Abstract
As the junction of floral development pathways, the FLOWERING LOCUS T (FT) protein called ‘florigen’ plays an important role in the process of plant flowering through signal integration. We isolated four transcripts encoding different isoforms of a FT orthologous gene CmFTL1, from Chrysanthemum morifolium cultivar ‘Jimba’. Sequence alignments suggested that the four transcripts are related to the intron 1. Expression analysis showed that four alternative splicing (AS) forms of CmFTL1 varied depending on the developmental stage of the flower. The functional complement experiment using an Arabidopsis mutant ft-10 revealed that the archetypal and AS forms of CmFTL1 had the function of complementing late flower phenotype in different levels. In addition, transgenic confirmation at transcript level showed CmFTL1 and CmFTL1ast coexist in the same tissue type at the same developmental stage, indicating a post-transcriptional modification of CmFTL1 in Arabidopsis. Moreover, ectopic expression of different AS forms in chrysanthemum resulted in the development of multiple altered phenotypes, varying degrees of early flowering. We found that an alternative splicing form (CmFTL1-astE134) without the exon 2 lacked the ability causing the earlier flower phenotype. The evidence in this study indicates that complex alternative processing of CmFTL1 transcripts in C. morifolium may be associated with flowering regulation and hold some potential for biotechnical engineering to create early-flowering phenotypes in ornamental cultivars.
Similar content being viewed by others
Introduction
Alternative splicing (AS) produces multiple mRNA transcripts or isoforms from a single gene through the assembly of exons in different ways using alternative splice sites in the precursor mRNA (pre-mRNA).1 AS is known to contribute to gene regulation and proteome diversity, and in plants AS is involved in many important biological processes including stress responses,2,3 development4–6 and reproductive growth.7,8 On the basis of EST/cDNA evidence in Arabidopsis, it has been predicted that more than 61% of genes are alternatively spliced under normal conditions, and the number will increase under stress.9,10
In Arabidopsis, the FLOWERING LOCUS T (FT) gene has been demonstrated to encode florigen, which integrates environmental and endogenous signals and triggers flowering.11,12 The florigen is a small protein (FT) and belongs to a large group of plant proteins with structural similarities to phosphatidylethanol-amine-binding protein. In Arabidopsis, FT proteins move to the shoot apical meristem via the phloem, where they interact with a basic leucine zipper (bZIP) transcription factor, FD to form a transcriptional complex. This complex directly activates floral regulator genes such as APETALA 1 (AP1) and FRUITFULL (FUL) to promote flowering.13,14 In general, FT orthologous genes consist of four exons and three introns and have a high level of sequence homology and similar functions in evolutionarily distant plant species.15–17 AS of FT orthologous genes was previously observed in London plane (Platanus acerifolia Willd), maize (Zea mays) and sunflower (Helianthus annuus). In the Zea mays, two orthologous FT genes ZCN18 and ZCN26 (Zea mays CETRORADIALIS), have developmentally specific patterns with alternatively spliced transcripts.18 The alternative splicing of PaFT in Platanus acerifolia was more complicated in terms of temporal and spatial expression patterning, suggesting it may be associated with multiple regulatory roles in different stages of growth.19 In sunflower, a frameshift mutation in the third exon of HaFT1 affects flowering time through interference with the function of another paralog.20
Most chrysanthemum (Chrysanthemum morifolium) cultivars are typical short-day plants where the flowering time is subject to the short-day conditions. Stable year-round flower production is costly and difficult. In recent years, deeper understanding of the mechanisms of floral transition at the molecular level and further developments in chrysanthemum are a point of interest to researchers. Three FT-like (FTL) genes have been isolated from a wild diploid chrysanthemum, Chrysanthemum seticuspe. CsFTL3 acts as a floral stimulator under SD conditions while the functions of other two CsFTLs are still not clear.21 Currently, AS activity in FT orthologous genes have never been reported in chrysanthemum. In the present work, CmFTL1 and four isoforms resulting from alternative splicing were isolated from C. morifolium cultivar ‘Jimba’. Detailed sequencing, splicing sites and types of CmFTL1 alternatively spliced forms were studied. Ectopic expression of CmFTL1 and four AS transcripts were examined for complementation of the Arabidopsis ft-10 mutants. The phenotypes of chrysanthemum that overexpressed CmFTL1 and CmFTL1asts showed varying degrees of early flowering. The results provided strong evidences of highly functional conservation of FT orthologs in both Arabidopsis and chrysanthemum.
Materials and methods
Plant material and growth conditions
Chrysanthemum morifolium cultivars ‘Jimba’ were used for the experiments. Plants were grown in a greenhouse and maintained at 20 °C with an 8 h photoperiod (SD conditions) and the light intensity of 1200 μmol m−2 s−1. To analyse the expression patterns of CmFTL1 and CmFTL1asts under natural conditions, leaf samples were initially collected at an early developmental stage and at 3-day intervals until visual colour stage, then samples were collected at 5-day intervals until flowers opened. All leaf samples were collected at 16:00 hours21–23 on each time point.
For genetic transformation experiments, transformed Arabidopsis plants were grown in a greenhouse with a 16 h photoperiod (LD condition) and a 25 °C/18 °C day/night temperature. Transgenic chrysanthemum plants were grown under the condition described above.
CmFTL1 and CmFTL1asts cloning
Total RNA was extracted from the third fully expanded leaf (counting from the apex) of ‘Jimba’ plants using the RNAiso reagent (TaKaRa, Dalian, China) and treated with RNase-free DNase (TaKaRa) according to the manufacturer’s instructions. The cDNA synthesized from a 1 μg aliquot of total RNA by using the TaKaRa RNA PCR Kit (AMV) v2.1 (TaKaRa) was used as a template to isolate both the CmFTL1 and CmFTL1ast sequences by performing a PCR primed by the oligomer pair 5′-TAATGCCGAGGGAAAGGGAT-3′/5′-GTCATGATTGATTCAGTTAAAGTGT-3′. Full-length versions of the cDNAs were derived by applying both 5′- and 3′-RACE PCR using a GeneRacer kit (Invitrogen, Carlsbad, CA, USA). The amplicons (~500 bp in length) were inserted into the pMD19-T vector (TaKaRa) for sequencing. Genomic DNA was extracted from juvenile leaves using a modified CTAB method.24 The first intron (intron#1) sequence of CmFTL1 was amplified by a PCR based on the primer pair 5′-GCGTAATAGGGGATGTTCTTGATAG-3′/5′-ACTAGGACTTGGAGCATCAGGAT-3′. The resulting amplicon was introduced into pMD19-T for sequencing.
CmFTL1 and splicing sites analysis
The sequence of the CmFTL1 gene was submitted to NCBI and analysed with BLAST (http://blast.ncbi.nlm.nih.gov/Blast.cgi). The genomic DNA sequence of CmFTL1 exon 1–intron 1–exon 2 was submitted to the Alternative Splice Site Predictor (http://wangcomputing.com/assp/) to enable a prediction of the AS sites present. The full-length mRNA of CmFLT1 was submitted to NCBI (http://blast.ncbi.nlm.nih.gov/Blast.cgi) for BLASTing, and the sequences of homologous genes in other species with high identities were identified. The multiple sequences alignment was performed using the software BioXM2.6 (Nanjing Agricultural University, Nanjing, China) and DNAMAN (Lynnon LLC., San Ramon, CA, USA).
Expression analysis by quantitative real-time PCR and semi-quantitative PCR
The abundance of CmFTL1 transcripts was determined by semi-quantitative PCR and quantitative real-time PCR (qRT-PCR). Total RNA was extracted using the RNAiso reagent (TaKaRa) followed by RNase-free DNase treatment (TaKaRa). A 1 μg aliquot of total RNA was reverse-transcribed with SMART MMLV Reverse Transcriptase (Clontech, Palo Alto, CA, USA) according to the manufacturer’s instructions. The resulting cDNA was diluted fivefold of its original concentration, and a 5 μL aliquot was used as template in a 15 μL qPCR SYBR Premix Ex Taq (TaKaRa). All reactions were performed in triplicates. The reaction was initially denatured (95 °C/2 min), then cycled 40 times through 95 °C/15 s, 55 °C/15 s, 72 °C/20 s. The chrysanthemum EF1a sequence (AB679278) was chosen as the reference. The ‘calibrator sample’ was designated as the most highly expressed time point for each gene of interest and therefore represented the highest relative transcription level of 1.0. The data showed the mean and s.e. of at least three biological replicates. The qRT-PCR template was a 1 μL aliquot of cDNA (1000 ng μL−1); the reaction conditions were given in Supplementary Table 2.
Arabidopsis and chrysanthemum transformation
Both the A. thaliana FT loss-of-function mutant ft-10 and ‘Jimba’ were transformed with various CmFTL1 constructs.
The A. thaliana FT promoter (6977 nt) from the pENTR4-AtFT vector was sub-cloned into pHGW (Invitrogen) to obtain pHGW-proAtFT 25 via LR recombination. The CmFTL1 open reading frame was amplified from the cDNA described above with the primer pair (Supplementary Table 1), the product was then inserted into pHGW-proAtFT to form pHGW-proAtFT-CmFTL1. This construct was used to test for complementation of the A. thaliana ft-10 mutant. To construct the CmFTL1 overexpression vector, the open reading frame sequence of CmFTL1 and CmFTL1asts was amplified using a pair of primers with SalI and NotI restriction enzymes sites (Supplementary Table 1), and the resulting PCR product was digested with corresponding restriction enzymes and inserted into pENTR1A, then sub-cloned into the GATEWAY vector pMDC4326 via LR recombination. Arabidopsis plants were transformed using the floral dip method,27 while transgenic lines of C. morifolium ‘Jimba’ were achieved using a modified Agrobacterium-mediated transformation protocol.28
Result
Molecular cloning and sequence analysis of CmFTL1 and CmFTL1asts from ‘Jimba’
We used RACE strategies to clone the CmFTL genes of C. morifolium cultivar ‘Jimba’. The products of CmFTL1 were electrophoresed through 2% agarose gels and cloned into the sequencing vector. Different sizes of CmFTL1 cDNA products were obtained. Analysis of the sequences revealed that the CmFTL1 had at least five different alternatively spliced transcripts. Compared with the normal full-length CmFTL1 transcript, two of the alternatively spliced products (CmFTL1astIn1-1, CmFTL1astIn1-2) contained the different lengths of the intron and two transcripts (CmFTL1astE134 and CmFTL1astIn1-3) lacked exons (Figure 1a).
The nucleotide sequences of CmFTL1 and its AS products. (a) Sequence alignment of CmFTL1 cDNA and its AS products; asterisks indicate separation of the four exons. (b) The genomic sequence of a segment of CmFTL1. Exons are shown boxed. The letters highlighted in grey indicate the AS sites. The bold letters indicate the initiation (ATG) and the termination (TAA) codons. (c) Processing of CmFTL1. Horizontal lines indicate the 5′ and 3′ UTRs. The coding regions are shown as grey-stippled boxes and the introns as white boxes; dotted lines indicate unknown introns. The vertical lines in the white boxes indicate the AS sites.
To confirm the CmFTL1 alternative splicing sites, the intron1 was cloned from gDNA and sequenced (Supplementary Figure 1). The prediction of AS sites by ASSP showed the sites at boundary of intron1 are not only the normal GT-AG constitutive splicing sites, but also had the alternatively spliced potential. The intron1of CmFTL1 had two latent alternative acceptor sites (Figure 1b). According to reports,29 there are five distinct types of alternative splicing events: exon skip type (ES type), intron retention type (RI type), alternative donor type (AD type), alternative acceptor type (AA type) and mutually exclusive exons (MEE type). Besides the archetypal spliced form (CmFTL1), four other CmFTL1 spliced forms were detected (Figure 1c). Among the four spliced forms, CmFTL1-astE134 and CmFTL1-astIn1-3 were resulted from ES type events, while CmFTL1-astIn1-1 and CmFTL1-astIn1-3 were derived from RI type events. CmFTL1-astIn1-2 had only an AD type event. Comparison of the various CmFTL1 splice forms showed that alternative splicing events tended to occur more frequently in the first intron than in the second or third intron.
The polypeptide sequence of CmFTL1 and CmFTL1ast transcripts
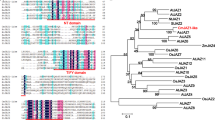
The archetypal CmFTL1 transcript encodes a 174 residue protein which shares 72% sequence identity with the A. thaliana FT protein, 90% with lettuce LsFT and 89% with rice RFT1 (Table 1). The residues Tyr85 and Gln140, both of which are critical for the functionality of AtFT,30,31 were present in CmFTL1, at positions 84 and 139, respectively (Figure 2a). Those AS transcripts in which the number of nucleotides inserted/deleted was not a multiple of three are expected to encode a frame shift product, while those truncated by the creation of a premature termination codon are expected to have lost residues in exon 4 essential for FT-like activity30,31 (Figure 2b).
Analysis of the amino-acid alignments of CmFTL1 and CmFTL1asts. Multiple alignment on GENEBANK: AtFT (AB027504.1), AtTFL1 (NM_120465.2), AtTSF (GQ884981.1), BvFT1 (HM448910.1), BvFT2 (HM448912.1), HaFT1 (GQ884981.1), HaFT2 (GQ884987.1), HaFT4 (GU985573.1), LsFT (BAK14368.1), OsHd3a (JX472280), OsRFT1 (AB426873.1). (a) Partial amino-acid sequence alignment of the phosphatidylethanol-amine-binding protein (PEBP) family members. Asterisks indicate the residues Tyr85 (Y)/Gln140 (Q) and His88 (H)/Asp144 (D) contributing to FT and TFL1 functioning, respectively.30, 31 The conserved segment region B in the fourth exon, corresponding to the external loop of the PEBP family proteins, is boxed. (b) Alignment of amino-acid sequence of CmFTL1 and its alternative splicing transcripts, the asterisks separate the exons into Exon1, Exon2, Exon3 and Exon4.
Alternative splicing plays an important role in increasing the diversity of proteomes in both animals and plants. AS forms encode variant proteins, leading to completely different or multiple biological functions. However, the number of nucleotides inserted/deleted in CmFTL1 alternative splicing forms were not in multiples of 3, resulting in frame-shifts that truncated the CmFTL1asts protein and losing the amino acids that are critical for FT or TFL1-like protein function and the regions that are essential for FT-like activity in exon 430,31 (Figure 2b). In addition, premature termination codons (PTCs) may be the targets for nonsense-mediated mRNA decay (NMD),32,33 however many PTC-containing transcripts also exist that remain functional.34,35 Thus, further research of the protein encoded by CmFTL1asts is required to confirm whether the PTC-containing transcripts lead to functional truncated proteins or to NMD.
Phenotypes of transgenic Arabidopsis expressing ectopic CmFTL1 and CmFTL1asts
In order to gain the functional divergence of CmFTL1 and its alternative splicing isoforms, we attempted to rescue the loss-of-function A. thaliana mutant (ft-10) by CmFTL1 and CmFTL1asts, respectively. The vectors carrying AtFTpro: CmFTL1 and 35S: CmFTL1asts were separately introduced into A. thaliana by Agrobacterium-mediated transformation (Supplementary Figure 2).
The effect on phenotype of the various transgenes was assessed by comparing the number of rosette leaves formed at the flowering stage between the transgenic lines constructed in both a wild-type and a ft-10 mutant background. Under the growing conditions imposed, wild-type plants produced 15 rosette leaves at flowering and the ft-10 mutants 36. On the basis of the number of rosette leaves produced at the flowering stage, we classified the recovery phenotypes of transgenic lines as strong (<18), moderate (19–24) and weak (>25). Among 19 proAtFT::CmFTL1 ft-10 transformants analysed, 5 produced strong phenotype, while the other 14 produced moderate phenotypes (Figure 3a). Thus the CmFTL1 transgene appeared able to at least partially overcome the delay in flowering induced by the loss of AtFT (Figure 3b). Three of the four CmFTL1ast sequences (the exception was CmFTL1astE134) were able to partially rescue the ft-10 mutant. Among 25 transformants with the 35S:CmFTL1-ast In1-1 in ft-10 mutant, 8 lines displayed strong phenotype, 11 lines with moderate phenotype and the remaining 6 lines displayed weak recovery (Figure 3c). Among a total of 15 transgenic lines with 35S:CmFTL1-ast In1-2 transgenes, 4 lines showed strong changes in flowering time (13–18 rosette leaves), 9 lines showed moderate changes and the rest 2 lines showed weak recovery change in flowering time (Figure 3d). Among 15 of the 35S:CmFTL1-ast In1-3 transgenic lines, only 1 line showed strong phenotype, 12 lines showed the moderate phenotype and the rest 8 lines showed weak phenotype (Figure 3e).
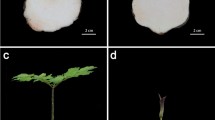
The phenotypic effect of expressing CmFTL1-derived transgenes in A. thaliana. (a) Rosette leaf number at the flowering stage of transgenic and non-transgenic A. thaliana. Plants grown under long day conditions. (b–f) The phenotype of transgenic plants either harbouring or not harbouring AtFT, and constitutively expressing (b) CmFTL1, (c) CmFTL1astE134, (d) CmFTL1astIn1-1, (e) CmFTL1astIn1-2 or (f) CmFTL1astIn1-3. (a) Strong complementation (13–18 rosette leaves) at flowering, (b) moderate complementation (19–24), (c) weak complementation (>25). Bar, 1 cm.
To verify that the the various CmFTL1-derived transgenes were successfully transcribed in A. thaliana, the putative transgenic lines were tested by qRT-PCR. In the lines harbouring an AS product, fragments corresponding to various sizes were detected (Supplementary Figure 3). Sequencing of the amplicons showed that the CmFTL1astIn1-1, 1-2 and 1-3 lines each produced AS as well as post-transcriptionally modified forms, suggesting that the intron1-containing AS products were processed post-transcriptionally to recreate the archetypal CmFTL1 transcript able to promote flowering.
Expression patterns of CmFTL1 and CmFTL1asts in chrysanthemum
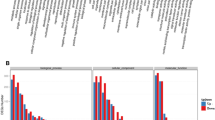
To investigate the potential function of CmFTL1 and CmFTL1asts in chrysanthemum, transgenic chrysanthemum plants with CmFTL1 and CmFTL1asts expressed under the control of the Cauliflower mosaic virus (CaMV) 35S promoter were generated, named by OX-CmFTL1 or OX-CmFTL1asts (Supplementary Figures 4–6). The phenotypes of transgenic lines were observed under natural conditions. The flower development of chrysanthemum involves six stages: before flower bud development stage, flower bud development stage (FBD), visible colour stage (VC) stage, earlier opening stage, opened flower stage and senescing flower stage.22 The FBD and the VC stage of OX-CmFTL1astIn1-1 and OX-CmFTL1astIn1-2 lines were both advanced by 5–8days, while OX-CmFTL1astIn1-3 lines were advanced only by 2–4 days. No significant alteration to flowering time was noted in the transgenic plants constitutively expressing CmFTL1astE134 and wild-type plants (Figure 4). The qRT-PCR and qPCR analyses showed that in wild-type plants, the abundance of CmFTL1 transcript peaked at the early flower bud development stage (Figure 5). Although the temporal pattern of the AS forms was similar to the wild type, their transcript abundance was much lower. However, the AS transcripts were higher than the CmFTL1 in the AS transgenic chrysanthemum plants (Figure 6). The results suggested that the CmFTL1 AS transcripts underwent post-transcriptional processing, similar to the CmFTL1ast A. thaliana transgenic lines, resulting in an increased abundance of the archetypal transcript.
The transcription of CmFTL1 and its AS products in the leaf of wild-type chrysanthemum plants grown under natural conditions. (a) The transcription of CmFTL1astsE134 and CmFTL1astIn1-1 was assayed using qRT-PCR. (b) The transcription of CmFTL1astIn1-2 and 1-3 was assayed using qPCR. Time points were days after transplanting. 98d: prior to the FBD stage; 101–104d: the FBD stage; 104–108d: the VC stage; 108–118d: the EO stage. Transcript abundances were normalized against that of CmEF1a.
The transcripts of CmFTL1 in its AS products transgenic chrysanthemum plants. (a) CmFTL1astsE134 line 49; (b) CmFTL1astIn1-1 line 63; (c) CmFTL1astIn1-2 line 4; (d) CmFTL1astIn1-2 line 10; (e) CmFTL1astIn1-3 line 4; (f) CmFTL1astIn1-3 line 25. Time points were days after transplanting. 98d: prior to the FBD stage; 101–104d: the FBD stage; 104–108d: the VC stage; 108–118d: the EO stage. The transcription of CmFTL1astsE134 and CmFTL1astIn1-1 was assayed using qRT-PCR, and that of CmFTL1astIn1-2 and 1-3 by qPCR.
Discussion
AS represents a significant strategy for regulating gene expression, through the generation of a diversity of proteins. The investigation of AS in A. thaliana over the past decade has revealed that almost half of the genes are affected.36 Transcriptomic analysis has suggested that >61% of A. thaliana genes may contain the intron-generated AS products. The proportion will likely rise as a greater range of developmental stages and growing conditions will be explored in the future.10,37 The product of FT and its various homologues is the key to regulate the timing of the transition from vegetative to reproductive growth. Here we demonstrated that the chrysanthemum FT homologue generated various forms of AS, as do the FT homologues in maize, sunflower and the London plane.18,19,38 The individual AS products of the maize genes ZCN18 and ZCN26 have not been defined, while the sunflower gene HaFT1 produces a transcript containing a part of its third intron, introducing premature stop codons. The London plane FT gene displays a rather complicated AS profile.
The AS products of CmFTL1 involved ES-, RI- and AD-type events. Four of the splicing points occurred in intron1 even though this sequence is just 118 nt in length. The equivalent intron in AtFT (817 nt) is much longer than this, so intron length is clearly not a decisive factor in determining AS frequency. Some experimental evidences gathered from rice and A. thaliana suggest that preferential combinations of AS events exist,25,39 but the data obtained here from chrysanthemum are insufficient to imply any such preference. The multiple alternative splicing forms of PaFT (Platanus acerifolia) show that the ES type is always associated with the AD type.19 Our data show that two of CmFTL1asts have the similar preference of alternative splicing types. The intron splice boundaries generally follow the rule of ‘GT…AG’.40 The alternative splicing sites tends to weaker than the constitutive splicing sites so that the sites in introns or exons that have alternative splicing events are not typical.41 The predicted position of the AS sites in intron 1 of CmFTL1 suggested that intron splice boundary sequences need not be the expected GT and AG, but can also have features of AS sites, suggesting that the AS sites in CmFTL1 are not randomly placed, but more likely reflect the outcome of directed evolutionary selection. The results would imply that the AS messages are functional. For example, in the case of the gene FLC, which encodes a repressor of flowering involved in the vernalization pathway, AS sites are shared between the two highly disparate species canola (oilseed rape, Brassica rapa) and trifoliate orange (Poncirus trifoliata).7,42 It is clear that the FT sequence are largely conserved during the evolution of the angiosperms, and that it tends not to experience frequent AS events. So far there were only three plant species that FT orthologs involved AS, suggesting that the selection of AS sites in CmFTL1 is an atypical FT orthologs.
In chrysanthemum, as in rice,23 FTL1 shows weak florigenic activity under short day conditions. The potential regulatory control of CmFTL1 over floral transition was explored by constitutively expressing the sequences corresponding to both the archetypal CmFTL1 transcript and its four AS forms in both A. thaliana and chrysanthemum. The ability of CmFTL1 to rescue the phenotype of the ft-10 mutation established that the chrysanthemum homologue has an equivalent functionality to the native AtFT. The partial rescue of the ft-10 mutant phenotype achieved by constitutively expressing three of the four AS sequences suggested that post-transcriptional processing was able to convert them into the archetypal CmFTL1 transcript. The behaviour of the transgenic chrysanthemum lines constitutively expressing CmFTL1 and its various AS products was similar to the transgenic A. thaliana plants. A possible scenario is that the CmFTL1 AS transcript encodes a truncated form of CmFTL1, which shows attenuated florigenic activity; alternatively, its product may induce the interaction of the CmFTL1 with CmFDL1, which induces flowering in both chrysanthemum and the A. thaliana ft-10 mutant. Because many AS transcripts also exist and remain functional,34,35 we hypothesize that the CmFTL1asts may have opportunities to occurred post-transcriptional modification when lacking of CmFTL1 at floral transition and formed the archetypal to complement the function of CmFTL1. And both A. thaliana ft-10 mutant and chrysanthemum that ectopically expressed CmFTL1astsE134 failed to show any florigentic activity, suggesting that the loss of exon2 in CmFLT1may influenced its function and then affected the regulation of flowering timing.
In non-transgenic chrysanthemum plants, the abundance of CmFTL1 transcript was substantially greater than that of the other four AS transcripts. This relationship was altered in the transgenic plants in which CmFTL1 or one of its AS products was constitutively expressed, suggesting that the AS transcripts were processed post-transcriptionally, as was the case in the transgenic A. thaliana plants. Thus, our data demonstrate that both CmFTL1 and its AS forms are involved in the regulation of floral transition. The florigenic activity of the CsFTL1 product is weaker than that of CsFTL3 when these genes are expressed in the chrysanthemum variety ‘Jimba’ under long days.21,23 Although residual CsFTL3 and increased CsFTL1 activity are likely to have a major impact on the flowering process, the phyB-mediated protein CsAFT is also known to be important in determining the flowering response to photoperiod in chrysanthemum.23 This may explain why CmFTL1 shows such a low level of florigen in ‘Jimba’.
In summary, this study showed that CmFTL1 is somewhat atypical of FT homologues, since its transcript can be modified prior to its translation. The participation in the flowering process of both CmFTL1 and its AS forms was demonstrated by studying the phenotypic consequences of expressing these genes constitutively in both A. thaliana and chrysanthemum. The representation of the various transcriptional forms of the gene was dependent on the developmental stage of the plant, but some post-transcriptional modification of the AS products narrowed the gap of transcription levels. The results demonstrate that manipulating CmFTL1 may have potential benefit in shortening the life cycle of the chrysanthemum plant.
References
Barbazuk WB, Fu Y, McGinnis KM . Genome-wide analyses of alternative splicing in plants: opportunities and challenges. Genome Res 2008; 18: 1381–1392.
Kwon YJ, Park MJ, Kim SG, Baldwin IT, Park CM . Alternative splicing and nonsense-mediated decay of circadian clock genes under environmental stress conditions in Arabidopsis . BMC Plant Biol 2014; 14: 136.
Feng J, Li J, Gao Z, Lu Y, Yu J, Zheng Q et al. SKIP confers osmotic tolerance during salt stress by controlling alternative gene splicing in Arabidopsis . Mol Plant 2015; 8: 1038–1052.
Shikata H, Hanada K, Ushijima T, Nakashima M, Suzuki Y, Matsushita T . Phytochrome controls alternative splicing to mediate light responses in Arabidopsis . Proc Natl Acad Sci USA 2014; 111: 18781–18786.
Wang HF, You CJ, Chang F, Wang Y, Wang L, Qi J et al. Alternative splicing during Arabidopsis flower development results in constitutive and stage-regulated isoforms. Front Genet 2014; 5: 25.
Zhang B, Liu ZX, Ma J, Song Y, Chen FJ . Alternative splicing of the AGAMOUS orthologous gene in double flower of Magnolia stellata (Magnoliaceae). Plant Sci 2015; 241: 277–285.
Yuan YX, Wu J, Sun RF, Zhang XW, Xu DH, Bonnema G et al. A naturally occurring splicing site mutation in the Brassica rapa FLC1 gene is associated with variation in flowering time. J Exp Bot 2009; 60: 1299–1308.
Lee JH, Ryu HS, Chung KS, Posé D, Kim S, Schmid M et al. Regulation of temperature-responsive flowering by MADS-box transcription factor repressors. Science 2013; 342: 628–632.
Filichkin SA, Priest HD, Givan SA, Shen R, Bryant DW, Fox SE et al. Genome-wide mapping of alternative splicing in Arabidopsis thaliana . Genome Res 2009; 20: 45–58.
Marquez Y, Brown JWS, Simpson C, Barta A, Kalyna M . Transcriptome survey reveals increased complexity of the alternative splicing landscape in Arabidopsis . Genome Res 2012; 22: 1184–1195.
Corbesier L, Vincent C, Jang S, Fornara F, Fan Q, Searle I et al. FT protein movement contributes to long-distance signaling in floral induction of Arabidopsis . Science 2007; 316: 1030–1033.
Notaguchi M, Abe M, Kimura T, Daimon Y, Kobayashi T, Yamaguchi A et al. Long-distance, graft-transmissible action of Arabidopsis FLOWERING LOCUS T protein to promote flowering. Plant Cell Physiol 2008; 49: 1645–1658.
Abe M, Kobayashi Y, Yamamoto S, Daimon Y, Yamaguchi A, Ikeda Y et al. FD, a bZIP protein mediating signals from the floral pathway integrator FT at the shoot apex. Science 2005; 309: 1052–1056.
Wigge PA, Kim MC, Jaeger KE, Busch W, Schmid M, Lohmann JU et al. Integration of spatial and temporal information during floral induction in Arabidopsis . Science 2005; 309: 1056–1059.
Fukuda M, Matsuo S, Kikuchi K, Kawazu Y, Fujiyama R, Honda I . Isolation and functional characterization of the FLOWERING LOCUS T homolog, the LsFT gene, in lettuce. J Plant Physiol 2011; 168: 1602–1607.
Hsu CY, Adams JP, Kim H, No K, Ma C, Strauss SH et al. FLOWERING LOCUS T duplication coordinates reproductive and vegetative growth in perennial poplar. Proc Natl Acad Sci USA 2011; 108: 10756–10761.
Li C, Zhang Y, Zhang K, Guo D, Cui B, Wang X et al. Promoting flowering, lateral shoot outgrowth, leaf development, and flower abscission in tobacco plants overexpressing cotton FLOWERING LOCUS T (FT)-like gene GhFT1 . Front Plant Sci 2015; 6: 454.
Danilevskaya ON, Meng X, Hou Z, Ananiev EV, Simmons CR . A genomic and expression compendium of the expanded PEBP gene family from maize. Plant Physiol 2007; 146: 250–264.
Zhang J, Liu G, Guo C, He Y, Li Z, Ning G et al. The FLOWERING LOCUS T orthologous gene of Platanus acerifolia is expressed as alternatively spliced forms with distinct spatial and temporal patterns. Plant Biol 2011; 13: 809–820.
Blackman BK . Interacting duplications, fluctuating selection, and convergence: the complex dynamics of flowering time evolution during sunflower domestication. J Exp Bot 2012; 64: 421–431.
Oda A, Narumi T, Li T, Kando T, Higuchi Y, Sumitomo K et al. CsFTL3, a chrysanthemum FLOWERING LOCUS T-like gene, is a key regulator of photoperiodic flowering in chrysanthemums. J Exp Bot 2012; 63: 1461–1477.
Yang Y, Ma C, Xu Y, Wei Q, Imtiaz M, Lan H et al. A zinc finger protein regulates flowering time and abiotic stress tolerance in chrysanthemum by modulating gibberellin biosynthesis. Plant Cell 2014; 26: 2038–2054.
Higuchi Y, Narumi T, Oda A, Nakano Y, Sumitomo K, Fukai S et al. The gated induction system of a systemic floral inhibitor, antiflorigen, determines obligate short-day flowering in chrysanthemums. Proc Natl Acad Sci USA 2013; 110: 17137–17142.
Doyle JJ . A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem Bull 1987; 19: 11–15.
Karimi M, Inzé D, Depicker A . GATEWAY vectors for Agrobacterium-mediated plant transformation. Trends Plant Sci 2002; 7: 193–195.
Curtis MD, Grossniklaus U . gateway cloning vector set for high-throughput functional analysis of genes in planta. Plant Physiol 2003; 133: 462–469.
Clough SJ, Bent AF . Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana . Plant J 1998; 16: 735–743.
Aida R, Ohira K, Tanaka Y, Yoshida K, Kishimoto S, Shibata M et al. Efficient transgene expression in chrysanthemum, Dendranthema grandiflorum (Ramat.) Kitamura, by using the promoter of a gene for chrysanthemum chlorophyll-a/b-binding protein. Breed Sci 2004; 54: 51–58.
Iida K, Fukami-Kobayashi K, Toyoda A, Sakaki Y, Kobayashi M, Seki M et al. Analysis of multiple occurrences of alternative splicing events in Arabidopsis thaliana using novel sequenced full-length cDNAs. DNA Res 2009; 16: 155–164.
Hanzawa Y, Money T, Bradley D . A single amino acid converts a repressor to an activator of flowering. Proc Natl Acad Sci USA 2005; 102: 7748–7753.
Ahn JH, Miller D, Winter VJ, Banfield MJ, Lee JH, Yoo SY et al. A divergent external loop confers antagonistic activity on floral regulators FT and TFL1 . EMBO J 2006; 25: 605–614.
Maquat LE . Nonsense-mediated mRNA decay: splicing, translation and mRNP dynamics. Nat Rev Mol Cell Biol 2004; 5: 89–99.
Isken O, Maquat LE . The multiple lives of NMD factors: balancing roles in gene and genome regulation. Nat Rev Genet 2008; 9: 699–712.
Kalyna M, Lopato S, Voronin V, Barta A . Evolutionary conservation and regulation of particular alternative splicing events in plant SR proteins. Nucleic Acids Res 2006; 34: 4395–4405.
Kalyna M, Simpson CG, Syed NH, Lewandowska D, Marquez Y, Kusenda B et al. Alternative splicing and nonsense-mediated decay modulate expression of important regulatory genes in Arabidopsis . Nucleic Acids Res 2012; 40: 2454–2469.
Wang BB, Brendel V . Genomewide comparative analysis of alternative splicing in plants. Proc Natl Acad Sci USA 2006; 103: 7175–7180.
Reddy AS, Marquez Y, Kalyna M, Barta A . Complexity of the alternative splicing landscape in plants. Plant Cell 2013; 25: 3657–3683.
Blackman BK, Strasburg JL, Raduski AR, Michaels SD, Rieseberg LH . The role of recently derived FT paralogs in sunflower domestication. Curr Biol 2010; 20: 629–635.
Iida K, Go M . Survey of conserved alternative splicing events of mRNAs encoding SR proteins in land plants. Mol Biol Evol 2006; 23: 1085–1094.
Modrek B, Lee C . A genomic view of alternative splicing. Nat Genet 2002; 30: 13–19.
Xia H, Bi J, Li Y . Identification of alternative 5'/3' splice sites based on the mechanism of splice site competition. Nucleic Acids Res 2006; 34: 6305–6313.
Zhang JZ, Li ZM, Mei L, Yao JL, Hu CG . PtFLC homolog from trifoliate orange (Poncirus trifoliata) is regulated by alternative splicing and experiences seasonal fluctuation in expression level. Planta 2009; 229: 847–859.
Acknowledgements
We thank Dr Yuehui He (Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences) for providing the pENTR4-AtFT promoter vector and seeds of the Arabidopsis ft-10 mutant. This work was supported by funds from the National Natural Science Foundation of China (31372100), ‘Programs of Innovation and Entrepreneurship Talents’ of Jiangsu Province, the Project Funded by the Priority Academic Program Development of Jiangsu Higher Education Institutions.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information for this article can be found on the Horticulture Research website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Mao, Y., Sun, J., Cao, P. et al. Functional analysis of alternative splicing of the FLOWERING LOCUS T orthologous gene in Chrysanthemum morifolium. Hortic Res 3, 16058 (2016). https://doi.org/10.1038/hortres.2016.58
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1038/hortres.2016.58