Abstract
Pseudomonas syringae is a plant pathogen well known for its capacity to grow epiphytically on diverse plants and for its ice-nucleation activity. The ensemble of its known biology and ecology led us to postulate that this bacterium is also present in non-agricultural habitats, particularly those associated with water. Here, we report the abundance of P. syringae in rain, snow, alpine streams and lakes and in wild plants, in addition to the previously reported abundance in epilithic biofilms. Each of these substrates harbored strains that corresponded to P. syringae in terms of biochemical traits, pathogenicity and pathogenicity-related factors and that were ice-nucleation active. Phylogenetic comparisons of sequences of four housekeeping genes of the non-agricultural strains with strains of P. syringae from disease epidemics confirmed their identity as P. syringae. Moreover, strains belonging to the same clonal lineage were isolated from snow, irrigation water and a diseased crop plant. Our data suggest that the different substrates harboring P. syringae modify the structure of the associated populations. Here, we propose a comprehensive life cycle for P. syringae—in agricultural and non-agricultural habitats—driven by the environmental cycle of water. This cycle opens the opportunity to evaluate the importance of non-agricultural habitats in the evolution of a plant pathogen and the emergence of virulence. The ice-nucleation activity of all strains from snow, unlike from other substrates, strongly suggests that P. syringae plays an active role in the water cycle as an ice nucleus in clouds.
Similar content being viewed by others
Introduction
The prevailing concepts about the life histories or life cycles of plant and animal pathogens have significant practical consequences for controlling or avoiding epidemics. Medical microbiologists long ago recognized that human pathogens can also reside and flourish in nonhost habitats, in which they can evolve and diversify independent of their activity as pathogens. This has led to the modern practices of water sanitation, surveillance of wild-animal populations, cleaning and maintenance of ventilation systems in buildings, limited entry of floral bouquets into hospitals etc. Plant pathologists have tended to focus more restrictively in defining the life cycles of plant pathogens principally by their agricultural contexts sensu stricto and in terms of the availability and response of host plants. Nonetheless, agricultural systems are open and most plant pathogens are not obligate parasites, suggesting that they might well be able to disperse and thrive elsewhere, yet the parts of their life cycles involving non-agricultural contexts have largely been ignored.
Pseudomonas syringae is an archetype among the plant pathogenic bacteria. Its biology, ecology and genetics have been studied extensively, but these studies are done almost exclusively in the context of the relationship of this bacterium with plants and agricultural habitats (Hirano and Upper, 2000; Sarkar and Guttman, 2004). Recently, strains of P. syringae, virulent on diverse species of crop plants, were isolated from epilithic biofilms from rivers in pristine regions outside the zones of agricultural production (Morris et al., 2007). This is one of the several observations suggesting that this plant pathogen is widely spread in non-agricultural niches. P. syringae has a strong net upward flux from plant canopies into the atmosphere (Lindemann et al., 1982), and there are several reports of isolation of P. syringae from clouds at several kilometers altitude (Sands et al., 1982; Amato et al., 2007). From clouds, P. syringae could eventually fall out with rain or snow to a variety of environments. Nevertheless, reports of this bacterium on wild plants have focused on weed or non-host plants in agricultural regions. P. syringae reported on thistle, oak, black locust (Lindemann et al., 1984), Arabidopsis thaliana (Jakob et al., 2002) and wild cherry (Vincente et al., 2004) were from plants in agricultural fields, woodland plantations or nurseries. Although we suspect that P. syringae is widely spread in the environment, the extent to which it is present in niches outside of agricultural contexts has not been established clearly.
In this work, we sought to find P. syringae in pristine habitats, which it might attain primarily via fall-out with rain and snow. Substrates investigated from these habitats included lake and stream water at altitudes above cultivated zones, snow pack or firn snow melting into these lakes and streams, and the nearby flora. We also examined rain from semirural zones and freshly fallen snow from diverse locations as indirect evidence of P. syringae falling out with precipitation. To avoid collecting P. syringae scrubbed from aerosols originating from nearby plant canopies, we sampled under conditions where population sizes of local air-borne P. syringae were likely to be low, such as in mid or late winter or above cement or gravelled areas. We also took sample plants in forests or prairies at the border of national parks, in private properties and other areas that had never been under cultivation.
Here, we present data on the densities of populations of P. syringae in a range of non-agricultural substrates and evidence confirming that these strains are bona fide P. syringae based on genotypic and phenotypic properties, including pathogenicity. Strains isolated from snow, irrigation water and a diseased crop that belonged to the same clonal lineage were identified in this work, thereby providing evidence for a link between agricultural and non-agricultural habitats. These results led us to propose an expanded life cycle or life history for this bacterium to encompass its dissemination via the water cycle and subsequent colonization of both agricultural and non-agricultural habitats. These findings are novel for a plant pathogen and are significant from the point of view of disease epidemiology, breeding for disease resistance and disease management practices such as quarantine and chemical and biological control.
Materials and methods
Isolation of putative P. syringae from environmental sources
The geographic origins and substrates from which P. syringae was isolated are presented in Table 1. All samples were collected in sterilized or disinfested containers, were kept in a cooler during transport and were stored in a refrigerator prior to processing. The methods used to process the different substrates depended on the nature of the substrate. In all cases, aliquots of the processed material were plated on a selective medium, as described below. Except for some of the wild-plant samples collected in the early part of this study, methods were quantitative, and also the number of P. syringae-like bacteria per gram of tissue, per leaf or per milliliter of water was determined.
For plants, several leaves from the same plant or from the same stand of plants were pooled, weighed and macerated in a stomacher (BagMixer, Interscience, St Nom-La-Breteche, France) for 1.5–3 min in 0.1 M phosphate buffer (8.75 g K2HPO4 and 6.75 g KH2PO4 per liter, pH 6.8). The volume of buffer used depended on the mass of tissue processed. All plants collected were apparently healthy.
Snow was collected where it had accumulated to at least several centimeters depth (to avoid snow that was in contact with the ground or with plants) and where there were no animal or human tracks. Some of the snow had fallen only a few hours before sampling. The top 1 cm was brushed away and the underlying snow was collected with sterile flour scoops and put into clean plastic bags. Enough snow was collected for each sample to render at least 1.5 l of melt water. The melt water was mixed thoroughly and 1 l was filtered across a sterile nitrous cellulose filter (pore diameter 0.22 μm). The filter was recovered and agitated with a sterile stir bar for about 3 min in 10 ml of the melt water filtrate. The 100-fold concentrated melt water was then transferred to a sterile plastic container. An aliquot of the crude melt water was also conserved.
Water from lakes and streams was collected in bottles at several centimetres below the surface. Volumes of 1.5–3 l were collected at each site. The water was mixed well and then filter-concentrated in the same manner as that for snow-melt water. The 100-fold concentrated samples and aliquots of the crude water were transferred to sterile plastic containers.
At the onset of storms, rainwater was collected in large plastic buckets (volume of 30 l or greater) lined with clean plastic bags. The buckets were placed in open areas to avoid run-off or splashing from plants and were placed on concrete surfaces or at sufficient heights (1.5–2 m) to avoid splashing from the ground. The rainwater was transferred to a sterile container and then filter-concentrated as described above. When at least 500 ml of rainwater was collected, the water was concentrated by a factor of 100. Otherwise, the water was concentrated by a factor of 10. Aliquots of the concentrated and the crude rainwater were stored into sterile plastic containers.
Methods for the collection and processing of epilithic biofilms are described by Morris et al. (2007). Homogenates of these biofilms were transferred to sterile plastic containers.
Processed samples were dilution-plated on KBC medium (Mohan and Schaad, 1987), a modification of King's medium B (King et al., 1954) (KB) supplemented with cephalexin and boric acid. We have previously used this medium for the detection of low densities of P. syringae in irrigation water retention basins (Riffaud and Morris, 2002). Some samples were also plated on 10% tryptic soy agar (per liter: 3 g tryptic soy broth (Difco, Detroit, MI, USA), 15 g agar) to estimate the total background mesophilic bacterial flora. Plates were incubated for up to 5 days at 22–25 °C. Production of fluorescent pigment on KBC was checked at 2 days after incubation, whereas definitive counts of fluorescent and total colonies were realized after 5 days of incubation. Fluorescent colonies were transferred to KB medium and the absence of cytochrome c oxidase was verified by streaking some of the 48 h culture with a sterile toothpick on filter paper imbibed with a fresh solution of 1% tetramethyl-ρ-phenylenediamine dihydrochloride (Gerhardt et al., 1981). Colonies lacking the cytochrome oxidase were purified and tested for their capacity to induce a hypersensitive reaction in tobacco (described below). Strains inducing this reaction were stored in 0.1 M phosphate buffer at 4 °C and in 40% glycerol at −80 °C for further characterization. A total of 140 strains were maintained for this study (Table 2). An additional 18 strains from epilithic biofilms previously described (Morris et al., 2007) were also included (strains originating from samples E-03 to E-06; Table 2).
Other bacterial strains
For comparison of phenotypes and genotypes of strains isolated in this study with other strains, we used the reference strains of P. syringae for which the genome has been sequenced (B728a pv. syringae, DC3000 pv. tomato and 1448A pv. phaseolicola) (Pseudomonas syringae Genome Resources: http://pseudomonas-syringae.org/home.html). Other strains of P. syringae from agricultural sources included the following strains: CC0001, CC0023, CC0024, CC0037, CC0094, CC0125, CC0206, CC0301, CC0354 and CC0440 from diseased cantaloupe in France and Morocco (Morris et al., 2000); CC0393, CC0403, CC0406 and CC0412 from water retention basins used for irrigation in southwestern France (Riffaud and Morris, 2002); and reference strains from the Collection Française de Bactéries Phytopathogènes (INRA-Angers, France) (P. syringae pv. syringae-type strain CFBP 1392, P. syringae pv. aptata strain CFBP 1906 and P. syringae pv. atrofaciens strain CFBP 2256). P. viridiflava strain PV612 isolated from diseased cantaloupe in Crete (Goumas and Chatzaki, 1998) was kindly provided by D Goumas. Other strains that were used simply to test the efficiency of the isolation medium are presented in Table 3.
Evaluation of host range and factors related to pathogenicity
The capacity of strains to induce a hypersensitive response was determined in tobacco by infiltrating fully developed leaves of plants of Nicotiania tobacum L. cv. Samsun at the 10-leaf stage (bacterial suspensions of 48 h cultures at approximately 1 × 108 CFU ml–1). Strains that induced the hypersensitive response within 48 h of incubation were further characterized for biochemical properties, production of syringomycin and virulence on three plant species.
Virulence of strains was determined on sugar beet (Beta vulgaris var. rapa L. cv. Sucrière), lettuce (Lactuca sativa L. cv. Mantila) and cantaloupe (Cucumis melo var. cantalupensis Naud. cv. Védrantais). For each strain, five plants of each species were inoculated at the two-leaf stage with a 50 μl aliquot of inoculum. The inoculum consisted of aqueous suspensions of 48-h bacterial cultures from KB plates adjusted to an A580 of 0.06 (∼ 3 × 108 CFU ml–1). The inoculum was infiltrated into the leaf blade of the oldest leaf near the junction with the petiole. Plants were incubated for 7 days in the greenhouse at ambient conditions of 17–25 °C. Strain CC0094 of P. syringae, pathogenic to all three plant species (Morris et al., 2000), was used as a positive control for all inoculation series. Sterile distilled water was used as a negative control. Symptoms were observed daily, and the reaction of the plant was scored at 7 days after inoculation as 0 (no obvious symptoms), 1 (hypersensitive-like reaction restricted to the point of inoculation), 2 (slight expansion of a necrotic zone of tissue away from the point of inoculation and/or necrosis and breaking of the petiole) or 3 (expansion of a necrotic zone of tissue away from the point of inoculation, leading to wilting or death of the entire leaf or the entire plant). Bacteria were re-isolated from selected plants to confirm that symptoms were caused by the inoculum.
Production of syringomycin was evaluated on SRM medium (Gross, 1985), according to the bioassay based on the sensitivity of Geotrichum candidum (Gross and DeVay, 1977). The distance between the border of the bacterial colony and the edge of the fungal lawn (zone of inhibition) was measured after 3 and 6 days of incubation.
Characterization of biochemical traits and ice-nucleation activity of strains
The presence of arginine dihydrolase, gelatinase and levan sucrase, the reduction of nitrate and the hydrolysis of esculin were tested as previously described by Lelliot et al. (1966). Ice-nucleation activity at −2 to −6 °C was determined for drops of aqueous bacterial suspensions containing a total of 107 cells as described previously (Morris et al., 2007).
Genotypic characterization of strains
Sequences of four genes of the core genome, rpoD, gyrB, cts and gapA were compared among strains isolated here and with well-described type strains of P. syringae pathovars. PCR amplifications were performed on aqueous suspensions of whole cells adjusted to 2 × 108 CFU ml−1 (OD580=0.12), from 48-h cultures on KB medium with the primers described by Yamamoto et al. (2000) for rpoD and gyrB and by Sarkar and Guttman (2004) for cts and gapA, and with reagents from the Qiagen multiplex PCR kit (Qiagen, Courtaboeuf, France) in DNA-Engine PTC200 thermal cyclers (MJ Research, Waltham, MA, USA). For each strain, 4 μl of bacterial suspension were added to 50 μl of reaction mixtures containing 25 μl of the kit's master mix, 160 pg of each of the forward and reverse primers (final concentration of 0.2 μM of each primer), 5 μl of the kit's Q-solution and sterile distilled water. For rpoD, 40 cycles of amplification were performed with template denaturation at 94 °C for 30 s, followed by annealing for 90 s at 63 °C for rpoD and gyrB and at 62 °C for cts and gapA, and extension at 72 °C for 1 min with the final extension lasting for 10 min. For gyrB, initial 20 cycles of amplification were performed with template denaturation at 94 °C for 30 s, while annealing started at 53 °C for 1 min and increased by 0.5 °C with every cycle for a final annealing temperature at 63 °C and extension at 72 °C for 2 min. This was followed by 25 additional cycles of amplification with denaturation at 94 °C for 30 s, annealing at 58 °C for 90 s and extension at 72 °C for 2 min with the final extension lasting for 7 min. For amplification of both genes, the initial cycle was preceded by 15 min of activation of the Taq polymerase at 96 °C.
Amplified products were electrophoresed on 2% agarose for 25 min at 80 V and purified by using the QIAquick kit (Qiagen), as per the manufacturer's instructions. Forward and reverse sequencing was conducted by Génome Express (Meylan, France) with the sequencing primers described previously (Yamamoto et al., 2000; Sarkar and Guttman, 2004).
Phylogenetic analysis
The DNA sequences obtained from the PCR products corresponding to the rpoD, gyrB, cts and gapA gene fragments of our strains were aligned and cut to the same size as that of the corresponding gene sequences deposited by Sarkar and Guttman (2004) in GenBank for strains 601, 301765, YM7902, N6801, FTRS _U7805, H5E1, K93001, KOZ8101, 301702, KN203, 301020, 6606, 302941, KN221, I_6 and 36_1. These strains represent the four genomic groups identified within P. syringae by Sarkar and Guttman (2004) based on an MLST analysis of seven gene fragments. Sequences for strains B728a, DC3000 and 1448A of P. syringae, for PAO-1 of P. aeruginosa and for Pf05 and Pf01 of P. fluorescens were obtained from GenBank and also cut to the same size. Seqman and Megalign (Lasergene, DNASTAR, Madison, WI, USA) were used for this step. The four fragments were concatenated and strains with identical sequences were identified using the nonredundant database program at http://pubmlst.org/cgi-bin/mlstanalyse/mlstanalyse.pl?site=pubmlst&page=nrdb&referer=usmirror1.pubmlst.org. The nonredundant alleles were then used to construct a Bayesian tree in MrBayes 3.1.2 (Huelsenbeck and Ronquist, 2001; Ronquist and Huelsenbeck, 2003), which uses the Markov chain Monte-Carlo (MCMC) method. The program was run for 2 600 000 generations with sample frequency of 10 and the s.d. of split frequencies was stabilized below 0.006. When summarizing the substitution model parameters and trees, 65 000 samples, representing the first 25% of all samples, were used for burn-in, that is, they were discarded to achieve an accurate estimate of the posterior probability distribution in the summary tree. All potential scale reduction factor (PSRF) values were close to 1.0. Two independent runs converged on the same tree.
Evaluation of selectivity and efficiency of the isolation medium
The relative efficiency of the isolation medium KBC for strains of P. syringae from the three different genomic groups proposed by Sawada et al. (1999) was compared to that of KB. Aqueous suspensions from 48-h cultures (25 °C on KB medium) of strains listed in Table 3 were dilution plated on both KB and KBC in three replicates per dilution. Total colonies were counted for up to 4 days of incubation at 25 °C.
Results
The strains isolated from non-agricultural substrates have properties consistent with those of pathogenic P. syringae from agricultural sources. All strains described here from non-agricultural substrates were fluorescent on KBC medium, they neither have cytochrome oxidase nor arginine dihydrolase, they hydrolyzed esculin and induced a hypersensitive reaction in tobacco. Furthermore, phylogenetic analyses based on sequences of gene fragments of the conserved housekeeping genes rpoD, gyrB, cts and gapA for a subset of the non-agricultural strains and known reference strains strongly support that strains from non-agricultural habitats are bona fide P. syringae (Figure 1). Hence, in the presentation of the results below, the non-agricultural strains will be referred to as P. syringae.
Phylogenetic tree constructed by Bayesian inference on the basis of the concatenated sequence of the four housekeeping gene fragments gapA, cts, gyrB and rpoD. The tree was rooted on P. aeruginosa PAO1. Branches that were shortened to fit the figure are indicated with /1/ and /2/. Posterior probabilities are indicated as percentages. The substrate of isolation for all P. syringae strains is indicated. In the case of strains with identical sequences, the source of the strain present in the tree is indicated followed by the source of any other identical strains (and by an indication of the number of identical strains from the same origin if there are more than one). There were six groups of strains with identical sequences for all four genes: (1) CFBP 1392 and 601; (2) KOZ8101 and 6606; (3) CC0037, CC0354, CC0440; (4) CC0094, CC0393, CC0403, CC0406, CC1475, CC1497 and CC1498; (5) CC0658 and CC0659 and (6) CC1448, CC1449, CC1450.
P. syringae is abundant in non-agricultural habitats
P. syringae was isolated from all substrates listed in Table 1. These included rain, snow, mountain lakes and streams, apparently healthy wild plants and epilithic biofilms. Many of these substrates were obtained in pristine areas above 2000 m altitude that could only be accessed on foot and were well above zones of agricultural production. In rain, in snow-melt water and in lakes and streams, P. syringae was present at concentrations generally between hundreds to several thousands bacteria per liter. However, in some cases, concentrations were as high as 104–105 bacteria per liter. Likewise, in association with wild plants, population sizes of P. syringae were several hundreds to 105 bacteria per gram or per leaf. P. syringae was not always found in the substrates analyzed. Nevertheless, with the exception of fresh snow, it was detected in over half of the samples of each type of substrate analyzed (Table 4).
P. syringae strains from non-agricultural substrates are pathogenic and ice-nucleation active; the frequency of these traits varies with substrate
A large proportion of the strains of P. syringae from non-agricultural sources was virulent on one or more of the hosts tested here (Table 2, Figure 2). Over 45% consistently induced compatible reactions on at least one of the three plant species tested, whereas 13% were virulent on all three species. Half of the strains also produced toxins (considered to be syringomycin-like) that inhibited G. candidum. The majority of strains (85%) were also ice-nucleation active. The proportion of strains with any particular phenotype, particularly ice-nucleation activity and toxin production, varied among the different substrates. For substrates from which at least 10 strains were characterized (wild plants, snow, water and epilithic biofilms), pair-wise, two-tailed Fisher's exact tests were used to determine whether the differences in proportions were significant. No significant differences (P⩽0.05) were observed in the frequency of strains virulent on at least one of the host plants tested among the different substrates (Figure 2). There were marked differences among substrates in the frequency of strains virulent on all three plant species tested (15% from wild plants, 6% from snow, 23% from water and 8% from epilithon), but these differences were not significant at P⩽0.05. However, for P⩽0.1, the frequency of strains virulent for all three species was significantly lower for snow strains than for water strains. Epilithic strains produced syringomycin less often than strains from snow (P=0.0004) and from wild plants (P=0.02). All strains from snow were ice-nucleation active, and this was significantly more frequent in strains from wild plants (P=0.0004) and from epilithon (P=0.0000) (Figure 2).
Frequency of ice-nucleation activity, of syringomycin-like toxin production and virulence in at least one or to all three host plants tested among strains from four non-agricultural substrates. Frequencies were calculated from results presented in Table 2. For syringomycin production, frequencies presented here concern strains with zones of inhibition ⩾2 mm. For virulence, frequencies were calculated in terms of strains producing compatible reactions on at least half of the plants tested per species (black squares in Table 2). The total number of strains characterized for each substrate is listed in parentheses below the substrate name. For each trait, the values represented by bars marked with a same letter are not significantly different (P⩽0.05) according to a two-tailed Fisher's exact test. The Fisher's test was conducted for all pair-wise comparisons of substrates. For virulence, there were no significant differences.
P. syringae strains from non-agricultural habitats are genetically very similar and in some cases indistinguishable from strains isolated from disease epidemics
Phylogenetic analyses were performed on the concatenated sequences of gene fragments from four housekeeping genes: rpoD (coding for sigma factor 70); gyrB (coding for DNA gyrase B); cts (also called gltA, coding for citrate synthase); and gapA (coding for glyceraldehyde-3-phosphate dehydrogenase). These four gene fragments (total length 1850 bp) were found to give reliable evolutionary relationships within the P. syringae species when compared with the results obtained with seven gene fragments (Hwang et al., 2005).
Bayesain inference was used to build a phylogenetic tree (Figure 1). The tree was rooted on P. aeruginosa PAO1, which has a DNA identity of only approximately 65% with our strains and thus represents a sufficiently divergent outgroup. Several P. syringae strains representing the diversity within the species, a strain of P. viridiflava (the species most closely related to P. syringae) and two P. fluorescens strains were also included in the tree to precisely determine the taxonomic position of our isolates.
The Bayesian tree reveals that all but one of our isolates fall into the same branch that also includes the type strain of P. syringae (CFBP1392) and the sequenced P. syringae pv. syringae strain B728a. This branch has a posterior probability of 99%, which indicates a very strong statistical support for this branch. DNA identity between isolates on this branch is at least 97% for any of the gene fragments, indicating that all isolates on this branch are very closely related (see also Table 5). Moreover, this branch is part of a larger cluster with 100% support that includes exclusively P. syringae reference strains. The isolate CC1427 is part of this larger cluster. More precisely, CC1427 is located on a branch with a posterior probability of 100%, together with the sequenced DC3000 strain isolated from tomato and three other pathogenic P. syringae strains isolated from crops. The closest relative of P. syringae, P. viridiflava 612, is located outside of the P. syringae cluster. Therefore, phylogenetic analysis using Bayesian inference very strongly supports the conclusion that all of our isolates are members of the P. syringae species.
Even if our isolates cluster with pathogenic P. syringae strains isolated from crops, they could still form their own separate subclusters composed of strains that adapted to life in non-agricultural habitats. However, most of our strains do not cluster away from crop isolates. Most revealing is the cluster containing CC0094, CC0658 and CC0301 (Figure 1). The strains CC0658 and CC0659 (with identical sequence) were isolated from a wild primula plant and CC0301 was isolated from cultivated cantaloupe, while the strains identical in all four gene fragments to the cantaloupe strain CC0094 were isolated from irrigation water and from freshly fallen snow, which strongly suggests that CC0094-like strains cycle through all three substrates.
The isolation medium permits isolation of strains representing all genomic groups of P. syringae but does not allow recovery of all strains at the same rate of efficiency
To evaluate the potential bias of the isolation medium for the types of P. syringae strains recovered from non-agricultural habitats, we determined the plating efficiency of the selective isolation medium KBC compared with KB medium on which all strains tested here can grow. KBC had a varying degree of selectivity for reference strains compared to KB. Percent recovery on KBC of the reference strains varied between just above 0% to over 100% relative to KB medium (Table 3). All of the three genomic groups identified by Sawada et al. (1999) had strains that could grow at recovery rates of over 50% of the total number of cells introduced onto the medium. On the other hand, numerous strains in each group were recovered at much lower rates. Only about 10% of the cells of DC3000 and 1448A that could grow on KB were recovered on KBC, whereas about 80% of the cells of B728a that grew on KB were recovered on KBC.
Discussion
Our results clearly demonstrate that plant pathogenic strains of P. syringae are ubiquitous in a wide range of non-agricultural substrates. Furthermore, we have also identified strains belonging to a same clonal lineage from both agricultural and non-agricultural substrates. Many of the non-agricultural substrates are non-plant substrates associated with alpine habitats or with the water cycle including snow, mountain lakes and streams, epilithic biofilms and rain. All species of wild plants sampled in this study also harbored pathogenic strains of P. syringae. The plant species examined here are all found in alpine and sub alpine zones. The Primulaceae, in particular, are also typically found in humid soils. Many of the samples collected here bordered lakes and streams. Silene acaulis forms dense mats on rocks without direct contact with the soil. Nevertheless, because of the altitude at which this perennial plant grows, it is likely to be repeatedly exposed to snow during its life.
Coupled to the current understanding of the ecology of P. syringae in agricultural systems and reports of its dissemination, our results lead us to suggest the life history or life cycle for P. syringae described in detail in Figure 3. This cycle was based on an initial proposition of Sands et al. (1982), suggested 25 years ago, for which we now provide significant corroborative evidence. We suggest that the water cycle is an important driver for dissemination of P. syringae, transporting it to a variety of permissive ecological niches, in addition to plants, in which it can survive. The high frequency (100%) of ice-nucleation active strains in snow samples relative to samples from other substrates corroborates the suggestion of Sands et al. (1982) that the ice-nucleation activity of P. syringae plays a role in this cycle. Nevertheless, this question will need to be further evaluated by an interdisciplinary approach combining microbiology, meteorology and atmosphere physics.
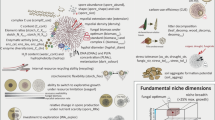
Hypothetical life cycle of P. syringae. Cells of this bacterium in clouds (Sands et al., 1982; Amato et al., 2007) and those scrubbed from the atmosphere are deposited with rain and snow. P. syringae is stocked in snow, but the duration of survival likely depends on environmental conditions and on the biotype of the bacterium. Rain and melting snow containing P. syringae feed into lakes and streams. These waters might be habitats for proliferation in the bulk water and/or in epilithic biofilms (Morris et al., 2007); streams are also vectors of dissemination. Water run-off from melting snow, rain, and streams and lakes transport P. syringae to wild plants and also wash bacteria from these plants and disseminate them elsewhere. These various niches harboring P. syringae (wild plants, snow, water and epilithic biofilms) each offer different selective pressures that likely have differential effects on the various biotypes of the P. syringae population. Some of these niches might contribute to the diminution of certain biotypes of P. syringae, whereas others might foster multiplication. Percolation of water underground as part of its trajectory down mountains will expose P. syringae to environmental conditions possibly differing in chemical concentrations, oxygen tension and other physical–chemical parameters compared to surface waters, thereby adding on additional selective pressures. Use of river waters for irrigation transports P. syringae to cultivated plants and to agricultural equipment. P. syringae will also come into contact with cultivated plants via rain or via irrigation systems based on collection of rain water (Riffaud and Morris, 2002) via aerial dissemination and via the other vectors previously described in studies of this bacterium in agricultural contexts. Certain biotypes of P. syringae might be better adapted than others to the stresses encountered in agricultural systems. P. syringae in the epiphytic phase on plants (wild or cultivated) are taken up by aerosols and transported to clouds in which they could be transported horizontally.
Water, in particular flowing water, has clearly been shown to harbor several species of plant pathogenic bacteria. For example, Ralstonia solanacearum can survive in aquatic habitats in association with plant roots (Elphinstone et al., 1998; Wenneker et al., 1999). Analysis of DNA from lake sediments in copper-mining regions also revealed the presence of R. solanacearum-like bacteria (Konstantinidis et al., 2003). Dickeya chrysanthemi (formerly Erwinia chrysanthemi) is thought to be a part of the indigenous microorganisms on weeds and in sediments in alpine streams in Australia (Cother and Gilbert, 1990). Numerous studies illustrated that Pectobacterium carotovorum (formerly E. carotovora) is present in a wide range of surface and underground waters including those used in irrigation (McCarter-Zorner et al., 1984; Harrison et al., 1987). Franc (1988) further revealed that this bacterium was present in at least 80% of samples of ocean water and rainwater collected on the west coast of the United States and in samples of aerosols along the coast. It was also present in about 5% of snow samples collected in the Rocky Mountains where precipitation occurs from air masses originating over the Pacific west coast of the United States. This led him to propose that P. carotovorum in ocean waters is taken up by aerosols, is transported inland by clouds and falls out inland with precipitation. This cycle also included the proposition that P. carotovorum enhanced cloud formation thereby enhancing rainfall due to the activity of its cells as cloud condensation nuclei (Franc and DeMott, 1998). In spite of the well-supported proposition of Franc and the other hypotheses concerning the role of upward flux in aerosols, of precipitation and of water flow in the long-distance dispersal of bacterial plant pathogens, plant pathology has remained relatively agrocentric in its consideration of the life cycles of pathogens. As far as we are aware, none of the life cycles of plant pathogenic bacteria presented in plant pathology textbooks and reviews takes into account the potential non-agricultural reservoirs to any significant extent. Here, we present a clearly illustrated hypothetical life cycle with the intent to promote further studies to validate and enrich our hypothesis for P. syringae, as well as to encourage new hypotheses for other plant pathogens.
The overall abundance of the different genotypes and phenotypes of P. syringae described in this study was likely influenced by the isolation medium used. Strains similar to B728a were the most abundant in the collection of non-agricultural strains examined here. This might be because bacteria related to B728a were more efficiently isolated on KBC medium than were those related to DC3000 and 1448A or because B728-like strains are in fact more abundant in non-agricultural substrates than DC3000-like and 1448A-like strains. Super-selectivity is a classic dilemma associated with the use of culture media in studies of population structure. Nevertheless, without this selective medium, which leads to a reduction of a factor of up to 10−4 of the background microflora (Riffaud and Morris, 2002), it would have been impossible to isolate P. syringae from certain substrates. Furthermore, the bias of KBC medium is likely identical for all substrates. Hence, although the absolute population structure of each substrate is likely biased, we feel that our observations of comparative trends in population structure are not significantly influenced by the isolation medium. The relative abundance of 1448A-like and DC3000-like strains compared to B728-like strains in the analyzed substrates will need to be determined in future studies possibly taking advantage of culture-independent metagenomic approaches (Committee on Metagenomics, 2007).
The substrates in which P. syringae was detected might simply offer means of survival and accumulation of bacteria immigrating with water flow. However, our data suggest that selection pressures associated with the different substrates lead to different biotype frequencies. This may be the result of differential growth and survival in these substrates or due to the need for certain phenotypes to attain access to the substrate. For example, P. syringae might need to be active as an ice nucleus to become incorporated into snow (Morris et al., 2004). This could explain why all strains from snow were ice-nucleation active, whereas significantly fewer strains from other sources were active as ice nuclei. Likewise, epilithic biofilms might offer a protective niche for strains that are not armed with syringomycin or related toxins, thereby allowing for a greater abundance of these deficient strains relative to other substrates. The different substrates also harbored varying frequencies of strains of P. syringae-like bacteria that did not induce HR in tobacco (data not presented here). The trends observed here should be confirmed with more extensive and completely randomized sampling of these substrates. Nevertheless, they suggest that non-plant substrates can play a role in structuring populations of P. syringae and therefore might be veritable habitats.
Our observations for P. syringae and those of Franc and co-workers concerning P. carotovorum evoke questions regarding sources and sinks in the flow of pathogens with the water cycle. Both of these plant pathogens can develop large populations when associated with plants, especially when plants become diseased. Based on current understanding of their abundance and population dynamics in agro-ecosystems, it is reasonable to propose that strains in non-agricultural habitats originate from plants, enter the water cycle via aerosol formation and then return to agricultural systems via rainfall or the use of irrigation waters containing these bacteria. It is currently unknown whether these pathogens proliferate to any significant extent in non-agricultural habitats, non-plant substrates in particular, and whether they can, in fact, be the sources of pathogens for epidemics. These features of the life cycle of these plant pathogens merit investigation.
Due to the current agrocentric view of the life cycles of plant pathogens, interest in the forces structuring populations of pathogens such as P. syringae has focused on cultivated plants (Hirano and Upper, 2000; Sarkar and Guttman, 2004; Guttman et al., 2006). Our results open the door to studies of the comparative roles of a range of niches in the evolution of P. syringae, in the emergence of various phenotypes and in the organization of its genome. Such studies will likely lead to re-evaluation of the importance of coevolution of P. syringae with plants and the impact of plants on the emergence of pathotypes. The abundance of this bacterium in diverse substrates provides the opportunity to formulate hypotheses on the role of these substrates in driving the evolution of P. syringae—the body of alternative hypotheses needed for a rigorous evaluation of the notion of coevolution. The life history that we propose for P. syringae is also a rich context for testing more recent hypotheses concerning the emergence of pathotypes. In particular, Arnold et al. (2007) have suggested that adaptation of bacteria to an ensemble of multiple stresses leads to the acquisition of virulence factors and to the emergence of pathogenic variants. Mechanisms of adaptation to varied niches or habitats might shed light on the enigma of broad host range. The development of cultural practices and plant breeding strategies for effective disease management critically relies on an understanding of the environmental factors to which plant pathogens adapt and on an overall comprehension of their ecology both within and outside of agro-ecosystems.
References
Amato P, Parazols M, Sancelme M, Paolo L, Mailhot G, Delort A-M . (2007). Microorganisms isolated from the water phase of tropospheric clouds at the Puy de Dôme: major groups and growth abilities at low temperatures. FEMS Microbiol Ecol 59: 242–254.
Arnold DL, Jackson RW, Waterfield NR, Mansfield J . (2007). Evolution of microbial virulence: the benefits of stress. Trends Genet 23: 293–300.
Committee on Metagenomics: Challenges and Functional Applications (2007). The New Science of Metagenomics: Revealing the Secrets of Our Microbial Planet. National Academies Press: Washington, DC.
Cother EJ, Gilbert RL . (1990). Presence of Erwinia chrysanthemi in two major river systems and their alpine sources in Australia. J Appl Bacteriol 69: 729–738.
Elphinstone JG, Stanford H, Stead DE . (1998). Survival and transmission of Ralstonia solanacearum in aquatic plants of Solanum ducamara and associated surface water. Bull OEPP 28: 93–94.
Franc GD . (1988). Long distance transport of Erwinia carotovora in the atmosphere and surface water. PhD thesis, Department of Plant Pathology, Colorado State University: Fort Collins, 131pp.
Franc GD, DeMott PJ . (1998). Cloud activation characteristics of airborne Erwinia carotovora cells. J Appl Meteorol 37: 1293–1300.
Gerhardt P, Murray RGE, Costilow RN, Nester EW, Wood WA, Krieg NR, Phillips GB (eds). (1981). Manual of Methods for General Bacteriology. American Society for Microbiology: Washington, DC.
Goumas DE, Chatzaki AK . (1998). Characterization and host range evaluation of Pseudomonas viridiflava from melon, blite, tomato, chrysanthemum and eggplant. Eur J Plant Pathol 104: 181–188.
Gross DC . (1985). Regulation of syringomycin synthesis in Pseudomonas syringae pv. syringae and defined conditions for its production. J Appl Bacteriol 58: 167–174.
Gross DC, DeVay JE . (1977). Population dynamics and pathogenesis of Pseudomonas syringae in maize and cowpea in relation to the in vitro production of syringomycin. Phytopathology 67: 475–483.
Guttman DS, Gropp SJ, Morgan RL, Wang PW . (2006). Diversifying selection drives the evolution of the type III secretion system pilus of Pseudomonas syringae. Mol Biol Evol 23: 2342–2354.
Harrison MD, Franc GD, Maddox DA, Michaud JE, McCarter-Zorner NJ . (1987). Presence of Erwinia carotovora in surface water in North America. J Appl Bacteriol 62: 565–570.
Hirano SS, Upper CD . (2000). Bacteria in the leaf ecosystem with emphasis on Pseudomonas syringae, a pathogen, ice nucleus, and epiphyte. Microbiol Mol Biol Rev 64: 624–653.
Huelsenbeck JP, Ronquist F . (2001). MrBayes: Bayesian inference of phylogenetic trees. Bioinformatics 17: 754–755.
Hwang MSH, Morgan RL, Sarkar SF, Wang PW, Guttman DS . (2005). Phylogenetic characterization of virulence and resistance phenotypes of Pseudomonas syringae. Appl Environ Microbiol 71: 5182–5191.
Jakob K, Goss EM, Araki H, Van T, Kreitman M, Bergelson J . (2002). Pseudomonas viridiflava and P. syringae—natural pathogens Arabidopsis thaliana. Mol Plant Microbe Interact 15: 1195–1203.
King EO, Ward MK, Raney DE . (1954). Two simple media for the demonstration of pyocyanin and fluorescein. J Lab Clin Med 44: 301–307.
Konstantinidis KT, Isaacs N, Fett J, Simpson S, Long DT, Marsh TL . (2003). Microbial diversity and resistance to copper in metal-contaminated lake sediment. Microb Ecol 45: 191–202.
Lelliot RA, Billing E, Hayward AC . (1966). A determinative scheme for the fluorescent plant pathogenic Pseudomonads. J Appl Bacteriol 29: 470–489.
Lindemann J, Arny DC, Upper CD . (1984). Epiphytic populations of Pseudomonas syringae pv. syringae on snap bean and nonhost plants and the incidence of bacterial brown spot disease in relation to cropping patterns. Phytopathology 74: 1329–1333.
Lindemann J, Constantinidiou HA, Barchet WR, Upper CD . (1982). Plants as source of airbone bacteria, including ice nucleation-active bacteria. Appl Environ Microbiol 44: 1059–1063.
McCarter-Zorner NJ, Franc GD, Harrison MD, Michaud JE, Quinn CE . (1984). Soft rot Erwinia bacteria in surface and underground waters in southen Scotland and in Colorado, United-States. J Appl Bacteriol 57: 95–105.
Mohan SK, Schaad NW . (1987). An improved agar plating assay for detecting Pseudomonas syringae pv. syringae and P. s. pv. phaseolicola in contaminated bean seed. Phytopathology 77: 1390–1395.
Morris CE, Georgakapolous D, Sands DC . (2004). Ice nucleation active bacteria and their potential role in precipitation. J Phys 121: 87–103.
Morris CE, Glaux C, Latour X, Gardan L, Samson R, Pitrat M . (2000). The relationship of host range, physiology, and genotype to virulence on cantaloupe in Pseudomonas syringae from cantaloupe blight epidemics in France. Phytopathology 90: 636–646.
Morris CE, Kinkel LL, Kun X, Prior P, Sands DC . (2007). A surprising niche for the plant pathogen Pseudomonas syringae. Infect Genet Evol 7: 84–92.
Riffaud CM-H, Morris CE . (2002). Detection of Pseudomonas syringae pv. aptata in irrigation water retention basins by immunofluorescence colony staining. Eur J Plant Pathol 108: 539–545.
Ronquist F, Huelsenbeck JP . (2003). MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19: 1572–1574.
Sands DC, Langhans VE, Scharen AL, de Smet G . (1982). The association between bacteria and rain and possible resultant meteorological implications. J Hungarian Meteorol Serv 86: 148–152.
Sarkar SF, Guttman DS . (2004). Evolution of the core genome of Pseudomonas syringae, a highly clonal, endemic plant pathogen. Appl Environ Microbiol 70: 1999–2012.
Sawada H, Suzuki F, Matsuda I, Saitou N . (1999). Phylogenetic analysis of Pseudomonas syringae pathovars suggests the horizontal gene transfer of argK and the evolutionary stability of hrp gene cluster. J Mol Evol 49: 627–644.
Vincente JG, Alves JP, Russell K, Roberts SJ . (2004). Identification and discrimination of Pseudomonas syringae isolates from wild cherry in England. Eur J Plant Pathol 110: 337–351.
Wenneker M, Verdel MSW, Groeneveld RMW, Kempenaar C, Van Beuningen AR, Janse JD . (1999). Ralstonia (Pseudomonas) solanacearum race 3 in surface water and natural weed hosts: first report on stinging nettle (Urtica dioica). Eur J Plant Pathol 105: 307–315.
Yamamoto S, Kaasai H, Arnold DL, Jackson RW, Vivian A, Harayama S . (2000). Phylogeny of the genus Pseudomonas: intrageneric structure reconstructed from the nucleotide sequences of gyrB and rpoD genes. Microbiology 146: 2385–2394.
Acknowledgements
We thank Steven Lindow (University of California, Berkeley) for critical reading and discussions of the paper. We also thank Hervé Lecoq, Cécile Desbiez and Jonathan Gaudin of INRA-Avignon for wild-plant samples and for help with plant identification, and Marie-Agnès Jacques (INRA-Angers), Marc Bardin and Philippe Nicot (INRA-Avignon) for contributing rain and snow samples. We are grateful to Roland Psenner and Johannes Fried (University of Innsbruck) who led us on the sampling excursion in the Austrian Alps and to the participation of Rose, Simon and Philippe Nicot in this excursion. We thank the Virginia Tech undergraduate students Chandler Douglas and Eric Hall for their help with DNA sequence analysis. This work was supported by INRA's Department of Plant Health and Environment.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Morris, C., Sands, D., Vinatzer, B. et al. The life history of the plant pathogen Pseudomonas syringae is linked to the water cycle. ISME J 2, 321–334 (2008). https://doi.org/10.1038/ismej.2007.113
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ismej.2007.113
Keywords
This article is cited by
-
Naïve Bayes Classifiers and accompanying dataset for Pseudomonas syringae isolate characterization
Scientific Data (2024)
-
Review of Pseudomonas species causing bacterial canker of Prunus species with emphasis on sweet cherry (Prunus avium) in New Zealand
European Journal of Plant Pathology (2024)
-
Bacillus spp. isolated from pepper leaves and their function and inhibition of the fungal plant pathogen Colletotrichum scovillei
Egyptian Journal of Biological Pest Control (2023)
-
Occurrence of plant pathogenic Pseudomonas syringae in the Danube River Basin: abundance and diversity assessment
Phytopathology Research (2023)
-
A critical review on bioaerosols—dispersal of crop pathogenic microorganisms and their impact on crop yield
Brazilian Journal of Microbiology (2023)