Abstract
Purpose
Noninvasive prenatal screening (NIPS) sequences a mixture of the maternal and fetal cell-free DNA. Fetal trisomy can be detected by examining chromosomal dosages estimated from sequencing reads. The traditional method uses the Z-test, which compares a subject against a set of euploid controls, where the information of fetal fraction is not fully utilized. Here we present a Bayesian method that leverages informative priors on the fetal fraction.
Method
Our Bayesian method combines the Z-test likelihood and informative priors of the fetal fraction, which are learned from the sex chromosomes, to compute Bayes factors. Bayesian framework can account for nongenetic risk factors through the prior odds, and our method can report individual positive/negative predictive values.
Results
Our Bayesian method has more power than the Z-test method. We analyzed 3,405 NIPS samples and spotted at least 9 (of 51) possible Z-test false positives.
Conclusion
Bayesian NIPS is more powerful than the Z-test method, is able to account for nongenetic risk factors through prior odds, and can report individual positive/negative predictive values.
Similar content being viewed by others
Introduction
In 1997, Dennis Lo and colleagues1 discovered the existence of fetal cell-free DNA in the plasma of the maternal peripheral blood. They used polymerase chain reaction to amplify a segment that is unique to the human Y chromosome; therefore, the polymerase chain reaction signal detected has to come from a male fetus. This landmark discovery laid the foundation for noninvasive prenatal screening (NIPS). More than a decade later, during which time massively parallel sequencing technologies made rapid progress, Stephen Quake’s and Dennis Lo’s groups2,3 reported independent successes in detecting fetal trisomy by sequencing cell-free DNA in maternal peripheral plasma. Following these reports, several clinical trials demonstrated more than convincingly the benefit of NIPS over traditional screening in detecting fetal trisomy.4,5,6 Consequently, next-generation sequencing–based NIPS has been rapidly integrated into prenatal care.
In light of compelling new evidence, professional societies endorsed NIPS over the traditional trisomy screen.7,8,9,10,11 Particularly, the American College of Medical Genetics and Genomics recently revised their early position of restricting NIPS to high-risk patients12 to recommend that “NIPS can replace conventional screening for Patau, Edwards, and Down syndromes across the maternal age spectrum, for a continuum of gestational age beginning at 9–10 weeks, and for patients who are not significantly obese.”11 The recommendation for gestational age and maternal weight is to ensure that the fetal fraction (the proportion of cell-free DNA that is originated from the fetus) is large enough for NIPS to be effective. The American College of Medical Genetics and Genomics emphasizes the importance of the fetal fraction and recommends that all laboratories “include a clearly visible fetal fraction on NIPS reports.” Multiple studies have pointed out that the fetal fraction plays a crucial role in the effectiveness of NIPS to detect trisomy.13,14,15,16 The consensus for the detecting limit of the fetal fraction appears to be 4%, although theoretical studies have suggested that 2% also works,14 and more optimistic authors have suggested that as long as GC bias is accounted for and sequence depth is unlimited, NIPS can be effective for an arbitrarily small fraction of fetal DNA.17
The prevailing method used to analyze NIPS data sets, first outlined in ref.3, is the Z-test method, which calculates a Z-score that measures deviation of a chromosomal dosage from a set of euploid control samples. The Z-test pays no special attention to the fetal fraction, other than that the deviation approximately equals the fetal fraction for a trisomy fetus. (For a euploid fetus, the deviation is approximately 0.) A chromosomal dosage can be estimated reliably from low-coverage sequencing reads (Supplementary Materials and Methods online). NIPS uses euploid control samples to define a baseline. Their chromosomal dosages are estimated, the sample mean μ and the sample standard deviation σ are computed, where μ is expected to be 2, but varies slightly from chromosome to chromosome, and different chromosomes have markedly different σ estimates (Supplementary Figure S1). Suppose a euploid mother carries a fetus, and denote c the estimated dosage of an autosome. Let h be the fetal fraction, then c is expected to be μ + h when the fetus is trisomy, and μ − h when the fetus is monosomy. We have either h = c−μ (for trisomy) or −h = c−μ (for monosomy); combined together, h = ∣c−μ∣. Define x = c−μ, which is the centered chromosomal dosage, then ∣x∣ is an unbiased estimator of h for a trisomy or monosomy fetus. (Note when the fetus is euploid, x is expected to be 0 and irrelevant to h.) The Z-score is defined as Z = x/σ. Obviously, a larger h tends to produce a more significant Z-score.
We identify three inadequacies of the Z-test method in NIPS. First, the Z-score is measured relative to σ. A euploid control sample may have an x estimate that is small enough to appear by chance, but large enough relative to σ such that a significant Z-score is obtained, resulting in a false positive. Second, it is well known that fetal trisomy has an increased risk with respect to maternal age.18 The Z-test method has difficulty incorporating such information. Third, with the Z-score one has to specify a threshold to call positives, negatives, and no calls. But such a threshold varies for different investigators3,19 and varies even for the same investigators over time.3,4 Here we develop a Bayesian method and demonstrate its advantage over the Z-test method through analyzing a real data set. The Bayesian method allows us to emphasize the fetal fraction through informative priors. The informative priors effectively down-weight a Z-score whose corresponding ∣x∣ is small. This alleviates the first inadequacy of the Z-test method. The test statistic produced by the Bayesian method is the Bayes factor.20 Bayes factor is the change of odds (of trisomy) in light of the data, and we can compute posterior odds of trisomy by multiplying Bayes factor by prior odds of trisomy. The prior odds can be the age-adjusted prevalence of trisomy of an autosome. This addresses the second inadequacy of the Z-test method. From the posterior odds, we can compute and report the positive predictive value (PPV) and the negative predictive value (NPV). Because both PPV and NPV are probabilities, they are easily interpretable and more informative than a Z-score. This mitigates the third inadequacy of the Z-test method.
Materials and methods
In the Supplementary Materials and Methods we documented how patients were recruited and data collected. We also provided details on read quality control; how we decided the optimal bin sizes when binning reads together; the hidden Markov model we used to remove maternal copy-number variants, as well as the regions harboring copy-number variants at the population level due to reference bias; how we accounted for GC bias; how we inferred fetal fractions from sex chromosomes; how we fit candidate probability densities to empirical distribution of fetal fractions and the densities we obtained; how we computed Bayes factor; and how we simulated chromosomal dosages to compare Bayes factors against Z-scores.
Code availability
R-code implementing methods described in the paper are available upon request.
Results
Fetal fractions
We first inferred fetal fractions (denoted by h) for each sample using sex chromosomes. The empirical distributions of these fetal fractions were used to formulate our informative priors for Bayesian analysis. For a female fetus the expected chromosome X and Y dosages are 2 and 0 respectively, and fetal fraction of a female fetus estimated from sex chromosomes is expected to be 0. For a male fetus, the expected X dosage is 2−h and the expected Y dosage is h. Thus, both X and Y dosages are informative to the fetal fraction of a male fetus. Our data suggested that the Y dosage is more informative than the X dosage (Figure 1a). We estimated the fetal fractions for all samples using both X and Y dosages (Supplementary Materials and Methods). Figure 1b shows the histograms of the inferred fetal fraction (on the log scale). The two modes correspond to putative female and male fetuses. To avoid arbitrariness in specifying a threshold, we used the K-means method to divide the log fetal fractions into two groups. The inferred threshold of fetal fraction is 0.028. Of 3,405 samples analyzed, 1,693 (49.7%) had fetal fractions greater than 0.028 and thus were carrying putative male fetuses, and the remaining 1,712 (50.3%) samples were carrying putative female fetuses.
Fetal fractions estimated from sex chromosomes. (a) Plot shows that both X and Y chromosome contribute to fetal fraction estimates, as expected, and the Y is more informative than the X. (b) Plot shows histogram of the observed fetal fractions and fitted beta density (solid line) and LogN density (dashed line). The x-axis is on the log10 scale.
Taking fetal fraction estimates from samples carrying putative male fetuses, we digressed to investigate how maternal age, gestational age, and maternal weight affect the log fetal fraction. Because these investigations relied on the linear regression, we used the log fetal fraction instead of fetal fraction, as the former was a better fit to a normal distribution, which is the basic assumption for linear regression. Maternal age in our sample followed a bimodal distribution, with two modes at 27 and 35 years of age (Supplementary Figure S2). Evidently, women in their late childbearing years took advantage of the new “two-child policy” in China effective 1 January 2016. Simple linear regression suggested an association between maternal age and log fetal fraction (P=6.0 × 10−10), but the association disappeared after controlling for gestational age and maternal weight (P = 0.12). This nonassociation agrees with a previous study using American samples of European descents.21 The gestational age is positively associated with the log fetal fraction (P = 7.2 × 10−24) and maternal weight is negatively associated with the log fetal fraction (P = 4.0 × 10−49). Both agree with earlier studies.2,13 Gestational age accounts for 4.2% variation of log fetal fraction and maternal weight accounts for 13.9%, and combined they account for 15.4%.
Prior specification
The computation of the Bayes factor requires one to integrate the likelihood, which contains data and the parameter (fetal fraction here), over the prior distributions of the parameter, separately for the null and alternative models (Supplementary Materials and Methods). Thus, we need to specify and justify priors for both null (euploidy) and alternative (trisomy) models. Indeed, the art of Bayesian methods is prior specification. (The toil of Bayesian methods is computation, but for our application the computation is rather simple.) The expected fetal fraction estimated from the sex chromosomes for a putative female fetus is 0, and the observed variation of the estimates can be regarded as the variation under the null model. The fetal fractions of male fetuses can be reliably estimated from the sex chromosomes (in the absence of mosaicism), and the observed variation can be regarded as the variation under the alternative model. After visual inspection of the histograms, we elected to use beta and log-normal (LogN) distributions to fit the empirical distributions of the fetal fractions. With fetal fractions of putative female fetuses, the fitted density was denoted by g0*; with those of putative male fetuses, g1*; and with those of female and male fetuses combined, g2*, where * can be either B (for beta) or L (for LogN). The parameters of the fitted distributions were determined by matching the means and the variances of an empirical distribution and a candidate distribution. The explicit forms of the fitted distributions can be found in the Supplementary Materials and Methods, and Figure 1b demonstrates the goodness of fit.
Using the sharp null is common practice in Bayesian methods. In our context, the sharp null specifies the null distribution as a point mass h = 0, as opposed to assuming h following a distribution that has its density concentrating near 0 but with variations. The sharp null, however, is a choice often made for convenience rather than merits.22 In our application, the sharp null produced much-inflated Bayes factors (Supplementary Figure S3), which would favor the alternative model to produce false positives. We thus avoided the sharp null, and instead used g0* as informative prior for the null model. To choose between B (g0 B) and LogN (g0 L), we simulated chromosomal dosages (Supplementary Materials and Methods) and computed Bayes factors (with a prior for the alternative model to be specified below). Supplementary Figure S4 shows that the two sets of Bayes factors were almost identical to each other. We chose g0 B as the null prior in our data analysis because its computation is easier.
To specify priors under the alternative model, it was tempting to use g1* as an informative prior. Standard theory states that the marginal likelihood of a composite hypothesis is the weighted average of the likelihood over all constituent point hypotheses, where the prior serves as the weight.23 This gave rise to our first concern of using g1* as prior for the alternative model: g1* has almost no weights at or near 0, which risks being overly against the null model. The second but related concern was that g1 B placed 10 times more weight at fatal fraction 0.04 than at 0.02, and for g1 L the weight ratio between the two fetal fractions is 150. Another choice of the prior for the alternative model was g2*. We examined the posterior mean of each candidate prior (in light of the Z-test likelihood) and used this to guide our choice of prior for the alternative model. Figure 2 plotted the observed fetal fractions and their posterior mean estimates under g1* and g2*. Priors g1* overestimated the posterior mean for small observed fetal fractions, particularly in region (0.01,0.05). Priors g2* performed desirably, with B slightly better than LogN. We therefore chose g2 B as prior for the alternative model in our data analysis. This prior places 0.7 units of weight at fetal fraction 0.04 for each unit weight at 0.02. (Supplementary Figure S5 compares g1 B and g2 B quantitatively using simulated chromosomal dosages.)
Bayesian analysis
Data processing produced a pair (x, σ) for each autosome of each sample, where x is a centered chromosomal dosage, and σ is the sample standard deviation of chromosomal dosages of euploid controls. A Z-score can be computed via z = x/σ. Treating ∣x∣ as a one-sample estimate of fetal fraction h, we obtained a normal likelihood, which is the natural likelihood associated with the Z-test. Combining the likelihood with the priors for h under the null and alternative models we can compute Bayes factors (Supplementary Materials and Methods). Bayes factor (BF) is the change of odds in light of data:

It is well known that the prior odds of fetal trisomy increase with the maternal age.18 For example, at age 25 the odds (or the risk/prevalence) of a woman having a child with Down syndrome is 1/1,300; at age 35, the odds increase to 1/365; and at age 45, 1/30. With Bayesian inference this age-dependent risk can be conveniently incorporated into our analysis. From the posterior odds, the posterior probability of trisomy (τ) can be computed by τ = ω/(1 + ω). Because we choose between the null and the alternative models, the posterior probability of euploidy is 1 − τ. For a given sample, if it is called positive, then the individual PPV is τ. This is because PPV is a ratio between number of true positives—which is τ in our one-sample situation—and number of all positive calls—which is 1. Similarly, if it is called negative then the individual NPV is 1 − τ.
We compared our Bayesian method with the Z-test method by down-sampling reads and examining the consistency between the original data and the down-sampled data. The counts of positives/negatives were obtained by varying threshold of test statistics. For log10BF the threshold ranges from 0.67 to 89.60 and for Z-score, from 3.41 to 21.89. We counted the number of positives (n+) and number of negatives (n−) in the down-sampled data set that were positive in the original data set. Assuming the positives called in the original data sets were the truth, a more powerful method was expected to have a larger n+ (which mimic true positives) and a smaller n− (which mimic false negatives). Figure 3 demonstrates that, compared with the Z-test, our Bayesian method produced more “true positives” and fewer “false negatives.” In other words, the results were more consistent between the original data and the down-sampled data when our Bayesian method was used, and less consistent when the Z-test method was used.
Power comparison between Bayes factor (solid line) and Z -score (dashed line). Left panel: plot of counts of positives in the original data sets versus counts of positives in the down-sampled data sets that are also positives in original data set. Right panel: plot of positives in the original data sets versus counts of negatives in the down-sampled data sets but positives in original data set.
For each autosome of each individual, we calculated a centered chromosomal dosage, estimated the fetal fraction, and computed a Z-score and a Bayes factor. Figure 4 plots Z-score against fetal fraction. Those dots whose log10BF > 1 are in black with their sizes proportional to log10BF, and the rest are in gray with a fixed size. Dots are aligned on lines of different slopes, and dots associated with chromosomes 21, 18, and 13 are shown in red, green, and blue, respectively, to confirm this observation. This was expected because Z-scores associated with the same chromosome shared a common denominator sample standard deviation, and sample standard deviations varied with the chromosomal sizes and the coverage (Supplementary Figure S1). Figure 4 also shows that larger Z-scores and fetal fractions tend to associate with bigger Bayes factors, and most importantly, different dots may have the same Z-scores but different fetal fractions. While the Z-test treated two tests of the same Z-score equally, our Bayesian method assigned them different Bayes factors after taking into account the fetal fractions.
Z -values, fetal fractions, and Bayes factors. On the x-axis is the Z-value, and y-axis the fetal fraction. The points whose log10 Bayes factors are >1 are black and their sizes are proportional to log10 Bayes factor; the rest are in light gray and of the same size. Points that are associated with chromosomes 13, 18, and 21 are marked in blue, green, and red, respectively.
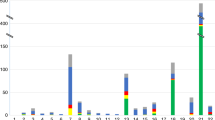
Our analysis produced 51 positive calls by the Z-test (Z-scores > 4). We examined whether and how our Bayesian method would call differently. Figure 5 plots fetal fractions estimated from sex chromosomes against those estimated from the (putative) trisomy autosomes, with the size of each dot proportional to the log10BF. The 28 (of 51) dots representing putative female fetuses are gray, and the remaining 23 dots representing putative male fetuses are black. For a male fetus if the autosome trisomy is true, the fetal fraction estimated from the autosome will match that estimated from the sex chromosomes. Thus we focus on the male fetus below. Of the 23 dots, 14 aligned closely along the diagonal line, and 11 of the 14 were confirmed positives. These suggested that the fetal fractions estimated from the sex chromosomes agreed well with those estimated from the trisomy autosomes. Of these 14 dots, 13 had Bayes factors greater than 1,412, and the one exception had a Bayes factor of 19 (represented by the dot in a gray triangle). The chromosome in question was chromosome 20 and trisomy 20 is rare,24 which means the prior odds are small, say 1/2,000. Then the posterior odds are <1/100, so this one was called negative by our Bayesian method. There were 9 dots that deviated from the diagonal line. Of these, 7 (in the gray circle) had Bayes factors smaller than 20, and were called negative by our Bayesian method. Interestingly, none of these 7 were associated with chromosomes 13, 18, or 21. The 8th abnormal sample (in the gray square) had the largest difference in fetal fraction estimates between the sex chromosomes and the autosomes. Its Bayes factor was 186 and it was associated with chromosome 3. This one was also called negative by our Bayesian method because the prior odds for trisomy 3 with mosaicism were extremely small. The 9th abnormal dot is enclosed by a gray diamond. This one was called a true-positive trisomy 13 with mosaicism because the Bayes factor was extraordinarily large, more than 33 million. Our results appear to be at odds with other studies, considering the near-perfect true-positive rates reported by others.4,5,6 One possible explanation is that other near-perfect true-positive reports focused on trisomy 13, 18, and 21, while the 9 suspected false positives in our analysis all relate to other chromosomes.
Annals of Z -test positive calls. Each dot correspond to a significant Z-test (Z-score > 4), whose x-axis is fetal fraction estimated from sex chromosome, y-axis is fetal fraction estimated from trisomy, and whose size is proportional to its log10 Bayes factor. Gray dots are putative female fetus. Black dots are putative male fetus. See the main text for description of dots in triangle, circle, square, and diamond.
Discussion
We developed a Bayesian method to analyze a NIPS data set to detect fetal trisomy such as Down syndrome, and demonstrated that our Bayesian method is more powerful than the traditional Z-test. One source of power gain of our Bayesian method is the reduced false positives. A euploid fetus can have an estimate of centered chromosomal dosage x of, say, 0.012, just by chance. (Recall ∣x∣ estimates fetal fraction h, and a female fetus, whose h estimate from the sex chromosomes is expected to be 0, can have an h estimate as large as 0.028.) If the corresponding σ = 0.003, which is realistic according to Supplementary Figure S1, we have Z = 4, which would be (falsely) called positive by the Z-test method. On the other hand, Bayes factor is likely to remain small because the prior under the null favors small x. In fact, plugging x = 0.012 and σ = 0.003 into our Bayes factor program, we have a Bayes factor of 0.001. Another (undemonstrated) source of power gain is the reduced false negatives. Our Bayesian method can incorporate age-adjusted prior odds into decision making, which is more likely to call out true positives among high-risk pregnancies. Currently NIPS is being developed to detect microduplications (and microdeletions), which loosely is equivalent to detecting trisomy (or monosomy) of a much smaller chromosome; we expect our Bayesian method to make valuable contributions.
We used Z-test (normal) likelihood to compute Bayes factors. This perhaps was not the best likelihood to use. At the minimum, since σ was not known but estimated, the correct distribution of Z-scores should be the t instead of the normal. We can easily incorporate other forms of likelihood into our Bayes factor calculation. For example, we can use the 100-kb bin as a unit to produce a composite likelihood for each chromosome (by multiplying the normal likelihood of each bin), which might perform better than the normal likelihood at a chromosomal level, because the bin-based likelihood accounts for the different variances among bins.
Our Bayesian method pays extra attention to the fetal fraction. We learned distributions of the fetal fractions from the sex chromosomes to formulate priors under the null and alternative models, and incorporated these informative priors into our Bayesian method to test for fetal autosomal trisomy. This agnostic approach to specify informative priors avoided subjective bias on prior specification that often drew critiques. The parallel nature of the genetic data affords us an opportunity to specify informative priors for our Bayesian analysis. To be specific, the data collected from the sex chromosomes can be viewed as independent of the data collected from the autosome. We estimated the fetal fractions from the sex chromosomes, learned their distributions, and applied these distributions as priors to perform Bayesian analysis of autosomes. In doing so, we did not “use the data twice,” which would violate the first principles of Bayesian analysis. Rather, we picked the informative part of the data (sex chromosomes) to learn about the parameter, and then applied the learned knowledge to analyze other—nonoverlapping—parts of the data (autosomes).
The learned priors provided us an opportunity to avoid the sharp null. The sharp null hypotheses often used in the literature are seldom exactly true. They exist because they are simple and often a good enough approximation.22 The testing of a null hypothesis can usually be made more realistic by “spreading” the hypothesis over a small region.22 It is noted that in testing a normal mean the sharp null is a good approximation to an interval null as long as the width of the interval is less than about one-half of the standard error of the sample mean.20,25 The genetic data sets we analyzed here, however, suggested that using a sharp null favors the alternative model (Supplementary Figure S3). This is because the variation of the fatal fraction under the null (estimated from the putative female fetuses) is substantial compared with the sample standard deviations of the autosomal chromosomal dosages from euploid control samples.
Our method can conveniently incorporate, through prior odds, factors such as maternal age and the odds change obtained from the traditional first-trimester screen. Using posterior probability, our method can directly compute individual PPV and NPV, which makes our method more useful to clinicians and genetic counselors. Both PPV and NPV are of important clinical utilities but difficult to obtain from the Z-test. This is because the Z-test is a classical null-hypothesis significance test, which does not allow researchers to state evidence for the null hypothesis, while NPV is a confidence measurement for the null hypothesis.
References
Lo YMD, Corbetta N, Chamberlain PF et al. Presence of fetal DNA in maternal plasma and serum. Lancet. 1997;350:485–487.
Fan HC, Blumenfeld YJ, Chitkara U, Hudgins L, Quake SR. Noninvasive diagnosis of fetal aneuploidy by shotgun sequencing DNA from maternal blood. Proc Natl Acad Sci USA 2008;105:16266–16271.
Chiu RWK, Chan KCA, Gao Y et al. Noninvasive prenatal diagnosis of fetal chromosomal aneuploidy by massively parallel genomic sequencing of DNA in maternal plasma. Proc Natl Acad Sci USA 2008;105:20458–20463.
Chiu RWK, Akolekar R, Zheng YWL et al. Non-invasive prenatal assessment of trisomy 21 by multiplexed maternal plasma DNA sequencing: large scale validity study. BMJ 2011;342http://www.bmj.com/content/342/bmj.c7401.long.
Bianchi DW, Parker RL, Wentworth J et al. DNA sequencing versus standard prenatal aneuploidy screening. N Engl J Med 2014;370:799–808.
Norton ME, Jacobsson B, Swamy GK et al. Cell-free DNA analysis for noninvasive examination of trisomy. N Engl J Med 2015;372:1589–1597.
Committee opinion no. 640: cell-free DNA screening for fetal aneuploidy. Obstet Gynecol 2015;126:e31–7.
Society for Maternal-Fetal Medicine (SMFM) Publications Committee. #36. Prenatal aneuploidy screening using cell-free DNA. Am J Obstet Gynecol 2015;212:711–6.
Benn P, Borrell A, Chiu RW et al. Position statement from the Chromosome Abnormality Screening Committee on behalf of the Board of the International Society for Prenatal Diagnosis. Prenat Diagn 2015;35:725–34.
Dondorp W, De WG, Bombard Y et al. Non-invasive prenatal testing for aneuploidy and beyond: challenges of responsible innovation in prenatal screening. Eur J Hum Genet 2015;23:1438–50.
Gregg AR, Skotko BG, Benkendorf JL et al. Noninvasive prenatal screening for fetal aneuploidy, 2016 update: a position statement of the American College of Medical Genetics and Genomics. Genet Med. 2016;18:1056–1065.
Grody WW, Thompson BH, Gregg AR, Schneider A et al. ACMG position statement on prenatal/preconception expanded carrier screening. Genet Med. 2013;15:482–483.
Canick JA, Palomaki GE, Kloza EM, Lambert-Messerlian GM, Haddow JE. The impact of maternal plasma DNA fetal fraction on next generation sequencing tests for common fetal aneuploidies. Prenat Diagn. 2013;33:667–674.
Benn P, Cuckle H. Theoretical performance of non-invasive prenatal testing for chromosome imbalances using counting of cell-free DNA fragments in maternal plasma. Prenat Diagn. 2014;34:778–783.
Suzumori N, Ebara T, Yamada T et al. Fetal cell-free DNA fraction in maternal plasma is affected by fetal trisomy. J Hum Genet 2016;61:647–652.
Fiorentino F, Bono S, Pizzuti F et al. The importance of determining the limit of detection of non-invasive prenatal testing methods. Prenat Diagn. 2016;36:304–311.
Fan HC, Quake SR. Sensitivity of noninvasive prenatal detection of fetal aneuploidy from maternal plasma using shotgun sequencing is limited only by counting statistics. PLoS One. 2010;5:1–7.
Newberger DS. Down syndrome: prenatal risk assessment and diagnosis. Am Fam Physician 2000;62:825–32.
Bianchi DW, Platt LD, Goldberg JD, Abuhamad AZ, Sehnert AJ, Rava RP. Genome-wide fetal aneuploidy detection by maternal plasma DNA sequencing. Obstet Gynecol 2012;119:890–901.
Kass RE, Raftery AE. Bayes factor. J Am Stat Assoc 1995;90:773–795.
Rava RP, Srinivasan A, Sehnert AJ, Bianchi DW. Circulating fetal cell-free DNA fractions differ in autosomal aneuploidies and monosomy X. Clin Chem. 2013;60:243–250.
Good IJ. C420. The existence of sharp null hypotheses. J Stat Comput Simul 1994;49:241–242.
Rouder JN, Speckman PL, Sun D, Morey RD, Iverson G. Bayesian t tests for accepting and rejecting the null hypothesis. Psychon Bull Rev 2009;16:225–237.
Mavromatidis G, Dinas K, Delkos D, Vosnakis C, Mamopoulos A, Rousso D. Case of prenatally diagnosed non-mosaic trisomy 20 with minor abnormalities. J Obstet Gynaecol Res 2010;36:866–868.
Berger JO, Delampady M. Testing precise hypotheses. Statist Sci 1987;2:317–335.
Acknowledgments
H.B. and Y.G. were supported in part by the US Department of Agriculture/Agriculture Research Service under contract 6250-51000-057.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of Interest
YG is a consultant for Beijing USCI Medical Laboratory and owns its stock options. Beijing USCI Medical Laboratory contributed to fund the study, but played no role in designing experiment, analyzing data, and interpreting results.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Xu, H., Wang, S., Ma, LL. et al. Informative priors on fetal fraction increase power of the noninvasive prenatal screen. Genet Med 20, 817–824 (2018). https://doi.org/10.1038/gim.2017.186
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2017.186
Keywords
This article is cited by
-
Estimation of cell-free fetal DNA fraction from maternal plasma based on linkage disequilibrium information
npj Genomic Medicine (2021)