Abstract
Purpose
We conducted a systematic literature review to summarize the current health economic evidence for whole-exome sequencing (WES) and whole-genome sequencing (WGS).
Methods
Relevant studies were identified in the EMBASE, MEDLINE, Cochrane Library, EconLit and University of York Centre for Reviews and Dissemination databases from January 2005 to July 2016. Publications were included in the review if they were economic evaluations, cost studies, or outcome studies.
Results
Thirty-six studies met our inclusion criteria. These publications investigated the use of WES and WGS in a variety of genetic conditions in clinical practice, the most common being neurological or neurodevelopmental disorders. Study sample size varied from a single child to 2,000 patients. Cost estimates for a single test ranged from $555 to $5,169 for WES and from $1,906 to $24,810 for WGS. Few cost analyses presented data transparently and many publications did not state which components were included in cost estimates.
Conclusion
The current health economic evidence base to support the more widespread use of WES and WGS in clinical practice is very limited. Studies that carefully evaluate the costs, effectiveness, and cost-effectiveness of these tests are urgently needed to support their translation into clinical practice.
Similar content being viewed by others
Introduction
The first human genome sequence was produced in 2003, and its production cost between $500 million and $1 billion. For many years, the cost of sequencing an entire genome remained prohibitively expensive for routine use in clinical practice, with an estimated cost of $20–25 million in 2006.1 However, since 2008, when next-generation sequencing (NGS) approaches entered the research setting, there has been a significant decline in sequencing costs. These approaches allow either the whole genome (via whole-genome sequencing (WGS) or parts of it (via whole-exome sequencing (WES) or targeted panels) to be sequenced faster, at great depth and increasing sensitivity, and this decrease in costs has made the clinical application of WES and WGS more feasible.2 Conventional molecular testing of patients with genetic disorders has hitherto relied primarily on single-gene or panel testing or microarrays. However, estimates suggest that up to 50% of patients fail to receive a molecular diagnosis after such testing and embark on a diagnostic odyssey which is both slow and costly for health-care providers.3
Given this, several studies have investigated the benefits arising from the application of WES and WGS in the clinic. These studies have primarily focused on intellectual learning disability4 or Mendelian conditions more broadly,3,5,6,7,8,9 consistently reporting diagnostic yields between 25 and 30%. In addition, WES and WGS have often informed prognoses or treatment of patients, for example in inflammatory bowel disease10 and epilepsies.4,9
Considerable evidence therefore exists that applying WES or WGS in the clinic will improve the diagnosis and (in some cases) treatment of genetic disease, which may improve patient health outcomes and facilitate the more efficient use of health-care resources. However, demand is increasing for evidence on the incremental costs and health outcomes associated with these technologies compared with those used in current practice to support these assertions and to ensure that these new technologies are not merely an expensive add-on to patient care.2 Much of the recent demand for evidence is driven by national sequencing programs such as the Genomics England 100,000 Genomes Project and the Genome Canada initiative. Before these programs were launched, sequencing capacity was concentrated in individual research or clinical laboratories that purchased their own equipment and ran their own sequencing pipelines, buying reagents as required. However, these national sequencing initiatives have challenged suppliers to offer a turnkey service to meet a volume requirement of whole genomes, and this has resulted in the development of prototype sequencing machines optimized to sequence at scale. Despite the substantial reductions in sequencing costs following these developments, there is still little information available on the costs of bioinformatics analysis and clinical interpretation (and on what the costs of sequencing are, if not done at scale), as documented in earlier literature reviews.11,12,13 It is therefore important to understand what economic evaluation evidence is currently available in the literature on clinical applications of WES and WGS, and where there are gaps in the evidence base.
This paper reports the results of a literature review that summarizes the current evidence in the health-economic literature on the use of WES and WGS in a clinical setting. We highlight the areas where good evidence exists and the areas characterized by a paucity of data, and we explore the methodological and applied economic considerations associated with the more widespread use of NGS approaches in clinical practice in cancer, rare diseases, and pathogens.
Materials and methods
The aim of this literature review was to summarize the current health-economic evidence base for WES and WGS. The review focused specifically on WES and WGS, as these are the newest NGS technologies, with potentially the highest costs. The main objective was to identify economic evaluations of these technologies. Economic evaluation has been defined as “the comparative analysis of alternative courses of action in terms of both their costs and consequences,” and this approach can be used to quantify and compare the costs and outcomes of health-care interventions with a view to identifying those that represent the best value for money.14 For the purposes of this review no restrictions were placed on the comparisons made within these economic evaluations: studies comparing WGS with WES were eligible for inclusion, as were studies comparing these testing approaches against other forms of testing, such as single-gene tests. This review therefore also aimed to identify publications reporting data on either the costs of WES or WGS testing pathways, or on clinically relevant outcome measures for these tests. Publications reporting narrowly defined sequencing costs (e.g. cost per megabase/run) that are not linked to specific clinical applications were not included, as an earlier review summarizes these costs for different sequencing platforms.13
Search strategies and study selection
Search strategies were designed for EMBASE, MEDLINE, the Cochrane Library, EconLit, and the University of York Centre for Reviews and Dissemination (CRD) Database. These strategies included both index terms and free-text words (see Supplementary Appendix S1 online). Key words were identified by examining relevant references in the literature and the Medical Subject Headings (MeSH) used by EMBASE and MEDLINE (http://www.nlm.nih.gov/mesh/). Table 1 describes the study selection criteria. Search strategies were restricted to publications written in English or German (the first author is a native German speaker) between January 2005 and July 2016. This date range reflects the novelty of the sequencing technologies under consideration: older publications either do not exist or would be outdated owing to technological developments. No restrictions were placed on study length as it was anticipated that relevant data might be presented in conference abstracts.
The literature search was undertaken in July 2016 by K.S. Following deduplication, publications were scanned for relevance by title and abstract, and clearly irrelevant publications were excluded. The full text of the remaining publications was then evaluated for relevance by both K.S. and S.W.. Any uncertainties were resolved in consultation with J.B. The reference lists of retrieved publications and the publications of recognized authors in this field were also reviewed to identify additional studies that met the inclusion criteria.
Data extraction
Relevant material was extracted from the included publications using a bespoke proforma (see Supplementary Appendix S2). This proforma was initially piloted on three publications. Following minor revisions, all publications were then assessed using this proforma. The extracted data included bibliographic details and information on the study objective, disease context, patient population, and sequencing methods. Information was also extracted on the study perspective, assumptions, methods, outcomes, costs, main results, and author-reported study limitations.
Classification of publications
Publications were categorized as economic evaluations, cost studies, or outcome studies. Publications were defined as economic evaluations only if data on both costs and outcomes were presented for WES or WGS and a comparison with at least one alternative testing option (which could be no testing) was undertaken. Publications that only included data on the costs of testing pathways or were budget-impact analyses, were classified as cost studies. Publications that only included clinically relevant outcome data were classified as outcome studies.
Publications were further categorized by type of economic evaluation, with five approaches considered. All five approaches measure health-care costs in monetary terms, but they differ in the measurement of health outcomes. Two of these approaches were classified as partial economic evaluations. In cost-consequence analysis, costs and health outcomes are reported separately, with multiple outcome measures presented. This allows decision makers to combine this information in different ways to inform health-care resource-allocation decisions. In cost-minimization analysis, it is assumed that health outcomes are identical for the interventions under consideration and these alternatives are therefore compared in terms of their relative costs.
The remaining three approaches were classified as full economic evaluations. In cost-effectiveness analysis (CEA) outcomes are measured in health-related terms using natural units such as life-years gained. Cost-utility analysis (CUA) uses quality-adjusted life years (QALYs) as an outcome measure, and these combine information on longevity and quality of life. In both CEA and CUA studies results are expressed as ratios of incremental costs to incremental outcomes and compared with a threshold incremental cost-effectiveness ratio to determine whether an intervention is cost-effective. Finally, in cost-benefit analysis (CBA), health (and sometimes nonhealth) outcomes are valued in monetary terms using approaches such as willingness-to-pay questionnaires and discrete choice experiments. CBAs may be more informative for some genomic tests than CEAs or CUAs, particularly when a test does not offer an obvious survival benefit.15 CBA results are commonly expressed as a ratio of costs to benefits, or as a sum representing the net benefit of one intervention compared with another.
Data analysis
The data were summarized in several ways. Counts were calculated for study characteristics including type of study and form of economic evaluation and diagnostic yield estimates were extracted from all publications. Studies with “difficult to diagnose” cohorts were also highlighted. These studies either explicitly stated that probands were difficult to diagnose or focused on patients who had previously undergone multiple unsuccessful diagnostic procedures. All cost estimates for WES and WGS testing were extracted from the included studies, including both costs and commercial prices. Where the cost year was not stated, the latest date at which the costing must have been conducted was used (e.g. date of manuscript submission). Cost estimates were converted to British pounds and US dollars based on purchasing-power parities.16 Purchasing-power parities reflect the cost of purchasing a standardized set of goods and services in different countries and are subject to fewer fluctuations than exchange rates. Costs were then inflated to 2016 values using a UK health-care inflation index.17 Finally, minimum and maximum costs were calculated for each sequencing approach.
Results
The database search identified 1,277 publications, of which 302 were duplicates (Figure 1). After titles and abstracts were screened and full-text publications assessed for eligibility, 14 publications were identified that fulfilled the inclusion criteria. A review of the reference lists of retrieved publications and the publications of recognized authors in this field identified 22 additional relevant publications (7 of which were identified after July 2016, the planned end date for literature screening18,19,20,21). Overall, 36 relevant publications were identified.
Study characteristics
Table 2 summarizes the characteristics of these 36 publications. Detailed descriptions are provided in Supplementary Appendix S3. The publications investigated the use of WES and WGS in a variety of conditions with a genetic background. The most common conditions were neurological or neurodevelopmental disorders (7 publications), with 13 (36%) publications investigating WES and WGS exclusively in children or newborns. The study sample size varied from a single child to a cohort of 2,000 patients.
The studies reported in the following nine publications (22%) did not have a patient population. Bennette et al.22 used a decision analytic model to evaluate the cost-effectiveness of returning incidental findings from WES and WGS in the United States. Buchanan-Hughes et al.23 constructed a decision tree to calculate the cost utility of bacterial WGS in the diagnostic pathway of urinary tract infections. Chrystoja and Diamandis24 presented a discussion of the challenges and opportunities associated with the use of WGS as a diagnostic test. The report by the Foundation for Genomics and Population Health25 described the costs of WES for colorectal cancer in three UK National Health Service laboratories. Plöthner et al.26 investigated the costs associated with executing WGS in German clinical practice. Sabatini et al.27 conducted a bottom-up cost study of WES in Canada and compared the findings with the costs of the traditional clinical pathway for diagnosing neurodevelopmental disorders, using payer cost-impact models. Van Nimwegen28 constructed a decision model to examine the cost-effectiveness of WES in clinical practice in the Netherlands. Finally, Tsiplova et al.20,21 calculated the cost-effectiveness of WES and WGS compared with chromosomal microarray analysis (CMA) in three hypothetical testing scenarios.
Twenty-one economic evaluations were identified, of which 8 were full economic evaluations18,19,20,21,22,23,28,29,30 and 13 were partial economic evaluations.3,31,32,33,34,35,36,37,38,39,40,41,42 Seven studies presented data on the costs of WES or WGS testing pathways,24,25,26,27,43,44,45 and eight studies presented data on clinically relevant outcome measures for these tests.5,6,8,9,10,46,47,48 Of the eight full economic evaluations, two were CUAs22,23 and six were CEAs, published between 2014 and 2017 in Australia (2), the United States (1), the UK (1), the Netherlands (1), and Canada (1).18,19,20,21,28,29,30 Of these publications, the study by Soden et al.29 did not directly report WES costs but estimated the cost-effectiveness threshold for WES in pediatric neurodevelopmental disorders by calculating the cost of the current diagnostic pathway.
All 13 partial economic evaluations were cost-consequence analyses. These studies were published between 2013 and 2016 in the Netherlands (4), Australia (3), Canada (2), France (1), the United States (2) and the United Kingdom (1). Eleven of these publications investigated WES, with the study by Pankhurst et al.42 evaluating bacterial WGS and the study by Shashi et al.3 evaluating several potential NGS approaches. While all 13 publications reported cost estimates for WES or WGS, only 3 stated the methods and sources underlying these estimates.35,37,42
Of the seven studies that presented data on the costs of WES or WGS testing pathways, four evaluated WGS. Chrystoja and Diamandis24 reviewed the potential of WGS and summarized cost data extracted from previously published scientific studies and commercial sources. Dewey et al.43 estimated the costs of WGS in the United States as well as the costs associated with curation and clinical follow-up. Weymann et al.45 estimated the costs of using WGS to inform treatment decisions in patients with advanced cancers. Towne et al.44 estimated the cost of using WES to achieve a diagnosis in rare presentations in a pediatric population. The studies by the Foundation for Genomics and Population Health,25 Sabatini et al.,27 and Plöthner et al.26 have been described previously.
Eight studies, published between 2011 and 2016, presented data on clinically relevant outcome measures for WES and WGS. Seven of these studies investigated WES;5,6,8,9,10,46,47 the eighth evaluated WGS.48 Four publications used the traditional care pathway in the investigated condition as a comparator.6,10,46,48 Three publications were retrospective analyses6,47,48 and three were diagnostic studies, estimating the diagnostic yield of WES in a variety of conditions.5,8,9 The final two studies were case studies on single probands and families.10,46
Twenty-four (67%) of the publications that met the inclusion criteria focused on WES, with five (14%) focusing on WGS, five (14%) focusing on both WGS and WES, and two (6%) evaluating bacterial WGS. The most common study setting was the United States (36% of publications). The earliest two studies were published in 2011, with all other studies published between 2013 and 2017.
WES and WGS costs
Table 3 summarizes the cost estimates for NGS approaches, with detailed information provided in Supplementary Appendix S4. Twenty-nine studies reported cost estimates, of which 18 reported costs for WES. Estimates ranged from £382 ($555)32 to £3,592 ($5,169)34,38 for a single WES test. The highest estimate for a single test (£3,592) was based on commercial prices. The highest estimate of the actual costs of a single WES test (i.e. not commercial prices) was £1,808 ($2,602).37 Cost estimates for a trio ranged from £2,658 ($3,825)44 to £6,466 ($9,304).38 Thirteen publications stated that costs were estimated within the study; nine of these publications reported their costing approach.19,20,21,26,27,28,36,37,45 Many publications did not state which components were included in the cost estimate. The costs for reagents ranged from £291 ($420)35 to £1,171 ($1,685).31
Cost estimates varied little over time. The lowest estimate for a single WES test was £736 ($1,060)44 in 2013 and £736 ($1,070)28 in 2017. In terms of country-level costs, the lowest estimate (£382; $555)32 was reported in an Australian study from 2015, and the highest estimate (£3,592; $5,169)36,38 was reported in two Canadian studies from 2014. There were no regional differences: the lowest cost estimates in North America (£736; $1,060)44 and the rest of the world (£382; $555)32 were similar, but far less with the higher cost estimates (£3,592; $5,169 in North America,36,38 £3,401; $4,907 in the rest of the world).40,41
Six studies estimated the cost of WGS, four of which used data from commercial sources.24,26,28,43 Cost estimates ranged from £1,312 ($1,906) for sequencing using the HiSeq X in Germany26 to £17,243 ($24,810) for an unspecified platform in Canada.24 Four studies used a transparent bottom-up approach to estimate the cost of WGS.20,21,26,28,45 There was limited evidence of a reduction in the cost of WGS over time, with the lowest estimate declining from £10,497 ($15,146)43 in 2013 to £1,312 ($1,906)26 in 2017. However, this is based on a small sample. The two cost estimates for bacterial WGS were considerably lower than those for WES or WGS in humans.
Finally, of the 16 studies that calculated test costs rather than applying an assumed figure or using a commercial price, only 10 described the cost calculation in a transparent manner.
WES and WGS outcomes
A variety of outcomes were assessed in the publications that met the inclusion criteria, including successful diagnoses, diagnostic yield, sensitivity and specificity, quality-adjusted life years (QALYs), time to diagnosis, change in clinical management, acute clinical usefulness, mortality/survival, parent satisfaction, frequencies of disease subtypes, mode of inheritance, spectrum of genetic events, reporting of incidental findings, target capture, and prediction of bacterium species and drug susceptibility. Diagnostic yield was the most common outcome measure (18 publications). Table 4 summarizes diagnostic yield estimates by sequencing approach, with detailed information provided in Supplementary Appendix S5. The lowest diagnostic yield for WES (3%) was estimated in a patient group with colorectal cancer.37 The highest rate for WES (79%) was reported for individuals with childhood-onset muscle disorders.18 Around a third of the included publications investigated WES or WGS in a population group that was difficult to diagnose.
WES and WGS cost-effectiveness
Eight studies estimated the cost-effectiveness of WES or WGS. Sagoo et al.30 estimated the cost-effectiveness of WES compared with usual testing practice (genetic tests and disease gene panel tests) in a variety of disease contexts. When WES was introduced later in the testing pathway, the incremental cost per additional positive diagnosis was £3,213 ($4,670). When WES was used as a near first-line test the incremental cost per additional positive diagnosis was £2,230 ($3,242).
Van Nimwegen28 compared the costs and outcomes associated with WES (using the HiSeq 4000) with those of conventional diagnosis for pediatric neurological disorders (which includes magnetic resonance imaging scans, electroencephalography, and muscle biopsies). In the author’s first analysis, WES was treated as a last-resort test, with an incremental cost per additional diagnosis of £8,319 ($12,092). The second analysis treated WES as a first-line test, and in many plausible scenarios this resulted in cost savings.
Soden et al.29 estimated that WES would be cost-effective in pediatric neurodevelopmental disorders, compared with existing nongenomic investigative approaches (including laboratory tests, radiologic procedures, and electromyograms) on a cost-per-diagnosis basis if the cost of WES was no more than £2,123 ($3,063) per individual.
Schofield et al.18 reported that WES offered a cost saving per diagnosis of £6,483 ($9,342) compared with traditional investigations (muscle biopsy, histological and biochemical analyses, Sanger sequencing) for the diagnosis of pediatric muscle diseases.
Stark et al.19 examined the cost-effectiveness of three scenarios for implementing WES as a routine clinical test for infants with suspected monogenic disorders. Integrating WES after standard investigations (including biochemical investigations, imaging, and neurophysiological studies) cost £3,830 ($5,518) per additional diagnosis, replacing a subset of existing investigations with WES cost £1,238 ($1,784) per additional diagnosis, and implementing WES as a first-line test to replace most investigations saved £1,030 ($1,484) per additional diagnosis.
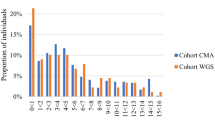
Tsiplova et al.20,21 evaluated the cost-effectiveness of strategies involving WES and WGS compared with CMA in autism spectrum disorder. Adding WES to CMA cost £13,912 ($20,046) per additional diagnosis, whereas implementing WGS instead of CMA cost £32,219 ($46,424) per additional diagnosis using a HiSeq 2500 sequencer, and £14,219 ($20,488) using a HiSeq X sequencer. Implementing WGS compared with a WES-plus-CMA approach cost £106,590 ($153,587) per additional diagnosis using a HiSeq 2500 sequencer, and £15,464 ($22,283) using a HiSeq X sequencer.
Buchanan-Hughes et al.23 investigated the cost-effectiveness of bacterial WGS to guide targeted antibiotic selection in urinary tract infections, finding that bacterial WGS was more expensive than methods employed in current clinical practice, with poorer health outcomes.
Finally, Bennette et al.22 estimated the cost-effectiveness of generating information on incidental findings using genomic sequencing. The cost per QALY gained for cardiomyopathy patients was estimated to be £32,187 ($46,313), for colorectal cancer patients it was £82,623 ($118,883), and for healthy individuals it was £42,102 ($60,578). Generating information on incidental findings for cardiomyopathy patients and healthy individuals was therefore cost-effective, compared with a threshold of $100,000 per QALY gained.
Discussion
This review summarizes the current health-economic evidence base for WES and WGS. We found that the costs of a single test ranged from £382 to £3,592 per patient for WES, from £1,312 to £17,243 per patient for WGS in humans, and from £40 to £487 for bacterial WGS. There was no evidence that the cost of WES was falling over time, and only limited evidence that the cost of WGS was decreasing, contrary to the widely held belief that genomic sequencing costs are falling rapidly in clinical settings. There were also few regional differences in costs. The most commonly utilized outcome measure was diagnostic yield, with few studies using outcome measures recommended for use in economic evaluations, such as survival or quality of life. Furthermore, in only a few individual patients was the provision of an accurate molecular diagnosis from WES/WGS followed through to the concomitant changes in clinical investigations and treatment, where significant health-economic impact may be anticipated. Most of the identified publications concluded that WES and WGS were superior in economic terms to other testing methods, but many of these analyses were based on small patient samples and only eight publications were full economic evaluations. Of these eight publications, only five produced evidence that WES or WGS may represent a cost-effective use of limited health-care resources. As the cost-effectiveness of WES and WGS is probably dependent on clinical context, study timing, patient population, and other health system factors, we are unable to make a broad statement about the cost-effectiveness of these technologies.49 Overall, the results of this review indicate that the current health-economic evidence base to support the more widespread use of WES and WGS in clinical practice is very limited.
The absence of economic evaluations of WES and WGS in the literature was anticipated before this review was undertaken, in light of the novelty of these technologies. We therefore also identified publications reporting data on either costs or clinically relevant outcome measures for WES and WGS. Even when we broadened the scope of our review, robust evidence that could inform health-care resource-allocation decisions remained scarce. Few publications described how cost estimates were calculated or whether the costing perspective was that of a health-care provider, or a broader societal perspective. It was also unclear in many studies what items included in the cost estimates represented resource use. In many cases cost estimates either reflected commercial prices or were hypothetical threshold costs at which WES or WGS could become cost-effective; there were few high-quality detailed microcosting studies. There was also overlap between studies, with several applying cost estimates from early studies to new clinical contexts. In terms of health outcomes, around a third of the identified publications investigated WES or WGS in “difficult-to-diagnose” population groups that had previously undergone a variety of unsuccessful diagnostic tests. Given this, it was difficult to estimate an overall diagnostic yield for genomic sequencing, as study results varied depending on the clinical context and on whether probands were tested alone or as trios. Long-term data on costs and health outcomes were particularly scarce. Furthermore, publication dates ranged from 2011 to 2017. In this rapidly evolving field of research, it is likely that some of these cost and health outcomes estimates are already outdated. Given this heterogeneity in methods and study quality, we have not reported estimates of mean costs and diagnostic yield: such measures would mask variations by sequencing platform and clinical context and would be of limited use in informing budgetary decisions regarding the use of WES and WGS in health care. Overall, these findings suggest that the cost, health-outcome, and cost-effectiveness results presented in the identified papers are not generalizable to different clinical and country settings, and are of limited use in informing health-care resource-allocation decisions.
This is the first literature review that has summarized the economic evidence in the literature related to the use of next-generation genomic sequencing approaches such as WES and WGS in clinical settings. However, this review has two key limitations. First, over half of the included publications were identified from other sources, which could indicate that the search strategies were not sufficiently sensitive. A key challenge when specifying the search strategies was that sequencing methods and health-economic evidence are commonly misclassified in the literature. Broad search terms were therefore combined to ensure that all forms of NGS and all types of economic evidence were captured. Consequently, this approach identified several studies that evaluated tests other than WES or WGS, or that mentioned the potential cost-effectiveness of genomic sequencing, but did not present any evidence. This highlights the need for clear definitions and methods for economic evaluations.
Second, the literature search did not specifically aim to identify clinical outcomes, focusing only on outcome measures typically used in economic evaluations (e.g. QALYs, life years gained). However, studies including other outcome measures were included if they were identified. This decision was made because the inclusion of search terms such as “successful diagnosis” would have resulted in a prohibitively large increase in irrelevant papers. Publications similar to the included outcome studies might therefore have been excluded from this review. However, when the references of articles summarizing outcome data were reviewed, the same sources were cited in most cases. This suggests that key publications in this field were not overlooked.
Given the pace of change in genomic research, there is probably a lag between published health-economic data and current practice in laboratories and hospitals. That said, the data presented in this review provide a useful starting point to explore the potential methodological and applied economic considerations associated with the more widespread use of NGS approaches in clinical practice. A key concern is the paucity of studies utilizing those outcome measures that are recommended for use in economic evaluations by major health technology assessment agencies such as the National Institute for Health and Care Excellence in the United Kingdom and the Pharmaceutical Benefits Advisory Committee in Australia. Outcome measures such as life years or QALYs are recommended by such institutions as this enables comparisons to be made between interventions when health-care resource-allocation decisions are being considered. The widespread use of “diagnostic yield” as an outcome measure in the studies identified in this review is therefore problematic for health technology assessment agencies. In the short to medium term, studies evaluating the effectiveness and cost-effectiveness of WES and WGS could better inform the allocation of health-care resources by presenting data on health outcomes using metrics such as life years and QALYs. Studies investigating the link between diagnostic yield and patient survival and quality of life would also contribute to this literature. In the longer term, health technology assessment agencies may wish to consider whether focusing solely on health benefits is appropriate for interventions such as WES and WGS. Although this is the correct focus when resource-allocation decisions are being made with respect to a national health-care budget, it is possible that important nonhealth benefits arising from the use of WES and WGS (such as the inherent value of diagnostic information) are not captured when adopting this approach.15 Widening the recommended perspective (often permitted when evaluating public-health interventions) may therefore improve overall population health and well-being.
A second concern is that most of the studies identified in this review utilized small patient samples. Related to this, the estimates of costs, health outcomes and cost-effectiveness reported in these studies varied widely. This highlights the importance in future studies of genomic sequencing of collecting such data at scale, prospectively, with coverage of a diverse range of patient groups. Large national sequencing initiatives such as the Genomics England 100,000 Genomes Project are well placed to generate such estimates, providing essential evidence to inform the translation of WES and WGS into clinical practice. Research into the health economics of NGS approaches should be a key component of such initiatives.
We have four additional practical recommendations. First, few studies reported all of the cost components of the overall cost of either WES or WGS. It was therefore difficult in many cases to identify key cost drivers. Future studies should report costs by stage of testing for WES and WGS and highlight particularly notable costs. Second, resource use and unit costs were often not reported in a disaggregated manner. Again, this hinders interpretation and makes it difficult to identify the cost components that researchers and commissioners should focus on in order to reduce the cost of testing. Third, several studies did not calculate the actual cost of WES or WGS, instead using prices derived from commercial sources. These prices do not necessarily reflect the economic value associated with WES or WGS, and do not capture the opportunity cost of using limited health-care resources to implement genomic testing in routine clinical practice. Economic evaluations that use such prices may therefore recommend incorrect and inefficient adoption decisions. Future studies evaluating the cost-effectiveness of WES or WGS should use calculated costs instead of prices. Finally, several studies did not include a health economist as an author. This probably contributed to heterogeneity in methods and study quality. Future studies of the cost-effectiveness of WES and WGS should include trained health economists as coinvestigators.
This review has demonstrated that the current health-economic evidence base supporting the more widespread use of WES and WGS in clinical practice is very limited. A key consequence of this finding is that it is currently difficult for health technology assessment agencies to efficiently allocate scarce health-care resources where WES and WGS approaches are concerned. Studies that carefully evaluate the costs, effectiveness, and cost-effectiveness of these interventions in a transparent manner are urgently required, to enable more informed decision making in this context.
References
National Human Genome Research Institute. The Cost of Sequencing a Human Genome. https://www.genome.gov/sequencingcosts/. Accessed 17 May 2017.
Davies SC. Annual Report of the Chief Medical Officer 2016, Generation Genome. Department of Health: London, 2017.
Shashi V, McConkie-Rosell A, Rosell B et al. The utility of the traditional medical genetics diagnostic evaluation in the context of next-generation sequencing for undiagnosed genetic disorders. Genet Med. 2014;16:176–182.
de Ligt J, Willemsen MH, van Bon BW et al. Diagnostic exome sequencing in persons with severe intellectual disability. N Engl J Med 2012;367:1921–1929.
Lee H, Deignan JL, Dorrani N et al. Clinical exome sequencing for genetic identification of rare Mendelian disorders. JAMA. 2014;312:1880–1887.
Sawyer SL, Hartley T, Dyment DA et al. Utility of whole-exome sequencing for those near the end of the diagnostic odyssey: time to address gaps in care. Clin Genet 2016;89:275–284.
Taylor JC, Martin HC, Lise S et al. Factors influencing success of clinical genome sequencing across a broad spectrum of disorders. Nat Genet. 2015;47:717–726.
Yang Y, Muzny DM, Reid JG et al. Clinical whole-exome sequencing for the diagnosis of mendelian disorders. N Engl J Med 2013;369:1502–1511.
Yang Y, Muzny DM, Xia F et al. Molecular findings among patients referred for clinical whole-exome sequencing. JAMA 2014;312:1870–1879.
Worthey EA, Mayer AN, Syverson GD et al. Making a definitive diagnosis: successful clinical application of whole exome sequencing in a child with intractable inflammatory bowel disease. Genet Med 2011;13:255–262.
Beale S, Sanderson D, Sanniti A, Dundar Y, Boland A. A scoping study to explore the cost-effectiveness of next-generation sequencing compared with traditional genetic testing for the diagnosis of learning disabilities in children. Health Technol Assess 2015;19:1–90.
Canadian Agency for Drugs and Technologies in Health. Next Generation DNA Sequencing: A Review of the Cost Effectiveness and Guidelines. Ottawa, Canada. 2014.
Frank M, Prenzler A, Eils R, Graf von der Schulenburg JM. Genome sequencing: a systematic review of health economic evidence. Health Econ Rev 2013;3:29.
Drummond MF, Sculpher MJ, Claxton KP, Stoddart GL, Torrance GW. Methods for the Economic Evaluation of Health Care Programmes, 4th edn. Oxford University Press: Oxford, UK, 2015.
Buchanan J, Wordsworth S, Schuh A. Issues surrounding the health economic evaluation of genomic technologies. Pharmacogenomics 2013;14:1833–1847.
Organisation for Economic Co-operation and Development. Prices and purchasing power parities (PPP). http://www.oecd.org/std/prices-ppp/. Accessed 17 May 2017.
Curtis L. Unit Costs of Health, Social Care. Personal Social Services Research Unit, University of Kent, 2016.
Schofield D, Alam K, Douglas L et al. Cost-effectiveness of massively parallel sequencing for diagnosis of paediatric muscle diseases. NPJ Genom Med 2017;2:4.
Stark Z, Schofield D, Alam K et al. Prospective comparison of the cost-effectiveness of clinical whole-exome sequencing with that of usual care overwhelmingly supports early use and reimbursement. Genet Med. 2017;19:867–874.
Tsiplova K, Zur RM, Marshall CR et al. A microcosting and cost-consequence analysis of clinical genomic testing strategies in autism spectrum disorder. Genet Med 2017;19:1268–1275.
Tsiplova K, Zur RM, Ungar WJ A microcosting and cost-consequence analysis of genomic testing strategies in autism spectrum disorder. Technology Assessment at SickKids, Hospital for Sick Children: Toronto, Canada, 2016. http://www.sickkids.ca/pdfs/Research/TASK/microcosting/69711-Microcosting-FULL-REPORT.pdf.
Bennette CS, Gallego CJ, Burke W, Jarvik GP, Veenstra DL. The cost-effectiveness of returning incidental findings from next-generation genomic sequencing. Genet Med. 2015;17:587–595.
Buchanan-Hughes AM, Griffiths A, Evans J, Slater D, Eddowes LA. Investigating the cost-effectiveness of bacterial whole-genome sequencing for enabling targeted antibiotic selection in urinary tract infections. Value Health 2015;18:A510.
Chrystoja CC, Diamandis EP. Whole Genome Sequencing as a Diagnostic Test: Challenges and Opportunities. Clin Chem 2014;60:724.
Foundation for Genomics and Population Health. Next Steps in the Sequence: The Implications of Whole Genome Sequencing for Health in the UK. 2011. https://www.ncbi.nlm.nih.gov/pubmed/27380512.
Plöthner M, Frank M, von der Schulenburg JMG. Cost analysis of whole genome sequencing in German clinical practice. Eur J Health Econ 2017;18:623–633.
Sabatini LM, Mathews C, Ptak D et al. Genomic sequencing procedure microcosting analysis and health economic cost-impact analysis: a report of the Association for Molecular Pathology. J Mol Diagn 2016;18:319–328.
van Nimwegen KJ. Health Technology Assessment of Next-Generation Sequencing. Radboud University: Nijmegen, The Netherlands, 2017.
Soden SE, Saunders CJ, Willig LK et al. Effectiveness of exome and genome sequencing guided by acuity of illness for diagnosis of neurodevelopmental disorders. Sci Transl Med 2014;6:265ra168.
Sagoo G, Norbury G, Mohammed S, Kroese M. The Budget Impact and Cost-Effectiveness of Introducing Whole-Exome Sequencing-Based Virtual Gene Panel Tests into Routine Clinical Genetics. PHG Foundation: Cambridge, UK, 2017.
Bonnefond A, Philippe J, Durand E et al. Highly sensitive diagnosis of 43 monogenic forms of diabetes or obesity through one-step pcr-based enrichment in combination with next-generation sequencing. Diabetes Care. 2014;37:460.
Ghaoui R, Cooper ST, Lek M et al. Use of whole-exome sequencing for diagnosis of limb-girdle muscular dystrophy: outcomes and lessons learned. JAMA Neurol 2015;72:1424–1432.
Lee E-J, Dykas D, Bale A et al. Whole exome sequencing in evaluation of thrombophilia: a novel 33-gene panel. Blood 2015;126:3529.
McDonell LM, Warman Chardon J, Schwartzentruber J et al. The utility of exome sequencing for genetic diagnosis in a familial microcephaly epilepsy syndrome. BMC Neurol. 2014;14:22.
McInerney-Leo AM, Harris JE, Leo PJ et al. Whole exome sequencing is an efficient, sensitive and specific method for determining the genetic cause of short-rib thoracic dystrophies. Clin Genet 2015;88:550–557.
McInerney-Leo AM, Marshall MS, Gardiner B et al. Whole exome sequencing is an efficient and sensitive method for detection of germline mutations in patients with phaeochromcytomas and paragangliomas. Clin Endocrinol 2014;80:25–33.
Neveling K, Feenstra I, Gilissen C et al. A post-hoc comparison of the utility of Sanger sequencing and exome sequencing for the diagnosis of heterogeneous diseases. Hum Mutat. 2013;34:1721–1726.
Sawyer SL, Schwartzentruber J, Beaulieu CL et al. Exome sequencing as a diagnostic tool for pediatric-onset ataxia. Hum Mutat 2014;35:45–49.
Schieving JH. PP05.5 – 3064: The role of exome sequencing in daily pediatric neurology practice. Eur J Paediatr Neurol 2015;19:S47.
van Nimwegen KJ, Schieving JH, Willemsen MA et al. The diagnostic pathway in complex paediatric neurology: a cost analysis. Eur J Paediatr Neurol 2015;19:233–239.
Monroe GR, Frederix GW, Savelberg SM et al. Effectiveness of whole-exome sequencing and costs of the traditional diagnostic trajectory in children with intellectual disability. Genet Med. 2016;18:949–956.
Pankhurst LJ, Del Ojo Elias C, Votintseva AA et al. Rapid, comprehensive, and affordable mycobacterial diagnosis with whole-genome sequencing: a prospective study. Lancet Respir Med 2016;4:49–58.
Dewey FE, Grove ME, Pan C et al. Clinical interpretation and implications of whole-genome sequencing. JAMA. 2014;311:1035–1045.
Towne MC, Beggs AH, Agrawal PB Efficiency of whole exome/genome sequencing for achieving a diagnosis in rare presentations. Annual meeting of the American Society of Human Genetics, Boston, MA, 22–26 October 2013.
Weymann D, Laskin J, Roscoe R et al. The cost and cost trajectory of whole‐genome analysis guiding treatment of patients with advanced cancers. Mol Genet Genomic Med 2017;5:251–260.
Bettencourt C, López-Sendón JL, García-Caldentey J et al. Exome sequencing is a useful diagnostic tool for complicated forms of hereditary spastic paraplegia. Clin Genet 2014;85:154–158.
Valencia CA, Husami A, Holle J et al. Clinical impact and cost-effectiveness of whole exome sequencing as a diagnostic tool: a pediatric center’s experience. Front Pediatr 2015;3:67.
Willig LK, Petrikin JE, Smith LD et al. Whole-genome sequencing for identification of Mendelian disorders in critically ill infants: a retrospective analysis of diagnostic and clinical findings. Lancet Respir Med 2015;3:377–387.
Hatz MHM, Schremser K, Rogowski WH. Is individualized medicine more cost-effective? a systematic review. Pharmacoeconomics. 2014;32:443–455.
Acknowledgments
This study was funded by the UK Health Innovation Challenge Fund (a scheme supported in equal measure by the Department of Health and the Wellcome Trust). We also acknowledge support through Wellcome Trust Centre for Human Genetics Wellcome Trust Core Award Grant 090532/Z/09/Z. S.W. and J.C.T. are partly funded by the National Institute for Health Research Oxford Biomedical Research Centre. K.S. is partly funded by the Erasmus Plus scheme of the University Duisburg-Essen.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclosure
The authors declare no conflict of interest.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Schwarze, K., Buchanan, J., Taylor, J.C. et al. Are whole-exome and whole-genome sequencing approaches cost-effective? A systematic review of the literature. Genet Med 20, 1122–1130 (2018). https://doi.org/10.1038/gim.2017.247
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2017.247