Abstract
We report a family case of type II early-onset Alzheimer’s disease (AD) inherited over three generations. None of the patients in the family had mutations in the genes believed to be the major risk factors for AD, such as APP, presenilin 1 or 2. Targeted exome sequencing of 249 genes that were previously reported to be associated with AD revealed a rare mutation in hemochromatosis (HFE) gene known to be associated with hemochromotosis. Compared to previous studies, we show that HFE mutation can possess the risk of AD in transferrin-, APOE- and APP-normal patients.
Similar content being viewed by others
Alzheimer’s disease (AD) is a serious brain disorder that appears through memory loss and cognitive impairments. Age is the most prominent risk factor, and the number of affected individuals rises dramatically with an aging population. There are two types of AD: type I—with late onset (65 years and later), and type II—with early onset (before 65). The last one has an autosome-dominant character of inheritance and such cases make <5% of all cases of AD.
Initially four genes have been definitively implicated in the etiology of AD. Mutations of the genes encoding β-amyloid precursor protein (APP) and presenilin 1 and 2 (PSEN1, PSEN2) cause a rare, type II form of the disease,1,2 though some studies reported mutations in these genes in families with late-onset AD. APOE gene (allele 4) has been established as a susceptibility gene of the type I form of AD.3,4
Multiple genome-wide association studies (GWAS) suggested that mutations in genes such as ARSB, CAND1, GRN, MAPT, MICA, PLAU, PICALM and many others may also affect AD development.5 Currently the AD database includes >500 genes that could be associated with the risk of AD development;6 however, for almost half of them the data are contradictive and come to different conclusions.7
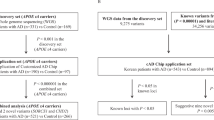
In this study we tried to reveal the genetic variation that caused AD in a Russian family by analyzing 249 genes that demonstrated positive association with AD according to the AlzGene database.6 Russian family N from Oryol region is characterized by appearance of AD cases in at least three generations. It is known that a woman from the second generation of this family, as well as her mother, suffered from a severe form of dementia and died at the age of 72. One of her sons and daughter also died from AD at the age of 79 and 78, respectively; two other sons were diagnosed with mild cognitive impairment (MCI). None of the fourth-generation family members have AD or MCI yet (Figure 1 and Supplementary Table S1).
Genealogy tree of the studied family. Rectangles indicate male individuals, circles indicate female individuals. Symbols representing the patients diagnosed with AD are filled with black, symbols representing the patients diagnosed with MCI are filled with gray. The code names are shown for the patients analyzed in the study.
Two hundred and forty-nine Alzheimer-associated genes were chosen for the analysis (Supplementary Information). Capturing was performed using the SureSelect approach involving the coding regions of the genes surrounded by 100-bp margins, 5′ promoter regions (up to 1,500 bp upstream of the transcription start site) and 3′ noncoding regions (up to 1,500 bp downstream of the mRNA polyadenylation site). Synthesis of biotinylated RNA oligos complementary to selected regions of human genome was ordered from Agilent (Agilent Technologies, Santa Clara, CA, USA).
SureSelect Target Enrichment Cupturing was performed according to the standard protocol. All seven samples were sequenced with SOLiD 4 platform and four samples (members of the third generation) were sequenced with SOLiD 4 and Illumina GAIIx platforms. Fifty-base pair reads were generated on both platforms.
The sequencing data generated by GAIIx were mapped to NCBI37 reference human genome (the version used for 1000 Genomes project) with BWA aligner. The sequencing data originating from SOLiD platform were mapped to the same reference genome with Bioscope software.
For SNV- and indel-calling, we performed a standard ‘best-practices’ GATK approach as described in Van der Auwera et al.8 Briefly, we first marked the duplicates potentially originating from the same DNA fragment due to PCR amplification, that is, the reads that appeared to have the same start and end coordinates after mapping. We next performed indel realignment aimed to improve the original mapping by BWA/Bioscope. Both SNVs and indels were called simultaneously by the GATK HaplotypeCaller tool for all samples. The quality of obtained SNVs was assessed by GATK quality recalibration procedure based on the reference databases of confirmed SNVs and control sets of all potential variants. Only the variants that successfully passed the filter were taken for further analysis.
We proved that none of the patients had polymorphisms in APOE, PSEN1, PSEN2 and APP genes, which are known to be the largest risk factors for AD.
In total, 14,819 SNVs and indels passed the GATK quality control procedure and had non-reference genotype in at least one sample according to at least one sequencing platform. As all the diseased patients from the third generation (P3A, P3B, P3C, P3D) were genotyped by both Illumina and SOLiD platforms, only the SNPs having the same genotype according to both sequencing technologies in every patient were considered. Among those we considered 1,403 variants that had non-reference genotype in all the samples diagnosed with AD (P3A, P3B, P3C, P3D, all four representing the third generation of the family). The variants were annotated based on their localization with respect to genes and their influence on protein product (sense, nonsense, missense) with ANNOVAR software.9 We considered only nonsynonymous variants affecting the sequence of the protein and discarded variants in introns, splice sites and promoters that might also be of interest. For the variants found in dbSNP we assigned appropriate rs numbers. The variants that had been previously discovered in 1000 Genomes project were annotated with their frequencies in the European population. The data on clinical significance of SNVs were taken from clinvar and GWAS databases.10,11
Among the 1,403 variants, 167 variants with relatively low (<0.2) frequency in the European population were considered. We next filtered 11 nonsynonymous variants. We did not consider the variants appearing in the genes often harboring deleterious mutations in healthy individuals, like HLA genes. The variants known to be associated with clinical phenotype were considered to be of high interest. The variants of interest were validated with Sanger sequencing. The results of the described annotations are shown in Table 1.
We found a rare SNP in the hemochromatosis (HFE) gene in all diseased patients. The variant is known to be associated with hemochromatosis according to clinvar database. The distribution of genotypes within the studied family corresponded to the AD status of all the patients: particularly, P4B and P4C were diagnosed as healthy and lacked the variant allele. The latest investigations emphasize that mutations in transferrin (TF) and HFE genes involved in iron metabolism may enhance AD development.12,13 It has been reported that mutations in these genes can be associated with iron accumulation in specific brain areas of patients with AD.14 HFE and TF mutations were shown to interact genetically, increasing the risk of AD in patients having both mutations.15
Here we report a case of AD with HFE gene mutated while TF or APOE genes were not affected. The genotypes were checked in the NGS data (HFE and APOE) and validated with Sanger sequencing (HFE, APOE and TF). It was shown recently that HFE C282Y (rs1800562) mutation on its own does not increase the chances of AD: the fraction of AD-affected individuals does not differ significantly between HFE C282Y-positive and wild-type cohorts.15 As HFE mutations were shown to cause AD cooperatively with mutations in TF or APOE, our data suggest that the family might have a different affected gene that increases the risk of AD. The variant might affect a gene out of the scope of our targeted sequencing or appear in a non-coding DNA locus (splice site or promoter). Thus, broader exome sequencing might reveal a variant causing AD cooperatively with the mutation in HFE. As TF is involved in iron transfer, other genes participating in iron transfer, particularly, TF receptor (TFRC) gene, are the most interesting candidates for further sequencing.
References
Rogaev EI, Sherrington R, Rogaeva EA, Levesque G, Ikeda M, Liang Y et al. Familial Alzheimer’s disease in kindreds with missense mutations in a gene on chromosome 1 related to the Alzheimer’s disease type 3 gene. Nature 1995; 376: 775–778.
Cruchaga C, Chakraverty S, Mayo K, Vallania FLM, Mitra RD, Faber K et al., for the NIA-LOAD/NCRAD Family Study Consortium. Rare variants in APP, PSEN1 and PSEN2 increase risk for AD in late-onset Alzheimer’s disease families. PLoS ONE 2012; 7: e31039.
Beecham GW, Martin ER, Li Y-J, Slifer MA, Gilbert JR, Haines JL et al. Genome-wide association study implicates a chromosome 12 risk locus for late-onset Alzheimer disease. Am J Hum Genet 2009; 84: 35–43.
Blom ES, Giedraitis V, Arepalli S, Hamshere ML, Adighibe O, Goate A et al. Further analysis of previously implicated linkage regions for Alzheimer’s disease in affected relative pairs. BMC Med Genet 2009; 10: 122.
Jin SC, Pastor P, Cooper B, Cervantes S, Benitez BA, Razquin C et al. Ibero-American Alzheimer Disease Genetics Group Researchers, Cruchaga C. Pooled-DNA sequencing identifies novel causative variants in PSEN1, GRN and MAPT in a clinical early-onset and familial Alzheimer’s disease Ibero-American cohort. Alzheimers Res Ther 2012; 4: 34.
Bertram L, McQueen MB, Mullin K, Blacker D, Tanzi RE . Systematic meta-analyses of Alzheimer disease genetic association studies: the AlzGene database. Nat Genet 2007; 39: 17–23.
Blomqvist ME-L, Reynolds C, Katzov H, Feuk L, Andreasen N, Bogdanovic N et al. Towards compendia of negative genetic association studies: an example for Alzheimer disease. Hum Genet 2006; 119: 29–37.
Van der Auwera GA, Carneiro MO, Hartl C, Poplin R, del Angel G, Levy-Moonshine A et al. From FastQ data to high-confidence variant calls: The genome analysis toolkit best practices pipeline. Cur Protoc Bioinform. 2002; 43: 11.10.1–11.10.33.
Wang K, Li M, Hakonarson H . ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucl Acids Res 2010; 38: e164–e164.
Landrum MJ, Lee JM, Riley GR, Jang W, Rubinstein WS, Church DM et al. ClinVar: public archive of relationships among sequence variation and human phenotype. Nucl Acids Res 2013; 42: D980–D985.
Welter D, MacArthur J, Morales J, Burdett T, Hall P, Junkins H et al. The NHGRI GWAS Catalog, a curated resource of SNP-trait associations. Nucl Acids Res 2013; 42: D1001–D1006.
Mariani S, Ventriglia M, Simonelli I, Spalletta G, Bucossi S, Siotto M et al. Effects of hemochromatosis and transferrin gene mutations on peripheral iron dyshomeostasis in mild cognitive impairment and Alzheimer’s and Parkinson’s diseases. Front Aging Neurosci 2013; 5: 37.
Percy M, Somerville MJ, Hicks M, Garcia A, Colelli T, Wright E et al. Risk factors for development of dementia in a unique six-year cohort study. I. An exploratory, pilot study of involvement of the E4 allele of apolipoprotein E, mutations of the hemochromatosis-HFE gene, type 2 diabetes, and stroke. J Alzheimers Dis 2014; 38: 907–922.
Castellani RJ, Moreira PI, Perry G, Zhu X . The role of iron as a mediator of oxidative stress in Alzheimer disease. Biofactors 2012; 38: 133–138.
Lehmann DJ, Schuur M, Warden DR, Hammond N, Belbin O, Kölsch H et al. Transferrin and HFE genes interact in Alzheimer’s disease risk: the Epistasis Project. Neurobiol Aging 2012; 33: e1–13.
Acknowledgements
This work was supported by grants 11-04-12183 and 09-04-12206 from the Russian Foundation for Basic Research.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplemental Information for this article can be found on the Human Genome Variation website (http://www.nature.com/hgv)
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Artemov, A., Boulygina, E., Tsygankova, S. et al. Study of Alzheimer family case reveals hemochromotosis-associated HFE mutation. Hum Genome Var 1, 14004 (2014). https://doi.org/10.1038/hgv.2014.4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/hgv.2014.4