Abstract
Further development of haematopoietic stem cell (HSC) gene therapy will depend on enhancement of gene transfer safety: ad hoc improvement of vector design relating to each particular disease is thus a crucial issue for HSC gene therapy. We modified a previously described lentiviral vector by adding the Eμmar B-specific enhancer to a human CD19 promoter-derived sequence (Mol Ther 2004;10:45–56). We thus significantly improved the level of expression of the green fluorescent protein (GFP) reporter gene while retaining the specificity of expression in B-cell progeny of transduced human CD34+ progenitor cells obtained from cord blood or adult bone marrow. Indeed, GFP was strongly expressed from early medullary pro-B cells to splenic mature B cells whereas transgene expression remained low in transduced immature progenitors as in myeloid and T-lymphoid progeny retrieved from xenografted NOD/SCID/γcnull mice. Using this lentiviral vector, we further demonstrated the possibility to express a functional human BTK protein in long-term human CD34+ cell B-lymphoid progeny. This newly designed lentiviral vector fulfils one of the pre-requisites for the development of efficient and safe gene therapy for X-linked agammaglobulinaemia, the most common primary humoral immunodeficiency disorder.
Similar content being viewed by others
Introduction
Gene transfer into haematopoietic stem cells (HSC) represents a potential therapeutic approach for several inherited blood diseases. Until now, recombinant oncoretroviral vectors derived from the murine Moloney leukaemia virus have been used to genetically modify HSC.1, 2, 3, 4 However, the strong enhancer activity of native Moloney leukaemia virus sequences driving transgene expression in addition to the recently highlighted integration bias of these vectors towards gene regulatory regions near transcriptional start sites5, 6 are both associated with a major risk of insertional mutagenesis, which may challenge further clinical use of these vectors.7 Lentiviral vectors, mainly derived from human immunodeficiency virus type 1 (HIV-1), initially held attention by their capacity to transduce quiescent cells;8, 9 their ability to efficiently transduce HSC is now well demonstrated and proved superior compared to oncoretroviral vectors.10, 11 Moreover, lentiviral vectors offer more flexibility for inclusion of heterologous sequences like complex regulatory regions that allow tight control of transgene expression throughout haematopoietic differentiation.12 The efficient removal of most viral-coding sequences and native promoters in self-inactivating vectors leads to significant enhancement of gene transfer safety.13 Moreover, the risk of triggering oncogenic processes may be lower with lentiviral vectors since they do not target transcription start site for genomic integration as Moloney leukaemia virus vectors.6, 14 Indeed, HIV-1-derived lentiviral vectors are now moving to the clinic.15, 16
X-linked agammaglobulinaemia (XLA), or Bruton's disease, is a primary humoral deficiency that stands for about 85% of early B-cell defects.17 The disease results from an intramedullary block of B-cell differentiation at the pro-B-cell stage that causes agammaglobulinaemia. Mutations of the BTK gene (for Bruton's tyrosine kinase) was linked to XLA in 1993.18, 19 In mice, absence of functional Btk leads to the X-linked immunodeficiency (Xid) phenotype, which is less severe than human XLA.20, 21 Current treatment for Bruton's disease is based on regular intravenous Ig injections; it generalized since mid 80s and significantly increased the quality of life for XLA patients.22 Nevertheless, frequency of severe and handicapping diseases like disseminated enteroviral infections and chronic pulmonary diseases remains high and is still associated with increased morbidity for XLA individuals.23
As a definitive curative treatment, HSC gene therapy could represent an interesting alternative for XLA patients compared to substitutive intravenous Ig injection therapy.24 An important step towards XLA gene therapy feasibility by HSC gene transfer was recently achieved in the Btk/Tec−/− murine model using a retroviral vector that ubiquitously expressed the human BTK gene.25 Nevertheless, XLA gene therapy approval will strictly depend on improvement of risk–benefit ratio compared to standard intravenous Ig injection therapy; especially, risks of serious adverse effects associated with the use of ubiquitous oncoretroviral vector may not be acceptable in the context of Bruton's disease.
We recently developed a lentiviral vector carrying a promoter sequence derived from the human CD19 gene that allows efficient targeting of green fluorescent protein (GFP) transgene expression to the B-lymphoid progeny of genetically modified human CD34+ haematopoietic progenitors.26 This vector represents an interesting tool for XLA gene therapy; restriction of BTK transgene expression to the B-cell compartment, avoiding potentially detrimental ectopic expression in other lineages, is a rational approach to improve therapeutic safety.24 Moreover, human CD19 promoter-controlled BTK expression was proved safe and efficient to correct phenotypic defects in transgenic Xid mice.27
To improve the level of transgene expression, we looked for an enhancer sequence that could act specifically in B cells and increase CD19 promoter transcriptional activity. The Eμmar sequence comes from the JH–Cμ intronic region of the murine Ig heavy-chain locus where it physiologically acts as a transcriptional enhancer for heavy-chain μ gene expression in B cells.28 It is composed of a central enhancer sequence (Eμ) flanked by matrix attachment regions, which play a crucial role in sequence functionality.29 Indeed, matrix attachment regions act both as an enhancer of gene expression in B-lymphoid cells and as a repressor in other haematopoietic lineages and are notably able to locally remodel chromatin structure.30 These properties result from interactions with activating factors like Bright,31 the expression which is restricted to B cells, or matrix attachment region-BP1,32 and inhibiting factors like SATB1, expressed in non-B cells and especially in T cells,33 or NF-μNR.34 In the context of recombinant lentiviral vectors carrying the cytomegalovirus or phosphoglycerate kinase (PGK) ubiquitous promoters, Eμmar is able to specifically promote strong and position-independent GFP transgene expression in the B-lymphoid lineage.35
This work describes the development of a suitable gene transfer tool for future XLA gene therapy by improvement of the original CD19 promoter-based lentiviral vector by Eμmar sequence addition that allows efficient B-specific expression of the human BTK transgene throughout long-term human haematopoietic differentiation.
Results
B-specific enhancement of GFP expression by Eμmar addition
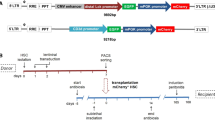
The murine Eμmar enhancer sequence was cloned upstream of the human CD19-derived promoter into our previously described B-specific lentiviral vector (Figure 1).26
Recombinant B-specific lentiviral vectors. Plasmid form of lentiviral vectors is represented. Promoter sequence is composed of the LCD19 fragment alone (L19) or combined with the Eμmar enhancer (EL19) to control expression of eGFP (GFP) or flagged human BTK (fBTK) transgenes. Size of non-viral exogenous sequences included in the original lentiviral backbone (pTRIPΔU3-EF1α-GFP, 3.6 kb) is shown: largest recombinant vector, namely EL19-fBTK, is 7 kb long. GFP, green fluorescent protein; LTR, long terminal repeat; RRE, Rev-responsive element; TRIP, central polypurine tract and central termination sequence (cPPT/CTS); Ψ, encapsidation sequence; ΔU3, deletion of HIV-1 native promoter/enhancer elements (Sin, self-inactivated vector).
Three cell lines of B-lymphoid (Nalm6), T-lymphoid (JA-16) and myeloid (U937) origin were transduced with the EF1-GFP ubiquitous lentiviral vector and with L19-GFP or EL19-GFP lentiviral vectors in parallel. As shown by flow cytometry analysis, the new EL19 composite promoter sequence led to strong and B-cell-specific enhancement of GFP expression intensity (MFI, mean fluorescence intensity) compared to the original L19 promoter (Figure 2a). The MFI per viral copy was about sixfold higher in the B-cell line with EL19 compared to L19 lentiviral vector (Figure 2b). In addition, intensity of transgene expression was more homogenous inside the GFP-positive cell population when the EL19 promoter was used; this is consistent with the previously described Eμmar capacity to allow independent transgene expression from random genomic viral integration sites.35
GFP expression in transduced human haematopoietic cell lines. Cell lines were transduced in parallel with control ubiquitous (EF1-GFP) and B-specific lentiviral vectors (L19-GFP and EL19-GFP). (a) GFP expression was detected 14 days post-transduction by flow cytometry. Transduction efficiency (vector copy number per cell in the whole-cell culture) is indicated in the right top corner of each dot plot. (b) Intensity of GFP expression (MFI) achieved per lentiviral copy in transduced cells. Evaluation of the viral copy number in transduced cells was calculated as follows: copies (in whole culture)/%GFP+ cells. Subsequently, the normalization has been done only in permissive conditions for transgene expression, that is, all EF1-transduced cells and Nalm6 L19- or EL19-transduced cells. GFP, green fluorescent protein; MFI, mean fluorescence intensity; ND, not done.
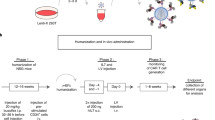
Human CD34+ cells from cord blood (CB) (n=4) or adult bone marrow (BM) (n=1) were transduced with EL19-GFP lentiviral vector and subjected to in vivo differentiation using the NOD/SCID (Non-Obese Diabetic–Severe Combined Immune Deficiency) murine xenograft model. GFP expression in differentiated human B cells (CD19+) and non-B cells (CD19−) recovered from mouse BM and spleen was measured by flow cytometry. GFP was preferentially expressed in B-lymphoid progeny with 46.9±7.1 and 12.1±3% GFP-positive cells for B and non-B cells respectively (Figure 3a). Importantly, the percentage of GFP-positive cells from each animal was always higher in B cells (mean ratio of 5.7±1). In addition, the MFI was significantly higher in B cells compared to non-B cells (591±110 vs 66±16 respectively, see Figure 3b). Results obtained with cells recovered from spleens were similar, though we observed slightly more GFP expression in the non-B-cell population (Figures 3a and b). Conversely, control transduction of CD34+ cells with the EF1-GFP ubiquitous vector produced homogenous GFP expression in all the progenies (Figure 3c). Molecular analyses of sorted B-cell and non-B-cell populations showed similar lentiviral transduction efficiencies, which demonstrated that preferential B-lymphoid GFP expression actually came from selective EL19 transcriptional activity in B cells (Figure 3d). Finally, addition of the Eμmar sequence strongly enhanced the transcriptional activity of the L19 promoter with a 12-fold increase of GFP expression level in BM CD19+ cells compared to the original vector, despite much lower viral genomic integration in CD34+ cells (Supplementary Figure S1 and Moreau et al.26). Intensity of transgene expression was even higher in EL19-GFP-transduced B cells compared to control EF1-GFP-transduced cells (591±110 vs 386±79 MFI respectively, Figure 3c).
GFP expression in human haematopoietic cells differentiated in vivo from EL19-GFP-transduced progenitors. CD34+ cells from cord blood (n=4 independent samples) or adult bone marrow (n=1) were transduced with the EL19-GFP lentiviral vector (4–12 μg p24 per 106 cells) and injected in NOD/SCID mice. Haematopoietic differentiation and GFP expression in human graft (hCD45+) were analysed by flow cytometry 8–12 weeks after CD34+ cell injection into tail vein. (a) Percentage of GFP-expressing cells in B-cell (CD19+) and non-B-cell (CD19−) populations recovered from mice bone marrow or spleen. Paired dots correspond to the parallel analysis of both cell populations from one animal. Analysis of GFP expression in splenic CD19− cells could not be performed for three mice since they virtually lacked such cells. Total number of analysed mice is indicated for each cell population. Mean values are represented by horizontal black lines. (b) Mean values±s.e. of normalized MFI (as defined in Materials and methods) in B-cell (CD19+) and non-B-cell (CD19−) GFP-positive populations retrieved from mice bone marrow or spleen. (c) Flow cytometry dot plots from two representative mice engrafted with EL19-GFP- or EF1-GFP-transduced CD34+ progenitors. (d) Transduction efficiency with EL19-GFP lentiviral vector in CD34+ progenitors and in vivo-differentiated progeny. Average lentiviral copy number per cell (±s.e.) was determined by quantitative PCR on total genomic DNA from whole CD34+ cells 7 days post-transduction and sorted population (CD19+ B cells and CD19− non-B cells) retrieved from mice haematopoietic organs 8–12 weeks after CD34+ cell injection. Number of total analysed animals from the five independent transduction protocols appears in brackets. (e) GFP expression along B-lymphoid differentiation. Flow cytometry analysis of a representative animal is shown. Five sub-populations of growing maturity were defined as followed: non-B progenitors (1, CD19−/CD34+), pro-B (2, CD19+/CD34+), pre-B and immature/mature B cells (3, CD19+/CD34−) in bone marrow; pre-B (4, CD19+/IgM−), immature/mature B cells (5, CD19+/IgM+) in spleen. GFP, green fluorescent protein; MFI, mean fluorescence intensity. Statistics: P<0.05 (*) by the nonparametric Wilcoxon signed ranks tests.
EL19 sequence promoter activity through B-lymphoid differentiation steps was then analysed by defining three sub-populations of growing maturity according to CD34 and IgM surface expression within total CD19+ B cells: the early pro-B (CD34+/IgM−), pre-B (CD34−/IgM−), and immature and mature B-cell (CD34−/IgM+) stages (Figure 3e). The percentage of GFP-positive cells was stable throughout B-differentiation steps, while intensity of GFP expression went up significantly from pro-B to pre-B cells in BM and remained high in mature splenic B cells (Figure 3e and Table 1). In contrast, percentage of GFP-positive cells and transgene expression intensity was significantly lower in CD34+ immature progenitors (Table 1).
Results from NOD/SCID mice transplanted with transduced CD34+ progenitors showed that transcriptional activity of the composite EL19 sequence was weak in the CD19-negative progeny. This population was mainly composed of CD33+ myeloid cells, in addition to few CD34+ immature progenitors in the BM (data not shown). We also conducted two independent experiments in NOD/SCID/γcnull mice36 that efficiently support thymic T-lymphocyte development. Transcriptional activity of the EL19 sequence was very low in the CD2+ T-cell progeny of CB- or BM-transduced progenitors retrieved from murine thymuses. Indeed, dim GFP expression was detected only in some rare T cells (Figure 4) while they were efficiently transduced (Figure 3d). Additional in vitro cultures also showed very low promoter activity of the EL19 sequence in myeloid, dendritic and erythroid lineages differentiated from transduced haematopoietic progenitors (Supplementary Figure S2).
GFP expression in T cells differentiated in NOD/SCID/γcnull mice thymus from EL19-GFP-transduced progenitors. CD34+ cells from cord blood (n=1) or adult bone marrow (n=1) were transduced with EL19-GFP lentiviral vector (8 and 10 μg P24 per 106 cells, respectively) and GFP expression was analysed by flow cytometry in human haematopoietic cells retrieved from NOD/SCID/γcnull mice thymus 10–12 weeks after CD34+ injection. (a) A representative dot plot from one animal showing B-lymphoid (CD19+) and T-lymphoid (CD2+) composition of human thymic graft. Histograms represent GFP expression in the CD2+ and CD19+ populations: level of GFP expression in negative control (untransduced cells) is shown with dotted lines. (b) Mean percentage (±s.e.) of GFP-positive cells (white bars) and average MFI (±s.e.) (black bars) detected within T-cell (CD2+) and non-T-cell (CD2−) populations from four analysed animals (two animals from each protocol). GFP, green fluorescent protein; MFI, mean fluorescence intensity.
Efficient human BTK expression with enhanced B-specific lentiviral vectors
We replaced the GFP sequence by the Flag-tagged human BTK (fBTK) into L19- and EL19-GFP vectors (Figure 1).
Haematopoietic cell lines of different origins were transduced in parallel with the L19- and EL19-fBTK lentiviral vectors as well as with the ubiquitous EF1-fBTK vector as positive control. Sixteen days after transduction, transduction efficiency was checked by quantitative PCR on genomic DNA and transgenic fBTK protein expression was analysed by immunoblotting; as expected, fBTK was detected in all cell types when the EF1-fBTK vector was used (Figure 5a). In contrast, expression of fBTK was specifically induced in B-cell lines with the EL19-fBTK vector, while L19-fBTK transduction did not result in efficient transgenic protein expression even in B cells. Since BTK is activated by tyrosine phosphorylation upon B-cell receptor (BCR) ligation,37 fBTK-transduced Raji B cells were stimulated by IgM antibodies. Following fBTK immunoprecipitation, an antiphosphotyrosine immunoblot was performed showing that fBTK could be tyrosine phosphorylated after 5–10 min of BCR cross-linking (Figure 5b). Only low level of fBTK tyrosine phosphorylation was observed in the context of EL19-fBTK transduction; since it resulted in weak transgenic protein expression, low phosphorylation efficiency could come from competition with the endogenous BTK protein. Nevertheless, as clearly shown with EF1-fBTK transduction, addition of the Flag sequence did not impair BTK protein membrane recruitment at the signalosome upon BCR stimulation. Altogether, these data show that the EL19-fBTK lentiviral vector induces a specific and functional expression of human Btk in B-cell lines.
fBTK protein expression in transduced human haematopoietic cell lines. Human haematopoietic cell lines were transduced (0.7 μg p24 per 106 cells) with ubiquitous (EF1) or B-specific (L19, EL19) lentiviral vectors carrying the fBTK transgene, or kept non-transduced (NT). (a) Sixteen days after transduction, transgenic protein expression (76 kDa) was detected by anti-Flag immunoblotting; anti-Grb2 polyclonal antibodies were used on the same membranes as control for protein loading. Lentiviral copy number per cell in the whole population, as an indication of transduction efficiency, was determined by quantitative PCR on genomic DNA and is shown in italics under each lane. (b) Representative immunoblot of fBTK tyrosine phosphorylation after BCR cross-linking of transduced Raji cells (n=3). Raji cells were transduced in parallel with EF1- and EL19-fBTK lentiviral vectors: transduction efficiencies were measured at 0.2 vector copy per cell in both conditions. BCR cross-linking was performed in the transduced Raji cell line with anti-human IgM antibodies and stimulation stopped after 2, 5 or 10 min. Non-stimulated (NS) and pervanadate treated (PV) cells were used in parallel as phosphorylation controls. Tyrosine phosphorylation levels of immunoprecipitated fBTK transgenic protein was analysed by immunoblotting using 4G10 monoclonal antibody (pY). BCR, B-cell receptor; fBTK, Flag-tagged human BTK.
The EL19-fBTK lentiviral vector was subsequently used to transduce human CB CD34+ haematopoietic progenitors, and fBTK transgene expression was followed through long-term in vivo differentiation in NOD/SCID mice. The first transduction experiment was performed with high lentiviral vector titre (12 μg P24 per 106 cells). As expected, it led to multiple viral genomic integrations within differentiated cells as measured by quantitative PCR in sorted populations: 6.3±1.8 and 5.3±0.5 lentiviral copies per cell were detected in BM B cells and non-B cells respectively (Figure 6a, experiment 1). Quantitative reverse transcriptase-PCR (RT-PCR) analyses in the same two populations showed an efficient and preferential fBTK transgene expression in medullary and splenic B-lymphoid progeny: 2.7±0.8 and 1.1±0.3 transgene messengers per endogenous BTK messenger copy were detected, compared to 0.5±0.1 and 0.3±0.0 copies in non-B cells (Figure 6b, experiment 1).
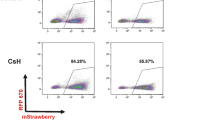
fBTK expression in human haematopoietic cells differentiated in vivo from EL19-fBTK-transduced progenitors. CD34+ cells from CB were transduced with the EL19-fBTK lentiviral vector and subjected to in vivo differentiation in NOD/SCID mice. Lentiviral transduction was done with 12 μg p24 per 106 CD34+ cells in the first experiment and with 3 μg p24 per 106 CD34+ cells in the next three ones (subsequently pooled for analysis). Haematopoietic differentiation and fBTK transgene expression in human graft (hCD45+) were studied by flow cytometry, cellular and molecular analyses 8–12 weeks after CD34+ cell injection into mouse tail vein. (a) Transduction efficiency. Mean values±s.e. of lentiviral copies per cell in sorted populations: haematopoietic progenitors injected into mice (input CD34+, grey bars), B cells (CD19+, white bars) and non-B cells (CD19−, black bars) retrieved from engrafted mice BM or Sp. Total number of analysed mice for each cell sub-population is indicated in brackets. (b) fBTK transgene messenger expression. Mean values (±s.e.) of fBTK transgene cDNA copies per endogenous BTK cDNA copy measured by real-time quantitative RT-PCR (see Materials and methods) in sorted populations as described above. Total number of analysed mice for each cell sub-population is indicated in brackets. Statistics: P=0.125 (*) by the non-parametric Wilcoxon signed rank test. (c) fBTK transgene protein expression. Flow cytometry-sorted human cells from mouse BM (CD19+ B cells/CD19− non-B cells) were analysed for fBTK protein expression by immunofluorescence after fixation on glass slides and intracellular labelling with an anti-Flag monoclonal antibodies (see Materials and methods). Representative microscopic fields from two independent protocols (low viral titre transductions) initiated with BM- or CB-transduced CD34+ cells are shown. BM, bone marrow; CB, cord blood; fBTK, Flag-tagged human BTK; sp, spleen.
A second set of experiments (experiments 2–4) was performed with lower lentiviral titres (3 μg p24 per 106 cells) to minimize viral genomic integrations. On average, we obtained less than 2 vector copies per cell in differentiated B cells and non-B cells: 1.3±0.3 and 1.4±0.2 copies respectively in BM (Figure 6a). In these conditions, fBTK messenger expression appeared weak and below the endogenous BTK level when molecular analysis of CD19-sorted populations was used (Figure 6b). However, since we were not able to determine transduced cell ratio among sorted cells, such global molecular analysis probably underestimates transgene expression in transduced cells. Eventually, fBTK transgenic protein was efficiently and preferentially detected in differentiated B cells by immunoflurorescence on glass slides (Figure 6c).
We analysed human haematopoietic engraftment features after injection of EL19-fBTK-transduced CD34+ progenitors into NOD/SCID mice. Even in high transduction conditions (experiment 1), composition of medullary and splenic human grafts was not modified by fBTK expression; in particular, no significant effect was noticed on B-lymphoid differentiation (Supplementary Figure S3).
Discussion
We have developed a B-specific HIV-1-derived lentiviral vector that generates strong preferential long-term transgene expression throughout B-lymphoid differentiation of genetically modified human haematopoietic progenitors. The recombinant expression cassette is based on association of our previously described minimal human CD19-derived promoter26 with the murine Eμmar B-specific enhancer sequence. Several viral vectors targeting transgene expression to the B-cell compartment have been described concomitantly with our work: a Moloney leukaemia virus γ-retroviral encompassing a similar human CD19 promoter-derived sequence,38 lentiviral vectors carrying first a combination of Eμmar and ubiquitous promoters,35 or more recently a minimal promoter derived from the murine Ig-kappa gene associated with 3′ and 5′ enhancers from the murine Ig heavy-chain locus (the latter corresponds to the Eμmar sequence).39 However, none of these recombinant vectors have been evaluated for the expression of another transgene than GFP.
Compared to our original vector encompassing the minimal CD19 promoter alone, upstream addition of the Eμmar sequence finally results in a 12-fold increase of GFP transgene expression in B-lymphoid cells even with fewer lentiviral integrations per cell. In the meantime, B-preferential expression of the transgene is not impaired; it remains significantly lower to absent in haematopoietic precursors, myeloid, erythroid, dendritic and T-lymphoid cell progenies. If small percentages of GFP-positive cells are still usually detected in non-B cells, the level of transgene expression remains weak in these populations compared to CD19+ B cells; importantly, GFP-expression intensity differential between B cells and non-B cells is significantly more marked with the EL19 sequence compared to the L19 promoter alone.26 Residual long terminal repeat promoter activity of self-inactivating lentiviral vectors could also significantly contribute to the unspecific-background GFP expression detected in non-B cells.40
We further demonstrate that enhanced B-specific recombinant lentiviral vector allows efficient long-term BTK transgene expression in transduced human CD34+ B-cell progeny. In high transduction efficiency conditions, strong and preferential fBTK transgene expression was readily detected in medullar and splenic B cells compared to non-B-cell counterparts, mainly myeloid cells. With limited viral integrations per cell, transgenic BTK protein expression in the B-cell progeny was detected by immunofluorescence. However, using RT-PCR analysis, only low fBTK transgene expression was measured that remains below endogenous level; in addition, B specificity seemed to be absent in these conditions. However, it is likely that fBTK transgene expression was underestimated here, taking into account our global molecular analysis strategy that works on sorted populations that are composed of a mix of transduced and non-transduced cells. Moreover, the observed low B specificity of fBTK transgene expression does not reflect high transcriptional activity of the EL19-fBTK expression cassette in non-B cells, since endogenous BTK level, taken as a reference, is about fourfold lower in latter compared to B cells (data not shown).
BTK is widely expressed in haematopoietic cells, mainly in myeloid and B-lymphoid lineages, but is notably absent from T-lymphoid and plasma cells.41 BTK is involved in multiple signalling pathways and takes part in regulation of various critical biological processes inside haematopoietic cells, including proliferation, differentiation and apoptosis.42, 43 In the context of the development of XLA gene therapy, transcriptional targeting of BTK transgene expression to the B-cell compartment thus appears as a rational strategy to avoid unwanted biological effects that could result from ectopic BTK expression.24 Moreover, since B-cell defect is the main cause of XLA, B-lymphoid-restricted BTK expression should provide full correction of the disease, as already shown with transgenic Xid mice.27 In this context, transgene expression pattern controlled by our B-specific lentiviral vector is well adapted to XLA gene therapy. First, as demonstrated with GFP, transgene expression begins early at the pro-B-cell stage and remains high and constant from pre-B to circulating mature B cells; strong BTK expression beginning at the pre-B cell stage proved notably important for phenotypic correction in Xid mice.44 Moreover, the EL19 promoter sequence does not lead to transgene expression in T-lymphoid progeny; transgene expression in plasma cells could not be analysed as such cells do not differentiate in NOD/SCID mice, and CD19-negative terminally differentiated plasma cells are hardly obtained with existing culture systems.45, 46 Analysis of fBTK transgene expression pattern could not be conducted as precisely as with the GFP transgene; nevertheless, it is unlikely that significant modification of EL19 promoter activity arises from fBTK cDNA substitution in place of the GFP reporter gene.
Until now, therapeutic potential and safety of BTK gene complementation were mainly evaluated in murine models.24 However, BTK involvement in signalling pathways could vary between humans and mice, as reported for TLR signalling in monocytes47 and suggested by major differences in XLA and Xid phenotypes. Therefore, the induction of BTK transgenic expression in human cells is the main concern regarding development of XLA gene therapy. Designing XLA gene therapy pre-clinical studies with human cells currently remains limited by the poor availability of HSC-enriched primary XLA samples and the need for pertinent experimental models. Only one study has reported that the transduction with BTK-expressing vaccinia virus into XLA-derived B-cell lines permits the recovering of BCR-induced Ca2+ mobilization.48
X-linked agammaglobulinaemia represents a delicate situation in which the development of gene therapy as an alternative to currently available substitute treatment will necessitate a much improved risk–benefit ratio, when compared to most diseases for which it has been attempted, including X-SCID, HIV disease or various forms of cancers. In this context, the achievement of efficiently targeted BTK expression through haematopoietic differentiation to the B-lymphoid lineage with our improved lentiviral vector represents the first step towards this goal. Nevertheless, in the present form of our lentiviral vector, the strong enhancer activity of Eμmar still represents a potential risk of random endogenous gene activation through viral genomic integration. In clinical perspectives, further vector development should include the use of insulator sequences to achieve confinement of the enhancer activity.16 Moreover, the use of tumour-prone animal models could be useful to assess the mutagenic potential of this lentiviral vector.14 Finally, the development of biological assays and models that would allow evaluation of BTK gene-transfer efficiency and safety directly in human XLA HSC will be critical to progress towards safe XLA gene therapy.
Materials and methods
Primary cells and cell lines
Cord blood and adult BM samples were obtained after individual informed consent for research use from our own institute or external biological resource centre (EFS, Besançon, France). Samples were used as fresh material or after thawing and overnight culture in RPMI, 10% fetal calf serum, 1% penicillin/streptomycin supplemented with DNAseI (100 U ml−1; Roche Applied Science, Mannheim, Germany). CD34+ cells were subsequently enriched from CB and BM mononuclear fractions by immuno-selection as previously described.26
Human haematopoietic cell lines of B-lymphoid (Nalm6, Raji), myeloid (K562, U937) and T-lymphoid (Jurkat subclone, JA16) origins were cultivated in RPMI medium (Cambrex, Viviers, Belgium) with 10% fetal calf serum (Gibco/Invitrogen, Paisley, UK) and 1% penicillin/streptomycin (100 U ml−1; Invitrogen). Human 293T epithelial cells were cultivated in Dulbecco's modified Eagle's medium (DMEM; Cambrex) with 10% fetal calf serum and 1% penicillin/streptomycin.
In vitro and in vivo haematopoietic differentiation assays
B-lymphoid and myeloid parallel differentiation was performed on murine MS5 stromal cell line as previously described.26 For dendritic differentiation, 2 × 104 CD34+ cells per well (24-well plate) were cultured for 2 weeks (with ¼ changeover after 1 week) in RPMI, 1% penicillin/streptomycin, 10% fetal calf serum, supplemented with human recombinant stem cell factor (20 ng ml−1 a gift from Amgen, Thousand Oaks, CA, USA), granulocyte-macrophage colony-stimulating factor (GM-CSF, 100 ng ml−1; Leucomax, Novartis, Rueil Malmaison, France), TNFα (2.5 ng ml−1) and Flt3-L (20 ng ml−1). The protocol for erythroid differentiation was adapted from Giarratana et al.49
NOD/LtSz-scid mice and NOD/SCID/γcnull mice were initially obtained from Charles River Laboratories (L'Arbresle, France). Experiments and animal care are performed in accordance with institutional guidelines. Transplantation of human CD34+ cells was performed as in previously published work,26 except the absence of treatment with anti-CD122 antibody for NOD/SCID/γcnull mice.
Flow cytometry analyses
Expression of GFP gene and phenotypic markers of human haematopoietic differentiation in cell lines and primary cells differentiated in vitro and in vivo were analysed by flow cytometry using an LSR2 cytometer and FacsDiva software (Becton Dickinson Immunocytometry Systems (BDIS), San Jose, CA, USA). Cell sub-population sorting was performed with an Aria cytometer (BDIS).
A large panel of fluorochrome-conjugated murine monoclonal antibodies (from Becton Dickinson/Bioscience, San Jose, CA, USA or Beckman Coulter/Immunotech, Miami, FL, USA) directed against human haematopoietic lineage markers was used in combination. Four different antigens in addition to GFP expression could be analysed on a single cell using in parallel R-phycoerythrin-, phycoerythrin-cyanin5-, allophycocyanin- and phycoerythrin-cyanin7)-conjugated antibodies. Dead cells were excluded by gating out 4′,6-diamidino-2-phenylindole (Molecular Probes, Eugene, OR, USA)-positive elements and further analyses were done gating on human CD45-positive cells.
Construction and production of recombinant lentiviral vectors
All lentiviral constructions were derived from the initial self-inactivating lentiviral vector backbone TripΔU3-EF1α-GFP (a kind gift of Dr Charneau, Institut Pasteur, Paris, France)50 that was modified by insertion of the LCD19 promoter sequence (L19) in place of the EF1α ubiquitous promoter as previously described.26 Original L19-GFP lentiviral vector was next modified by addition of the Eμmar murine enhancer sequence at the 5′ MluI restriction site directly upstream of LCD19 promoter (EL19-GFP vector, Figure 1). The Eμmar fragment (1023 bp) was generated by PCR from mouse fibroblast genomic DNA (blood quick pure; Macherey Nagel, Hoerdt, France) with the following oligonucleotidic primers (MluI recognition sequences are underlined): Eμmar-Fo (5′-gttacgcgttagagaggtctggtggagcct) and Eμmar-Re (5′-gttacgcgtccaaccagcatgttcaactga); correct orientation of Eμmar was checked by PCR after cloning of MluI into viral vector. Flag-BTK cDNA (fBTK) was obtained by RT-PCR on total RNA (nucleospin RNAII; Macherey Nagel) from Nalm6 pre-B cell line using FlagBTK-Fo (5′-ggatcc atggattacaaggacgacgatgacaaggatatcgccgcagtgattctgga, BamHI restriction site is underlined and Flag sequence showed in italics) and BTK-Re (5′-ctcgagtcaggattcttcatccatgacatcta, XhoI restriction site is underlined) primers. Generated fragment (2010 bp) was cloned (TA cloning kit; Invitrogen) and sequence integrity (Genome Express, Meylan, France) verified against Ensembl reference BTK sequence (ENST00000308731). The fBTK cDNA was ultimately inserted in place of the GFP gene between BamHI and XhoI restriction sites; lentiviral constructions were subsequently named L19-fBTK and EL19-fBTK according to upstream promoter sequence (Figure 1). Reliability of regulating and coding regions was finally checked comparatively to reference sequences.
Lentiviral vector production and transduction into cell lines or primary cells were performed as described in Moreau et al.26 However, lentiviral vector titres were calculated by determining HIV-1 p24 protein level in concentrated viral supernatants (Alliance HIV-1 P24 Antigen ELISA Kit; Perkin Elmer, Waltham, MA, USA) that were subsequently used at less than 1 or 3–12 μg p24 per 106 cells for cell line and primary cells respectively.
Analysis of transduction efficiency and transgene expression
Transduction efficiency was measured by quantitative PCR on total genomic DNA (nucleospin RNAII/DNA buffer set; Macherey Nagel) from cell cultures or sorted human cells engrafted in NOD/SCID mice. Samples were analysed in duplicate for endogenous EPO-R gene (Forward (Fo): 5′-ctgctgccagctttgagtacacta, Reverse (Re): 5′-gagatgccagagtcagataccacaa, Probe: 5′-accccagctcccagctcttgcgt) and lentiviral RRE sequence (Fo: 5′-ccttgggttcttgggagca, Re: 5′-gcctgtaccgtcagcgtcat, Probe: 5′-aggaagcactatgggcgcagcg) parallel quantification and reaction performed as described in Moreau et al.26
Quantitative RT-PCR was performed to detect fBTK transgene expression. Real-time PCR was done with cDNA from total cell RNA using a LightCycler apparatus and SYBRgreen fluorescent reagent (LightCycler 2.0/fast-start SYBRgreen; Roche) with the following primers: FLAG-Fo 5′-atggattacaaggacgacgatgac and BTK-Re 5′-ggaaccactgtttcaacacaagtg (236 bp fBTK amplification); BTKfull-Fo 5′-tcccttatcccttccaggt and BTKfull-Re 5′-aatttggcagcccatagc (209 bp fBTK plus endogenous BTK amplification); GAPDH-Fo 5′-gtcatccctgagctagacgg and GAPDH-Re 5′-gggtcttactccttggaggc (356 bp GAPDH amplification); GFP-Fo 5′-gcaccatcttcttcaaggacga and GFP-Re 5′-cttgatgccgttcttctgc (200 bp GFP amplification). Parallel amplification of plasmid standards (101–106 targeted cDNA copies) was used for quantification of cDNA samples. PCR specificity was controlled by determining the size of amplicons and checking the absence of fBTK or GFP amplification in non-transduced samples. Ratio of transgenic fBtk relative to endogenous BTK expression was determined by quantification of fBTK and total BTK (transgene+endogenous) cDNA in the same sample.
Green fluorescent protein expression was detected by flow cytometry and intensity of expression was quantified by MFI in positive cells: expressed values are normalized MFI that correspond to increasing factor of fluorescence intensity in GFP-positive transduced cells compared to GFP-negative control cells (=MFI(GFP+)/MFI(GFP−)).
Transgenic fBTK protein expression was detected in haematopoietic cell lines by western blot analysis on total protein lysate (106 cells) using a rabbit anti-Flag polyclonal antibody (F7425; Sigma Aldrich, St Louis, MO, USA) and secondary anti-rabbit horseradish peroxidase-coupled antibody. fBTK protein was also detected by immunoflurescence (IF) with a mouse anti-Flag monoclonal antibody (clone M2; Sigma Aldrich) and secondary labelling with Alexa Fluor 488-coupled anti-mouse IgG antibody (Invitrogen).
Unexpectedly, intracytoplasmic fBtk protein could neither be detected by flow cytometry, even in high-expressing cell lines, after cell permeabilization and direct labelling with allophycocyanin-coupled monoclonal anti-Flag antibody (surelight allophycocyanin; Perkin Elmer) nor by indirect labelling with primary biotinylated anti-Flag monoclonal antibody (clone M2; Sigma Aldrich) and various fluorescent secondary reagents (allophycocyanin-coupled anti-mouse antibody or streptavidin).
Analysis of fBTK tyrosine phosphorylation
B-cell receptor-positive B-cell line (Raji, Burkitt's lymphoma) was transduced with EF1-fBTK or EL19-fBTK lentiviral vectors (1 μg p24 per 106 cells). After 7 days, BCR is triggered by sheep anti-human IgM polyclonal antibodies (5 μg per 107 cells; Upstate, Dundee, Great Britain) at 37 °C; pervanadate treatment (100 mM for 5 min) was used as positive phosphorylation control. Transgenic fBTK protein was subsequently immunoprecipitated from whole-cell lysate (1% NP40 lysis buffer, 1 mM orthovanadate) using a mouse anti-Flag monoclonal antibody (clone M2; Sigma Aldrich) and protein G-agarose beads (fast flow; Sigma Aldrich). Tyrosine phosphorylation was detected by western blot with an antiphosphotyrosine monoclonal antibody (clone 4G10; Upstate); immunoprecipitated fBtk was detected with an anti-BTK polyclonal antibody (Btk/c-20; SantaCruz Biotechnology, Santa Cruz, CA, USA).
Conflict of interest
The authors declare no competing financial interests.
References
Hacein-Bey-Abina S, Le Deist F, Carlier F, Bouneaud C, Hue C, De Villartay JP et al. Sustained correction of X-linked severe combined immunodeficiency by ex vivo gene therapy. N Engl J Med 2002; 346: 1185–1193.
Gaspar HB, Parsley KL, Howe S, King D, Gilmour KC, Sinclair J et al. Gene therapy of X-linked severe combined immunodeficiency by use of a pseudotyped gammaretroviral vector. Lancet 2004; 364: 2181–2187.
Aiuti A, Slavin S, Aker M, Ficara F, Deola S, Mortellaro A et al. Correction of ADA-SCID by stem cell gene therapy combined with nonmyeloablative conditioning. Science 2002; 296: 2410–2413.
Ott MG, Schmidt M, Schwarzwaelder K, Stein S, Siler U, Koehl U et al. Correction of X-linked chronic granulomatous disease by gene therapy, augmented by insertional activation of MDS1-EVI1, PRDM16 or SETBP1. Nat Med 2006; 12: 401–409.
Wu X, Li Y, Crise B, Burgess SM . Transcription start regions in the human genome are favored targets for MLV integration. Science 2003; 300: 1749–1751.
Hematti P, Hong BK, Ferguson C, Adler R, Hanawa H, Sellers S et al. Distinct genomic integration of MLV and SIV vectors in primate hematopoietic stem and progenitor cells. PLoS Biol 2004; 2: 2183–2190.
Cavazzana-Calvo M, Fischer A . Gene therapy for severe combined immunodeficiency: are we there yet? J Clin Invest 2007; 117: 1456–1465.
Naldini L, Blomer U, Gallay P, Ory D, Mulligan R, Gage FH et al. In vivo gene delivery and stable transduction of nondividing cells by a lentiviral vector. Science 1996; 272: 263–267.
Reiser J, Harmison G, Kluepfel-Stahl S, Brady RO, Karlsson S, Schubert M . Transduction of nondividing cells using pseudotyped defective high-titer HIV type 1 particles. Proc Natl Acad Sci USA 1996; 93: 15266–15271.
Uchida N, Sutton RE, Friera AM, He D, Reitsma MJ, Chang WC et al. HIV, but not murine leukemia virus, vectors mediate high efficiency gene transfer into freshly isolated G0/G1 human hematopoietic stem cells. Proc Natl Acad Sci USA 1998; 95: 11939–11944.
Case SS, Price MA, Jordan CT, Yu XJ, Wang L, Bauer G et al. Stable transduction of quiescent CD34(+)CD38(−) human hematopoietic cells by HIV-1-based lentiviral vectors. Proc Natl Acad Sci USA 1999; 96: 2988–2993.
Logan AC, Lutzko C, Kohn DB . Advances in lentiviral vector design for gene-modification of hematopoietic stem cells. Curr Opin Biotechnol 2002; 13: 429–436.
Miyoshi H, Blomer U, Takahashi M, Gage FH, Verma IM . Development of a self-inactivating lentivirus vector. J Virol 1998; 72: 8150–8157.
Montini E, Cesana D, Schmidt M, Sanvito F, Ponzoni M, Bartholomae C et al. Hematopoietic stem cell gene transfer in a tumor-prone mouse model uncovers low genotoxicity of lentiviral vector integration. Nat Biotechnol 2006; 24: 687–696.
Levine BL, Humeau LM, Boyer J, Macgregor RR, Rebello T, Lu X et al. Gene transfer in humans using a conditionally replicating lentiviral vector. Proc Natl Acad Sci USA 2006; 103: 17372–17377.
Bank A, Dorazio R, Leboulch P . A phase I/II clinical trial of beta-globin gene therapy for beta-thalassemia. Ann NY Acad Sci 2005; 1054: 308–316.
Conley ME, Broides A, Hernandez-Trujillo V, Howard V, Kanegane H, Miyawaki T et al. Genetic analysis of patients with defects in early B-cell development. Immunol Rev 2005; 203: 216–234.
Vetrie D, Vorechovsky I, Sideras P, Holland J, Davies A, Flinter F et al. The gene involved in X-linked agammaglobulinaemia is a member of the src family of protein-tyrosine kinases. Nature 1993; 361: 226–233.
Tsukada S, Saffran DC, Rawlings DJ, Parolini O, Allen RC, Klisak I et al. Deficient expression of a B cell cytoplasmic tyrosine kinase in human X-linked agammaglobulinemia. Cell 1993; 72: 279–290.
Thomas JD, Sideras P, Smith CI, Vorechovsky I, Chapman V, Paul WE . Colocalization of X-linked agammaglobulinemia and X-linked immunodeficiency genes. Science 1993; 261: 355–358.
Rawlings DJ, Saffran DC, Tsukada S, Largaespada DA, Grimaldi JC, Cohen L et al. Mutation of unique region of Bruton's tyrosine kinase in immunodeficient XID mice. Science 1993; 261: 358–361.
Howard V, Greene JM, Pahwa S, Winkelstein JA, Boyle JM, Kocak M et al. The health status and quality of life of adults with X-linked agammaglobulinemia. Clin Immunol 2006; 118: 201–208.
Winkelstein JA, Marino MC, Lederman HM, Jones SM, Sullivan K, Burks AW et al. X-linked agammaglobulinemia: report on a United States registry of 201 patients. Medicine (Baltimore) 2006; 85: 193–202.
Moreau T, Calmels B, Barlogis V, Michel G, Tonnelle C, Chabannon C . Potential application of gene therapy to X-linked agammaglobulinemia. Curr Gene Ther 2007; 7: 284–294.
Yu PW, Tabuchi RS, Kato RM, Astrakhan A, Humblet-Baron S, Kipp K et al. Sustained correction of B-cell development and function in a murine model of X-linked agammaglobulinemia (XLA) using retroviral-mediated gene transfer. Blood 2004; 104: 1281–1290.
Moreau T, Bardin F, Imbert J, Chabannon C, Tonnelle C . Restriction of transgene expression to the B-lymphoid progeny of human lentivirally transduced CD34+ cells. Mol Ther 2004; 10: 45–56.
Maas A, Dingjan GM, Grosveld F, Hendriks RW . Early arrest in B cell development in transgenic mice that express the E41K Bruton's tyrosine kinase mutant under the control of the CD19 promoter region. J Immunol 1999; 162: 6526–6533.
Forrester WC, van Genderen C, Jenuwein T, Grosschedl R . Dependence of enhancer-mediated transcription of the immunoglobulin mu gene on nuclear matrix attachment regions. Science 1994; 265: 1221–1225.
Oancea AE, Berru M, Shulman MJ . Expression of the (recombinant) endogenous immunoglobulin heavy-chain locus requires the intronic matrix attachment regions. Mol Cell Biol 1997; 17: 2658–2668.
Jenuwein T, Forrester WC, Fernandez-Herrero LA, Laible G, Dull M, Grosschedl R . Extension of chromatin accessibility by nuclear matrix attachment regions. Nature 1997; 385: 269–272.
Herrscher RF, Kaplan MH, Lelsz DL, Das C, Scheuermann R, Tucker PW . The immunoglobulin heavy-chain matrix-associating regions are bound by Bright: a B cell-specific trans-activator that describes a new DNA-binding protein family. Genes Dev 1995; 9: 3067–3082.
Zong RT, Scheuermann RH . Mutually exclusive interaction of a novel matrix attachment region binding protein and the NF-muNR enhancer repressor. Implications for regulation of immunoglobulin heavy chain expression. J Biol Chem 1995; 270: 24010–24018.
Dickinson LA, Joh T, Kohwi Y, Kohwi-Shigematsu T . A tissue-specific MAR/SAR DNA-binding protein with unusual binding site recognition. Cell 1992; 70: 631–645.
Scheuermann RH, Chen U . A developmental-specific factor binds to suppressor sites flanking the immunoglobulin heavy-chain enhancer. Genes Dev 1989; 3: 1255–1266.
Lutzko C, Senadheera D, Skelton D, Petersen D, Kohn DB . Lentivirus vectors incorporating the immunoglobulin heavy chain enhancer and matrix attachment regions provide position-independent expression in B lymphocytes. J Virol 2003; 77: 7341–7351.
Ishikawa F, Yasukawa M, Lyons B, Yoshida S, Miyamoto T, Yoshimoto G et al. Development of functional human blood and immune systems in NOD/SCID/IL2 receptor {gamma} chain(null) mice. Blood 2005; 106: 1565–1573.
Aoki Y, Isselbacher KJ, Pillai S . Bruton tyrosine kinase is tyrosine phosphorylated and activated in pre-B lymphocytes and receptor-ligated B cells. Proc Natl Acad Sci USA 1994; 91: 10606–10609.
Werner M, Kraunus J, Baum C, Brocker T . B-cell-specific transgene expression using a self-inactivating retroviral vector with human CD19 promoter and viral post-transcriptional regulatory element. Gene Therapy 2004; 11: 992–1000.
Laurie KL, Blundell MP, Baxendale HE, Howe SJ, Sinclair J, Qasim W et al. Cell-specific and efficient expression in mouse and human B cells by a novel hybrid immunoglobulin promoter in a lentiviral vector. Gene Therapy 2007; 14: 1623–1631.
Logan AC, Haas DL, Kafri T, Kohn DB . Integrated self-inactivating lentiviral vectors produce full-length genomic transcripts competent for encapsidation and integration. J Virol 2004; 78: 8421–8436.
Smith CI, Baskin B, Humire-Greiff P, Zhou JN, Olsson PG, Maniar HS et al. Expression of Bruton's agammaglobulinemia tyrosine kinase gene, BTK, is selectively down-regulated in T lymphocytes and plasma cells. J Immunol 1994; 152: 557–565.
Yang WC, Collette Y, Nunes JA, Olive D . Tec kinases: a family with multiple roles in immunity. Immunity 2000; 12: 373–382.
Lindvall JM, Blomberg KE, Valiaho J, Vargas L, Heinonen JE, Berglof A et al. Bruton's tyrosine kinase: cell biology, sequence conservation, mutation spectrum, siRNA modifications, and expression profiling. Immunol Rev 2005; 203: 200–215.
Drabek D, Raguz S, De Wit TP, Dingjan GM, Savelkoul HF, Grosveld F et al. Correction of the X-linked immunodeficiency phenotype by transgenic expression of human Bruton tyrosine kinase under the control of the class II major histocompatibility complex Ea locus control region. Proc Natl Acad Sci USA 1997; 94: 610–615.
Kikuchi K, Lian ZX, He XS, Ansari AA, Ishibashi M, Miyakawa H et al. Appearance of human plasma cells following differentiation of human B cells in NOD/SCID mouse spleen. Clin Dev Immunol 2003; 10: 197–202.
Tarte K, De Vos J, Thykjaer T, Zhan F, Fiol G, Costes V et al. Generation of polyclonal plasmablasts from peripheral blood B cells: a normal counterpart of malignant plasmablasts. Blood 2002; 100: 1113–1122.
Perez de Diego R, Lopez-Granados E, Pozo M, Rodriguez C, Sabina P, Ferreira A et al. Bruton's tyrosine kinase is not essential for LPS-induced activation of human monocytes. J Allergy Clin Immunol 2006; 117: 1462–1469.
Fluckiger AC, Li Z, Kato RM, Wahl MI, Ochs HD, Longnecker R et al. Btk/Tec kinases regulate sustained increases in intracellular Ca2+ following B-cell receptor activation. EMBO J 1998; 17: 1973–1985.
Giarratana MC, Kobari L, Lapillonne H, Chalmers D, Kiger L, Cynober T et al. Ex vivo generation of fully mature human red blood cells from hematopoietic stem cells. Nat Biotechnol 2005; 23: 69–74.
Sirven A, Ravet E, Charneau P, Zennou V, Coulombel L, Guetard D et al. Enhanced transgene expression in cord blood CD34(+)-derived hematopoietic cells, including developing T cells and NOD/SCID mouse repopulating cells, following transduction with modified trip lentiviral vectors. Mol Ther 2001; 3: 438–448.
Acknowledgements
This work was supported in part by Institut Paoli-Calmettes, by Association Française contre les Myopathies (AFM) and by Inserm (Centre d'Investigations Cliniques en Biothérapie de Marseille). TM is a recipient of a fellowship from Ministère de l'Education Nationale de l'Enseignement Supérieur et de la Recherche, and from Association Française contre les Myopathies (AFM). JAN is supported by an Institut National du Cancer (INCa) Grant PL-06-026. We thank all the staff at the Beauregard and Bouchard maternity hospitals and at the Cell Therapy Facility and the cord blood bank for collection and access to cord blood samples. We thank Anne Marie Imbert, Marilyne Dijon and Gwladys Zabouo for permanent discussions and advices. We also thank Patrick Gibier and Jean Christophe Orsoni at the UMR599 campus animal facility; Claude Alzieux and Claude Constant at the IPC Radiation Therapy department; Jean-Rémy Galindo and Marc Barad for flow cytometry cell sorting at UMR599 and CIML.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on Gene Therapy website (http://www.nature.com/gt)
Rights and permissions
About this article
Cite this article
Moreau, T., Barlogis, V., Bardin, F. et al. Development of an enhanced B-specific lentiviral vector expressing BTK: a tool for gene therapy of XLA. Gene Ther 15, 942–952 (2008). https://doi.org/10.1038/gt.2008.17
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gt.2008.17
Keywords
This article is cited by
-
Gene transfer into hematopoietic stem cells as treatment for primary immunodeficiency diseases
International Journal of Hematology (2014)
-
Engineering humanized mice for improved hematopoietic reconstitution
Cellular & Molecular Immunology (2012)
-
Development of B-lineage Predominant Lentiviral Vectors for Use in Genetic Therapies for B Cell Disorders
Molecular Therapy (2011)
-
Correction of B-cell development in Btk-deficient mice using lentiviral vectors with codon-optimized human BTK
Leukemia (2010)