Abstract
The sulfidic Frasassi cave system affords a unique opportunity to investigate niche relationships among sulfur-oxidizing bacteria, including epsilonproteobacterial clades with no cultivated representatives. Oxygen and sulfide concentrations in the cave waters range over more than two orders of magnitude as a result of seasonally and spatially variable dilution of the sulfidic groundwater. A full-cycle rRNA approach was used to quantify dominant populations in biofilms collected in both diluted and undiluted zones. Sulfide concentration profiles within biofilms were obtained in situ using microelectrode voltammetry. Populations in rock-attached streamers depended on the sulfide/oxygen supply ratio of bulk water (r=0.97; P<0.0001). Filamentous epsilonproteobacteria dominated at high sulfide to oxygen ratios (>150), whereas Thiothrix dominated at low ratios (<75). In contrast, Beggiatoa was the dominant group in biofilms at the sediment–water interface regardless of sulfide and oxygen concentrations or supply ratio. Our results highlight the versatility and ecological success of Beggiatoa in diffusion-controlled niches, and demonstrate that high sulfide/oxygen ratios in turbulent water are important for the growth of filamentous epsilonproteobacteria.
Similar content being viewed by others
Introduction
Sulfidic caves form in limestone (CaCO3) rocks where sulfide-rich groundwater interacts with oxygen at the water table. The Frasassi cave system hosts a rich, sulfur-based lithoautotrophic microbial ecosystem (Sarbu et al., 2000; Vlasceanu et al., 2000; Macalady et al., 2006, 2007; Jones et al., 2008). Previous studies of the geochemistry of the cave waters have revealed that they are mixtures of slightly salty, sulfidic groundwater diluted 10–60% by oxygen-rich, downward-percolating meteoric water (Galdenzi et al., 2007). Initial observations of the abundant biofilms in cave streams and pools suggested that they respond dynamically to seasonal and episodic hydrologic changes. In particular, we noted that changes in specific conductivity (tracking freshwater dilution) and water flow characteristics correspond with morphological differences in the biofilms. Our initial observations motivated a systematic, multi-year study of the population structure of biofilms collected from cave waters with a wide range of hydrological and geochemical characteristics. The goal of this study was to identify the ecological niches (defined as the range of environmental conditions that permit growth) of major biofilm-forming populations.
Unraveling the effects of changing hydrologic conditions on microbial growth within the cave system is not trivial because dilution of the sulfidic groundwater has multiple effects relevant to microbial metabolism. The input of meteoric water increases water depth and flow rates, dilutes dissolved species in the sulfidic aquifer and adds dissolved oxygen. Water flow conditions that increase turbulence also increase sulfide degassing from the water and oxygen transport into the water from the oxygenated cave air. We investigated microbial populations using a full-cycle rRNA approach. Environmental conditions were measured using microelectrode voltammetry and bulk water geochemical analyses. We found that both sulfide/oxygen ratios and physical water flow characteristics are important for determining the distributions of sulfur-oxidizing groups.
Materials and methods
Field site, sample collection and geochemistry
The Grotta Grande del Vento-Grotta del Fiume (Frasassi) cave system is actively forming in Jurassic limestone in the Appennine Mountains of the Marches Region, Central Italy. The waters of the cave system are near neutral (pH 6.9–7.4) and have specific conductivities ranging from 1200 to 3500 μS cm−1, or roughly 4–5% of average marine salinity. The major ions are Na+, Ca2+, Cl−, HCO3− and SO42− (Galdenzi et al., 2007). Concentrations of electron donors and acceptors other than sulfur species and oxygen are as follows: bicarbonate (5.3–7.1 mM), ammonium (30–175 μM), methane (1–20 μM), dissolved iron (<0.1 μM), dissolved manganese (<0.04 μM), nitrate (not detected, <0.7 μM) and nitrite (not detected, <2.0 μM). Organic carbon concentrations range between 0.16 and 4.5 mg l−1.
Biofilms from cave springs and streams were collected from 5 to 40 cm water depth at sample locations shown in Figure 1 in May (wet season) and August (dry season) in 2005, 2006 and 2007. Biofilms were harvested using sterile plastic transfer pipettes into sterile tubes, stored on ice and processed within 4–6 h of collection. Subsamples for fluorescence in situ hybridization (FISH) were fixed in 4% (w/v) paraformaldehyde and stored at −20 °C. Samples for clone library construction were preserved in four parts RNAlater (Ambion/Applied Biosystems, Foster City, CA, USA) to one part sample (v/v). Water samples were filtered (0.2 μm) into acid-washed polypropylene bottles and stored at 4 °C until analyzed. Conductivity, pH and temperature of the waters were measured in the field using sensors attached to a 50i multimeter (WTW, Weilheim, Germany). Dissolved sulfide and oxygen concentrations were measured in the field using a portable spectrophotometer (Hach Co., Loveland, CO, USA) using the methylene blue and indigo carmine methods, respectively. Duplicate sulfide analyses were within 1% of each other. Replicate oxygen analyses were within 20% of each other. Nitrate, nitrite, ammonium and sulfate were measured at the Osservatorio Geologico di Coldigioco Geomicrobiology Lab using a portable spectrophotometer within 12 h of collection according to the manufacturer's instructions (Hach Co.). Light microscopy was performed on live samples within 8 h of collection on a Zeiss Model 47-30-12-9901 optical microscope ( × 1250) at the Osservatorio Geologico di Coldigioco Geomicrobiology Lab.
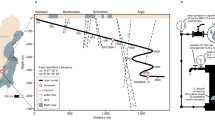
Map of the Frasassi cave system showing sample locations (open circles). The small inset shows the location of the Frasassi cave system in Italy. Major named caves in the Frasassi system are shown in different shades of gray. Topographic lines and elevations in meters refer to the surface topography. Base map courtesy of the Gruppo Speleologico CAI di Fabriano.
Microelectrode voltammetry
Voltammetric signals are produced when dissolved or colloidal species interact with the surface of a gold amalgam working electrode. Electron flow resulting from redox half-reactions at a 100 μm diameter tip is registered as a current that is proportional to concentration (Skoog et al., 1998; Taillefert and Rozan, 2002). The gradients associated with microbial metabolism in biofilms are thus readily measured using this technique. Aqueous and colloidal species that are electroactive at gold amalgam electrode surfaces include: H2S, HS−, S8, polysulfides, S2O32−, S4O62−, HSO3−, Fe2+, Fe3+, FeS(aq), Mn2+, O2 and H2O2 (Luther et al., 1991, 2001; Xu et al., 1998; Taillefert et al., 2000; Druschel et al., 2003, 2004; Glazer et al., 2004, 2006). Voltammetric analyses of dissolved sulfur, iron and other species in the field were accomplished using a DLK-60 potentiostat powered with a 12 V battery and controlled with a GETAC ruggedized computer. To protect the electrochemical system from drip waters and high humidity, the potentiostat was contained inside a storm case containing dryrite humidity sponges and modified with rubber stripping to allow the communication ribbon cable and electrode cables to go outside the case while keeping the inside sealed. Electrodes were constructed as described in Brendel and Luther (1995).
Voltammetric analyses in the cave system involved placing working (gold amalgam) electrodes into a narishege three-axis micromanipulator with a two-arm magnetic base on a steel plate. At each sampling location, the reference and counter electrodes were placed in the flowing water near the biofilm. The working electrode was lowered to the air–water interface, then to the biofilm–water interface and subsequently lowered in increments to profile the biofilm. We used both cyclic voltammetry between −0.1 and −1.8 V (vs Ag/AgCl) at scan rates from 200 to 2000 mV s−1 with a 2 s conditioning step, and square wave voltammetry between −0.1 and −1.8 V (vs Ag/AgCl) at scan rates from 200 to 1000 mV s−1, with a pulse height of 25 mV. Analyses were carried out in sets of at least 10 sequential scans at each sampling point in space, with the first three scans discarded.
Clone library construction
Environmental DNA was obtained using phenol–chloroform extraction as described in Bond et al. (2000) using 1 × buffer A (100 mM Tris, 100 mM NaCl, 25 mM EDTA, 1 mM sodium citrate, pH 8.0) instead of phosphate-buffered saline for the first washing step. Small subunit ribosomal RNA genes were amplified by PCR from the bulk environmental DNA. Libraries were constructed from four samples using the bacteria-specific primer set 27f and 1492r. Each 50 μl reaction mixture contained environmental DNA template (1–150 ng), 1.25 U ExTaq DNA polymerase (TaKaRa Bio Inc., Shiga, Japan), 0.2 mM each dNTPs, 1 × PCR buffer, 0.2 μM 1492r universal reverse primer (5′-GGTTACCTTGTTACGACTT-3′) and 0.2 μM 27f primer (5′-AGAGTTTGATCCTGGCTCAG-3′). A universal library was constructed from sample FS06-12 using universal forward primer 533f (5′-GTGCCAGCCGCCGCGGTAA-3′) and 1492r. Thermal cycling was as follows: initial denaturation 5 min at 94 °C, 25 cycles of 94 °C for 1 min, 50 °C for 25 s and 72 °C for 2 min followed by a final elongation at 72 °C for 20 min. PCR products were cloned into the pCR4-TOPO plasmid and used to transform chemically competent OneShot MACH1 T1 Escherichia coli cells as specified by the manufacturer (TOPO TA cloning kit; Invitrogen, Carlsbad, CA, USA). Colonies containing inserts were isolated by streak plating onto Luria–Bertoni agar containing 50 μg ml−1 kanamycin. Plasmid inserts were screened using colony PCR with M13 primers (5′-CAGGAAACAGCTATGAC-3′ and 5′-GTAAAACGACGGCCAG-3′). Colony PCR products of the correct size were purified using the QIAquick PCR purification kit (Qiagen Inc., Valencia, CA, USA) following the manufacturer's instructions. Full-length sequences for 70–80 clones from each bacterial library were obtained, in addition to 60 sequences from the universal library constructed from sample FS06-12.
Sequencing and phylogenetic analysis
Clones were sequenced at the Penn State University Biotechnology Center using T3 and T7 plasmid-specific primers. Sequences were assembled with Phred base calling using CodonCode Aligner v.1.2.4 (CodonCode Corp., Dedham, MA, USA) and manually checked for ambiguities. The nearly full-length gene sequences were compared against sequences in public databases using BLAST (Altschul et al., 1990) and submitted to the online analyses CHIMERA_CHECK v.2.7 (Cole et al., 2003) and Bellerophon 3 (Huber et al., 2004). Putative chimeras were excluded from subsequent analyses. Non-chimeric sequences were aligned using the NAST aligner (DeSantis et al., 2006), added to an existing alignment containing >150 000 nearly full-length bacterial sequences in ARB (Ludwig et al., 2004), and manually refined. Alignments were minimized using the Lane mask (1286 nucleotide positions) (Lane, 1991). Phylogenetic trees were computed using neighbor joining (general time-reversible model) with 1000 bootstrap replicates. Neighbor-joining trees were compared with maximum likelihood trees (general time-reversible model, site-specific rates and estimated base frequencies). Both analyses were computed using PAUP* 4.0b10 (Swofford, 2000).
Probe design and FISH
Probes were designed and evaluated as described in Hugenholtz et al. (2001), including checks against all publicly available sequences using megaBLAST searches of the non-redundant databases at National Center for Biotechnology Information (NCBI). FISH experiments were carried out as described in Amann (1995) using the probes listed in Table 1. Oligonucleotide probes were synthesized and labeled at the 5′ ends with fluorescent dyes (Cy3, Cy5 and FLC) at Sigma-Genosys (St Louis, MO, USA). Cells were counterstained after hybridization with 4′,6′-diamidino-2-phenylindole (DAPI), mounted with Vectashield (Vectashield Laboratories Inc., Burlingame, CA, USA) and viewed on a Nikon E800 epifluorescence microscope. Images were collected and analyzed using NIS Elements AR 2.30, Hotfix (Build 312) image analysis software. The object count tool was used to measure areas covered by cells hybridizing with specific probes. Ten images were collected for each sample, taking care to represent the sample variability, and a total DAPI-stained area of approximately 3 × 104 μm2 (equivalent to the area of 5 × 104 E. coli cells) was analyzed for quantitation.
Statistical analyses
The program MINITAB (Minitab Inc., State College, PA, USA) was used for all statistical analyses. A two-sided Student's t-test was used to compare sulfide/oxygen ratios associated with Thiothrix and epsilonproteobacteria, and correlations between parameters were analyzed using the Pearson method.
Nucleotide sequence accession numbers
The 16S rRNA gene sequences determined in this study were submitted to the GenBank database under accession numbers EF467442–EF467519 and EU101023–EU101289.
Results
Field observations and geochemistry
Two common biofilm morphologies were collected from a variety of sample sites in the cave system (Figure 1) over a 3-year period: (1) rock-attached streamers and (2) biofilms developed on the surface of fine sediment. Streamers 1–5 mm thick and 5–20 cm long were attached to rocks in quickly flowing or turbulent water (Supplementary Figure 1, upper panel). Sediment surface biofilms <1 mm thick were present in eddies or stream reaches with slower flow, at the interface between the water column and fine gray sediment (Supplementary Figure 1, lower panel). We commonly observed the two biofilm types coexisting in a patchwork corresponding to spatial variations in water flow. All of the biofilms contained abundant elemental sulfur particles, as evidenced by microscopic observations of the particles under polarized light and by rapid dissolution of the particles in ethanol (Nielsen et al., 2000).
Dissolved ion concentrations, including concentrations of electron donors and acceptors such as sulfides, ammonium and sulfate, were strongly correlated with specific conductivity (0.89 <r<0.98; P<0.0001). This result is consistent with previous work showing that the cave water geochemistry is controlled primarily by physical mixing processes rather than biological activity (Galdenzi et al., 2007). In contrast, dissolved oxygen concentrations were not strongly correlated with specific conductivity (r=−0.39; P=0.04). Total dissolved sulfide and oxygen concentrations for each biofilm sample collected in the study are plotted in Figure 2.
Dissolved oxygen and total sulfide concentrations for waters hosting Frasassi biofilm samples. Concentration field for Lower Kane Cave (LKC) (Engel et al., 2003) is shown in gray for comparison. Symbols are colored if more than 50% of the biofilm cell area is composed of a single population or group based on FISH. Colored squares with error bars show the mean±1 s.d. for each major biofilm type. Samples analyzed by 16S rDNA cloning are circled. The open diamond symbol represents a filamentous epsilonproteobacterial biofilm from LKC reported in Engel et al. 2004. The range of conditions inside the RS05-21 (Thiothrix) biofilm based on voltammetry is indicated by the blue dashed box.
Voltammetric microelectrodes were used to investigate the spatial variability of sulfide and other redox-active species within biofilms and the surrounding bulk water. Iron species were not detected (Fe2+ and Fe3+ <5 μM, FeS <∼0.5 μM as FeS monomer), consistent with total dissolved Fe concentrations <0.1 μM measured using inductively coupled plasma mass spectrometry. Oxygen concentrations were at or below 15 μM (detection limit) for all waters analyzed, as expected based on values obtained from spectrophotometric tests at the same sites. Microelectrode profiles of sulfide concentrations within biofilms (Figure 3) are discussed below.
Vertical sulfide concentration profiles measured using voltammetric microelectrodes. (a) Beggiatoa biofilm developed in the sediment and at the sediment–water interface. Zero depth on the y axis corresponds to the upper surface of the biofilm. Dissolved oxygen concentrations were <15 μM (detection limit) for all points. Marine sediment curve is from a Beggiatoa mat described in Jorgensen and Revsbech (1983). (b) Filamentous epsilonproteobacterial mat (‘streamers’) developed in turbulent water. The biofilm (gray box) is attached at the upstream end to a rock several centimeters above the sediment–water interface. Zero depth on the y axis corresponds to the upper surface of the biofilm. Stream water flows both above and below the biofilm.
16S rDNA clone libraries
Clone libraries were constructed to investigate evolutionary relationships among the most abundant biofilm populations and to facilitate the evaluation of 16S rRNA probes. We recently described the phylogeny of clones from two Frasassi stream biofilms dominated by Beggiatoa species (Macalady et al., 2006). Four additional biofilms were selected for 16S rDNA cloning to capture a wide range of geochemical conditions and biofilm morphologies (Figure 2, cloned samples indicated by large circles). Libraries were constructed using bacteria-specific primers because FISH analyses indicated that the biofilms contained few archaea (described below). Sample FS06-12 was also cloned using universal primers in an attempt to retrieve archaeal sequences, but we only obtained bacterial sequences. Bacterial and universal primer sets yielded similar clones, with sequences most closely related to ‘Thiobacillus baregensis’ accounting for 91% in the universal library and 86% in the bacterial library.
The taxonomy of clones from each library is summarized in Figure 4 and Supplementary Table 1. Between 25 and 97% of the clones in each library were associated with known or putative sulfur-oxidizing clades within gamma-, beta- and epsilonproteobacteria (Figure 4). Gammaproteobacterial clones (Figure 5) include representatives of sulfur-oxidizing groups Beggiatoa (86–92% identity), Thiothrix (92–99% identity) and an unnamed clade containing ‘Thiobacillus baregensis’ (94–99% identity) and the recently described sulfur-oxidizing lithoautotroph Thiovirga sulfuroxydans (86–93% identity) (Ito et al., 2005). Beggiatoa clones were retrieved from four sample locations (Figure 4) and form a coherent clade most closely related to non-vacuolate, freshwater Beggiatoa strains (Ahmad et al., 2006). Most betaproteobacterial clones were related to species of the sulfur-oxidizing genera Thiobacillus (>97% identity) or Thiomonas (90–99% identity).
Distribution of 16S rDNA clones in Frasassi stream biofilms. Shaded wedges represent known or putative sulfur-oxidizing clades. Deltaproteobacteria associated with sulfate-reducing clades are shown in black. White wedges include all other clones (see Supplementary Table 1). GS02-zEL and GS02-WM clone libraries are described in Macalady et al. (2006). Clones from both bacterial and universal libraries are combined for sample FS06-12.
Neighbor-joining phylogenetic tree showing gamma(beta)proteobacteria. Frasassi clones are shown in bold followed by the number of clones represented in each phylotype. Neighbor-joining bootstrap values >50% are shown. Filled circles indicate nodes present in the maximum likelihood phylogeny. Sequences identical to the probe BEG811 are indicated by the dashed line.
Epsilonproteobacterial clones (Figure 6) were phylogenetically related to Arcobacter species or to members of the Sulfurovumales, Sulfuricurvales and 1068 groups, which have few or no cultivated representatives. The majority of the clones were associated with the Sulfurovumales clade (Figure 4) and were distantly related to cultivated strains, including the named species Sulfurovum lithotrophicum (88–94% identity). Sulfuricurvales group clones were rare and shared 96–97% identity with Sulfuricurvum kujiense. Frasassi clones in both Sulfurovumales and Sulfuricurvales were most closely related to clones from other sulfidic caves and springs (98–99% identity), including filaments from Lower Kane Cave (LKC) groups I and II (Engel et al., 2003). Arcobacter clones were diverse and only distantly related to the closest cultivated strains (91–94% identity). The 1068 group has no cultivated representatives and contains clones from deep subsurface igneous rocks, sulfidic caves and springs, groundwater and wetland plant rhizospheres. Frasassi clones associated with the 1068 group included phylotypes that shared less than 92% identity with each other, and add significantly to the known diversity within this clade. There was support in both neighbor-joining and maximum likelihood phylogenies for the placement of the 1068 group at the base of the epsilonproteobacteria (Figure 6).
Neighbor-joining phylogenetic tree showing epsilonproteobacteria. Frasassi clones are shown in bold followed by the number of clones represented in each phylotype. Neighbor-joining bootstrap values >50% are shown. Filled circles indicate nodes present in the maximum likelihood phylogeny. Clades which hybridize with probe EP404 are indicated by the dashed line.
Biofilm morphology and population structure
Twenty-eight biofilms, including those selected for 16S rDNA cloning, were homogenized and examined using epifluorescence microscopy after FISH. Probes and hybridization conditions are listed in Table 1. Probe BEG811 was designed to bind specifically to Beggiatoa populations in environmental samples from Frasassi (Macalady et al., 2006), and matches new Beggiatoa clones retrieved in this study (Figure 5). Probe EP404, targeting epsilonproteobacteria, has no mismatches with Frasassi clones from this or previous studies (n>120), with the exception of seven clones within the Arcobacter and 1068 groups (Figure 6). The EP404 probe does not match any publicly available sequences outside the epsilonproteobacteria.
FISH experiments revealed three major biofilm types, as shown in Figure 2. The dominant group in each biofilm sample accounted for more than 50% of the total DAPI cell area (Figure 2, colored symbols) with one exception (PC05-11). Sediment surface biofilms (n=15) were dominated by 5–8 μm diameter Beggiatoa filaments with abundant large sulfur inclusions and gliding motility. Streamers (n=13) were dominated either by 1.5 μm diameter gammaproteobacterial filaments with holdfasts and sulfur inclusions (Thiothrix), or by filamentous epsilonproteobacteria with holdfasts and no sulfur inclusions (1–2.5 μm diameter). Non-filamentous cells targeted by EP404 made up less than 5% of the EP404-positive cell area in each sample. As reported previously (Macalady et al., 2006), the 23S rDNA probe GAM42a produces no signal from Frasassi Beggiatoa filaments at 35% (v/v) formamide concentration. GAM42a-positive filaments with holdfasts and sulfur inclusions did not bind with probes EP404 or Delta495a. Given these observations, GAM42a-positive filaments could be assumed to be members of the Thiothrix clade. Archaeal cells in the biofilms were rare or not detected using the probe ARCH915. Consistent with this result, bacterial cell area measured using the EUBMIX probe was consistently within 15% of the area measured using the nucleic acid stain DAPI, which stains both archaeal and bacterial DNA. Representative FISH photomicrographs of the three major biofilm types are shown in Supplementary Figure 2.
Discussion
Niches of sulfur-oxidizing populations
Niche differentiation is defined as the tendency for coexisting populations to have different niches or environmental requirements. This concept is rarely explored in field studies of microorganisms but is one of the processes commonly invoked to explain the enormous complexity of natural microbial communities. This study reveals that at least three niche dimensions (sulfide, oxygen and water flow characteristics) are critical for niche differentiation among major groups of sulfur-oxidizing bacteria present in the cave waters. Filamentous epsilonproteobacteria dominate in waters with high sulfide and low oxygen, while Thiothrix dominate in waters with low sulfide and high oxygen. A similar pattern was suggested by 16S rDNA clone frequencies in a study of LKC (Engel et al., 2004), but has not been demonstrated until now. Figure 2 also shows that either sulfide or oxygen concentrations alone are poor predictors of biofilm compositions. All three of the dominant sulfur-oxidizing groups tolerate very low oxygen concentrations (<5 μM). Furthermore, Beggiatoa-dominated biofilms colonize the entire range of sulfide and oxygen concentrations measured in the cave waters, but only in locations where the shear stress caused by the flowing water is low enough to permit the accumulation of fine sediment.
The role of sulfide and oxygen concentrations in determining the composition of the biofilms is most clearly demonstrated in Figure 7, showing biofilm community composition plotted against sulfide/oxygen ratios. We observed a strong linear correlation between filamentous epsilonproteobacterial area % and the sulfide/oxygen ratio of water hosting streamers (r=0.97; P<0.0001). Correlations between epsilonproteobacterial area % and either sulfide or oxygen concentrations alone were weaker, with r values of 0.82 (P=0.002) and −0.80 (P=0.003), respectively. In sharp contrast to streamer populations, Beggiatoa filaments colonized the entire range of sulfide, oxygen and sulfide/oxygen ratios observed in the cave waters (Figures 2 and 7). Consistent with this result, Beggiatoa biofilms were observed immediately adjacent to both Thiothrix and filamentous epsilonproteobacterial streamers in situ, always in less turbulent or more slowly flowing water.
Sulfide/oxygen ratios for Frasassi biofilms analyzed using fluorescence in situ hybridization (FISH). The upper panel shows a linear correlation (P<0.0001) between sulfide/oxygen ratio and the microbial composition of streamers. The open diamond (not included in correlation) represents a biofilm from Lower Kane Cave (Engel et al., 2004) and assumes 0.2 μM O2 (detection limit). The dashed arrow shows how the sulfide/oxygen ratio for the sample would change assuming an O2 concentration of 0.1 μM. The lower panel shows the distribution of biofilm types compared based on sulfide/oxygen ratios. Colored boxes and associated bars show the mean±1 s.d. for each major biofilm type. Mean sulfide/oxygen ratios associated with Thiothrix and filamentous epsilonproteobacteria habitats are significantly different (P=0.0004).
Although changes in turbulence and water depth have the potential to alter gas exchange and therefore water chemistry, our data suggest that physical effects of water flow are more significant under the range of conditions present in the cave system. Because Beggiatoa filaments lack holdfasts, they can be washed out by flows that are too strong to allow the accumulation of fine sediment. Likewise, non-motile Thiothrix or epsilonproteobacteria filaments may become buried below the zone where oxidants are available in stream reaches that are accumulating sediment.
Our results support the idea that morphological and behavioral adaptations to physical constraints are responsible for the separate niches colonized by large, filamentous bacteria (Schulz and Jorgensen, 2001; Preisler et al., 2007). Similar to Beggiatoa described from other environments, Beggiatoa at Frasassi inhabit diffusion-controlled sediments, and can respond to changing geochemical conditions by gliding vertically in the sediment column. Sulfide concentration profiles through Beggiatoa mats reflect diffusion-controlled transport, although Frasassi sediments differ from typical marine or lacustrine sediments in that sulfide diffuses both from water above and sediment below the biofilms (Figure 3). Non-motile filaments with holdfasts (Thiothrix, filamentous epsilonproteobacteria) colonized niches with strong currents and a narrower supply ratio of turbulently mixed sulfide and oxygen (Figure 7). Interestingly, vacuolated marine Beggiatoa with holdfasts have recently been identified at cold seeps (Kalanetra et al., 2004). Frasassi clones are only distantly related (∼88% identity) to Beggiatoa species with holdfasts (Ahmad et al., 2006), and we did not observe Beggiatoa with holdfasts in any of the cave samples.
Among streamer populations, we observed a strong niche differentiation between Thiothrix and filamentous epsilonproteobacteria based on sulfide/oxygen ratios of bulk water. Microbial activity within biofilms modifies sulfide/oxygen ratios on a submillimeter scale due to oxygen consumption at the biofilm surface and sulfide production from sulfate reduction or sulfur disproportionation deeper in the biofilm. Sulfide concentrations based on microelectrode voltammetry varied up to several fold with depth in individual biofilms, typically reaching the highest values in the center. A representative sulfide concentration profile through epsilonproteobacterial biofilm sample PC06-110 is shown in Figure 3b. Sulfide concentrations within Thiothrix streamers could not be profiled due to their small size and rapid motions in the stream flow. The sulfide concentration just outside biofilm RS05-21 was approximately 200 μM, compared to 350 μM in the interior. On the basis of these values, the approximate range of conditions inside the Thiothrix biofilm are shown as a blue dashed box in Figure 2, and clearly overlap with those permitting the growth of filamentous epsilonproteobacteria. Nonetheless, Thiothrix-dominated biofilms contained at most 3.6 area % epsilonproteobacterial filaments. The lack of epsilonproteobacteria in the interior of Thiothrix biofilms with appropriate sulfide/oxygen ratios can be explained by factors such as competition for other limiting resources, or antagonistic interactions such as antibiotic production.
Both Thiothrix and filamentous epsilonproteobacteria tolerate extremely low oxygen (<3 μM) and low sulfide (<50 μM) concentrations. However, Thiothrix-dominated biofilms do not occur at sulfide concentrations above 210 μM, suggesting that sulfide toxicity may play a role in excluding them from high-sulfide environments. The absence of epsilonproteobacterial filaments in waters with oxygen concentrations above 3 μM is also striking, suggesting that oxygen toxicity may be limiting to epsilonproteobacteria. Functional genomic studies provide some evidence that epsilonproteobacteria are uniquely sensitive to oxygen among proteobacteria inhabiting sulfidic and microoxic environments due to electron transport proteins with the potential to produce millimolar levels of superoxide anions during oxidative stress (St Maurice et al., 2007). Filamentous epsilonproteobacteria are also apparently unable to store elemental sulfur intracellularly, an attribute that may limit their ability to consume toxic levels of oxygen in the absence of high sulfide concentrations. The inability to store intracellular So may also be a disadvantage in permanently or transiently low-sulfide environments such as those where Thiothrix thrive.
Filamentous epsilonproteobacteria have previously been described from LKC, Wyoming (Engel et al., 2003). Frasassi sequences differ significantly from LKC clones, and do not hybridize with previously published oligonucleotide probes LKC59 and LKC1006 targeting environmental groups (Engel et al., 2003). LKC waters have an order of magnitude lower sulfide concentrations than those hosting filamentous epsilonproteobacteria at Frasassi (Figure 2). Nonetheless, Figure 7 shows that the LKC epsilonproteobacteria colonize waters within the niche defined by high sulfide/oxygen supply ratios. As in LKC, no sulfur inclusions were observed in epsilonproteobacterial filaments, suggesting that this is a consistent physiological attribute.
Frasassi biofilms host a wide variety of other known or putatitive sulfur-oxidizing taxa as shown in Figure 4. Close relatives of ‘Thiobacillus baregensis’ are present in all clone libraries analyzed to date, sometimes comprising the majority of clones (for example, sample FS06-12). Circumstantial evidence suggests that members of this novel clade are sulfur oxidizers (Elshahed et al., 2003; Ito et al., 2005). This study is the first to our knowledge to retrieve a significant percentage of Arcobacter clones from a freshwater environment. A marine strain (Candidatus Arcobacter sulfidicus) that grows attached to solid substrates via filamentous sulfur strands in high-flow, microoxic, sulfidic environments has recently been described (Wirsen et al., 2002). Sulfide concentrations for growth of the marine Arcobacter (400–1200 μM) are broadly consistent with the Frasassi environment. Further work will be required to evaluate the ecological roles of Arcobacter and other sulfur cycling populations in Frasassi cave waters.
Accession codes
References
Ahmad A, Kalanetra KM, Nelson DC . (2006). Cultivated Beggiatoa spp. define the phylogenetic root of morphologically diverse, noncultured, vacuolate sulfur bacteria. Can J Microbiol 52: 591–598.
Amann RI . (1995). In situ identification of microorganisms by whole cell hybridization with rRNA-targeted nucleic acid probes. Mol Microb Ecol Manual 3: 1–15.
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ . (1990). Basic local alignment search tools. J Mol Bio 215: 403–410.
Bond PL, Smriga SP, Banfield JF . (2000). Phylogeny of microorganisms populating a thick, subaerial, predominantly lithotrophic biofilm at an extreme acid mine drainage site. Appl Environ Microbiol 66: 3842–3849.
Brendel PJ, Luther GW . (1995). Development of a gold amalgam voltammetric microelectrode for the determination of dissolved Fe, Mn, O2, and S(-Ii) in porewaters of marine and fresh-water sediments. Env Sci Technol 29: 751–761.
Campbell BJ, Engel AS, Porter ML, Takai K . (2006). The versatile ɛ-proteobacteria: key players in sulphidic habitats. Nature Rev Microbiol 4: 458–468.
Cole JR, Chai B, Marsh TL, Farris RJ, Wang Q, Kulam SA et al. (2003). The Ribosomal Database Project (RDP-II): previewing a new autoaligner that allows regular updates and the new prokaryotic taxonomy. Nucleic Acids Res 31: 442–443.
Daims H, Brühl A, Amann R, Schleifer K-H, Wagner M . (1999). The domain-specific probe EUB338 is insufficient for the detection of all Bacteria: development and evaluation of a more comprehensive probe set. Syst Appl Microbiol 22: 434–444.
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K et al. (2006). Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72: 5069–5072.
Druschel GK, Hamers RJ, Luther GW, Banfield JF . (2003). Kinetics and mechanism of trithionate and tetrathionate oxidation at low pH by hydroxyl radicals. Aquatic Geochem 9: 145–164.
Druschel GK, Sutka R, Emerson D, Luther GW, Kraiya C, Glazer B . (2004). Voltammetric investigation of Fe–Mn–S species in a microbially active wetland. In: Wanty RB, Seal RR (eds). Eleventh International Symposium on Water–Rock Interaction WRI-11. Balkema: Saratoga Springs, NY, USA, pp 1191–1194.
Elshahed MS, Senko JM, Najar FZ, Kenton SM, Roe BA, Dewers TA et al. (2003). Bacterial diversity and sulfur cycling in a mesophilic sulfide-rich spring. Appl Environ Microbiol 69: 5609–5621.
Engel AS, Lee N, Porter ML, Stern LA, Bennett PC, Wagner M . (2003). Filamentous ‘Epsilonproteobacteria’ dominate microbial mats from sulfidic cave springs. Appl Environ Microbiol 69: 5503–5511.
Engel AS, Porter ML, Stern LA, Quinlan S, Bennett PC . (2004). Bacterial diversity and ecosystem function of filamentous microbial mats from aphotic (cave) sulfidic springs dominated by chemolithoautotrophic ‘Epsilonproteobacteria’. FEMS Microbiol Ecol 51: 31.
Galdenzi S, Cocchioni M, Morichetti L, Amici V, Scuri S . (2007). Sulfidic groundwater chemistry in the Frasassi caves, Italy. J Cave Karst (in press).
Glazer BT, Luther GW, Konovalov SK, Friederich GE, Nuzzio DB, Trouwborst RE et al. (2006). Documenting the suboxic zone of the Black Sea via high-resolution real-time redox profiling. Deep Sea Res II 53: 1740–1755.
Glazer BT, Marsh AG, Stierhoff K, Luther GW . (2004). The dynamic response of optical oxygen sensors and voltammetric electrodes to temporal changes in dissolved oxygen concentrations. Analyt Chim Acta 518: 93–100.
Huber T, Faulkner G, Hugenholtz P . (2004). Bellerophon; a program to detect chimeric sequences in multiple sequence alignments. Bioinformatics 20: 2317–2319.
Hugenholtz P, Tyson GW, Blackall LL . (2001). Design and evaluation of 16S rRNA-targeted oligonucleotide probes for fluorescent in situ hybridization. In: de Muro MA, Rapley R (eds). Gene Probes: Principles and Protocols. Humana Press: London. pp 29–42.
Ito T, Sugita K, Yumoto I, Nodasaka Y, Okabe S . (2005). Thiovirga sulfuroxydans gen. nov., sp. nov., a chemolithoautotrophic sulfur-oxidizing bacterium isolated from a microaerobic waste-water biofilm. Int J Syst Evol Microbiol 55: 1059–1064.
Jones DS, Lyon EH, Macalady JL . (2008). Geomicrobiology of sulfidic cave biovermiculations. J Cave Karst (in press).
Jorgensen BB, Revsbech NP . (1983). Colorless sulfur bacteria, Beggiatoa spp. and Thiovulum spp., in O2 and H2S microgradients. Appl Environ Microbiol 45: 1261–1270.
Kalanetra KM, Huston SL, Nelson DC . (2004). Novel, attached, sulfur-oxidizing bacteria at shallow hydrothermal vents possess vacuoles not involved in respiratory nitrate accumulation. Appl Environ Microbiol 70: 7487–7496.
Lane DJ . (1991). 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds). Nucleic Acid Techniques in Bacterial Systematics. Wiley: New York. pp 115–175.
Ludwig W, Strunk O, Westram R, Richter L, Meier H, Yadhukumar et al. (2004). ARB: a software environment for sequence data. Nucleic Acids Res 32: 1363–1371.
Luther GW, Ferdelman TG, Kostka JE, Tsamakis EJ, Church TM . (1991). Temporal and spatial variability of reduced sulfur species (FeS2, S2O32−) and porewater parameters in salt-marsh sediments. Biogeochem 14: 57–88.
Luther GW, Glazer BT, Hohmann L, Popp JI, Taillefert M, Rozan TF et al. (2001). Sulfur speciation monitored in situ with solid state gold amalgam voltammetric microelectrodes: polysulfides as a special case in sediments, microbial mats and hydrothermal vent waters. J Environ Monitoring 3: 61–66.
Macalady JL, Jones DS, Lyon EH . (2007). Extremely acidic, pendulous microbial biofilms from the Frasassi cave system, Italy. Environ Microbiol 9: 1402–1414.
Macalady JL, Lyon EH, Koffman B, Albertson LK, Meyer K, Galdenzi S et al. (2006). Dominant microbial populations in limestone-corroding stream biofilms, Frasassi cave system, Italy. Appl Environ Microbiol 72: 5596–5609.
Manz W, Amann R, Ludwig W, Wagner M, Schleifer K-H . (1992). Phylogenetic oligodeoxynucleotide probes for the major subclasses of Proteobacteria: problems and solutions. Syst Appl Microbiol 15: 593–600.
Nielsen PH, de Muro MA, Nielsen JL . (2000). Studies on the in situ physiology of Thiothrix spp. present in activated sludge. Environ Microbiol 2: 389–398.
Preisler A, de Beer D, Lichtschlag A, Lavik G, Boetius A, Jorgensen BB . (2007). Biological and chemical sulfide oxidation in a Beggiatoa inhabited marine sediment. ISME J 1: 341.
Sarbu S, Galdenzi S, Menichetti M, Gentile G . (2000). Geology and biology of the Frasassi caves in central Italy: an ecological multi-disciplinary study of a hypogenic underground karst system. In: Wilkens H (ed). Ecosystems of the World. Elsevier: New York, NY. pp 359–378.
Schulz HN, Jorgensen BB . (2001). Big bacteria. Annu Rev Microbiol 55: 105–137.
Skoog D, Holler F, Neiman T . (1998). Principles of Instrumental Analysis, 5. Brooks/Cole–Thomas Learning. Pacific Grove: CA.
Stahl DA, Amann R . (1991). Development and application of nucleic acid probes. In: Stackebrandt E, Goodfellow M (eds). Nucleic acid techniques in bacterial systematics. John Wiley & Sons Ltd: Chichester, England, pp 205–248.
St Maurice M, Cremades N, Croxen MA, Sisson G, Sancho J, Hoffman PS . (2007). Flavodoxin:quinone reductase (FqrB): a redox partner of pyruvate:ferredoxin oxidoreductase that reversibly couples pyruvate oxidation to NADPH production in Helicobacter pylori and Campylobacter jejuni. J Bacteriol 189: 4764–4773.
Swofford DL . (2000). PAUP*: Phylogenetic Analysis Using Parsimony and Other Methods (Software). Sinauer Associates: Sunderland, MA.
Taillefert M, Bono AB, Luther GW . (2000). Reactivity of freshly formed Fe(III) in synthetic solutions and (pore)waters: voltammetric evidence of an aging process. Env Sci Technol 34: 2169–2177.
Taillefert M, Rozan TF . (2002). Electrochemical methods for the environmental analysis of trace elements biogeochemistry. In: Taillefert M, Rozan TF (eds). Environmental Electrochemistry: Analyses of Trace Element Biogeochemistry. American Chemical Society: Washington DC, pp 3–14.
Vlasceanu L, Sarbu SM, Engel AS, Kinkle BK . (2000). Acidic cave wall biofilms located in the Frasassi Gorge, Italy. Geomicrobiol J 17: 125–139.
Wirsen CO, Sievert SM, Cavanaugh CM, Molyneaux SJ, Ahmad A, Taylor LT et al. (2002). Characterization of an autotrophic sulfide-oxidizing marine Arcobacter sp. that produces filamentous sulfur. Appl Environ Microbiol 68: 316–325.
Xu K, Dexter SC, Luther GW . (1998). Voltammetric microelectrodes for biocorrosion studies. Corrosion 54: 814–823.
Acknowledgements
This paper was improved by the comments of three anonymous reviewers. We thank A Montanari for logistical support and the use of facilities and laboratory space at the Osservatorio Geologico di Coldigioco (Italy) and S Mariani, S Galdenzi and S Cerioni for expert advice and field assitance. We also thank P D’Eugenio, M Mainiero, S Recanatini, R Hegemann, H Albrecht, K Freeman and R Grymes for assistance in the field. We thank B Thomas and J Moore for water analyses, and L Albertson and T Stoffer for laboratory assistance. E Fleming provided valuable comments on the paper. DE contributed to this research as an undergraduate student and was supported in 2006 by a Barrett Foundation scholarship. This work was supported by grants to JLM from the Biogeosciences Program of the National Science Foundation (EAR 0311854 and EAR 0527046) and NASA NAI (NNA04CC06A). GKD acknowledges support from the American Chemical Society Petroleum Research Fund (43356-GB2) and NSF-EPSCoR-VT (EPS 0236976).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on The ISME Journal website (http://www.nature.com/ismej)
Rights and permissions
About this article
Cite this article
Macalady, J., Dattagupta, S., Schaperdoth, I. et al. Niche differentiation among sulfur-oxidizing bacterial populations in cave waters. ISME J 2, 590–601 (2008). https://doi.org/10.1038/ismej.2008.25
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2008.25
Keywords
This article is cited by
-
Thiothrix and Sulfurovum genera dominate bacterial mats in Slovak cold sulfur springs
Environmental Microbiome (2023)
-
Thiobacillus as a key player for biofilm formation in oligotrophic groundwaters of the Fennoscandian Shield
npj Biofilms and Microbiomes (2023)
-
Dissecting Light Sensing and Metabolic Pathways on the Millimeter Scale in High-Altitude Modern Stromatolites
Microbial Ecology (2023)
-
Nutrient-limited subarctic caves harbour more diverse and complex bacterial communities than their surface soil
Environmental Microbiome (2022)
-
Global patterns of diversity and metabolism of microbial communities in deep-sea hydrothermal vent deposits
Microbiome (2022)