolation, physical barrier, dispersal history and past environmental heterogeneity (EH)), which can result in genetic divergence of microbial assemblages. The final hypothesis is that the differences in microbial composition and diversity, similar to those of macroorganisms, reflect the influences of both past events and contemporary environmental variations (Martiny et al., 2006).
Chinese Ecosystem Research Network (CERN) is a long-term research project established in 1988 by the Chinese Academy of Sciences (CAS) to carry out comprehensive ecological monitoring and research on diverse ecosystems. We collected 212 soil samples that had been solely applied with chemical fertilizer N (N), P, K or in combination with organic manure (OM) for about 18 years from three CERN stations across the Northern and Southern China (at about 1000 km distance). Here, the applications of different fertilizers were considered as contemporary environmental disturbances, and the sampling locations and soil profiles could be seen as proxy assemblage of past evolutionary and ecological events whose legacies had been maintained because of the spatial dissimilarity. We explored the relative influence of contemporary environmental disturbances and historical contingencies on soil bacterial diversity by culture-independent molecular approaches and advanced statistical analyses. The objectives of this study were to test the hypotheses that were similar to those documented for macroorganisms (Willis and Whittaker, 2002; Ricklefs, 2004; Qian et al., 2007), the differences in soil bacterial diversity are driven by both contemporary disturbances and historical contingencies, and their relative importance differs depending on the different spatial scales over which diversity is measured and compared.
Materials and methods
Description of the experimental site and sampling
In total, 212 soil samples associated with different fertilization treatments, soil profiles and sampling times (January and July 2006) were collected from Fengqiu (FQ), Taoyuan (TY) and Qiyang (QY), three CERN stations across the Northern and Southern China (Supplementary Table S1). FQ (35 °00′N, 114 °24′E), TY (28 °55′N, 111 °27′E) and QY (26 °45′N, 111 °53′E) sites are located along a latitude gradient with distinct temperature, annual rainfall and soil types. In detail, FQ had a temperate monsoon climate, with a mean annual temperature of 13.9 °C and a mean annual rainfall of 605 mm; TY and QY were subtropical monsoon climate, with mean annual temperature of 16.5 and 18.1 °C, and mean annual rainfall of 1437 and 1288 mm, respectively. The soil of FQ, derived from alluvial sediments of the Yellow River and classified as aquic inceptisols with a sandy loam texture, was distinctly different from the soils of TY and QY. Both of the soils of TY and QY were derived from Quaternary red earth but developed with different soil types of paddy soil and red soil (agri-udic ferrosols with silty clay texture) due to different agricultural practices.
These sampling stations with distinct characteristics, as well as soil profiles, may thus be seen as proxy assemblage of past evolutionary and ecological events such as spatial isolation, physical barrier, dispersal history and past EH. Despite the distinct characteristics between different experimental stations, the climate and soil characteristics of different experimental plots were generally the same at the same station before the experiments of fertilization with chemical fertilizer N, P, K or in combination with OM in a randomized block design for about 18 years, making the fertilization treatments (generally three or four replicates in each treatment) an appropriate variable to represent the contemporary environmental disturbances. Therefore, the examination of the responses of soil microbial diversity to the different fertilization treatments among the sampling locations and profile depths offers an appropriate opportunity to explore the relative importance of contemporary disturbances and historical contingencies on soil bacterial diversity. Furthermore, to examine whether seasonal variation was an important factor for bacterial diversity variation, an additional set of soil samples was collected 6 months after the initial collection.
Soil samples were collected by taking 10 soil cores (approximately 5 cm in diameter) from each plot and mixing them to form one composite sample for each replicate. Each sample was placed in a sterile plastic bag, sealed and transported to the laboratory on ice. All samples were passed through a 2.0 mm sieve and stored at −80 °C.
Soil DNA extraction, PCR and DGGE analyses
Soil DNA was extracted using the UltraClean Soil DNA Isolation Kits (Mo Bio Laboratories, Solana Beach, CA, USA) according to the manufacturer's instruction. The 16S rRNA gene was PCR amplified using the extracted DNA as template and universal bacterial primers 954f (GCACAAGCGGTGGAGCATGTGG) with a GC clamp and 1369r (GCCCGGGAACGTATTCACCG) (Yu and Morrison, 2004). Amplicons (about 200 ng) were analyzed by DGGE using 6% (w/v) acrylamide/bisacrylamide (37.5:1, mass:mass) gels containing a 35%–60% linear gradient of formamide and urea (100% denaturing solution contained 40% (v/v) formamide and 7 M urea). The electrophoresis was run for 6 h at 120 V and a constant temperature of 60 °C, using a DCode Universal Mutation Detection System (Bio-Rad Laboratories, Hercules, CA, USA). The gels were stained with SYBR Gold Nucleic Acid Gel Stain (1:10 000; Invitrogen-Molecular Probes, Eugene, OR, USA) for 30 min, scanned by the Gel Documentation System (Syngene, Frederick, MD, USA) and analyzed using the software Quantity One (Bio-Rad Laboratories).
Operational taxonomic unit definition and diversity indices calculation
The genotypic diversity of each of the soil bacterial communities was determined by DGGE profile data. DGGE can be considered as a high-throughput method at the cost of relatively low taxonomic resolution. However, the estimate of the relative variation trend of the soil bacterial diversity remains valid if the examined richness proportionally changes among the samples. Each detected band was defined as an operational taxonomic unit (OTU), and the number of bands was defined as the genotypic richness of each sample (Bell et al., 2005a; Reche et al., 2005). The pixel intensity for each band was detected by Quantity One software and was expressed as the relative abundance (pi) (Reche et al., 2005). Shannon index (H') and Simpson index (D), the most widely used diversity indices, were calculated using the richness and relative abundance data based on the following equation:


where pi=ni/N, ni is the abundance of the ith OTU and N the total abundance of all OTUs in the sample. Richness, Shannon index and Simpson index were selected to reflect the bacterial diversity properties (Supplementary Table S1).
Multivariate regression tree analysis
It was not possible to sample all potential categorical variable combinations because many combinations did not exist in the field experiment. As a result our sampling design was not factorial. Despite the potential for confounding interactions between incomplete factor levels, the single and combined responses of bacterial diversity estimates to categorical factors still can be explored by multivariate regression tree (MRT) analysis, as MRT analysis is well suited for complex ecological datasets with missing treatment combinations, missing values and high-order interactions (De'ath, 2002). In fact, we compared the results of MRT using whole data and a partial set of data with extracted symmetrical variable combinations and achieved similar results except that unsymmetrical variable was pruned in the latter (compare Figure 1 and Supplementary Figure S1). In this study, therefore, we focused on describing the results of MRT using whole datasets. For an MRT analysis, dissimilarity among the response variables is defined as the total sum of squares of the response variable values (for example, richness, Shannon index and Simpson index), and the least sum of squares criterion is used to repeatedly split data into two groups based on one of the classification variables (for example, sampling locations, soil profiles, sampling times and different fertilization treatments). That is to say, each split tends to minimize the dissimilarity within the two resulting groups and maximize the dissimilarity between the two resulting groups based on a single environmental classification by comparing all the potential splits in the data.
Multivariate regression tree of the soil bacterial diversity data associated with different sampling locations (FQ, QY and TY), fertilization treatments (sole or conjoint application of N, P, K and OM, present (PRES) and absent (ABSE) were used to indicate if certain fertilizer was applied), soil profile depth (horizon A (0–20 cm) and B (20–40 cm)) and sampling times (first (FIRS) and second (SECO)). The standardized diversity estimates were used to construct MRT. Bar plots show the multivariate means of diversity estimates at each branch, and the numbers of samples included in that splits are shown under bar plots. FQ, Fengqiu; MRT, multivariate regression tree; OM, organic manure; QY, Qiyang; TY, Taoyuan.
Each split in an MRT was represented graphically as a branch in a tree. Each branch in the tree was labeled with the levels of the classification variable that were placed in that branch (for example, TY and QY versus FQ, Figure 1). The multivariate means of response variables (for example, richness, Shannon index and Simpson index) are shown as bar plot (Figure 1). The number of samples included in that branch is shown under the bar plot. Diversity indices were standardized to the same mean before performing MRT analysis. A 10 cross-validation process was used to decrease the structure complexity of MRT to highlight the main relationship between biological data and environmental variables while the predictive ability was only marginally worse than that of the whole tree, as the cross-validated relative error decreased to a plateau in that split. Cross-validated relative error varies from zero for a perfect tree to close to one for a poor tree (De'ath, 2002). MRT analysis was carried out by using the package ‘mvpart’ within the ‘R’ statistical programming environment.
Aggregated boosted tree analysis
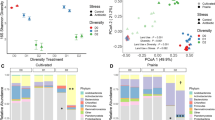
Aggregated boosted tree (ABT) analysis is a statistical learning method that aims to attain both accurate prediction and explanation. A major strongpoint of ABT analysis is that it can increase the accuracy of prediction relative to other methods such as boosted trees, bagged trees, random forests and generalized additive models. Furthermore, the reasons we use ABT analysis are because (1) it can deal with many types of response variables (numeric, categorical and censored) and environmental variables (numeric, categorical) (De'ath, 2007), which is well suited for our data; (2) it can quantitatively and visually evaluate the relative influence of environmental variables on the soil bacterial diversity variation (Figure 2) and (3) interactions between predictors (environmental variables) can also be quantified and visualized (Figure 3). The optimizing of the ABT model is necessary to attain accurate prediction and explanation during ABT analysis. A major estimate of how close model predictions are to their true values on average is the predictive error (PE) of a statistical model (De'ath, 2007). A statistical model with lower PE value means a higher accuracy of prediction. For an ABT analysis, comparison of PE of trees based on different interaction levels can determine the level of interaction influence between the environmental variables. The single response of each diversity estimate (for example, richness, Shannon index and Simpson index) to environmental variables and the relative influence of different factors on the soil bacterial diversity variation can be quantitatively evaluated and visualized as partial dependency plots and relative importance plot (Figure 2). ABT analysis was carried out by using the package ‘gbmplus’ within the ‘R’ statistical programming environment.
Relative importance and partial dependency plots of extracted predictors for richness (a) Shannon index (b) and Simpson index (c) by optimized ABT model considering the main effect and two-way interaction effect and using predictors of sampling location, soil profile, sampling time, OM and P. N and K were not included in the optimized ABT model because their effects on the diversity estimates were negligible as determined by a series of pre-ABT analysis. ABT, aggregated boosted tree; FQ, Fengqiu; OM, organic manure; QY, Qiyang; TY, Taoyuan.
Partial dependency plots of the Shannon index show the interaction effects of sampling location with OM and P. For the QY soil, applications of OM and P showed a distinct positive effect on the Shannon index comparing to no OM and P application; but for the FQ and TY soils, applications of OM and P showed no substantial effects on the Shannon index. FQ, Fengqiu; OM, organic manure; QY, Qiyang; TY, Taoyuan.
Results
Multivariate regression trees
The MRT analysis explains the relationship of diversity estimates and environmental variables in a visualized tree with nine splits based on sampling locations, soil profiles, sampling times and fertilizers P and OM (Figure 1). The tree explained 78.9% of the variance of the standardized diversity indices. In this tree, diversity estimates were first split by sampling locations. Data from TY and QY were placed into one group and data from FQ were placed into a separate group. This split produced two groups of data, each more homogenous than the original group. This single split in the data accounted for 29.1% of the variation in the original dataset. Each of the two groups was split further by soil profile depth, which explained 17.6% and 1.9% of the variation, respectively. The diversity estimates of 0–20 cm soils from QY and TY were then split by sampling location again into two groups of TY and QY. This split accounted for 11.7% of the variation in the data. The following left group was split by sampling time accounting for 11.7% of the variation and the right group was split one by one according to OM, P and sampling time, which explained 3.7%, 1.8% and 1.4% of the variation, respectively. To summarize, diversity estimates were mainly distinguished by sampling location with a total explanation of 40.8% of the variation, followed by soil profile (19.5%), sampling time (13.1%), OM (3.7%) and P (1.8%). The contributions of N and K to the variation were less than 1.4% and were not used in structuring the MRT.
Bar plots at the nine nodes of MRT effectively illustrated the overall distributions of diversity estimates throughout the tree. Furthermore, the single regression tree was also useful to describe the distribution of single diversity estimate throughout different groups (Supplementary Figure S2). Overall, the soil bacterial diversity was higher in FQ than in QY and TY, higher in surface soil than in subsurface soil and under some constrained conditions, soil bacterial diversity was higher in first sampling than in second sampling (for example, TY surface soils) and higher in soils fertilized with OM and P than in soils without OM and P application (for example, QY surface soil). When we compared paired samples for the same sampling location, sampling time and fertilization treatment but different soil profiles, a strong correlation between soil bacterial diversity and the sampling depth was observed (Figure 4).
Because MRT is a form of constrained clustering, it is possible to explore whether there are still other unrecorded important environmental factors responsible for the unexplained diversity variation by comparing the MRT solution to unconstrained clustering solution. The unconstrained cluster analysis accounts for substantially more of the diversity variation than the MRT analysis (Figure 5).
Comparison of the variance of soil bacterial diversity explained by MRT analysis (constrained clustering) and cluster analysis (unconstrained clustering) across the classification size. The unconstrained cluster analysis accounts for substantially more of the diversity variation than the MRT analysis, which indicates that some unrecorded factors are responsible for the difference in explaining the diversity variation. MRT, multivariate regression tree.
Aggregated boosted trees
It is necessary to optimize the ABT model by determining the levels of interaction influences and which environmental variable should be used in analysis before using ABT analysis to achieve the final results. The optimizing of the ABT model of richness, Shannon index and Simpson index performed the same process. Hence, we considered Shannon index as an example to show the process of optimizing the ABT model.
To determine the levels of interaction influences of environmental variables on Shannon diversity estimates, seven ABTs were fitted by considering the interaction effect from one way to seven way and using all seven predictors. The PE of the ABT model considering just main effect abruptly reduced from a main effect of 2.51% to a PE of 1.40% of the ABT model, considering both main effect and two-way interaction effect and then leveled off, which indicates that it is enough to just consider main effects and two-way interaction effects of seven environmental variables on the diversity estimates. Similarly, to determine whether all seven predictors should be used to evaluate their effects on Shannon index, seven further ABTs were fitted by considering the main effect and two-way interaction effect and dropping out one predictor for every ABT. The PE of ABT dropping out sampling location, soil profile, sampling time, OM and P increased, whereas the PE of ABT dropping out N and K decreased. This result suggests that, differing from sampling location, soil profile, sampling time, OM and P, the effects of N and K on Shannon index were negligible.
As a result, the optimized ABT model, just considering the main effect and two-way interaction effect and using predictors of sampling location, soil profile, sampling time, OM and P, was used to explore (1) the relative importance of different environmental variables in influencing the soil bacterial diversity, (2) the responses of diversity estimates to different environmental variables and (3) the 10 possible two-way interaction effects. The relative importance of environmental variables on three diversity estimates shows generally the same trend, with a strong effect for sampling location, moderate effects for soil profile and sampling time, and weak effects for the applications of OM and P (Figure 2). The partial dependency plots of the single predictor showed a predominant increase in soil bacterial diversity in FQ, moderate increase in diversity in surface soil and the first sampling soil, and relative weak increase in the diversity in the soils with OM and P applications (Figure 2). The response of the Simpson index to environmental variables (Figure 2c) was opposite to the response of the Shannon index (Figure 2b) and richness (Figure 2a) to environmental variables. The results of ABT were consistent with the results of MRT.
The relative importance, partial influence and interactions for the five extracted predictors were used to identify that of the 10 possible two-way interactions were important. The results showed that, although two-way interaction effects generally showed as plus effect and determined by higher important predictors, still some pair interactions showed contrasting pattern. For example, the applications of OM and P had a strong effect on soil bacterial diversity at QY, but little effect at FQ and TY (Figure 3).
Discussion
Differences in bacterial diversity
The MRT and optimized ABT showed that the soil bacterial diversity was substantially higher in FQ than in QY and TY, which indicated the existence of horizontal spatial variation of the bacterial diversity. However, because the number of our sampling locations to evaluate the spatial variation of bacterial diversity is very limited, we could not definitely conclude that the bacterial diversity distribution is along a horizontal spatial gradient (for example, a latitude or climate gradient represented by the three sampling locations). Although there was a study showing that soil microbial diversity decreased with increasing latitude, and correlated positively with measures of atmospheric temperature and pH (Staddon et al., 1998), other studies argued that bacterial diversity was unrelated to site temperature and latitude, but differed by ecosystem type, and these differences could largely be explained by soil pH (Fierer and Jackson, 2006). In our study, the soil pH in FQ was also distinctly higher than that in QY and TY (data not shown). Therefore, although there are some superficial contradictions among the studies as mentioned above, a common factor of soil pH could still be used to explain the results. Moreover, although a definite horizontal spatial gradient was not found in our study, at least the observation that the microbial diversity is substantially different between locations is true but not the ‘noise’ among the three locations that can be easily considered as the presentation of historical changes and events.
In our study, another substantial variation was also observed that the soil bacterial diversity was higher in surface soil than in subsurface soil, which indicated the vertical spatial variation of soil bacterial diversity. Similarly, other research reported that the surface bacterial communities had diversity index (reciprocal of Simpson index) values that were 2–3 orders of magnitude greater than those for the subsurface communities in low-carbon soils (Zhou et al., 2002). The same trend of bacterial diversity was also observed in meromictic lake ecosystem (Øvreås et al., 1997). All of these studies, including our work, indicate that the distribution of bacterial diversity exists with a vertical spatial pattern that bacterial diversity decreases from the surface to the subsurface soil.
The soil bacterial diversity also showed seasonal variation, but the responses to seasonal variation (sampling time) were different in different sampling locations. For example, two sets of surface soil samples collected in January and July 2006 in TY showed that samples collected in January had an estimated bacterial richness 30%–86% higher than the corresponding samples with the same fertilization treatments collected in July (Supplementary Table S1). However, the two sets of surface soil samples from QY showed no definite variation trend between the paired samples. Considering the agricultural practice that the TY soil was water saturated with paddy rice in July but relatively dry in January, it is not surprising that the soil bacterial diversity in TY distinctly decreased in July because of unfavorable anaerobic environment for bacteria. Despite the distinct variation trend in TY or the unsystematic variation in QY, the seasonal variation of soil bacterial diversity seems under the control of sample location.
The applications of different fertilizers were a group of the least influencing factors among all the factors considered in this study. In some constrained situations, the application of OM and P also showed distinct influences on soil bacterial diversity. For example, the application of OM and P can distinctly increase the soil bacterial diversity in surface soil in QY (Figures 1, 3). Here, comparing with the historical evolution factors, the applications of different fertilizers were thought to be an appropriate proxy of contemporary environmental disturbances. A large body of other studies has also showed that microbial diversity can be affected by different environmental disturbances, including heavy metal (Gans et al., 2005), organic pollutant (Stephen et al., 1999) and herbicide (el Fantroussi et al., 1999).
As the DGGE fingerprinting method underestimates the true bacterial richness (Muyzer et al., 1993), we cannot estimate the absolute bacterial diversity changes across the environmental variables. However, the estimate of the relative variation trend of the soil bacterial diversity remains valid if the examined richness proportionally changes among the samples, which is expected especially when the samples are taken from a common habitat (Bell et al., 2005a). Similarly, although sampling effort affects richness estimates, alterations to the sampling effort are not expected to significantly alter the relative estimates so long as they remain constant among the samples.
Historical contingencies and contemporary disturbances
This study is the first quantitative examination of the relative importance of contemporary disturbances and historical contingencies in influencing large-scale soil bacterial diversity using a large set of manipulated field-based data by the combination of culture-independent molecular techniques and advanced statistical analyses. A large body of previous studies have shown that microbial assemblages can be affected by different environmental disturbances (el Fantroussi et al., 1999; Stephen et al., 1999; Gans et al., 2005). However, almost all of this kind of work has been site-specific, limiting our understanding of the factors that structure soil bacterial communities across regions. At the same time, more and more evidence supports the idea that free-living microorganisms vary in abundance, distribution and diversity, across various taxonomic and spatial scales (McArthur et al., 1988; Cho and Tiedje, 2000; Crump et al., 2004; Fierer and Jackson, 2006). Taxa–area relationships have been repeatedly reported in microbial communities, in both contiguous and island habitats (Green et al., 2004; Horner-Devine et al., 2004; Bell et al., 2005a), which provided further evidence for microbial biogeography. However, most of such studies just exclude the hypothesis that microbial assemblages are spatially random. They still did not answer the question that how much of the spatial variation in microbial distributions and assemblages is due to the contemporary environmental variations or historical contingencies. In fact, if studies are not directed and driven within a theory framework, the field of microbial biogeography will probably become merely phenomenological description, instead of exploring the mechanisms that generate the patterns (Martiny et al., 2006; Prosser et al., 2007). Some recent studies intensively explored the relative importance of historical (spatial) factors and contemporary environmental (local) factors to microbial assemblages in aquatic environment (Yannarell and Triplett, 2005; Langenheder and Ragnarsson, 2007; Vyverman et al., 2007), whereas similar research in soil environment was scarce. A recent study examined the responses of Burkholderia ambifaria intraspecific diversity to spatial distance and EH in a patch soil ecosystem and showed that whole-genome similarities may reflect the simultaneous effects of both spatial distance and EH in microbial populations, whereas the pure effects of each factor only contributed to <2% of the total genetic variation (Ramette and Tiedje, 2007). This study was still performed in a small spatial scale. Therefore, it is necessary to explore the relative influence of contemporary disturbances and historical contingencies on the soil bacterial diversity variation in a large spatial scale (for example, regional scale), which we have done in this study.
Both MRT and ABT analyses showed that soil bacterial diversity can be distinguished by most pre-defined categorical factors and that the relative importance of different categorical factors on soil microbial diversity variation was ranked as sampling location, soil profile, sampling time, OM and P. N and K seemed to have no substantial effect on the differences in the soil bacterial diversity. The significance of our results is that the absolutely stronger driving factor of historical contingencies on soil bacterial diversity variation was determined. MRT analysis indicates that 60.3% of the variation in soil bacterial diversity can be attributed to the historical contingencies and 5.5% can be explained by environmental disturbances (OM and P application). Whereas, other studies investigating the influence of environmental and historical parameters on bacterial assemblages using multivariate methods and molecular fingerprinting methods just achieved low fractions of explanation and were hard to definitely distinguish a more important factor (Langenheder and Ragnarsson, 2007; Ramette and Tiedje, 2007), although we do recognize that it may be more difficult to distinguish the effect of historical or spatial factor over relatively small spatial scale. Furthermore, different from the previous studies exploring the influence of environmental and historical factors on bacterial assemblages, the seasonal variation of soil bacterial diversity was considered and examined in this study, which make our examinations to soil bacterial diversity more comprehensive but not just snapshots.
The strong effects of distinct geographic location and soil profile on soil bacterial diversity indicate a strong biogeographic provincialism, in which differences in bacterial composition and diversity are due to past evolutionary events (for example, spatial isolation, physical barrier, dispersal history and past EH) rather than present attributes of the environment. In fact, both horizontal distinct location and vertical soil profile did not directly affect soil microbial diversity per se. Location and depth are just related to the possibility that past divergence and diversification of microbial assemblages, whether due to neutral genetic drift or adaptation to the past environments, are inherited by the genetic isolation because of spatial separation (Borcard and Legendre, 1994).
Our results also showed that some, not all, of the fertilization treatments caused soil bacterial diversity variation at a small spatial scale (for example, the application of OM and P can distinctly increase the soil bacterial diversity in surface soil in QY). This result indicates that contemporary environmental disturbances can also influence soil bacterial diversity under the control of the overall pattern of provincialism caused by historical contingencies. That is to say, although present environmental disturbances cannot rapidly and completely erase the effects of past evolutionary and ecological events, they also labeled their effects on soil bacterial composition and diversity in a local spatial scale because of the enormous dispersal capabilities of the microorganisms.
The major novel result of this study is that we observed distinct scale-dependence of the effect of historical contingencies and contemporary environmental disturbances on soil bacterial diversity in a sole well-conducted experiment, which is very similar to those documented in macroorganisms (Willis and Whittaker, 2002; Ricklefs, 2004; Qian et al., 2007). Martiny et al. (2006) also suggested that the relative influence of historical and environmental factors seems to be related to the scale of sampling, but their conclusion arrived from the comparison with several studies over different spatial scale. For example, in intercontinental-scale (tens of thousands of kilometers) studies, microbial assemblages could be significantly distinguished by distance but did not correlate with many environmental factors measured (Papke et al., 2003; Whitaker et al., 2003; Vyverman et al., 2007). In small-scale (a few kilometers) studies, some just found the significant environmental effects (Kuske et al., 2002; Horner-Devine et al., 2004) and some found both the environmental and spatial effects (Langenheder and Ragnarsson, 2007; Ramette and Tiedje, 2007). At intermediate scales (10–3000 km), some studies found a significant distance effect (Green et al., 2004; Reche et al., 2005; Yannarell and Triplett, 2005), and environmental conditions also seemed to influence the composition at this spatial scale (Rohwer et al., 2002; Green et al., 2004; Hewson and Fuhrman, 2004; Yannarell and Triplett, 2005).
Although MRT explained most of the diversity variation, still 21.1% of the diversity variation cannot be explained by MRT in this study. The comparison of the MRT solution to unconstrained clustering solution showed that the unconstrained cluster analysis accounts for substantially more of the diversity variation than MRT analysis. This indicates that unobserved factors, additional to the explanatory variables of the tree analysis, are responsible for the difference in explaining the diversity variation. These unrecorded factors may be unmeasured environmental and spatial variability, sampling effects and neutral ecological drift (Ramette and Tiedje, 2007). In fact, the existence of unexplained variation may more fit into the actual situation, because it is impossible to record all of the possible factors. However, it is still a substantial observation that the effect of historical contingencies on microbial diversity was stronger than that of contemporary disturbances, as MRT analysis totally explained 78.9% of the diversity variation and historical contingencies (both sampling location and soil profile) can be used to explain 60.3% of the diversity variation.
This study combined molecular techniques and advanced statistical analyses to examine the relative importance of contemporary disturbances and historical contingencies on the soil bacterial diversity variation based on a large set of manipulated field-based data at a regional spatial scale. To our knowledge, no previous study has performed such work. Our results clearly show that historical contingencies could be the dominant factor to drive the bacterial diversity variation across a regional spatial scale (about 1000 km), whereas some of the contemporary disturbances also caused soil bacterial diversity variation at a local spatial scale, which exclusively demonstrated the hypothesis that the influence patterns of contemporary disturbances and historical contingencies on soil bacterial diversity are fundamentally similar to the patterns observed for plants and animals. This observation indicates that there are some aspects of biogeography that might be common to all life, which would extend our understanding of the biogeography of organisms.
References
Balvanera P, Pfisterer AB, Buchmann N, He J-S, Nakashizuka T, Raffaelli D et al. (2006). Quantifying the evidence for biodiversity effects on ecosystem functioning and services. Ecol Lett 9: 1146–1156.
Bell T, Ager D, Song J-I, Newman JA, Thompson IP, Lilley AK et al. (2005a). Larger islands house more bacterial taxa. Science 308: 1884.
Bell T, Newman JA, Silverman BW, Turner SL, Lilley AK . (2005b). The contribution of species richness and composition to bacterial services. Nature 436: 1157–1160.
Borcard D, Legendre P . (1994). Environmental control and spatial structure in ecological communities: an example using oribatid mites (Acari, Oribatei). Environ Ecol Stat 1: 37–61.
Cho J-C, Tiedje JM . (2000). Biogeography and degree of endemicity of fluorescent Pseudomonas strains in soil. Appl Environ Microbiol 66: 5448–5456.
Crump BC, Hopkinson CS, Sogin ML, Hobbie JE . (2004). Microbial biogeography along an estuarine salinity gradient: combined influences of bacterial growth and residence time. Appl Environ Microbiol 70: 1494–1505.
De'ath G . (2002). Multivariate regression trees: a new technique for modeling species–environment relationships. Ecology 83: 1105–1117.
De'ath G . (2007). Boosted trees for ecological modeling and prediction. Ecology 88: 243–251.
el Fantroussi S, Verschuere L, Verstraete W, Top EM . (1999). Effect of phenylurea herbicides on soil microbial communities estimated by analysis of 16S rRNA gene fingerprints and community-level physiological profiles. Appl Environ Microbiol 65: 982–988.
Ettema CH, Wardle DA . (2002). Spatial soil ecology. Trends Ecol Evol 17: 177–183.
Fierer N, Jackson RB . (2006). The diversity and biogeography of soil bacterial communities. Proc Natl Acad Sci USA 103: 626–631.
Fierer N, Schimel JP, Holden PA . (2003). Variations in microbial community composition through two soil depth profiles. Soil Biol Biochem 35: 167–176.
Finlay BJ . (2002). Global dispersal of free-living microbial eukaryote species. Science 296: 1061–1063.
Gans J, Wolinsky M, Dunbar J . (2005). Computational improvements reveal great bacterial diversity and high metal toxicity in soil. Science 309: 1387–1390.
Green JL, Holmes AJ, Westoby M, Oliver I, Briscoe D, Dangerfield M et al. (2004). Spatial scaling of microbial eukaryote diversity. Nature 432: 747–750.
He JZ, Shen JP, Zhang LM, Zhu YG, Zheng YM, Xu MG et al. (2007). Quantitative analyses of the abundance and composition of ammonia-oxidizing bacteria and ammonia-oxidizing archaea of a Chinese upland red soil under long-term fertilization practices. Environ Microbiol 9: 2364–2374.
He JZ, Xu ZH, Hughes J . (2005). Analyses of soil fungal communities in adjacent natural forest and hoop pine plantation ecosystems of subtropical Australia using molecular approaches based on 18S rRNA genes. FEMS Microbiol Lett 247: 91–100.
He JZ, Xu ZH, Hughes J . (2006). Molecular bacterial diversity of a forest soil under residue management regimes in subtropical Australia. FEMS Microbiol Ecol 55: 38–47.
Hewson I, Fuhrman JA . (2004). Richness and diversity of bacterioplankton species along an estuarine gradient in Moreton Bay, Australia. Appl Environ Microbiol 70: 3425–3433.
Horner-Devine MC, Lage M, Hughes JB, Bohannan BJM . (2004). A taxa–area relationship for bacteria. Nature 432: 750–753.
Kuske CR, Ticknor LO, Miller ME, Dunbar JM, Davis JA, Barns SM et al. (2002). Comparison of soil bacterial communities in rhizospheres of three plant species and the interspaces in an arid grassland. Appl Environ Microbiol 68: 1854–1863.
Langenheder S, Ragnarsson H . (2007). The role of environmental and spatial factors for the composition of aquatic bacterial communities. Ecology 88: 2154–2161.
Lefranc M, Thenot A, Lepere C, Debroas D . (2005). Genetic diversity of small eukaryotes in lakes differing by their trophic status. Appl Environ Microbiol 71: 5935–5942.
Lozupone CA, Knight R . (2007). Global patterns in bacterial diversity. Proc Natl Acad Sci USA 104: 11436–11440.
Madsen EL . (2005). Identifying microorganisms responsible for ecologically significant biogeochemical processes. Nat Rev Microbiol 3: 439–446.
Martiny JBH, Bohannan BJM, Brown JH, Colwell RK, Fuhrman JA, Green JL et al. (2006). Microbial biogeography: putting microorganisms on the map. Nat Rev Microbiol 4: 102–112.
McArthur JV, Kovacic DA, Smith MH . (1988). Genetic diversity in natural populations of a soil bacterium across a landscape gradient. Proc Natl Acad Sci USA 85: 9621–9624.
Muyzer G, de Waal E, Uitterlinden A . (1993). Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl Environ Microbiol 59: 695–700.
Naeem S, Li S . (1997). Biodiversity enhances ecosystem reliability. Nature 390: 507–509.
Øvreås L, Forney L, Daae FL, Torsvik V . (1997). Distribution of bacterioplankton in meromictic lake Saelenvannet, as determined by denaturing gradient gel electrophoresis of PCR-amplified gene fragments coding for 16S rRNA. Appl Environ Microbiol 63: 3367–3373.
Papke RT, Ramsing NB, Bateson MM, Ward DM . (2003). Geographical isolation in hot spring cyanobacteria. Environ Microbiol 5: 650–659.
Prosser JI, Bohannan BJM, Curtis TP, Ellis RJ, Firestone MK, Freckleton RP et al. (2007). The role of ecological theory in microbial ecology. Nat Rev Microbiol 5: 384–392.
Qian H, White PS, Song J-S . (2007). Effects of regional vs ecological factors on plant species richness: an intercontinental analysis. Ecology 88: 1440–1453.
Rajaniemi TK, Goldberg DE, Turkington R, Dyer AR . (2006). Quantitative partitioning of regional and local processes shaping regional diversity patterns. Ecol Lett 9: 121–128.
Ramette A, Tiedje JM . (2007). Multiscale responses of microbial life to spatial distance and environmental heterogeneity in a patchy ecosystem. Proc Natl Acad Sci USA 104: 2761–2766.
Reche I, Pulido-Villena E, Morales-Baquero R, Casamayor EO . (2005). Does ecosystem size determine aquatic bacterial richness? Ecology 86: 1715–1722.
Ricklefs RE . (2004). A comprehensive framework for global patterns in biodiversity. Ecol Lett 7: 1–15.
Rohwer F, Seguritan V, Azam F, Knowlton N . (2002). Diversity and distribution of coral-associated bacteria. Mar Ecol Prog Ser 243: 1–10.
Sandaa R-A, Torsvik V, Enger O, Daae FL, Castberg T, Hahn D . (1999). Analysis of bacterial communities in heavy metal-contaminated soils at different levels of resolution. FEMS Microbiol Ecol 30: 237–251.
Staddon WJ, Trevors JT, Duchesne LC, Colombo Ca . (1998). Soil microbial diversity and community structure across a climatic gradient in western Canada. Biodivers Conserv 7: 1081–1092.
Stephen JR, Chang Y-J, Gan YD, Peacock A, Pfiffner SM, Barcelona MJ et al. (1999). Microbial characterization of a JP-4 fuel-contaminated site using a combined lipid biomarker/polymerase chain reaction-denaturing gradient gel electrophoresis (PCR-DGGE)-based approach. Environ Microbiol 1: 231–241.
Torsvik V, Salte K, Sørheim R, Goksøyr J . (1990). Comparison of phenotypic diversity and DNA heterogeneity in a population of soil bacteria. Appl Environ Microbiol 56: 776–781.
Vyverman W, Verleyen E, Sabbe K, Vanhoutte K, Sterken M, Hodgson DA et al. (2007). Historical processes constrain patterns in global diatom diversity. Ecology 88: 1924–1931.
Whitaker RJ, Grogan DW, Taylor JW . (2003). Geographic barriers isolate endemic populations of hyperthermophilic archaea. Science 301: 976–978.
Willis KJ, Whittaker RJ . (2002). Species diversity—scale matters. Science 295: 1245–1248.
Yannarell AC, Triplett EW . (2005). Geographic and environmental sources of variation in lake bacterial community composition. Appl Environ Microbiol 71: 227–239.
Yu Z, Morrison M . (2004). Comparisons of different hypervariable regions of rrs genes for use in fingerprinting of microbial communities by PCR-denaturing gradient gel electrophoresis. Appl Environ Microbiol 70: 4800–4806.
Zhou J, Xia B, Treves DS, Wu LY, Marsh TL, O'Neill RV et al. (2002). Spatial and resource factors influencing high microbial diversity in soil. Appl Environ Microbiol 68: 326–334.
Acknowledgements
This work was partially supported by the National Basic Research Program of China (2005CB121105), the Natural Science Foundation of China (40571082, 50621804) and the Chinese Academy of Sciences (KZCX2-YW-408). The assistance of staff at the Fengqiu, Taoyuan and Qiyang experimental stations in soil sampling are sincerely appreciated.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The ISME Journal website (http://www.nature.com/ismej)
Supplementary information
Rights and permissions
About this article
Cite this article
Ge, Y., He, Jz., Zhu, Yg. et al. Differences in soil bacterial diversity: driven by contemporary disturbances or historical contingencies?. ISME J 2, 254–264 (2008). https://doi.org/10.1038/ismej.2008.2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2008.2
Keywords
This article is cited by
-
Nitrogen deposition mediates more stochastic processes in structuring plant community than soil microbial community in the Eurasian steppe
Science China Life Sciences (2024)
-
Biogeographic patterns of meio- and micro-eukaryotic communities in dam-induced river-reservoir systems
Applied Microbiology and Biotechnology (2024)
-
Elevated temperature and CO2 strongly affect the growth strategies of soil bacteria
Nature Communications (2023)
-
A Small Study on Clostridioides difficile in Spinach Field Soil and the Chemical and Microbial Factors that may Influence Prevalence
Current Microbiology (2023)
-
Geographic Scale Influences the Interactivities Between Determinism and Stochasticity in the Assembly of Sedimentary Microbial Communities on the South China Sea Shelf
Microbial Ecology (2023)