Abstract
Purpose
Part of the hidden genetic variation in heterogeneous genetic conditions such as inherited retinal diseases (IRDs) can be explained by copy-number variations (CNVs). Here, we explored the genomic landscape of IRD genes listed in RetNet to identify and prioritize those genes susceptible to CNV formation.
Methods
RetNet genes underwent an assessment of genomic features and of CNV occurrence in the Database of Genomic Variants and literature. CNVs identified in an IRD cohort were characterized using targeted locus amplification (TLA) on extracted genomic DNA.
Results
Exhaustive literature mining revealed 1,345 reported CNVs in 81 different IRD genes. Correlation analysis between rankings of genomic features and CNV occurrence demonstrated the strongest correlation between gene size and CNV occurrence of IRD genes. Moreover, we identified and delineated 30 new CNVs in IRD cases, 13 of which are novel and three of which affect noncoding, putative cis-regulatory regions. Finally, the breakpoints of six complex CNVs were determined using TLA in a hypothesis-neutral manner.
Conclusion
We propose a ranking of CNV-prone IRD genes and demonstrate the efficacy of TLA for the characterization of CNVs on extracted DNA. Finally, this IRD-oriented CNV study can serve as a paradigm for other genetically heterogeneous Mendelian diseases with hidden genetic variation.
Similar content being viewed by others
Introduction
Inherited retinal diseases (IRDs) are characterized by a wide spectrum of nonsyndromic and syndromic phenotypes all involving visual impairment.1 In addition to their clinical heterogeneity, IRDs are characterized by extensive genetic heterogeneity.2
Despite the large number of identified IRD genes, the genetic basis of IRD remains unknown in 20–50% of cases albeit analyzed by whole-exome sequencing (WES).3, 4, 5 Part of the potential missing genetic variation that may be causative for IRD can be attributed to mutations in novel, yet unidentified disease genes.6, 7 It is anticipated that another fraction of unsolved cases can be caused by noncoding sequence variations in known IRD genes, examples of which have been increasingly reported.8, 9, 10
Several studies using different approaches demonstrated that structural variations including copy-number variations (CNVs) represent an additional mutation type significantly contributing to the missing genetic variation in IRD.11, 12, 13, 14 In 2016, Van Cauwenbergh et al. developed a high-resolution array, arrEYE, for targeted CNV analysis of known and candidate IRD genes.15 Despite different algorithms developed for CNV identification using next-generation sequencing (NGS) data,16, 17 most NGS studies still fail to detect them, leading to an underestimation of CNVs as underlying cause of IRD. Although several IRD genes like EYS,18 USH2A,19 and PCDH1520 are known to be prone to CNV formation, little is known about the contribution of CNVs to IRD in general.
Here, we aimed to identify genomic features that are correlated with CNV occurrence in IRD genes, and to propose a ranking of CNV-prone IRD genes. Furthermore, we demonstrated the efficacy of targeted locus amplification (TLA) for the characterization of complex CNVs on extracted DNA.
Materials and methods
Literature mining
RetNet is a summary of genes that have been associated with isolated and syndromic IRD. An extensive literature mining of virtually all reported CNVs in IRD genes listed in RetNet has been performed by manual text mining (Supplementary Table S1 and Supplementary Text S1 online).
Search for CNV-prone IRD genes
The genomic landscape of RetNet genes has been investigated by listing the following genomic features: gene size, number of associated Database of Genomic Variants (DGV) variants,21 total number of associated repeats, Alu repeats, long interspersed nuclear elements (LINE), long terminal repeat elements (LTR), and segmental duplications (SDs), using the UCSC Table Browser (Supplementary Table S2 and Supplementary Text S1).
The rank correlation analysis was performed using the Kendall rank correlation coefficient or tau, which works on ranked, nonparametric data. Calculation of the statistical difference of the values of the different genomic features between the group of genes with one or fewer CNVs and the group of genes with at least two CNVs was performed using a classic t-test. Correlations between the different genomic features were calculated using Pearson correlation coefficients (Supplementary Figure S1 and Supplementary Text S1).
CNV analysis in an IRD cohort
The patient study adhered to the tenets of the Declaration of Helsinki. CNV analysis was performed in a longitudinal way in a diagnostic IRD cohort that underwent different analyses: multiplex ligation-dependent probe amplification (MLPA), arrEYE,15 single-nucleotide polymorphism (SNP) chip analysis, conventional polymerase chain reaction (PCR) to assess nonamplification and real-time PCR (qPCR). Patients P6, P10, and P11 underwent WES prior to CNV screening. In case of a heterozygous CNV in USH2A, the mutation on the second allele was identified by targeted NGS (Supplementary Text S1).
Characterization of identified CNVs
Based on the genomic coordinates obtained from CNV analysis, further delineation of the breakpoint regions of each CNV was performed by designing conventional PCR or qPCR assays in the breakpoint regions for the homozygous and heterozygous CNVs, respectively, and by performing iterative delineation rounds. When both breakpoint regions were sufficiently delineated, junction PCR was performed by conventional PCR, followed by Sanger sequencing of the junction product (Supplementary Text S1).
CNVs with highly repetitive breakpoint regions or for which no specific junction product could be obtained were further delineated using TLA (Cergentis, Utrecht, the Netherlands).22 For each CNV one or more inverse PCR primers (Supplementary Table S3) were designed in a specific anchor region. For a deletion this is the region immediately upstream or downstream of the deletion breakpoints that is not deleted with certainty. For a duplication it is the region close to one of both breakpoints that is duplicated with certainty (Supplementary Text S1).
Assessing the underlying mechanisms of the identified CNVs
For each CNV that has been delineated at the nucleotide level, an extensive bioinformatics analysis was performed on the breakpoint regions to try to unravel the underlying mechanism, as previously described (Supplementary Text S1).23
Assessing the cis-regulatory effect of deleted noncoding elements in the EYS and PCDH15 regions
To identify putative cis-regulatory elements in the EYS and PCDH15 regions, an assay for transposase-accessible chromatin with high-throughput sequencing (ATAC-seq) was performed according to established protocols24 on adult postmortem human retinal tissue (T.J.C., unpublished data). To determine that transposase-accessible regions represented active promoters or enhancers, chromatin immunoprecipitation with high-throughput sequencing (ChIP-Seq) was performed for the H3K27ac and H3K4me2 histone marks and transcription factors CRX, OTX2, NRL, RORB, and MEF2D using previously described protocols.25 RNA-seq was performed using standard methods on RNA extracted from nuclei, to enrich for enhancer RNAs (Supplementary Text S1).26
Human PCDH15-associated CRX-bound region (CBR) 1 and 2 were PCR amplified from a healthy control individual (Supplementary Table S3). CBR1 was cloned in a dsRed expressing vector without basal promoter. CBR2 was cloned in another dsRed expressing vector, upstream of a Rho-basal promoter. DNA cocktails containing the PCDH15 reporter vectors and a pCAG-GFP vector as an electroporation control were coelectroporated into isolated retinas of P0 mice. After 8 days of in vitro organ/tissue culture, retinas were harvested, fixed, and imaged. The detailed protocol used for the electroporation assays has been described previously (Supplementary Text S1).27
Results
Mapping the CNV landscape in IRD genes
First, we attempted to characterize the CNV landscape in IRD by an exhaustive literature mining of CNVs in all RetNet genes.2 This revealed 1,345 patients with a reported CNV in one of the known IRD disease genes, more specifically 317 unique CNVs in 81 different genes (Supplementary Table S1). The discrepancy between the total number of CNVs and the number of unique CNVs can be explained by the presence of common CNVs in certain IRD genes. Supplementary Table S4 lists the 15 top-ranked genes according to the total CNV count, together with the number of unique CNVs in the corresponding gene. For example, CLN3 and NPHP1 have high total but very low unique CNV counts, which can be attributed by a common deletion of exons 7 and 8 of CLN3,28 and a whole-gene deletion of NPHP1.29, 30 Apart from genes with common CNVs, several genes like USH2A, OPA1, and EYS are characterized by many different CNVs, contributing to their high CNV load. Finally, ABCC6 is characterized by both a common exon 23–29 deletion31 and a high number of other less frequent CNVs.
Mapping the genomic landscape of IRD genes
Next, we assessed different genomic features for all RetNet genes (Supplementary Table S2).2 The top-ranked genes according to each of these factors are listed in Table 1, together with the respective number of CNVs reported in literature and DGV. The output of gene size shows that EYS, PCDH15, and USH2A are the largest IRD genes, which is in agreement with the large number of reported CNVs (Supplementary Table S4). While there are large genes for which no disease-related CNVs have been reported up to now, for most of them a significant number of CNVs have been reported in DGV. The other outputs according to the number of total and specific type of repeats in the genomic region are different from the output based on gene size. Although these new top-ranked genes span a smaller genomic region, they contain a higher absolute number of repeats, or are specifically enriched in specific types of repeats.
To assess which of these genomic features could be used to identify and prioritize CNV-prone IRD genes, we first performed a rank correlation analysis, comparing the rank of the genes according to the different genomic features and their respective rank in the literature and DGV lists (Supplementary Figure S1). This revealed that the strongest correlation is obtained for gene size, both according to literature and DGV, followed by LINE and LTR repeats, and total repeats. Importantly, the Alu repeats and SDs do not seem to correlate with the number of CNVs. To confirm these results the different IRD genes were divided into two groups based on the absence or presence of at least two CNVs in a gene according to the DGV list. For each of the features we evaluated whether there was a significant difference between the two groups (Supplementary Figure S1). This analysis confirmed the previous results, showing gene size as the feature with the most significant difference between both groups. There was no significant difference in the number of Alu repeats and SDs. Finally, we investigated if the different studied genomic features are correlated with each other. This revealed strong correlations of LINE and LTR repeats and total repeats with gene size, while both Alu repeats and SDs do not seem to correlate with gene size.
Expanding the CNV mutation spectrum in IRD genes
Apart from the mapping of the CNV spectrum in IRD genes by the literature mining, we expanded the CNV spectrum in IRD by the identification of 30 CNVs in 29 unrelated families, 21 of which are unique and 13 of which are novel (Table 2).
Patients P6, P10, and P11 underwent WES prior to CNV screening. In P6 this led to the identification of an apparently homozygous nonsense mutation in MERTK: c.2323C>T, p.(Arg775*). Segregation analysis demonstrated a recessive carrier state in the mother but not in the biologically proven father, indicating pseudohomozygosity. This was confirmed by qPCR analysis, revealing a heterozygous whole-gene deletion of MERTK in both the index patient and his father (Supplementary Figure S2). In patients P10 and P11 both arrEYE analysis and WES coverage data analysis revealed two apparently identical homozygous deletions of PDE6G (Supplementary Table S5). Comparison of SNP chip data in the region surrounding this deletion revealed an identical haplotype in both patients (Supplementary Table S6).
The main contribution to the CNVs identified in this study comes from the USH2A gene, with 15 CNVs out of 29 patients. For all but one USH2A-mutated patients the CNV is heterozygous and the mutation on the second allele was initially identified by targeted NGS. This highlights the importance of CNV screening in patients carrying only one heterozygous pathogenic mutation in disease-associated genes known to harbor mutations inherited under a recessive paradigm, like USH2A.
Characterization of identified CNVs and assessment of their underlying mechanisms
All but two of the 21 unique CNVs were further characterized to identify the exact breakpoints. In total, we were able to fully characterize 14 out of 19 investigated CNVs at the nucleotide level (Table 2 and Supplementary Figure S3). A conventional delineation strategy based on junction PCR led to the complete characterization of eight CNVs at the nucleotide level (Figure 1), leaving 11 CNVs not fully delineated. The reason for this was the presence of SDs in both breakpoint regions of the MERTK deletion, while for the 10 remaining CNVs there were highly repetitive breakpoint regions. For all but one sample we moved on to TLA to further characterize the CNVs in a hypothesis-neutral manner. This approach allowed the identification of the CNV breakpoints in six out of nine patients (Figure 2), including a more complex CNV in patient P3 (Figure 3).
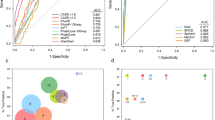
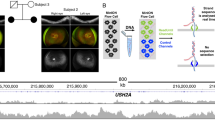
Identification and characterization of a noncoding PCDH15 deletion. (a) Location of the deletion. The homozygous 1.3-Mb deletion identified in P9 is depicted in red, removing the first noncoding exon of the PCDH15 gene, the MTRNR2L5 gene with unknown function and two PCDH15-associated CRX-bound regions (CBR). CBR1 is situated in the promoter region, while CBR2 is located approximately 100 kb upstream. (b) Delineation of the deletion. Further refinement of the deletion breakpoint regions by conventional PCR. Black, orange and blue shaded boxes and connecting lines indicate nondeleted exon and introns, deleted exon, intron and upstream region, and nondeleted upstream region, respectively. Short grey horizontal lines correspond to designed PCR amplicons, used to delineate the deletion. The black dotted line indicates the junction product. (c) Sanger sequencing of the junction product. Delineation of the deletion at the nucleotide level, chr10: g.56478660_57777934del. (d) Electroporation assays. In order to assess the cis-regulatory effect of CBR1 and CBR2, electroporation reporter assays were performed in mouse retinal explants. The first construct consisted of CBR2 cloned in a dsRed expressing vector, upstream of a Rho-basal promoter, as CBR2 is located in a more distant regulatory region. For the second construct, CBR1 was cloned in a dsRed expressing vector without basal promoter, as it is located in the promoter region. Cis-regulatory activity could be demonstrated for the CBR2 construct, while CBR1 seems to fail in driving dsRed expression on its own. (e) Cis-regulatory landscape of PCDH15. Epigenomic marks and transcription factor binding were assessed in human adult retina, shown here for the PCDH15 locus: ATAC-seq; ChIP-seq for H3K27ac, H3K4me2, CRX, OTX2, NRL, RORB and MEF2D; and RNA-seq. The region of the deletion is marked with a shaded rectangle and the putative active promoter and enhancer regions included in the deleted region are in lighter shading.
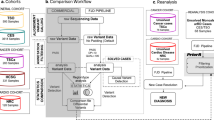
Identification and characterization of a BEST1 deletion using targeted locus amplification (TLA) on extracted genomic DNA. (a) Location of the deletion. The heterozygous 8-kb deletion identified in P1 is depicted in red, removing the first two exons of the BEST1 gene. (b) Primer design TLA. Initial real-time polymerase chain reaction (qPCR) delineation of the deletion was hampered by highly repetitive breakpoint regions, followed by further delineation using TLA. The minimal deleted region based on the qPCR delineation is depicted in red; the gray dotted line corresponds to the 5′ and 3′ breakpoint regions and the surrounding genomic region in black. In case of a deletion, the outward-oriented TLA primers indicated in orange need to be designed in an anchor region that is not deleted, here in the downstream surrounding genomic region. (c) TLA read mapping. The amplified PCR products are sequenced by next-generation sequencing (NGS), followed by mapping of the resulting reads showing one distinct peak surrounding the anchor region. (d) Delineation of the deletion. Detailed analysis of the reads in the breakpoint regions led to the identification of the deletion at nucleotide level: chr11:g.61711373_61719810del. The upper, middle, and lower lines represent the sequences of the 5′ wild-type (wt) region, the deletion junction product, and the 3′ wt region, respectively. Microhomology of 3 bp is indicated in gray. The two boxes show the mapped reads in both breakpoint regions. Green sequences are forward reads; red sequences are reverse reads. The fully colored reads are wt reads, while the reads containing shaded sequences are junction reads spanning the deletion breakpoints. The turquoise and orange boxes highlight the 5′ and 3′ sequences, separated by the 3 bp of microhomology.
Identification and characterization of a complex noncoding EYS deletion using targeted locus amplification (TLA). (a) Location of the deletion. The homozygous 16-kb deletion identified in P3 is depicted in red, located upstream of the EYS gene. (b) Delineation of complex deletion. Due to problems using conventional polymerase chain reaction (PCR) as an initial delineation strategy, TLA was used for further characterization of the deletion at nucleotide level. Analysis of the resulting TLA reads revealed a complex copy-number variation (CNV) consisting of a deletion of chr6: 66,631,318-66,647,641 (hatched region), together with an insertion of a 455-bp fragment consisting of a 277-bp sequence (chr6: 66,628,168-66,628,444) located upstream of the deletion (turquoise region) joined to an inverted 178-bp fragment (chr6: 66,642,009-66,642,186) situated in the deleted region (orange region). The upper part of the figure represents the wt sequence, while the lower part shows the resulting mutated sequence. The underlying mutational mechanism, represented in the middle part, is most probably a replication-based mechanism. The numbers correspond to different template switches, based on the observed microhomology in every breakpoint region. (c) Microhomology at the breakpoints. Comparison of the sequences at the three breakpoint junctions shows microhomology at every junction, supporting microhomology-mediated break-induced replication as the underlying mechanism. (d) Cis-regulatory landscape of EYS. Epigenomic marks and transcription factor binding were assessed in human adult retina, shown here for the upstream EYS region: ATAC-seq; ChIP-seq for H3K27ac, H3K4me2, CRX, OTX2, NRL, RORB and MEF2D; and RNA-seq. The region of the deletion is marked with a shaded rectangle and the putative active promoter and enhancer region included in the deleted region are in lighter shading, for both the deletion presented in this figure (insertions/inversion indicated by black vertical lines) and another regulatory EYS deletion identified in P2. For the latter, both the promoter and an enhancer region coincide with the CNV, whereas for the complex CNV no cis-regulatory epigenomic marks could be seen in the affected region.
In patient P5 conventional mutation screening of KCNV2 by PCR and Sanger sequencing revealed nonamplification of exon 2, suggesting a homozygous deletion. After delineation of the deletion, segregation analysis was performed in the family using the junction PCR primers. Although both children of the index patient are obligate carriers of this deletion, segregation of the junction product could only be demonstrated in one of the children. Subsequent qPCR of exon 2 of KCNV2 in the daughter demonstrated a heterozygous deletion of this exon. The most probable explanation for this finding is the occurrence of two different overlapping heterozygous deletions, only one of which was identified by the initial delineation. This was corroborated using qPCR, revealing a second larger deletion overlapping with the smaller one found in the index case and in the daughter (Supplementary Figure S4).
To unravel the underlying mechanism of the formation of the CNVs identified in this study, a bioinformatics analysis was performed on both breakpoint regions of the 14 molecularly characterized CNVs (Supplementary Table S7). Short stretches of microhomology (1–3 bp) were identified at eight CNV junctions. Three CNVs showed a perfect transition at their junction, accompanied by an insertion of seven nucleotides for one of them. The three remaining CNVs display more complex insertions, which can be explained by multiple iterative template switches based on short stretches of microhomology (Figure 3).32 None of the CNVs has similar repetitive elements in the corresponding breakpoint regions, except for the MERTK deletion, where the presence of SDs points to nonallelic homologous recombination (NAHR) as a surmised underlying mechanism. The presence of previously described structural variation–associated sequence motifs in all studied breakpoint regions can result in genomic instability and facilitate the formation of CNVs.32, 33 Visualization of microhomology and repetitive elements and an overview of the identified sequence motifs can be found in Supplementary Figures S5–S7.
Assessing the effect of noncoding CNVs in the EYS and PCDH15 loci
Three CNVs were found in noncoding regions: two involving the promoter region and the first exon of EYS (P4) and PCDH15 (P9), respectively, previously reported as pathogenic CNVs.20, 34 The third homozygous CNV is located 214 kb upstream of EYS (P3). In order to assess a potential regulatory effect on their neighboring genes, a cis-regulatory data set was evaluated for the PCDH15 and EYS loci in human adult retina. For the promoter deletions of EYS (P4) and PCDH15 (P9), several cis-regulatory elements coincide with the CNVs, likely being disrupted by the deletions and affecting gene expression. For the complex CNV upstream of EYS (P3), no cis-regulatory, epigenomic marks could be identified in the CNV region.
The potential regulatory effect of the large upstream deletion of PCDH15 (P9) was further investigated by electroporation assays in mouse retinal explants. The deleted region harbors two CBRs, one partially overlapping with the first noncoding exon and included in the promoter region of the gene (CBR1), the other one located approximately 100 kb upstream (CBR2) (Figure 1). Therefore, the human CBR1 sequence was cloned immediately upstream of the dsRed reporter gene, while CBR2 was cloned upstream of a dsRed reporter gene carrying a minimal basal promoter. Electroporation of both constructs in mouse retinal explants demonstrated cis-regulatory activity of CBR2, while CBR1 seems to fail in driving dsRed expression on its own (Figure 1).
Discussion
Exploring the CNV and genomic landscape of IRD genes reveals CNV-prone IRD genes
The major aim of this study was to map the genomic landscape of IRD genes in order to prioritize those genes prone to CNVs. We conducted an extensive literature mining of reported CNVs in all known IRD genes listed in RetNet and retrieved 1,345 CNVs in 81 different genes from 300 different publications, more than half of which were published in the past six years.
We investigated genomic features of all known IRD disease genes to find susceptibility factors for CNV formation and to establish rankings of CNV-prone IRD genes (Table 1). The correlation between gene size and number of repeats is reflected by the presence of large genes such as EYS, PCDH15, and USH2A in the different rankings. Interestingly, in the output of the number of associated repeats, several genes such as CLN3, CRX, PNPLA6, and FAM161A can be found at a considerably higher position compared to the output list of gene size, meaning that they are located in genomic regions highly enriched with repeats. Because Alu repeat density has previously been associated with recurrent exonic loss35 and nonrecurrent intragenic deletions,36 this specific type of repeat was also taken into account. Based on this output, genes such as ZNF408, CLUAP1, ROM1, and PRPF8 have the largest increase in rank compared with gene ranking based on total number of associated repeats and have high Alu repeat percentages. As NAHR between low-copy repeats such as SDs is an important mechanism of recurrent genomic rearrangements,32 the presence of SDs was also assessed, resulting in many highly ranked genes not present in the other outputs. As expected, several of these genes, like NPHP1, CA4, BBS4, CIB2, CDHR1, RGR, and CNNM4 overlap with the NAHR-prone regions proposed by Bujakowska et al. in 2016.13 Finally, the occurrence of LINE and LTR repeats was investigated, as they also have the potential to mediate NAHR,37, 38 leading to a list of genes highly similar to the gene size output.
Correlation analysis demonstrated that gene size has the strongest correlation with the presence of a CNV in that specific gene, followed by the number of associated LINE and LTR repeats and the number of total repeats. However, these last two features are both highly correlated with gene size. Although Alu repeats and SDs have been shown to play a role in the formation of CNVs,32 the presence of these repeats does not seem to correlate with the occurrence of CNVs. The absence of a significant association of CNVs with Alu repeats was previously proposed, suggesting that Alu elements do not currently play an important role in the formation of CNVs, while they did so in the past. This is supported by the observation of highly significant colocalization of older SDs, which can be seen as fixed CNVs in the population, with Alu repeats, while this correlation is decreasing rapidly for younger SDs and is totally absent for CNVs. This study also demonstrated that CNVs co-occur with SDs, but that the correlation is much smaller than may be expected, suggesting an important contribution of other mechanisms in the formation of CNVs.39
We correlated the results of the literature mining to the output list of CNV-prone IRD disease genes and found discrepancies for genes for which no associated CNVs could be found in literature, despite a high number of CNVs in DGV. One possible explanation may be that a CNV in an IRD gene leading to loss of function is not necessarily pathogenic, while a dominant pathogenic missense mutation in the same gene causing a gain of function has a pathogenic effect. In addition, some of these genes have been discovered only recently as IRD genes, and are less prevalent in the context of IRD. A last and important reason is that CNV screening has not been routinely implemented, likely leading to an underestimation of the CNV mutational load in these genes. In this respect, the occurrence of CNVs could be expected in ADAMTS18, CERKL, CNGB3, CRB1, DHDDS, FAM161A, and IMPG1, as they are highly ranked as CNV-prone genes according to gene size or total repeats and as mutations in these genes are well known causes of IRD. In order to increase CNV detection in a clinical setting, we recommend performing routine targeted CNV screening in the most prevalent 30 top-ranked IRD genes according to genomic length, such as EYS, PCDH15, USH2A, CDH23, ALMS1, CRB1, CNGB3, and CHM, especially for monoallelic patients in case of autosomal recessive inheritance. The latter statement is corroborated by the high percentage of CNVs (i.e., 30%; 15/50) in the monoallelic USH2A cohort of IRD patients screened in the current study.
Expanding the CNV mutation spectrum in IRD genes
The proposed list of CNV-prone IRD genes was further validated by the identification of 21 different CNVs in 29 families, affecting 10 different genes. The initial CNV analyses in this study were performed in subcohorts with IRD, rather than in one general IRD cohort. Therefore, the denominators for these different subcohorts with IRD are the following: (i) heterozygous CNVs in patients with vitelliform macular dystrophy screened for BEST1 mutations by sequencing of the coding region and MLPA (2/140; 1.4%); (ii) heterozygous CNVs in patients with optic atrophy screened for OPA1 mutations by sequencing of the coding region and MLPA (2/234; 0.43%); (iii) heterozygous CNVs in IRD cases with monoallelic USH2A mutation, identified by MLPA (15/50; 30%); and (iv) homozygous CNVs identified by homozygosity mapping in an IRD cohort of consanguineous origin (6/99; 6%).
It is striking that most CNVs were found in USH2A, which is the third largest IRD gene. Other genes that are highly ranked according to gene size are EYS, MERTK, OPA1, and PCDH15. Moreover, MERTK is located in one of the recently identified NAHR-prone regions.13 PRPH2 and PDE6G are smaller genes, but highly enriched in Alu repeats. CNVs in BEST1, PRPH2, and SPATA7 have previously been described only once,40, 41, 42 and for PDE6G this is the first CNV reported in IRD. To the best of our knowledge, this is the second PDE6G genetic defect reported in IRD after the initial identification as an IRD gene.43
Interestingly, we found two cases of pseudohomozygosity in two different nonconsanguineous families: a homozygous nonsense mutation in MERTK c.2323C>T, p.(Arg775*) in an arRP family and a homozygous exon 2 deletion of KCNV2 in a family with arCRD. In the first family we found a heterozygous whole-gene deletion of MERTK, overlapping with the nonsense mutation. In the second family we showed two different overlapping heterozygous KCNV2 deletions. These cases of pseudohomozygosity illustrate the importance of segregation analysis, CNV screening, and delineation in patients with apparent homozygous mutations without consanguineous background.
Apart from 18 coding CNVs, we identified three CNVs located in noncoding regions. Two of these are a deletion of the first noncoding exon and the associated promoter region of EYS and PCDH15, respectively, most probably affecting transcription by the disruption of retina-specific cis-regulatory elements. This was supported by integration with a cis-regulatory data set generated in human adult retina and by in vivo studies in mouse retinal explants. For the third noncoding deletion there is no evidence for overlap with any of the retina-specific cis-regulatory elements. According to DGV, several CNVs overlap with this region, as is the case for the rest of the EYS gene, including the coding parts. Altogether, although we could not provide evidence for a pathogenic effect of this CNV, an effect on the chromatin conformation of the EYS region and on EYS transcription cannot be excluded.
Characterization of CNVs in a hypothesis-neutral manner and identification of replicative mechanisms as major underlying contributor
As the CNV identification study was performed in a longitudinal setting, different targeted CNV analysis strategies were used, including a customized microarray.15 Several recent studies have reported WES as a tool to identify both homozygous and heterozygous CNVs in IRD.11, 44, 45 Exact delineation of the CNVs using this approach is not possible, however, which can be overcome by the use of WGS.9, 14, 46
Apart from the conventional delineation strategies, we introduced TLA on extracted genomic DNA to characterize CNVs. Indeed, TLA has recently been described as a strategy to selectively amplify and sequence entire genetic loci on the basis of the crosslinking of physically proximal sequences.22 Advantage of the technique is that no detailed prior locus information is needed, as is the case for other targeted approaches. TLA has previously been shown to be effective when applied on living cells, while here, we demonstrated for the first time its efficacy on extracted human DNA.
In addition to the MERTK deletion that arose through NAHR of SDs located in both breakpoint regions, the detailed bioinformatics analyses performed for all delineated CNVs revealed three different groups according to putative underlying molecular mechanisms. The largest group consisting of eight CNVs with short stretches of microhomology may be explained by nonhomologous end joining or a replicative-based repair mechanism like microhomology-mediated break-induced replication, favoring the latter because an information scar, typical of nonhomologous end joining, was absent in all CNVs.32 Remarkably, the formation of the PDE6G CNVs identified in this study was not mediated by the presence of Alu repeats in the breakpoint regions, although this gene has the second highest Alu repeat percentage of all RetNet genes. The second group comprises three CNVs with no reported microhomology, and the presence of an information scar for one of them, pointing towards nonhomologous end joining as underlying mechanism. Finally, the last three CNVs are associated with more complex insertions, characterized by microhomology at all breakpoint junctions and most probably caused by microhomology-mediated break-induced replication, which has been proposed as the major mechanism for nonrecurrent structural variations.32 These results put replicative-based repair mechanisms forward as a major underlying contributor of the CNVs identified in this study.
General conclusion and perspectives
This comprehensive study investigated the role of CNVs in IRD. We mapped and correlated the CNV and genomic landscape of 256 IRD genes, revealing genomic features that point to CNV risk. We expanded the CNV mutation spectrum in IRD genes, characterized the identified CNVs in a hypothesis-neutral manner, and assessed their underlying mechanisms. Our study demonstrates the importance of the CNV mutational load in IRD and contributes to the elucidation of the hidden genetic variation in IRD. It will be useful for the interpretation of CNV data in future genomic studies of IRD. Finally, this IRD-oriented CNV study can be extrapolated to other genetically heterogeneous Mendelian diseases with hidden genetic variation.
Change history
08 October 2018
"The original version of this Article contained an error in the spelling of the author Anja K. Mayer, which was incorrectly given as Anja Kathrin Mayer. This has now been corrected in both the PDF and HTML versions of the Article."
References
den Hollander AI, Black A, Bennett J, Cremers FPM . Lighting a candle in the dark: advances in genetics and gene therapy of recessive retinal dystrophies. J Clin Invest 2010;120:3042–3053.
Daiger SP, Sullivan LS, Bowne SJ, Rossiter BJF. RetNet. Retinal Information Network. 1996. https://sph.uth.edu/retnet/. Last accessed on 3 June 2016.
Lee K, Berg J, Milko L et al. High diagnostic yield of whole exome sequencing in participants with retinal dystrophies in a clinical ophthalmology setting. Am J Ophthalmol 2015;160:354–363.
Beryozkin A, Shevah E, Kimchi A et al. Whole exome sequencing reveals mutations in known retinal disease genes in 33 out of 68 Israeli families with inherited retinopathies. Sci Rep 2015;5:13187.
Weisschuh N, Mayer AK, Strom TM et al. Mutation detection in patients with retinal dystrophies using targeted next generation sequencing. PLoS One. 2016;11:e0145951.
Arno G, Carss KJ, Hull S et al. Biallelic mutation of ARHGEF18, involved in the determination of epithelial apicobasal polarity, causes adult-onset retinal degeneration. Am J Hum Genet 2017;100:334–342.
Arno G, Agrawal SA, Eblimit A et al. Mutations in REEP6 cause autosomal-recessive retinitis pigmentosa. Am J Hum Genet 2016;99:1305–1315.
Coppieters F, Todeschini AL, Fujimaki T et al. Hidden genetic variation in LCA9-associated congenital blindness explained by 5′ UTR mutations and copy-number variations of NMNAT1. Hum Mutat 2015;36:1188–1196.
Small KW, DeLuca AP, Whitmore SS et al. North Carolina macular dystrophy is caused by dysregulation of the retinal transcription factor PRDM13. Ophthalmology 2016;123:9–18.
Liquori A, Vaché C, Baux D et al. Whole USH2A gene sequencing identifies several new deep intronic mutations. Hum Mutat 2016;37:184–193.
Khateb S, Hanany M, Khalaileh A et al. Identification of genomic deletions causing inherited retinal degenerations by coverage analysis of whole exome sequencing data. J Med Genet 2016;53:600–607.
Lindstrand A, Frangakis S, Carvalho CMB et al. Copy-number variation contributes to the mutational load of Bardet-Biedl syndrome. Am J Hum Genet 2016;99:318–336.
Bujakowska KM, Fernandez-Godino R, Place E et al. Copy-number variation is an important contributor to the genetic causality of inherited retinal degenerations. Genet Med 2017;19:643–651.
Carss KJ, Arno G, Erwood M et al. Comprehensive rare variant analysis via whole-genome sequencing to determine the molecular pathology of inherited retinal disease. Am J Hum Genet 2017;100:75–90.
Van Cauwenbergh C, Van Schil K, Cannoodt R et al. arrEYE: a customized platform for high-resolution copy number analysis of coding and noncoding regions of known and candidate retinal dystrophy genes and retinal noncoding RNAs. Genet Med 2017;19:457–466.
Tan R, Wang Y, Kleinstein SE et al. An evaluation of copy number variation detection tools from whole-exome sequencing data. Hum Mutat 2014;35:899–907.
Gambin T, Akdemir Z, Yuan B et al. Homozygous and hemizygous CNV detection from exome sequencing data in a Mendelian disease cohort. Nucl Acids Res 2017;45:1633–1648.
Pieras JI, Barragán I, Borrego S et al. Copy-number variations in EYS: a significant event in the appearance of arRP. Invest Ophthalmol Vis Sci 2011;52:5625–5631.
Steele-Stallard HB, Le Quesne Stabej P, Lenassi E et al. Screening for duplications, deletions and a common intronic mutation detects 35% of second mutations in patients with USH2A monoallelic mutations on Sanger sequencing. Orphanet J Rare Dis 2013;8:122.
Le Guédard S, Faugère V, Malcolm S, Claustres M, Roux A . Large genomic rearrangements within the PCDH15 gene are a significant cause of USH1F syndrome. Mol Vis 2007;13:102–107.
MacDonald JR, Ziman R, Yuen RKC, Feuk L, Scherer SW . The Database of Genomic Variants: a curated collection of structural variation in the human genome. Nucleic Acids Res 2014;42:D986–D992.
de Vree PJP, de Wit E, Yilmaz M et al. Targeted sequencing by proximity ligation for comprehensive variant detection and local haplotyping. Nat Biotechnol 2014;32:1019–1025.
Verdin H, D’haene B, Beysen D et al. Microhomology-mediated mechanisms underlie non-recurrent disease-causing microdeletions of the FOXL2 gene or its regulatory domain. PLoS Genet 2013;9:e1003358.
Buenrostro J, Wu B, Chang H, Greenleaf W . ATAC-seq: a method for assaying chromatin accessibility genome-wide. Curr Protoc Mol Biol 2015;109:21.29. 1-9.
Andzelm M, Cherry T, Harmin D et al. MEF2D drives photoreceptor development through a genome-wide competition for tissue-specific enhancers. Neuron 2015;86:247–263.
Kim T, Hemberg M, Gray JM et al. Widespread transcription at neuronal activity-regulated enhancers. Nature 2010;465:182–187.
Matsuda T, Cepko CL . Electroporation and RNA interference in the rodent retina in vivo and in vitro. Proc Natl Acad Sci USA 2004;101:16–22.
Lerner T, Boustany R, Anderson J et al. Isolation of a novel gene underlying Batten disease, CLN3. Cell 1995;82:949–957.
Caridi G, Murer L, Bellantuono R et al. Renal-retinal syndromes: association of retinal anomalies and recessive nephronophthisis in patients with homozygous deletion of the NPH1 locus. Am J Kidney Dis 1998;32:1059–1062.
Yuan B, Liu P, Gupta A et al. Comparative genomic analyses of the human NPHP1 locus reveal complex genomic architecture and its regional evolution in primates. PLoS Genet 2015;11:e1005686.
Ringpfeil F, Nakano A, Uitto J, Pulkkinen L . Compound heterozygosity for a recurrent 16.5-kb Alu-mediated deletion mutation and single-base-pair substitutions in the ABCC6 gene results in pseudoxanthoma elasticum. Am J Hum Genet 2001;68:642–652.
Carvalho CMB, Lupski JR . Mechanisms underlying structural variant formation in genomic disorders. Nat Rev Genet 2016;17:224–238.
Abeysinghe SS, Chuzhanova N, Krawczak M, Ball EV, Cooper DN . Translocation and gross deletion breakpoints in human inherited disease and cancer I: nucleotide composition and recombination-associated motifs. Hum Mutat 2003;22:229–244.
Eisenberger T, Neuhaus C, Khan AO et al. Increasing the yield in targeted next-generation sequencing by implicating CNV analysis, non-coding exons and the overall variant load: the example of retinal dystrophies. PLoS One 2013;8:e78496.
Boone P, Liu P, Zhang F et al. Alu-specific microhomology-mediated deletion of the final exon of SPAST in three unrelated subjects with hereditary spastic paraplegia. Genet Med 2011;13:582–592.
Boone PM, Yuan B, Campbell IM et al. The alu-rich genomic architecture of SPAST predisposes to diverse and functionally distinct disease-associated CNV alleles. Am J Hum Genet 2014;95:143–161.
Startek M, Szafranski P, Gambin T et al. Genome-wide analyses of LINE-LINE-mediated nonallelic homologous recombination. Nucleic Acids Res 2015;43:2188–2198.
Cardoso AR, Oliveira M, Amorim A, Azevedo L . Major influence of repetitive elements on disease-associated copy number variants (CNVs). Hum Genomics 2016;10:30.
Kim PM, Lam HYK, Urban A E et al. Analysis of copy number variants and segmental duplication in the human genome: evidence for a change in the process of formation in recent evolutionary history. Genome Res 2008;18:1865–1874.
Boon CJF, van den Born LI, Visser L et al. Autosomal recessive bestrophinopathy: differential diagnosis and treatment options. Ophthalmology 2013;120:809–820.
Boulanger-Scemama E, El Shamieh S, Démontant V et al. Next-generation sequencing applied to a large French cone and cone-rod dystrophy cohort: mutation spectrum and new genotype-phenotype correlation. Orphanet J Rare Dis 2015;10:85.
Mayer A, Mahajnah M, Zobor D, Bonin M, Sharkia R . Novel homozygous large deletion including the 5′ part of the SPATA7 gene in a consanguineous Israeli Muslim Arab family. Mol Vis 2015;65:306–315.
Dvir L, Srour G, Abu-Ras R, Miller B, Shalev SA, Ben-Yosef T . Autosomal-recessive early-onset retinitis pigmentosa caused by a mutation in PDE6G, the gene encoding the gamma subunit of rod cGMP phosphodiesterase. Am J Hum Genet 2010;87:258–264.
Watson CM, Crinnion LA, Berry IR et al. Enhanced diagnostic yield in Meckel-Gruber and Joubert syndrome through exome sequencing supplemented with split-read mapping. BMC Med Genet 2016;17:1.
Almoguera B, Li J, Fernandez-San Jose P et al. Application of whole exome sequencing in six families with an initial diagnosis of autosomal dominant retinitis pigmentosa: lessons learned. PLoS One 2015;10:e0133624.
Nishiguchi KM, Tearle RG, Liu YP et al. Whole genome sequencing in patients with retinitis pigmentosa reveals pathogenic DNA structural changes and NEK2 as a new disease gene. Proc Natl Acad Sci USA 2013;110:16139–16144.
Acknowledgements
This study was supported by Belspo (IAP project P7/43), Belgian Medical Genomics Initiative (E.D.B.), by the Ghent University Special Research Fund (BOF15/GOA/011) to E.D.B., Hercules Foundation AUGE/13/023 to E.D.B., by Funds for Research in Ophthalmology to K.V.S., and by the National Institutes of Health/National Institute of Neurological Disorders and Stroke (R37 NS028829) to M.E.G. S.N. is a PhD fellow of the Institute for Innovation by Science and Technology, integrated in the Research Foundation Flanders. S.V. is PhD fellow of the BOF (01D04716). M.B. is PhD fellow of the Research Foundation Flanders. F.C., H.V. and K.V.S. are postdoctoral fellows of the Research Foundation Flanders, and E.D.B. and B.P.L. are senior clinical investigators of the Research Foundation Flanders. The authors gratefully acknowledge the families who participated in this study.
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest. The sponsor or funding organization had no role in the design or conduct of this research.
Additional information
CNV Study Group James R. Lupski (Department of Molecular and Human Genetics, Baylor College of Medicine, Houston, Texas, USA), Claudia Carvalho (Department of Molecular and Human Genetics, Baylor College of Medicine, Houston, Texas, USA), Max van Min (Cergentis, Utrecht, The Netherlands), Eric Splinter (Cergentis, Utrecht, The Netherlands), Petra Klous (Cergentis, Utrecht, The Netherlands), Sarah De Jaegere (Center for Medical Genetics, Ghent University and Ghent University Hospital, Ghent, Belgium), and Sally Hooghe (Center for Medical Genetics, Ghent University and Ghent University Hospital, Ghent, Belgium).
Supplementary material is linked to the online version of the paper at
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Van Schil, K., Naessens, S., Van de Sompele, S. et al. Mapping the genomic landscape of inherited retinal disease genes prioritizes genes prone to coding and noncoding copy-number variations. Genet Med 20, 202–213 (2018). https://doi.org/10.1038/gim.2017.97
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2017.97
Keywords
This article is cited by
-
Cost-of-illness studies of inherited retinal diseases: a systematic review
Orphanet Journal of Rare Diseases (2024)
-
Whole genome sequencing enables new genetic diagnosis for inherited retinal diseases by identifying pathogenic variants
npj Genomic Medicine (2024)
-
Screening copy number variations in 35 unsolved inherited retinal disease families
Human Genetics (2024)
-
A comprehensive WGS-based pipeline for the identification of new candidate genes in inherited retinal dystrophies
npj Genomic Medicine (2022)
-
Clinical exome sequencing for inherited retinal degenerations at a tertiary care center
Scientific Reports (2022)