Abstract
Purpose: We report a male child who presented at 15 months with perianal abscesses and proctitis, progressing to transmural pancolitis with colocutaneous fistulae, consistent with a Crohn disease-like illness. The age and severity of the presentation suggested an underlying immune defect; however, despite comprehensive clinical evaluation, we were unable to arrive at a definitive diagnosis, thereby restricting clinical management.
Methods: We sought to identify the causative mutation(s) through exome sequencing to provide the necessary additional information required for clinical management.
Results: After sequencing, we identified 16,124 variants. Subsequent analysis identified a novel, hemizygous missense mutation in the X-linked inhibitor of apoptosis gene, substituting a tyrosine for a highly conserved and functionally important cysteine. X-linked inhibitor of apoptosis was not previously associated with Crohn disease but has a central role in the proinflammatory response and bacterial sensing through the NOD signaling pathway. The mutation was confirmed by Sanger sequencing in a licensed clinical laboratory. Functional assays demonstrated an increased susceptibility to activation-induced cell death and defective responsiveness to NOD2 ligands, consistent with loss of normal X-linked inhibitor of apoptosis protein function in apoptosis and NOD2 signaling.
Conclusions: Based on this medical history, genetic and functional data, the child was diagnosed as having an X-linked inhibitor of apoptosis deficiency. Based on this finding, an allogeneic hematopoietic progenitor cell transplant was performed to prevent the development of life-threatening hemophagocytic lymphohistiocytosis, in concordance with the recommended treatment for X-linked inhibitor of apoptosis deficiency. At >42 days posttransplant, the child was able to eat and drink, and there has been no recurrence of gastrointestinal disease, suggesting this mutation also drove the gastrointestinal disease. This report describes the identification of a novel cause of inflammatory bowel disease. Equally importantly, it demonstrates the power of exome sequencing to render a molecular diagnosis in an individual patient in the setting of a novel disease, after all standard diagnoses were exhausted, and illustrates how this technology can be used in a clinical setting.
Similar content being viewed by others
Main
Over the last year, a number of publications have reported the use of exome or genome sequencing in patients.1–6 Most of these studies made use of disease cohorts or families and do not report functional assays or a change in treatment. We report the use of whole exome sequencing to reach a clinical diagnosis and alter treatment in a single child with a life-threatening but previously undefined form of inflammatory bowel disease (IBD) (AHC [OMIM# 266600]).7
The patient is a male who initially presented at 15 months with poor weight gain and a perianal abscess. The abscess enlarged, drained spontaneously, but failed to close despite several rounds of oral, and then parenteral, antibiotics. He subsequently developed diarrhea and weight loss, despite supplemental enteral feedings, and his condition continued to deteriorate over a period of 6 months, with referral to our hospital at 30 months. He had a weight of 8.1 kg, length 81.2 cm, and body mass index of 12.7 (all <3 percentile), indicating severe stunting and malnutrition. Examination under anesthesia showed perineal fistulae and deep fissures. Initial endoscopy showed a rectal stricture and linear ulcers of the rectum; the sigmoid colon and proximal bowel were healthy. Biopsy showed focal active proctitis with ulceration. The child was treated with nasoenteric feeds and infiximab for a presumptive diagnosis of Crohn disease.
Despite treatment, the perineal fistulae persisted, and new ones developed threatening the scrotum. A diverting sigmoid colostomy was performed to divert fecal material and facilitate fistulae closure. The colostomy and mucus fistula failed to incorporate, and new fistulae developed. Although the perianal fistula and the mucosa of the defunctionalized distal limb recovered, the afferent limb became inflamed, eventually involving the entire colon, but not the terminal ileum or upper gastrointestinal (GI) tract. The patient was started on long-term total parenteral nutrition using a peripherally inserted central line. Within 6 weeks, the child developed bacterial sepsis, necessitating a 4-week intensive care unit course involving vasopressors and ventilatory support.
After months of local care, the colostomy healed while he was maintained on adalimumab, intravenous immunoglobulin, and intermittently on granulocyte colony-stimulating factor and prednisone. The child was lost to follow-up for 5 months, while treated at another medical center, but returned at 4 years of age, with malnutrition (9 kg, <3 percentile for age) and breakdown of the abdominal wall around the wound. New colocutaneous fistulae developed just above the level of the fascia with stool running between the fascia and the skin, creating a complex and life-threatening wound (Fig. 1A). Wound care was performed under general anesthesia on a daily basis. Endoscopy through the sigmoidostomy revealed severe mucosal inflammation with linear ulcerations (Fig. 1B). Axial imaging revealed persistence of transmural inflammation involving the entire colon.
Clinical presentation. A, Anterior abdomen showing the expanded abdominal wall defect containing colostomy and mucus fistula (arrow). Note formation of multiple new enterocutaneous fistulae (arrowheads). B, Endoscopic appearance of the colon before colectomy showing a large ulcer occupying approximately one third of the lumenal surface. C, Terminal ileum 3 weeks postcolectomy, normal mucosa. D, Terminal ileum during second postcolectomy flare. E, Terminal ileum 3 weeks after panel D, after initiation of bowel rest, nonabsorbed antibiotics, and tacrolimus.
After 4 months of hospitalization for nutritional optimization and complex wound care, the child underwent a total abdominal colectomy and end ileostomy. The mucus fistula was stapled off to create a Hartmann's pouch. Operative findings showed transverse colon fistulization through the muscles of the abdominal wall. Pathology revealed markedly active chronic colitis with large stretches of mucosal ulceration, granulation tissue, active cryptitis and occasional crypt abscesses, architectural crypt distortion, and transmural inflammation. Ileoscopy 3 weeks postcolectomy confirmed that the ileum was normal (Fig. 1B).
Within 6 weeks postcolectomy, there was clinical and radiologic evidence of a fistula developing between the incision and the ileum, which had been previously uninvolved; the ileostomy showed evidence of healing problems. Remission was induced using a combination of complete bowel rest with total parenteral nutrition, bowel decontamination with nonabsorbed antibiotics, and immunosuppression with tacrolimus. The fistula closed within 3 weeks, and the ileostomy sealed along the skin site. Although maintained on this therapy, the patient remained in remission for the next 4 months.
Recognizing that this was not a viable long-term regimen, we elected to treat the child with a high-dose cyclophosphamide regimen that has shown promise in inducing long-term remission of other autoimmune disorders, such as lupus and aplastic anemia.8 The patient did not experience any significant complications from this treatment, except localized varicella zoster. After the white blood cell count recovered from the chemotherapy, the patient received azathioprine for maintenance therapy and was supported by enteral and oral feedings with no parenteral nutrition. For 3 months postcyclophosphamide, he remained symptom-free and grew normally. Then, over the course of 2 weeks, the patient abruptly lost 10% of his body weight and developed severe ileitis with linear ulcerations (Fig. 1C) and recurrence of the enterocutaneous fistula. Bowel rest, gut decontamination, and immunosuppression with tacrolimus induced remission (Fig. 1D). This therapy was kept in place for 7 months, providing the longest period of remission since his initial presentation. Once again, however, this was not a viable long-term proposition.
The rarity of IBD in the first few years of life and the fact that congenital immune deficiency may present with IBD-like illness prompted a detailed immunologic and genetic evaluation for immune defects known to present with IBD.9,10 Immunological testing revealed the presence of antineutrophil antibodies and suboptimal neutrophil chemotaxis but a normal oxidative burst. CD15, CD16, and CD18 were normal. There was decreased natural killer (NK) cell cytotoxicity with reduced NK cell killing and increased T-cell activation by human leukocyte antigen-DR expression but no clinical or laboratory evidence of hemophagocytic lymphohistiocytosis (HLH). T-cell dysregulation was consistently noted with memory skewing of CD4 cells and inverted CD4/CD8 ratios. IL10 signaling was intact. As mentioned previously, a number of congenital immune deficiencies may present with IBD-like illness. Testing (performed both at our institution and at the second opinion location) for known defects including IKBKG/NEMO, NOD2, UNC13D, and PRF1 for familial HLH, FOXP3 for immunodysregulation polyendocrinopathy enteropathy X-linked syndrome, autoimmune lymphoproliferative syndrome panel for autoimmune lymphoproliferative disease, and signaling lymphocytic activation molecule-associated protein for X-linked lymphoproliferative syndrome type 1 (XLP1) was performed; results were normal. Copy number variant analysis identified no mutations. Although it is possible that continued single gene testing (for example, sequencing of known interacting partners of proteins already shown to be associated with immune deficiencies) may have eventually achieved a diagnosis in this child, no data existed to further guide this mode of testing; the team felt that this approach would be not only costly and time consuming but also unlikely to yield any additional information within an effective timeframe.
The hypothesis of an immune defect, the severity of the unchecked disease, and the significant long-term risks of the existing treatment regimen prompted discussion about the appropriateness of immune reconstitution. However, it was clear that the success of such an aggressive approach would be dependent on the exact underlying mechanism. We therefore sought to identify the causative mutation, providing the necessary information for management, through exome sequencing.
MATERIALS AND METHODS
This study was conducted under the oversight of an appropriate constituted institutional review board, and informed consent was obtained from all human subjects. Given the severity of the disease and the inability to make a diagnosis, the institutional review board approved the use of exome sequencing to facilitate a clinical diagnosis.
Exome analysis
Exome capture was performed using 5 μg of the patient's genomic DNA and a Roche Nimblegen HD2 sequence capture array (Roche Nimblegen, Madison, WI). The system targets approximately 180,000 human protein coding exons and 700 miRNA exons. The captured fragments were amplified and sequenced using a Roche 454 GS FLX instrument with Titanium reagents and protocol (Roche 454, Branford, CT). The sequence reads obtained were aligned to the human genome reference sequence (NCBI36/hg18), and variants (nucleotides from the patient that differed from the reference sequence including insertions, deletions, and substitutions) were identified using the Roche 454 gsMapper software tool supplied with the instrument. Additional information on the sequence capture, 454 sequencing, and analysis methodology is provided in the Supplemental Methods section, Supplemental Digital Content 1, http://links.lww.com/GIM/A143.
All gsMapper identified variants were subsequently annotated with information for identification of candidate mutations using an in-house purpose built software application, which derives and displays a variety of data for each variant. Data including the depth of coverage, conservation across species, percentage of reads with the variant, novelty, potential splice site alteration, known disease association, and likelihood that a variant is deleterious to the protein were extracted from reference data sets or computed in bulk for all variants. These data were stored in a custom Oracle database, and a front-end query tool was developed to allow sophisticated querying of the data. Filtering of the variants using a number of queries based on likely modes of inheritance (as outlined in the Results section) was performed to identify candidate mutations. Additional methodology for this variation annotation, including the thresholds used for filtering of variants, is provided in the Supplemental Methods section, Supplemental Digital Content 1, http://links.lww.com/GIM/A143.
Capillary sequencing validation
A small number of identified variants were validated in-house using ABI3730XL automated DNA Sanger sequencing. Additional information on the sequencing and interpretation methodology is provided in the Supplemental Methods section, Supplemental Digital Content 1, http://links.lww.com/GIM/A143. The causative mutation was confirmed by an external CLIA certified laboratory.
Activation-induced cell death
Peripheral blood mononuclear cells (PBMCs) from the patient and control samples were isolated from heparinized blood using Ficoll-hypaque (GE Healthcare, Piscataway, NJ) and resuspended in Roswell Park Memorial Institute supplemented with 1 mM l-glutamine (Invitrogen, Carlsbad, CA), 50 μM 2-ME, 100 U/mL penicillin (Invitrogen), 100 μg/mL streptomycin (Invitrogen), 10 mM sodium pyruvate (Invitrogen), 1 mM minimum essential medium, and 10% fetal calf serum (Hyclone, Logan, UT). Cells were stimulated for 2 days with 5 μg/mL phytohemagglutinin (Sigma-Aldrich Co., St. Louis, MO) followed by culture in media containing 100 U/mL interleukin (IL)-2 for an additional 3–4 days. On Day 7, phytohemagglutinin blasts were harvested and cultured on plates coated with antibodies to CD3 (OKT3). After 24 hours, cells were harvested and stained with Annexin V/propidium iodide (BD Biosciences, San Jose, CA) as per the manufacturer's instructions and then analyzed by flow cytometry.
NOD signaling
PBMCs were unstimulated or treated with IL-1β (10 ng/mL), muramyl dipeptide (20 μg/mL), or Tri-DAP (20 μg/mL) for 24 hours. Supernatants were then harvested and analyzed using an IL-8 enzyme-linked immunosorbent assay (R&D Systems, Minneapolis, MN). Analysis was carried out from two separately collected patient samples and six healthy controls.
RESULTS
The captured and eluted DNA was sequenced with an average read length of 378 nucleotides to an average depth of 34× coverage. In the final mapping, 99.61% of the high-quality sequence reads mapped to the human genome reference (hg18), with 76.16% of all uniquely mapped reads mapping within a region targeted by the NimbleGen exome array. A total of 99.1% of all targeted bases were covered by at least one sequence read, 95.7% were covered by at least five reads, and 85.9% were covered by at least 10 reads. Additional sequencing summary statistics are provided in Results Table, Supplemental Digital Content 2, http://links.lww.com/GIM/A144.
Whole exome sequencing identified 16,124 variants when compared with the human genome reference sequence; 14,597 of these variants had previously been identified, and 1,527 were novel (Table 1, category A). As expected, because of the sequence capture methodology used, the majority of these variants (approximately 99.31%) were located within genes; as the majority of targeted regions were protein coding exons, the majority of these genic variants were protein coding; 7158 of these changes were nonsynonymous (i.e., altering the amino acid encoded by the variant).
Manual inspection of a subset of >2000 variants confirmed approximately 0.65% as likely false positives; the majority of these were polynucleotide tract errors, and the remainder were missassemblies of reads to regions sharing high-sequence identity. Unsurprisingly, the majority of these misassemblies existed either in low-complexity regions or in regions highly conserved among members of protein families. Variants found in a small number of genes selected for their clinical significance were evaluated by Sanger sequencing. In all instances, the base calls were concordant across both technologies.
Table 1, category A, provides a summary of the total numbers and numbers of novel variations broken down by both location and variation class (insertions, deletions, and substitutions). As expected, the majority of variants identified were substitutions; insertions were the least common across all categories. A larger percentage of the novel variants were insertions or deletions rather than substitutions when compared with the previously identified variants. A small number of these nonsynonymous substitutions resulted in the production of stop codons; all homozygous examples were either previously identified or resulted in a stop in a protein commonly found in the population to harbor stop codons (Table 1, category A).
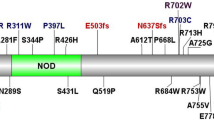
To reduce the search space, we hypothesized, based on the severity and unique clinical presentation, that this case was likely a recessive disorder caused by a hemizygous or homozygous mutation, or compound heterozygote. Sixty-six genes containing potential compound heterozygous mutations (where variants were nonsynonymous and predicted to be damaging by Polyphen, a tool that predicts the impact of an amino acid substitution on the structure and function of a human protein) were identified (Table 1, category B) and investigated; none of these candidates remained after exclusion based on novelty and sequence conservation. Seventy homozygous/hemizygous nonsynonymous variants were identified (Table 1C); eight were novel (when compared against all publicly available data sets) and predicted to be damaging by PolyPhen. Only two of these were highly conserved. One variant, in GSTM1, was excluded because this gene has a high null genotype frequency in the general population.11 The remaining variant was a G to A substitution at a highly conserved position in the X-linked inhibitor of apoptosis gene (XIAP), resulting in a hemizygous cysteine to tyrosine amino acid substitution (p.C203Y; Fig. 2). We did not find this variant at this position in a search of >2000 human control sequences or in orthologous genes from other species down to Drosophila. Confirmation of the variant in the child was carried out, and studies on mother confirmed the mutation (Fig. 3) and showed maternal skewed X-chromosome inactivation in NK, B, and T helper cell types.12
Phylogenetic conservation of the variant amino acid. A multiple sequence alignment of the region of the XIAP protein containing the variant. The human reference sequence is provided on the first line, with the patient's XIAP sequence listed directly below as “Var XIAP.” This is followed by the XIAP sequence from other species. The cysteine is conserved in all species identified. The C-Y substitution is visible in blue at position 231.
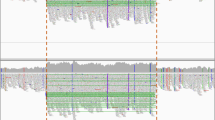
Clinical confirmation in the child and mother. The region of the XIAP gene surrounding the mutation in both the child and the mother was sequenced using the BigDye Terminator Cycle Sequencing kit and analyzed on an ABI3730XL automated DNA sequencer. The Sanger sequence trace from a normal human control is shown at the top. Hemizygosity at the candidate locus is confirmed in the child (middle panel). The mother is heterozygous at this locus (bottom panel).
The XIAP protein has a central role in the proinflammatory response, leading to activation of NFkB and subsequent activation of proinflammatory cytokines via the NOD signaling pathway, as well as a crucial role in mediating programmed cell death.13–15 Functionally testing these pathways in the patient's cells provided results consistent with aberrant XIAP function: muramyl dipeptide stimulation of patient's PBMCs was unable to induce the expression of IL-8 compared with controls (Fig. 4A), and CD3 stimulation of PBMC blasts resulted in enhanced apoptosis in the patient compared with controls (Fig. 4B). Although XIAP was not previously implicated in IBD, a detailed retrospect review of the original description of XLP2, a disorder caused by XIAP mutation, did describe colitis as an associated symptom in two patients.16
Functional validation of the mutation. A, NOD signaling pathway assay. Patient and control PBMCs were unstimulated or treated with IL-1β, MDP, or Tri-DAP, and activation was measured using an IL-8 ELISA. Values are representative data from two separately collected patient samples and six healthy controls. IL-8 production is shown on the y-axis. Data are represented as mean ± the standard error. Significant differences in production are highlighted: ** = P < 0.01, *** = P < 0.001. MDP stimulation of PBMCs from our patient was unable to induce the expression of IL-8 compared with controls. B, Apoptosis assay. Patient and control PBMC PHA blasts were stimulated with CD3 antibody, and activation-induced cell death was analyzed using flow cytometry after staining with Annexin V and propidium iodine. The top two panels are from control cells, with addition of CD3 in the right-hand side panel. The lower two panels are results from patient data, again with the CD3 stimulated cells on the right. CD3 stimulation of PHA blast resulted in enhanced apoptosis in this patient compared with controls. This is representative data from three separate experiments.
Since XIAP deficiency carries a high risk of death due to HLH, irrespective of potential benefit to the bowel disease, a decision was made that a hematopoietic progenitor cell transplant was indicated. With informed consent based on a comprehensive, multidisciplinary review of the risks, benefits, and alternative treatments, at the age of 5 years 8 months, the patient received myeloablative conditioning using busulfan at 0.8 mg/kg/dose × 16 doses with a target steady-state concentration of 800 ng/mL busulfan; fludarabine 40 mg/m2/dose × 4 doses; and equine antithymocyte globulin 30 mg/kg/dose × 3 doses. Graft versus host disease prevention was with tacrolimus and mycophenolic acid. To prevent mucositis, the patient received palifermin before and after the conditioning regimen, according to recommended dosing instructions. The patient received a 6/6 human leukocyte antigen matched cord blood with a total nucleated cell dose of 12e7/kg on day 0.
The patient tolerated the conditioning regimen well and received his hematopoietic progenitor cell infusion without complication. He developed an absolute neutrophil count above 500 for 3 days on day >19; his last red blood cell transfusion was on day >29; and his last platelet transfusion was on day >33. His postengraftment period has been complicated by a steroid-responsive engraftment syndrome/Grade I skin graft versus host disease on day >18 and asymptomatic adenovirus positive blood polymerase chain reaction (PCR) treated with cidofovir on day >20 with improvement in his viral load within a week. On day >27 after the transplant, he exhibited signs of encephalopathy with short-term memory loss initially noticed by his mother and subsequently objectively confirmed by repetitive physical examinations. On that same day, a spinal tap was performed that exhibited a positive PCR for Human Herpesvirus 6 (HHV6). A blood PCR was also positive for HHV6. Magnetic resonance imaging showed the classical hippocampal findings of posttransplant HHV6 encephalopathy. The patient was started on foscarnet on the first day of memory loss; his mental status has subsequently significantly improved and a repeat spinal tap 14 days later was negative for HHV6 by PCR.
The patient started trophic enteral feeds at day >31, started taking oral food at day >42 post-bone marrow transplant, and continues to enjoy a varied diet with no recurrence of his bowel disease. His Lansky score is 100%. Chimerism studies at this same time showed 100% donor chimerism in the CD33 compartment and CD3 compartment.
Prognosis with regard to preventing HLH is well established, but the anticipated effect of hematopoietic transplant on the intestinal disease was less certain. One reason for the lack of predictability is that the XIAP protein is expressed in all tissues and cell types, including the intestinal epithelium. Whether the residual XIAP deficiency in the epithelium will lead to disease recurrence is a matter of some concern, but there are no reports to our knowledge of patients with XIAP deficiency who are developing unusually severe GI tract disease posttransplant.
DISCUSSION
On the basis of data described in this report, this child has been diagnosed as having an XIAP mutation and resulting immunodeficiency. A second opinion by another institution not involved in this project concurred with the diagnosis. Mutations in XIAP are a known cause of XLP2, and although, as mentioned above, colitis has been described in patients with XLP2 mutations, the severe Crohn-like illness described here represents a novel manifestation of this genetic syndrome. Therefore, it is extremely unlikely that this association would have been made in the absence of genomic data. In fact, in our presequencing analysis of the literature, we generated a list of 2006 candidate genes based on the medical literature; XIAP was not identified as a potential candidate.
The length of this list and the fact that XIAP was not on it highlights one of the limitations of using single-gene sequencing testing; where the list of possible candidate genes is large (for example, where symptoms are shared among a number of disorders or in instances where there is atypical presentation), the cost of sequencing gene by gene may well outstrip the cost of whole exome (and shortly the cost of whole genome) sequencing. Furthermore, where single gene tests are ordered one by one, the time taken for this approach to provide actionable data may also exceed the time required for whole exome or genome sequencing and interpretation.
After a review of the findings, including the risks, benefits, and alternative treatments, an allogeneic hematopoietic progenitor cell transplant was recommended based on the mutation found in XIAP and performed in concordance with the suggested treatment for XIAP deficiency. At >42 days posttransplant, the child was able to eat and drink, and there has been no recurrence of GI disease, suggesting this mutation also drove the GI disease.
In addition to directing a course of treatment, our findings may also shed light on the mechanism of IBD pathogenesis in this patient; specifically, the means by which the NOD signaling pathway participates in regulation of the immune response to commensal organisms.17 Recently, NOD-induced NFkB activation has been shown to depend on XIAP via an indirect interaction between BIR2 and the NOD1/2-interacting protein RIP2.15 Given that this patient's mutation lies within this BIR2 domain, it is plausible that it may destabilize the NOD2/RIP2 complex, leading to defective signaling. Loss of NOD2 function can derepress toll-like receptor signaling,18 leading to loss of commensal organisms tolerance.19 Interestingly, XIAP also has been shown to be critical for toll-like receptor-modulated signaling in response to infection.20 The patient's ascending inflammation is consistent with a maladaptive response to commensal organisms. The diagnosis of IBD in a patient with a mutation in XIAP suggests a broader role for this gene in mucosal immune regulation. It is intriguing to note that there is a pronounced male bias in the incidence of Crohn disease in young children, pointing to mutations on the X-chromosome.
This work illustrates the value of exome sequencing in the care of patients, suggesting a model for diagnosis without the benefit of a cohort or informative pedigree. This approach required integration of clinical phenotyping, sequencing, bioinformatics, and functional studies by a multidisciplinary team. As sequencing costs fall and genomic data sets with phenotypes expand, we predict an increasing utilization of sequencing data in a clinical environment. In the interim, there is a need for development of multidisciplinary teams and large clinically useful data sets to fully use this approach as a clinical tool, as well as a need to build the infrastructure to enable this powerful tool to be practically implemented.
REFERENCES
Ng SB, Buckingham KJ, Lee C, et al. Exome sequencing identifies the cause of a mendelian disorder. Nat Genet 2010; 42: 30–35.
Ng SB, Turner EH, Robertson PD, et al. Targeted capture and massively parallel sequencing of 12 human exomes. Nature 2009; 461: 272–276.
Choi M, Scholl UI, Ji W, et al. Genetic diagnosis by whole exome capture and massively parallel DNA sequencing. Proc Natl Acad Sci USA 2009; 106: 19096–19101.
Roach JC, Glusman G, Smit AF, et al. Analysis of genetic inheritance in a family quartet by whole-genome sequencing. Science 2010; 328: 636–639.
Ashley EA, Butte AJ, Wheeler MT, et al. Clinical assessment incorporating a personal genome. Lancet 2010; 375: 1525–1535.
Lupski JR, Reid JG, Gonzaga-Jauregui C, et al. Whole-genome sequencing in a patient with Charcot-Marie-Tooth neuropathy. N Engl J Med 2010; 362: 1181–1191.
Lu Y, Bousvaros A . Healthcare burden of inflammatory bowel disease in the United States: more than pain and diarrhea. Inflamm Bowel Dis 2009; 15: 1767–1768.
Brodsky RA . High-dose cyclophosphamide for autoimmunity and alloimmunity. Immunol Res 2010; 47: 179–184.
Yamamoto-Furusho JK, Podolsky DK . Innate immunity in inflammatory bowel disease. World J Gastroenterol 2007; 13: 5577–5580.
Rahman FZ, Marks DJ, Hayee BH, Smith AM, Bloom SL, Segal AW . Phagocyte dysfunction and inflammatory bowel disease. Inflamm Bowel Dis 2008; 14: 1443–1452.
Lin HJ, Han CY, Bernstein DA, Hsiao W, Lin BK, Hardy S . Ethnic distribution of the glutathione transferase Mu 1–1 (GSTM1) null genotype in 1473 individuals and application to bladder cancer susceptibility. Carcinogenesis 1994; 15: 1077–1081.
Wengler GS, Parolini O, Fiorini M, et al. A PCR-based non-radioactive X-chromosome inactivation assay for genetic counseling in X-linked primary immunodeficiencies. Life Sci 1997; 61: 1405–1411.
Dubrez-Daloz L, Dupoux A, Cartier J . IAPs: more than just inhibitors of apoptosis proteins. Cell Cycle 2008; 7: 1036–1046.
Huang Y, Park YC, Rich RL, Segal D, Myszka DG, Wu H . Structural basis of caspase inhibition by XIAP: differential roles of the linker versus the BIR domain. Cell 2001; 104: 781–790.
Krieg A, Correa RG, Garrison JB, et al. XIAP mediates NOD signaling via interaction with RIP2. Proc Natl Acad Sci USA 2009; 106: 14524–14529.
Rigaud S, Fondanèche MC, Lambert N, et al. XIAP deficiency in humans causes an X-linked lymphoproliferative syndrome. Nature 2006; 444: 110–114.
Baumgart DC, Carding SR . Inflammatory bowel disease: cause and immunobiology. Lancet 2007; 369: 1627–1640.
Richardson WM, Sodhi CP, Russo A, et al. Nucleotide-binding oligomerization domain-2 inhibits toll-like receptor-4 signaling in the intestinal epithelium. Gastroenterology 2010; 139:904–917, e901–e906
Strober W, Kitani A, Fuss I, Asano N, Watanabe T . The molecular basis of NOD2 susceptibility mutations in Crohn's disease. Mucosal Immunol 2008; 1:S5-S9
Bauler LD, Duckett CS, O'Riordan MX . v regulates cytosol-specific innate immunity to Listeria infection. PLoS Pathog 2008; 4: e1000142.
Acknowledgements
This work was funded by the Children's Hospital of Wisconsin Foundation and the Medical College of Wisconsin. In addition, for no fee, Roche 454 Life Sciences carried out the sequence capture, library preparation, and ran the first of the sequencing runs. The authors thank the many devoted clinicians who contributed to the care of this patient, in particular, Dr. Michael Stephens (Pediatric Gastroenterology), Carrie Wachowiak, RN (Special Needs Service), Dr. Subra Kugathasan, and Dr. William Grossman, and the nurses and dieticians of the Pediatric Gastroenterology Section, Center-8, and East-8 units at the Children's Hospital of Wisconsin. The authors would also like to acknowledge the continued support of our Bone Marrow Transplant Program from the Midwest Athletes Against Childhood Cancer (MACC) fund. The authors would also thank donors to the Children's Hospital Foundation and the Foundation for helping to support this work. In addition, the authors would like to acknowledge the Bioinformatics team for development of the variant analysis tools: George Kowalski, Jeff DePons, Wesley Rood, Alexander Stoddard, Bradley Taylor, Greg McQuestion, Kent Brodie, and Stacey Zacher. Finally, the authors would like to acknowledge the family for their willingness to publish this case.
Author information
Authors and Affiliations
Corresponding author
Additional information
Disclosure: The authors declare no conflict of interest.
Supplemental digital content is available for this article. Direct URL citations appear in the printed text and are provided in the HTML and PDF versions of this article on the journal's Web site (www.geneticsinmedicine.org).
Rights and permissions
About this article
Cite this article
Worthey, E., Mayer, A., Syverson, G. et al. Making a definitive diagnosis: Successful clinical application of whole exome sequencing in a child with intractable inflammatory bowel disease. Genet Med 13, 255–262 (2011). https://doi.org/10.1097/GIM.0b013e3182088158
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1097/GIM.0b013e3182088158
Keywords
This article is cited by
-
Understanding the molecular mechanism of pathogenic variants of BIR2 domain in XIAP-deficient inflammatory bowel disease
Scientific Reports (2024)
-
Novel XIAP mutation with early-onset Crohn’s disease complicated with acute heart failure: a case report
BMC Cardiovascular Disorders (2023)
-
Lentiviral Gene Transfer Corrects Immune Abnormalities in XIAP Deficiency
Journal of Clinical Immunology (2023)
-
Using whole exome sequencing to identify susceptibility genes associated with nonsyndromic cleft lip with or without cleft palate
Molecular Genetics and Genomics (2023)
-
Monogenic inflammatory bowel disease-genetic variants, functional mechanisms and personalised medicine in clinical practice
Human Genetics (2023)