Abstract
Purpose
Mowat–Wilson syndrome (MWS) is a rare intellectual disability/multiple congenital anomalies syndrome caused by heterozygous mutation of the ZEB2 gene. It is generally underestimated because its rarity and phenotypic variability sometimes make it difficult to recognize. Here, we aimed to better delineate the phenotype, natural history, and genotype–phenotype correlations of MWS.
Methods
In a collaborative study, we analyzed clinical data for 87 patients with molecularly confirmed diagnosis. We described the prevalence of all clinical aspects, including attainment of neurodevelopmental milestones, and compared the data with the various types of underlying ZEB2 pathogenic variations.
Results
All anthropometric, somatic, and behavioral features reported here outline a variable but highly consistent phenotype. By presenting the most comprehensive evaluation of MWS to date, we define its clinical evolution occurring with age and derive suggestions for patient management. Furthermore, we observe that its severity correlates with the kind of ZEB2 variation involved, ranging from ZEB2 locus deletions, associated with severe phenotypes, to rare nonmissense intragenic mutations predicted to preserve some ZEB2 protein functionality, accompanying milder clinical presentations.
Conclusion
Knowledge of the phenotypic spectrum of MWS and its correlation with the genotype will improve its detection rate and the prediction of its features, thus improving patient care.
Similar content being viewed by others
Introduction
Mowat–Wilson syndrome (MWS) (OMIM # 235730) is characterized by distinctive facial appearance in association with variable moderate-to-severe intellectual disability (ID), epilepsy, Hirschsprung disease (HSCR), and multiple congenital anomalies, including genital anomalies (in particular hypospadias), congenital heart disease, agenesis of the corpus callosum, and eye defects. MWS is caused by deleterious de novo heterozygous variations in the ZEB2 gene. The majority of variants lead to haploinsufficiency through premature stop codons or large deletions.
MWS was first delineated by Mowat et al.1 as a syndromic condition characterized by distinctive facial phenotype, ID, and Hirschsprung disease in four of six reported individuals. The causative genetic defect was mapped to chromosome 2q21-q23 based on cytogenetic deletions in two patients,1,2 and narrowed to heterozygous mutations of the ZEB2 gene by subsequent reports.3,4 In 2002 Zweier et al.5 further delineated the phenotype of MWS with or without HSCR, invariably characterized by ZEB2 gene defects, and proposed that the condition be named Mowat–Wilson syndrome. More than 300 patients have been reported so far6,7,8,9,10,11,12,13,14,15,16,17 (additional reviewed articles are listed in Supplementary File S1 online).
We carried out a collaborative international study to further characterize and delineate the phenotype, natural history, and genotype–phenotype correlation of MWS.
We analyzed clinical data for 87 patients with a molecularly confirmed diagnosis of MWS, including 62 previously reported patients and 25 unpublished cases, and compared them with patients previously reported by other authors. We obtained these data through collaborations involving clinicians from various countries. Such primary data have never been collated from a large cohort of affected individuals.
In this article we present a comprehensive study of MWS features underlining a highly consistent phenotype for the disease, its genotype–phenotype correlations, and the phenotypic and clinical evolution taking place with age. The purpose of the article is to assist clinicians to identify the disease and to provide them with updated care recommendations for patient management.13
Materials and methods
Individuals affected by MWS and with a ZEB2 variation confirmed by appropriate methods (Supplementary File S2) were identified with the help of an international working group of collaborating clinicians and with the support of the Italian MWS Association. Through the respective clinicians, we invited them to fill in a highly detailed questionnaire. Informed consent for inclusion was obtained for all individuals involved in the study. Separate written informed consent was obtained for the publication of photographs. Approval for this study was granted by the Ethical Committee of Reggio Emilia.
Clinical and genetic data were transferred to an electronic database, and processed using the Statistical Package for the Social Sciences (SPSS, IBM, Chicago, IL), version 20.0. Group differences in phenotypes were compared using chi-square and Fisher’s exact tests. Post hoc analysis was applied with Bonferroni correction for all multiple comparisons. P<0.05 was considered statistically significant.
Results
A total of 87 patients are reported, ranging in age from 6 months to 36 years, including individuals not previously reported. The clinical diagnosis of MWS was confirmed by genetic testing in all of them. Patients originated from four continents and various ethnic backgrounds, although the vast majority were from Europe. The male-to-female ratio was 45:42. The mean age at the last clinical evaluation was 9 years and 10 months.
The main clinical features are presented in Table 1, in comparison with the literature. All numbers and percentages are related to the total number of patients that could be evaluated for a given feature. Figure 1 shows photographs of patients at various ages. Comprehensive clinical data are presented in Figure 2.
(a) Facial features, (b) skeletal features, (c) neurological phenotype, (d) developmental milestones, (e) congenital heart defects, (f) intestinal tract anomalies, (g) urogenital tract anomalies, (h) eye anomalies, (i) dental anomalies, (j) other features, (k) behavioral phenotype. *Normal developmental milestones range: sitting without support, 5–7 months; walking without support, 10–14 months; first words, 10–14 months. ASD, atrial septal defect; CHD, congenital heart disease; PDA, patent ductus arteriosus; SG, secondary generalization, VSD, ventricular septal defect; VUR, vesicoureteral reflux.
Family history
The average age of fathers and mothers at time of delivery was 34 years and 6 months and 31 years and 7 months respectively. No consanguinity was reported. Regarding two affected sisters presenting the same ZEB2 variant, germ-line mosaicism was considered because both parents had normal results on testing the mutation in blood.18 Germ-line mosaicism had already been considered in some previous reports.19,20
Birth history
The average gestational age was 39 weeks, with a range between 33 and 43. Weights at birth were within the normal range in both males and females. The average weight in males was 3,408 g, with a range between 2,140 g and 4,680 g; the average weight in females was 3,110 g, with a range between 1,240 g and 4,100 g (Supplementary Table S1). The birth weight of all males was between the 2nd and 100th centiles for gestational age (mean 51st centile); the females’ birth weight was between the 3rd and 100th centiles (mean 54th centile).
Length at birth was within the normal range for almost all the patients, both males and females. The average birth length in males was 49.9 cm, with a range between 45.5 and 54 cm (range 2nd and 99th centiles, mean 44th). The average birth length in females was 49.2 cm, with a range between 37 and 56 cm (range <1st and 100th centiles, mean 53rd).
Although the average occipital frontal circumference at birth was within the normal range in both genders, the majority of patients had circumferences under the 50th centile, with mean values of 27th centile for males and 31st for females. In males, the mean was 33.5 cm, with a range between 29.2 and 37 cm (<1st and 96th centiles); in females, the mean was 32.6 cm, with a range between 27 and 35 cm (<1st and 100th centiles).
Head circumference and growth
Microcephaly, i.e., occipital frontal circumference at least 2 SDs below the mean, was sometimes observed at birth, but more frequently tended to develop gradually throughout infancy but not all children were microcephalic (Figure 3a). Postnatally, microcephaly was noted in 5 of 14 patients (35.7%, mean 47.7±2.6 cm) in the age range from 0 to 5 years.
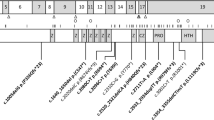
(a) Growth parameters at birth and at last clinical evaluation postnatally. (b) Distribution of the different ZEB2 defects found in our cohort. (c) Position of genetic defects identified in our cohort on a schematic representation of the ZEB2 transcript and protein. (d) Correlation between developmental milestones and broad categories of the underlying heterozygous ZEB2 variants. Absence of protein (synthesis), defective protein (synthesis), and full-gene deletion all refer to the variant allele. Bars represent mean values, boxes represent median with interquartile (25th–75th percentile) range, whiskers represent range, and dots represents outliers. Only one patient in the full-gene deletion group developed speech, therefore no box plot representation was possible. (e) Number of cardiac anomalies in relation to broad categories of the underlying ZEB2 variants. mRNA, messenger RNA; OFC, occipital frontal circumference, PN postnatally.
The majority of patients had a height below the 50th centile (68/80, 85%). Short stature was observed in 7 of 29 patients (24.13%, height 89.4±14.9 cm) in the age range from 0 to 6 years, in 8 of 25 patients (32%, height 125±13.4 cm) in the age range from 6 to 12 years, and in 16 of 25 patients (64%, height 150.8+16.8 cm) over 12 years. Adult height in males varied between 155 and 185 cm (n=9, mean 165.1 cm) and in females between 146 and 155 cm (n=2, mean 150.5 cm).
Most patients were of slender build, with normal body proportions. Body weight was <50th centile in 63 of 76 patients and <3rd centile in 25 of 76.
Overall, height (length) and weight were globally normal at birth, but often showed marked delay over time. Occipital frontal circumference, already under the 50th centile at birth, tended to fall progressively below normal values (Figure 3a).
Facial phenotype
All patients presented a distinctive facial appearance that renders the condition recognizable (Figure 1). A detailed list of the facial features is reported in Figure 2a.
Musculoskeletal anomalies
Musculoskeletal anomalies were frequent and various. A detailed list of skeletal anomalies reported in our cohort can be found in Figure 2b.
Neurodevelopment, brain anomalies, and epilepsy
MWS patients have moderate to severe ID. Hypotonia is often found in the first years of life and was detected in 79.1% of patients (Figure 2c). Developmental milestones such as sitting and walking were considerably delayed (Figure 2d): the mean age of sitting unsupported was 17.39 months, and the mean age of walking was 3 years and 9 months (range: 16 months to 11 years), although some individuals remain nonambulatory (33 of 86, 38.3%). A number of patients had a wide-based or ataxic-like gait, and sometimes held their arms up and flexed at the elbow, typical of individuals with Angelman syndrome. All milestones for fine motor skills were delayed. The majority of the oldest individuals (aged over 20 years) require help with dressing and other everyday activities. Speech rarely extends to more than a few words, with onset averaging at 4 years. Many patients have absence of speech (58 of 86 (67.4%), 10 of them still under 4 years of age), but show receptive language skills and communicate successfully using alternative methods, such as augmentative and alternative communication (AAC).
The neuroradiological phenotype in MWS patients, delineated recently,21 was characterized by anomalies of corpus callosum (79.6% of cases), hippocampal abnormalities (77.8%), enlargement of cerebral ventricles (68.5%), and white matter abnormalities (reduction of thickness 40.7%, localized signal alterations 22%). We also observed large basal ganglia and cortical and cerebellar malformations.
It was observed that epilepsy was one of the most frequently detected features of MWS. Of the 87 patients in our cohort, 83.9% were found to have had seizures (n=73/87) with a mean onset occurring at the age of 27.5 months. The most precocious onset in our cohort was registered at 1 month of age and the latest onset at 11 years of age. Our studies showed that 45% of patients have suffered fever-triggered seizures, 72.1% focal seizures, 50.8% generalized seizures, and 40% focal seizures with secondary generalization. In addition, 25.9% of patients had resistance to antiepileptic drugs (Figure 2c).
Cardiac anomalies
Congenital heart disease occurred in 60.5% of patients (52/86) (Figure 2e). The most frequent cardiac anomalies (>25% of cases) were not complex heart diseases and included patent ductus arteriosus, ventricular septal defect, and atrial septal defect. Pulmonary stenosis was present in almost 20% of patients and aortic coarctation in more than 10%. Other congenital heart defects with an incidence lower than 10% included bicuspid aortic valve, aortic valve stenosis, tetralogy of Fallot, pulmonary artery sling, proximal pulmonary artery stenosis, mitral valve prolapse, and pulmonary atresia.
Gastrointestinal tract
Constipation and Hirschsprung disease were frequent, occurring in 43.5% and 30.6% of patients, respectively (Figure 2f). Less common clinical features were inguinal hernia and pyloric stenosis.
Urogenital/renal anomalies
Urogenital anomalies were detected in 49 of 80 patients (61.25%; Figure 2g). The most frequent were hypospadias and cryptorchidism. Less common urogenital anomalies were webbed penis, microphallus, bifid scrotum, and vaginal septum. Renal anomalies included hydronephrosis, vesicoureteric reflux and, much more rarely, pelvic kidney.
Eye anomalies
The most frequent anomaly by far was strabismus (56.8%; Figure 2h). Quite common conditions were astigmatism, more common in females than in males (F:M=2.6:1, P=0.031), and myopia. Less frequent anomalies were nystagmus, palpebral ptosis, Axenfeld anomaly, microphthalmia, and irides heterochromia. Coloboma was present in only one patient.
Tooth anomalies
The two most frequent anomalies reported in our group were widely spaced teeth (50%) and delayed tooth eruption (47.7%) (Figure 2i). Other less frequent but still fairly common anomalies were malpositioned teeth (34.3%) and dental crowding (26.4%).
Other clinical features
Recurrent otitis media was frequent, detected in 34.6% of patients. This feature must be taken into account whenever children present with fever because affected children tend to have a high pain threshold. It can contribute to conductive hearing loss, which in our cohort was present in 9 of 77 patients (11.6%). Sensorineural hearing loss was less common (4/77, 5.2%).
Repeated vomiting attacks were fairly common, with 14 of 77 patients detected (18.2%).
Skin anomalies included hyperpigmentation (12/79, 15.2%), depigmentation (10/78, 12.8%), and accessory nipples (8/80, 10%).
Asplenia was fortunately rare (only 1 patient in our cohort), but it is important to investigate its presence, since it has been associated with serious infections (Figure 2j).
Behavioral phenotype
Some behavioral types were particularly frequent (present in over 60% of patients), including chewing or mouthing objects or body parts, underreaction to pain, and grinding of teeth. Other patterns, such as laughing for no obvious reason, fidgeting (flicking, tapping, and twirling of objects), unrealistic happiness or elation, switching lights on and off (more common in males than in females), rapid mood changes, standing close to others, and eating nonfood items, were also common (30–50%; Figure 2k).
Genotype
All the genetic defects in our cohort were heterozygous, including chromosome deletions encompassing the ZEB2 locus on 2q21-q23 (detected by comparative genomic hybridization and multiplex ligation-dependent probe amplification), exon deletions (detected by multiplex ligation-dependent probe amplification), and several different intragenic mutations (detected by direct sequencing).
Chromosome deletions (11.5%) involved an entire ZEB2 allele or its first exons, and ranged in length from a few hundred kilobases to 16.7 Mb. Additionally, we found two intragenic deletions: one no larger than 11 kb and comprising exons 8–9 and the other no larger than 28 kb, encompassing exons 5–8, and leading to an altered reading frame downstream of the deletion.
The majority of pathogenic defects were intragenic mutations, either nonsense mutations (37.9%), or small insertions/deletions/indels (46%) generating a frameshift and a premature stop codon (or, in one case, a longer protein with an altered C-terminus sequence).
Other variants included a deletion in the 3′ UTR region of ZEB2, probably altering transcript synthesis or regulation, and an intronic variation altering the splice acceptor site of exon 2, expected to generate a shorter protein with an altered N-terminus sequence by similarity to the variant observed by Zweier et al.22
No patients with missense mutations were detected.
Genetic defects in our cohort are summarized in Supplementary Table S2 online and Figure 3b.
Discussion
With more than 300 patients reported to date, MWS has become a well-known entity in clinical dysmorphology. Distinctive facial appearance is the most reliable clinical feature for recognizing this condition. Despite the fact that the facial gestalt is consistent with this diagnosis from an early age, a change in certain facial features with age has been reported.15 In childhood, earlobes seem to be uplifted, with a central depression, and eyebrows are large, medially flaring, and sparse in the middle. Facial features at this age also consist of a rounded skull, sparse and fine hair, puffy anterior neck, and excess skin on the rear of the neck. In childhood, the face is square with a high forehead, frontal bossing, epicanthal folds, hypertelorism, telecanthus, large and deep-set eyes, strabismus, a broad nasal bridge, a rounded nasal tip, a prominent columella, an open mouth with an M-shaped upper lip, and a prominent and triangular chin. In addition, children with MWS tend to smile frequently. In children of school age, the face is longer, with a prominent jaw. The eyebrows are broad, usually exhibiting sparseness in the middle part.
Beginning in adolescence, the eyebrows often become heavier, broad, and horizontal; the nasal profile is more convex; the nasal tip lengthens, becomes more depressed, and overhangs the philtrum; the face tends to lengthen; and a long, pointed, or “chisel-shaped” chin may be observed.
The uplifted earlobes do not alter significantly over time apart from the central depression, which is less noticeable in adults, and this feature is useful in confirming suspicion of this condition.
What is most useful for clinicians to identify this condition is the overall facial gestalt, which changes over time. Figure 2a lists the most frequent facial features in our cohort in order of frequency. Two patients presented 25 of 26 facial features; the average number was 15.8. The three most frequent features (uplifted earlobes, a rounded nasal tip, and a prominent columella) were present together in 40 patients.
Before the delineation of a facial phenotype, HSCR was considered the hallmark of MWS. This led to reports of a high proportion of MWS patients with HSCR and underdiagnosis of patients with MWS but without HSCR. Afterward, as more MWS patients without HSCR were identified based on other features, the frequency of HSCR began to decrease. After reviewing the literature and analyzing our cohort’s results, we concluded that the combined frequencies of HSCR and constipation were above 72%. The difference in HSCR frequency in our cohort and the literature (30.6% vs. 48.8%) reflects an enhanced knowledge of the clinical phenotype, which leads to the fact that patients are now not diagnosed on the basis of the presence of HSCR but rather on the basis of the facial gestalt. Because a considerable number of cases with severe constipation were not investigated by rectal biopsy due to risk of complications or lack of compliance, it is possible that the presence of HSCR is now underestimated. The data regarding the length of the aganglionic segment is not exhaustive in previously published cases, including our cohort. However, short and long segments have been reported both in males, in whom HSCR was more frequent, and in females.
Of note, although pulmonary artery sling with or without tracheal stenosis is quite rare (3.7%), in the general population this cardiac anomaly is present in 1 of 17,000 school-aged children (0.006%).23 This would make pulmonary artery sling more common in MWS patients, and we can therefore consider it a sign that may lead us to suspect MWS.
Ophthalmological findings have already been associated with MWS.24 Strabismus is the most common finding, followed by astigmatism, with a 2.6 times higher frequency in girls. Structural anomalies are rare, with Axenfeld anomaly and coloboma the most frequent.
Behavioral phenotypes in MWS have been delineated previously.25 According to our observations of the affected individuals, one feature requires special attention. Over 60% of patients (42 of 65) show underreaction to pain, which can be dangerous for these children. A recent study showed that the underreaction to pain results from a reduced responsivity to nociceptive stimulation rather than an inability to communicate discomfort.26
One of the major challenges posed by MWS is the recurrence of seizures.27 In 2013 Cordelli et al.28 hypothesized that a distinct electroclinical phenotype featuring focal and atypical absence seizures (which are often preceded by febrile seizures) and age-dependent electroencephalogram changes can be detected in the majority of patients with MWS.
Moreover, a recent paper21 provided no evidence of a significant correlation between the brain malformation phenotype and the epilepsy phenotype in our cohort. This reinforces the hypothesis put forward by Cordelli et al.29 that epilepsy in MWS has a genetic etiology.
The prevalence of MWS is low, which makes it still unknown to the majority of clinicians. This means that clinicians (pediatricians, neurologists, and/or clinical geneticists and genetic counselors) should be responsible for the coordination of patients’ care and management. As the number of adult patients is increasing, the role of neurologists and clinical geneticists tends to become even more important. Multidisciplinary follow-up is essential and should consist of regular evaluations performed by different specialists; in the first place, pediatricians, neurologists, cardiologists, nephrologists, and gastroenterologists, especially during early childhood. Because there is no cure at this time, the goal of follow-up is improving quality of life. We strongly recommend AAC as a key point for MWS patient care. AAC has been recognized as a valuable tool to stimulate patients’ communication skills, thus facilitating clinicians and parents in their compliance and determination of the behavioral phenotype. A detailed follow-up summary is reported in Table 2.
Genotype–phenotype correlation
We have grouped the genomic defects involving a single allele of ZEB2 into three major categories: deletions encompassing the whole gene, intragenic variants resulting in no protein synthesis/fast protein degradation, and intragenic variants resulting in synthesis of a defective protein.
The relatively small number of cases and the scarcity of functional data make it difficult to establish genotype–phenotype correlations and assess their statistical significance, particularly where intragenic mutations are concerned.
Only a few clinical features show some level of significance for genotype–phenotype correlation (Supplementary Table S3), and they consistently involve comparing full-gene deletion with another genotypic category. These locus deletions often span several megabases of DNA and include other genes and long noncoding RNAs, such as the ZEB2-antisense transcripts encoded in the 5′ portion of ZEB2 and involved in its transcriptional regulation.30 The variability and extent of these deletions may explain why they are usually associated with increased severity of some aspects, such as sitting age and onset of speech (Figure 3d). Full-gene deletion, or to a lesser extent the presence of a residual defective protein, also seems to negatively influence the number of cardiac defects, although their combined numbers are too small compared with cases of protein absence due to intragenic mutations (Figure 3e).
Transcripts from ZEB2 alleles with intragenic mutations are mostly predicted to undergo nonsense-mediated messenger RNA decay, or to generate very short and allegedly nonfunctional peptides. Here the variability of clinical presentations is likely due to a series of environmental and possibly genetic or epigenetic cofactors. Missense mutations of ZEB2 are a rare occurrence and were found in none of our cases. They have been described in the literature in association with mild presentations sharing some similarity with MWS,31,32 although it is debatable whether most of these atypical cases should actually be classified as MWS.
However, in addition to patients with missense mutations, there are a few cases also expected to behave differently than typical haploinsufficiency. Notably, the least severely affected of our patients (P23)—a 12-year-old girl with typical facial features but mild to moderate ID, no seizures, absence of HSCR—has a frameshift mutation predicted to disrupt only part of the C-terminal zinc-finger domain: c.3031delA, p.(S1011Afs*64). Our preliminary studies, involving ZEB2 recombinant constructs expressed in a HEK293T cell line, indicate that this variant protein correctly localizes in the nucleus and retains some repressor ability on the E-cadherin promoter in a dual luciferase assay (unpublished data), possibly explaining the favorable clinical phenotype. Another patient (P34) with a very mild clinical presentation has a splicing defect that alters only the N-terminal portion of the protein and abolishes the NuRD interacting motif, similarly to what was found in another patient.22 These observations seem to indicate that whenever a variant ZEB2 protein is predicted to preserve some of its functionality, we can expect a milder clinical presentation. The reverse might not always be the case, however; some mild cases have been found to have mutations abolishing ZEB2 protein function, suggesting the contribution of other genetic or environmental factors in shaping the MWS phenotype.
Conclusions
This paper analyzes data from the largest group of MWS patients reported to date.
Despite the variability of many features, MWS has a distinctive phenotype and is also genetically homogeneous, as all patients have a de novo mutation in a single gene, ZEB2, except for the rarest of cases in which a germ-line mosaicism has been proved in one of the parents.
The facial gestalt is easily recognizable, apart from the very rare patients with missense mutations. These should be discussed case by case as to whether a patient actually has MWS.
Since short stature is one of the most common features, we are currently collecting additional data to generate specific growth charts that will aid clinicians to improve the management and follow-up of children with MWS.
With the growing use of whole-exome sequencing for genes causing ID, we will undoubtedly identify ZEB2 mutations in patients who have a less obvious diagnosis of MWS. The study of these patients, as well as those with a milder phenotype and those with missense mutations, can help to define the boundaries of this condition. Better knowledge of the phenotypic spectrum of MWS and its genotype–phenotype correlations is likely to improve the detection rate of this condition and facilitate prediction of some MWS characteristics and a correct approach to some important features, such as epileptic seizures. This will hopefully aid in improving care for the patients and bringing much-needed support to their families.
References
Mowat DR, Croaker GD, Cass DT et al. Hirschsprung disease, microcephaly, mental retardation, and characteristic facial features: delineation of a new syndrome and identification of a locus at chromosome 2q22-q23. J Med Genet 1998;35:617–623.
Lurie IW, Supovitz KR, Rosenblum-Vos LS, Wulfsberg EA. Phenotypic variability of del(2) (q22-q23): report of a case with a review of the literature. Genet Couns. 1994;5:11–14.
Wakamatsu N, Yamada Y, Yamada K et al. Mutations in SIP1, encoding Smad interacting protein-1, cause a form of Hirschsprung disease. Nat Genet. 2001;27:369–370.
Cacheux V, Dastot-Le Moal F, Kaariainen H et al. Loss-of-function mutations in SIP1 Smad interacting protein 1 result in a syndromic Hirschsprung disease. Hum Mol Genet 2001;10:1503–1510.
Zweier C, Albrecht B, Mitulla B et al. “Mowat-Wilson” syndrome with and without Hirschsprung disease is a distinct, recognizable multiple congenital anomalies-mental retardation syndrome caused by mutations in the zinc finger homeo box 1B gene. Am J Med Genet 2002;108:177–181.
Yamada K, Yamada Y, Nomura N et al. Nonsense and frameshift mutations in ZFHX1B, encoding Smad-interacting protein 1, cause a complex developmental disorder with a great variety of clinical features. Am J Hum Genet 2001;69:1178–1185.
Wilson M, Mowat D, Dastot-Le Moal F et al. Further delineation of the phenotype associated with heterozygous mutations in ZFHX1B. Am J Med Genet A 2003;119A:257–265.
Zweier C, Temple IK, Beemer F et al. Characterisation of deletions of the ZFHX1B region and genotype-phenotype analysis in Mowat-Wilson syndrome. J Med Genet 2003;40:601–605.
Cerruti Mainardi P, Pastore G, Zweier C, Rauch A. Mowat-Wilson syndrome and mutation in the zinc finger homeo box 1B gene: a well defined clinical entity. J Med Genet 2004;41:e16.
Ishihara N, Yamada K, Yamada Y et al. Clinical and molecular analysis of Mowat-Wilson syndrome associated with ZFHX1B mutations and deletions at 2q22-q24.1. J Med Genet 2004;41:387–393.
Garavelli L, Cerruti-Mainardi P, Virdis R et al. Genitourinary anomalies in Mowat-Wilson syndrome with deletion/mutation in the zinc finger homeo box 1B gene (ZFHX1B). Report of three Italian cases with hypospadias and review. Horm Res. 2005;63:187–192.
Zweier C, Thiel CT, Dufke A et al. Clinical and mutational spectrum of Mowat-Wilson syndrome. Eur J Med Genet 2005;48:97–111.
Adam MP, Schelley S, Gallagher R et al. Clinical features and management issues in Mowat-Wilson syndrome. Am J Med Genet A 2006;140:2730–2741.
Dastot-Le Moal F, Wilson M, Mowat D, Collot N, Niel F, Goossens M. ZFHX1B mutations in patients with Mowat-Wilson syndrome. Hum Mutat. 2007;28:313–321.
Garavelli L, Zollino M, Mainardi PC et al. Mowat-Wilson syndrome: facial phenotype changing with age: study of 19 Italian patients and review of the literature. Am J Med Genet A 2009;149A:417–426.
Yamada Y, Nomura N, Yamada K et al. The spectrum of ZEB2 mutations causing the Mowat-Wilson syndrome in Japanese populations. Am J Med Genet A 2014;164A:1899–1908.
Coyle D, Puri P. Hirschsprung’s disease in children with Mowat-Wilson syndrome. Pediatr Surg Int 2015;31:711–717.
Cecconi M, Forzano F, Garavelli L et al. Recurrence of Mowat-Wilson syndrome in siblings with a novel mutation in the ZEB2 gene. Am J Med Genet A 2008;146A:3095–3099.
McGaughran J, Sinnott S, Dastot-Le Moal F et al. Recurrence of Mowat-Wilson syndrome in siblings with the same proven mutation. Am J Med Genet A 2005;137A:302–304.
Ohtsuka M, Oguni H, Ito Y et al. Mowat-Wilson syndrome affecting 3 siblings. J Child Neurol 2008;23:274–278.
Garavelli L, Ivanovski I, Caraffi SG et al. Neuroimaging findings in Mowat-Wilson syndrome: a study of 54 patients. Genet Med. 2017;19:691–700.
Zweier C, Horn D, Kraus C, Rauch A. Atypical ZFHX1B mutation associated with a mild Mowat-Wilson syndrome phenotype. Am J Med Genet A 2006;140:869–872.
Yu JM, Liao CP, Ge S et al. The prevalence and clinical impact of pulmonary artery sling on school-aged children: a large-scale screening study. Pediatr Pulmonol. 2008;43:656–661.
Bourchany A, Giurgea I, Thevenon J et al. Clinical spectrum of eye malformations in four patients with Mowat-Wilson syndrome. Am J Med Genet A 2015;167:1587–1592.
Evans E, Einfeld S, Mowat D, Taffe J, Tonge B, Wilson M. The behavioral phenotype of Mowat-Wilson syndrome. Am J Med Genet A 2012;158A:358–366.
Pradier B, Jeub M, Markert A et al. Smad-interacting protein 1 affects acute and tonic, but not chronic pain. Eur J Pain 2014;18:249–257.
Garavelli L, Mainardi PC. Mowat-Wilson syndrome. Orphanet J Rare Dis 2007;2:42.
Cordelli DM, Garavelli L, Savasta S et al. Epilepsy in Mowat-Wilson syndrome: delineation of the electroclinical phenotype. Am J Med Genet A 2013;161A:273–284.
Cordelli DM, Pellicciari A, Kiriazopulos D, Franzoni E, Garavelli L. Epilepsy in Mowat-Wilson syndrome: is it a matter of GABA? Epilepsia. 2013;54:1331–1332.
Beltran M, Puig I, Pena C et al. A natural antisense transcript regulates Zeb2/Sip1 gene expression during Snail1-induced epithelial-mesenchymal transition. Genes Dev. 2008;22:756–769.
Ghoumid J, Drevillon L, Alavi-Naini SM et al. ZEB2 zinc-finger missense mutations lead to hypomorphic alleles and a mild Mowat-Wilson syndrome. Hum Mol Genet 2013;22:2652–2661.
Heinritz W, Zweier C, Froster UG et al. A missense mutation in the ZFHX1B gene associated with an atypical Mowat-Wilson syndrome phenotype. Am J Med Genet A 2006;140:1223–1227.
Acknowledgments
The financial support of Telethon Italy (grant GEP 14131) is gratefully acknowledged. The authors thank the Galliera Genetic Bank, member of the Telethon Genetic Biobank Network (project GTB12001), funded by Telethon Italy, and the Associazione Italiana Mowat Wilson ONLUS (AIMW), for their assistance and kind collaboration. We also thank all cooperating family members for contributing the medical data necessary for this study. In addition, we are grateful to Luca Valcavi for helping with the design and adaptation of the figures and tables. We also thank the photographer Marco Bonazzi, the genetic nurse Maria Claudia Menozzi, and the nurse Margherita Raucci for their excellent work. B.C. is a senior clinical investigator of the Scientific Research Fund—Flanders. Written consent for publication of the clinical pictures was obtained from the patients’ parents.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclosure
The authors declare no conflict of interest.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copyof this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Ivanovski, I., Djuric, O., Caraffi, S. et al. Phenotype and genotype of 87 patients with Mowat–Wilson syndrome and recommendations for care. Genet Med 20, 965–975 (2018). https://doi.org/10.1038/gim.2017.221
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2017.221
Keywords
This article is cited by
-
Mowat-Wilson syndrome: unraveling the complexities of diagnosis, treatment, and symptom management
Egyptian Journal of Medical Human Genetics (2024)
-
Identification of the DNA methylation signature of Mowat-Wilson syndrome
European Journal of Human Genetics (2024)
-
“Liu-Liang-Chung” syndrome with multiple congenital anomalies and the distinctive craniofacial features caused by dominant ZEB2 gene gain mutation
BMC Pediatrics (2023)
-
Profile of copper-associated DNA methylation and its association with incident acute coronary syndrome
Clinical Epigenetics (2021)
-
Clinical characteristics of Polish patients with molecularly confirmed Mowat-Wilson syndrome
Journal of Applied Genetics (2021)