Abstract
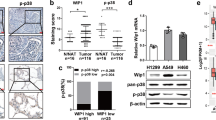
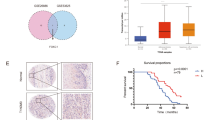
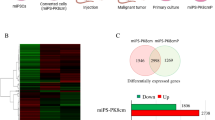
Non-small cell lung cancer (NSCLC) is the leading cause of cancer-related mortalities worldwide, yet this condition remains a poorly understood malignancy, and the subgroup of cancer stem cells (CSCs) leading to therapeutic resistance and adverse prognosis have not been well studied. CSCs frequently undergo symmetric division, which facilitates expansion of the stem cell pool, contributing to long-term relapse and therapy failure. CCAT1 could act as a miRNA sponge to influence downstream genes; however, its roles in NSCLC stem cell are unclear. We first identified activation of Wnt signalling in NSCLC. Analysis of the clinical data from a public database showed a significant decrease of the Wnt signalling repressor Let-7c. Using biological and informatics analyses, we hypothesized that CCAT1 stimulated the main factors of the Wnt signalling pathway, of which the three most deregulated genes were further confirmed by western blotting. Axitinib, a Wnt signalling inhibitor, effectively stimulated asymmetric division, similar to Let-7c. CCAT1 inhibition decreased the ratio of symmetric division of stem cells, and both Let-7c and Axitinib significantly abolished CCAT1 induction of symmetric division by inhibiting Wnt signalling. Restoration of Let-7c blocked the CCAT1 effects, forming the CCAT1/Let-7c/Wnt regulatory axis to control the division of lung cancer stem cells. Stimulation of stem cells to divide asymmetrically by delivering Let-7c or suppressive Axitinib could represent prospective strategies for curing lung cancer patients.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Chen W, Zheng R, Baade PD, Zhang S, Zeng H, Bray F et al. Cancer statistics in China, 2015. CA: A Cancer Journal for Clinicians 2016; 66: 115–132.
Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A . Global cancer statistics, 2012. CA: A Cancer Journal for Clinicians 2015; 65: 87–108.
Ma Z, Cai H, Zhang Y, Chang L, Cui Y . MiR-129-5p inhibits non-small cell lung cancer cell stemness and chemoresistance through targeting DLK1. Biochemical and Biophysical Research Communications 2017; 490: 309–316.
Sun X, Liu J, Xu C, Tang SC, Ren H . The insights of Let-7 miRNAs in oncogenesis and stem cell potency. J Cell Mol Med 2016; 20: 1779–1788.
Sun X, Jiao X, Pestell TG, Fan C, Qin S, Mirabelli E et al. MicroRNAs and cancer stem cells: the sword and the shield. Oncogene 2014; 33: 4967–4977.
Sun X, Xiao G, Zhang B, Meng J, Wang J, Xu C et al. miR-367 stimulates Wnt cascade activation through degrading FBXW7 in NSCLC stem cells. Cell Cycle 2017: 00–00.
Dey-Guha I, Alves CP, Yeh AC, Salony, Sole X, Darp R et al. A Mechanism for Asymmetric Cell Division Resulting in Proliferative Asynchronicity. Molecular Cancer Research 2015; 13: 223–230.
Pang Y, Liu J, Li X, Zhang Y, Zhang B, Zhang J, Du N, Xu C, Liang R, Ren H, Tang S, Sun X . Nano Let-7b sensitization of eliminating esophageal cancer stem-like cells is dependent on blockade of Wnt activation of symmetric division. International Journal of Oncology 2017; 51: 1077–1088.
Qu Y, Gharbi N, Yuan X, Olsen JR, Blicher P, Dalhus B et al. Axitinib blocks Wnt/β-catenin signaling and directs asymmetric cell division in cancer. Proceedings of the National Academy of Sciences 2016; 113: 9339–9344.
Pine SR, Liu W . Asymmetric Cell Division and Template DNA Co-Segregation in Cancer Stem Cells. Frontiers in Oncology 2014; 4: 226.
McHale PT, Lander AD . The Protective Role of Symmetric Stem Cell Division on the Accumulation of Heritable Damage. PLoS Comput Biol 2014; 10: e1003802.
Lerner RG, Petritsch C . A microRNA-operated switch of asymmetric-to-symmetric cancer stem cell divisions. Nat Cell Biol 2014; 16: 212–214.
Hwang W-L, Jiang J-K, Yang S-H, Huang T-S, Lan H-Y, Teng H-W et al. MicroRNA-146a directs the symmetric division of Snail-dominant colorectal cancer stem cells. Nat Cell Biol 2014; 16: 268–280.
Gómez-López S, Lerner RG, Petritsch C . Asymmetric cell division of stem and progenitor cells during homeostasis and cancer. Cell. Mol. Life Sci. 2014; 71: 575–597.
Williams SE, Beronja S, Pasolli HA, Fuchs E . Asymmetric cell divisions promote Notch-dependent epidermal differentiation. Nature 2011; 470: 353–358.
Chen ZZ, Huang L, Wu YH, Zhai WJ, Zhu PP, Gao YF . LncSox4 promotes the self-renewal of liver tumour-initiating cells through Stat3-mediated Sox4 expression. Nat Commun 2016; 7: 12598.
Yang X, Xie X, Xiao Y-F, Xie R, Hu C-J, Tang B et al. The emergence of long non-coding RNAs in the tumorigenesis of hepatocellular carcinoma. Cancer Letters 2015; 360: 119–124.
Meng J, Li P, Zhang Q, Yang Z, Fu S . A four-long non-coding RNA signature in predicting breast cancer survival. J Exp Clin Cancer Res 2014; 33: 84.
Lánczky A, Nagy Á, Bottai G, Munkácsy G, Szabó A, Santarpia L et al. miRpower: a web-tool to validate survival-associated miRNAs utilizing expression data from 2178 breast cancer patients. Breast Cancer Research and Treatment 2016; 160: 439–446.
Szasz AM, Lanczky A, Nagy A, Forster S, Hark K, Green JE et al. Cross-validation of survival associated biomarkers in gastric cancer using transcriptomic data of 1,065 patients. Oncotarget 2016; 7: 49322–49333.
Győrffy B, Bottai G, Lehmann-Che J, Kéri G, Őrfi L, Iwamoto T et al. TP53 mutation-correlated genes predict the risk of tumor relapse and identify MPS1 as a potential therapeutic kinase in TP53-mutated breast cancers. Molecular Oncology 2014; 8: 508–519.
Gao J, Aksoy BA, Dogrusoz U, Dresdner G, Gross B, Sumer SO et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci Signal 2013; 6: pl1.
Cerami E, Gao J, Dogrusoz U, Gross BE, Sumer SO, Aksoy BA et al. The cBio Cancer Genomics Portal: An Open Platform for Exploring Multidimensional Cancer Genomics Data. Cancer Discovery 2012; 2: 401.
Sun X, Xu C, Tang SC, Wang J, Wang H, Wang P et al. Let-7c blocks estrogen-activated Wnt signaling in induction of self-renewal of breast cancer stem cells. Cancer Gene Ther 2016; 23: 83–89.
Xu C, Sun X, Qin S, Zheng Z, Xu S, Luo G et al. Let-7a regulates mammosphere formation capacity through Ras/NF-kappaB and Ras/MAPK/ERK pathway in breast cancer stem cells. Cell Cycle 2015; 14: 1686–1697.
Sun X, Qin S, Fan C, Xu C, Du N, Ren H . Let-7: a regulator of the ERalpha signaling pathway in human breast tumors and breast cancer stem cells. Oncol Rep 2013; 29: 2079–2087.
Ma MZ, Chu BF, Zhang Y, Weng MZ, Qin YY, Gong W et al. Long non-coding RNA CCAT1 promotes gallbladder cancer development via negative modulation of miRNA-218-5p. Cell Death Dis 2015; 6: e1583.
Deng L, Yang S-B, Xu F-F, Zhang J-H . Long noncoding RNA CCAT1 promotes hepatocellular carcinoma progression by functioning as let-7 sponge. Journal of Experimental & Clinical Cancer Research 2015; 34: 18.
Yang J-H, Li J-H, Shao P, Zhou H, Chen Y-Q, Qu L-H . starBase: a database for exploring microRNA–mRNA interaction maps from Argonaute CLIP-Seq and Degradome-Seq data. Nucleic Acids Research 2011; 39 (suppl_1): D202–D209.
Li J-H, Liu S, Zhou H, Qu L-H, Yang J-H . starBase v2.0: decoding miRNA-ceRNA, miRNA-ncRNA and protein–RNA interaction networks from large-scale CLIP-Seq data. Nucleic Acids Research 2014; 42: D92–D97.
He X, Tan X, Wang X, Jin H, Liu L, Ma L . C-Myc-activated long noncoding RNA CCAT1 promotes colon cancer cell proliferation and invasion. Tumour Biol 2014; 35: 12181–12188.
Sun X, Tang S-C, Xu C, Wang C, Qin S, Du N et al. DICER1 regulated let-7 expression levels in p53-induced cancer repression requires cyclin D1. Journal of Cellular and Molecular Medicine 2015; 19: 1357–1365.
Acknowledgements
The authors acknowledge the Center for Translational Medicine of The First Affiliated Hospital of Xi’an Jiaotong University for technical assistance. The authors would like to thank Peijun Liu for assistance with experiments and technical guidance. The present study was financially supported through grants from the National Science Foundation for Young Scientists of China (grant No. 81602597) (Referred to Xin Sun). This work was also supported in part by Technical Research and Development Projects of Shaanxi Province (grant No. 2010K14-03) (Referred to Dapeng Liu), the National Natural Science Foundation of China (grant No. 81272418) (Referred to Hong Ren), and the Natural Science Foundation of Shaanxi Province (grant No. 2016JM8007) (Referred to Jing Zhang).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
All of the co-authors of the present study approved this article for publication. The authors declare that they have no conflicts of interest.
Additional information
Supplementary Information accompanies this paper on Gene Therapy website
Rights and permissions
About this article
Cite this article
Xu, C., Xiao, G., Zhang, B. et al. CCAT1 stimulation of the symmetric division of NSCLC stem cells through activation of the Wnt signalling cascade. Gene Ther 25, 4–12 (2018). https://doi.org/10.1038/gt.2017.98
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gt.2017.98
This article is cited by
-
Enhancement of TKI sensitivity in lung adenocarcinoma through m6A-dependent translational repression of Wnt signaling by circ-FBXW7
Molecular Cancer (2023)
-
A stem cell marker KLF5 regulates CCAT1 via three-dimensional genome structure in colorectal cancer cells
British Journal of Cancer (2022)
-
LncRNA in tumorigenesis of non-small-cell lung cancer: From bench to bedside
Cell Death Discovery (2022)
-
M6A associated TSUC7 inhibition contributed to Erlotinib resistance in lung adenocarcinoma through a notch signaling activation dependent way
Journal of Experimental & Clinical Cancer Research (2021)
-
The immunotherapy candidate TNFSF4 may help the induction of a promising immunological response in breast carcinomas
Scientific Reports (2021)