Abstract
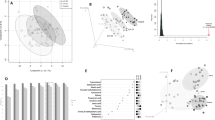
Obesity is a complex multifactorial disease involving genetic and environmental factors and influencing several different metabolic pathways. In this regard, metabonomics, that is the study of complex metabolite profiles in biological samples, may provide a systems approach to understand the global metabolic regulation of the organism in relation to this peculiar pathology. In this pilot study, we have applied a nuclear magnetic resonance (NMR)-based metabolomic approach on urinary samples of morbidly obese subjects. Urine samples of 15 morbidly obese insulin-resistant (body mass index>40; homeostasis assessment model of insulin resistance>3) male patients and 10 age-matched controls were collected, frozen and analyzed by high-resolution 1H-NMR spectroscopy combined with partial least squares-discriminant analysis. Furthermore, two obese patients who underwent bariatric surgery (biliopancreatic diversion and gastric bypass, respectively) were monitored during the first 3 months after surgery and their urinary metabolic profiles were characterized. NMR-based metabolomic analysis allowed us to identify an obesity-associated metabolic phenotype (metabotype) that differs from that of lean controls. Gut flora-derived metabolites such as hippuric acid, trigonelline, 2-hydroxyisobutyrate and xanthine contributed most to the classification model and were responsible for the discrimination. These preliminary results confirmed that in humans the gut microflora metabolism is strongly linked to the obesity phenotype. Moreover, the typical obese metabotype is lost after weight loss induced by bariatric surgery.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
McPherson K, Marsh T, Brown M . Modelling Future Trends in Obesity and the Impact on Health. Foresight Tackling Obesities: Future Choices. 2007, (http://www.foresight.gov.uk).
Cani PD, Delzenne NM . The role of the gut microbiota in energy metabolism and metabolic disease. Curr Pharm Des 2009; 15: 1546–1558.
Turnbaugh PJ, Ley RE, Mahowald MA, Magrini V, Mardis ER, Gordon JI . An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 2006; 444: 1027–1031.
Backhed F, Ding H, Wang T, Hooper LV, Koh GY, Nagy A et al. The gut microbiota as an environmental factor that regulates fat storage. Proc Natl Acad Sci USA 2004; 101: 15718–15723.
Goodacre R . Metabolomics of a superorganism. J Nutr 2007; 137: 259S–266S.
Li M, Wang B, Zhang M, Rantalainen M, Wang S, Zhou H et al. Symbiotic gut microbes modulate human metabolic phenotypes. Proc Natl Acad Sci USA 2008; 105: 2117–2122.
Nicholson JK . Global systems biology, personalized medicine and molecular epidemiology. Mol Syst Biol 2006; 2: 52.
Nicholson JK, Wilson ID . Understanding ‘global’ systems biology: metabonomics and the continuum of metabolism. Nat Rev Drug Discov 2003; 2: 668–676.
Bult MJF, van Dalen T, Muller AF . Surgical treatment of obesity. Eur J Endocrinol 2008; 158: 135–145.
Schneider BE, Mun EC . Surgical management of morbid obesity. Diabetes Care 2005; 28: 475–480.
Engelke UFH, Moolenaar SH, Hoenderop SMGC, Morava E, van der Graaf M, Heerschap A et al. Handbook of 1H-NMR spectroscopy in inborn errors of metabolism: body fluid NMR spectroscopy and in vivo MR spectroscopy. Heilbronn: SPS Verlagsgesellschaft, 2006.
Miccheli A, Marini F, Capuani G, Miccheli Tomassini A, Delfini M, Di Cocco ME et al. The influence of a sports drink on the post-exercise metabolism of elite athletes as investigated by NMR-based metabolomics. J Am Coll Nutr (in press).
Ross A, Schlotterbeck G, Dieterle F, Senn H . NMR spectroscopy techniques for application to metabolomics. In: Lindon JC, Nicholson JK, Holmes E (eds). The Handbook of Metabonomics and Metabolomics. Elsevier BV: Amsterdam, The Netherlands, 2007, pp 55–112.
Eriksson L, Johansson E, Kettaneh Wold N, Wold S . Multi and Megavariate Data Analysis. Umetrics AB: Umeå, 2001.
Nicholson JK, Holmes E, Wilson ID . Gut microorganisms, mammalian metabolism and personalized health care. Nat Rev Microbiol 2005; 3: 431–438.
Waldram A, Holmes E, Wang Y, Rantalainen M, Wilson ID, Tuohy KM et al. Top-down systems biology modeling of host metabotype-microbiome associations in obese rodents. J Proteome Res 2009; 8: 2361–2375.
Salek RM, Maguire ML, Bentley E, Rubtsov DV, Hough T, Cheeseman M et al. A metabolomic comparison of urinary changes in type 2 diabetes in mouse, rat, and human. Physiol Genomics 2007; 29: 99–108.
Williams RE, Lenz EM, Evans JA, Wilson ID, Granger JH, Plumb RS et al. A combined (1)H NMR and HPLC-MS-based metabonomic study of urine from obese (fa/fa) Zucker and normal Wistar-derived rats. J Pharm Biomed Anal 2005; 38: 465–471.
Shearer J, Duggan G, Weljie A, Hittel DS, Wasserman DH, Vogel HJ . Metabolomic profiling of dietary-induced insulin resistance in the high fat-fed C57BL/6J mouse. Diabetes Obes Metab 2008; 10: 950–958.
Holmes E, Loo RL, Stamler J, Bictash M, Yap IKS, Chan Q et al. Human metabolic phenotype diversity and its association with diet and blood pressure. Nature 2008; 453: 396–400.
Rezzi S, Ramadan Z, Martin FP, Fay LB, van Bladeren P, Lindon JC et al. Human metabolic phenotypes link directly to specific dietary preferences in healthy individuals. J Proteome Res 2007; 6: 4469–4477.
Sun J, Schnackenberg LK, Holland RD, Schmitt TC, Cantor GH, Dragan YP et al. Metabonomics evaluation of urine from rats given acute and chronic doses of acetaminophen using NMR and UPLC/MS. J Chromatogr B 2008; 871: 328–340.
Turnbaugh PJ, Hamady M, Yatsunenko T, Cantarel BL, Duncan A, Ley RE et al. A core gut microbiome in obese and lean twins. Nature 2009; 457: 480–484.
Turnbaugh PJ, Gordon JI . The core gut microbiome, energy balance and obesity. J Physiol 2009; 587 (Part 17): 4153–4158.
Matsuura F, Yamashita S, Nakamura T, Nishida M, Nozaki S, Funahashi T et al. Effect of visceral fat accumulation on uric acid metabolism in male obese subjects: visceral fat obesity is linked more closely to overproduction of uric acid than subcutaneous fat obesity. Metab Clin Exp 1998; 47: 929–933.
Buchwald H, Avidor Y, Braunwald E, Jensen MD, Pories W, Fahrbach K et al. Bariatric surgery: a systematic review and meta-analysis. JAMA 2004; 292: 1724–1737.
Zhang H, DiBaise JK, Zuccolo A, Kudrna D, Braidotti M, Yu Y et al. Human gut microbiota in obesity and after gastric bypass. Proc Natl Acad Sci USA 2009; 106: 2365–2370.
Maggard MA, Shugarman LR, Suttorp M, Maglione M, Sugerman HJ, Livingston EH et al. Meta-analysis: surgical treatment of obesity. Ann Intern Med 2005; 142: 547–559.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on International Journal of Obesity website
Rights and permissions
About this article
Cite this article
Calvani, R., Miccheli, A., Capuani, G. et al. Gut microbiome-derived metabolites characterize a peculiar obese urinary metabotype. Int J Obes 34, 1095–1098 (2010). https://doi.org/10.1038/ijo.2010.44
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ijo.2010.44
Keywords
This article is cited by
-
Gut bacteria-derived 3-phenylpropionylglycine mitigates adipocyte differentiation of 3T3-L1 cells by inhibiting adiponectin-PPAR pathway
Genes & Genomics (2023)
-
Urinary metabolite profiling and risk of progression of diabetic nephropathy in 2670 individuals with type 1 diabetes
Diabetologia (2022)
-
The Metabolic Signature of Cardiorespiratory Fitness: A Systematic Review
Sports Medicine (2022)
-
Effects of Sporisorium reiliana polysaccharides and Phoenix dactylifera monosaccharides on the gut microbiota and serum metabolism in mice with fructose-induced hyperuricemia
Archives of Microbiology (2022)
-
NEIL3-deficiency increases gut permeability and contributes to a pro-atherogenic metabolic phenotype
Scientific Reports (2021)