Abstract
Purpose
Fetal anomalies represent a poorly studied group of developmental disorders. Our objective was to assess the impact of whole-exome sequencing (WES) on the investigation of these anomalies.
Methods
We performed WES in 101 fetuses or stillborns who presented prenatally with severe anomalies, including renal a/dysgenesis, VACTERL association (vertebral defects, anal atresia, cardiac defects, tracheoesophageal fistula, renal anomalies, and limb abnormalities), brain anomalies, suspected ciliopathies, multiple major malformations, and akinesia.
Results
A molecular diagnosis was obtained in 19 cases (19%). In 13 of these cases, the diagnosis was not initially suspected by the clinicians because the phenotype was nonspecific or atypical, corresponding in some cases to the severe end of the spectrum of a known disease (e.g., MNX1-, RYR1-, or TUBB-related disorders). In addition, we identified likely pathogenic variants in genes (DSTYK, ACTB, and HIVEP2) previously associated with phenotypes that were substantially different from those found in our cases. Finally, we identified variants in novel candidate genes that were associated with perinatal lethality, including de novo mutations in GREB1L in two cases with bilateral renal agenesis, which represents a significant enrichment of such mutations in our cohort.
Conclusion
Our study opens a window on the distinctive genetic landscape associated with fetal anomalies and highlights the power—but also the challenges—of WES in prenatal diagnosis.
Similar content being viewed by others
Introduction
Birth defects are the main cause of mortality in infancy and represent an important source of morbidity at all ages, with a major proportion of cases being identified during pregnancy. The etiology of birth defects is complex. Copy-number variants (CNV) currently represent the most commonly recognizable cause of birth defects in fetuses and children. Indeed, the prevalence of submicroscopic pathogenic or likely pathogenic CNVs is 5.5–10.5% in fetuses with structural defects.1,2 The fact that CNVs represent a common cause of birth defects suggests that smaller genetic lesions, such as point mutations, may also contribute substantially to their pathogenesis.
Most pathogenic CNVs associated with birth defects occur de novo. A reciprocal relationship between the fitness effect of a mutation and the proportion of de novo mutations (DNMs) is observed in a dominantly inherited disorder. Accordingly, DNMs may explain a larger proportion of severe cases of birth defects compared with milder ones. For instance, the contribution of de novo CNVs appears greater in cases with multiple malformations than those with isolated malformations.1,2 Similarly, de novo truncating mutations are significantly enriched in syndromic but not isolated forms of congenital heart defects.3 One potential corollary of these observations would be that DNMs contribute more to birth defects in fetuses than in newborns or older children because the former group includes a subset of cases that are so severely affected they cannot survive beyond the perinatal period.
Classical studies have established that recessive mutations also explain a large fraction of lethal developmental defects in animal models.4 Thus, as for DNMs, severe forms of fetal malformations in humans may involve inherited recessive mutations in genes that would not be identified by studying newborns or older children with malformations.
Based on these considerations, we hypothesized that inherited and de novo point mutations, as is the case for CNVs, explain a substantial subset of fetal cases with severe anomalies. We further hypothesized that some of these lesions disrupt genes that are specifically associated with fetal phenotypes. To explore these hypotheses, we performed whole-exome sequencing (WES) in a series of fetuses and stillborns investigated prenatally because of severe anomalies. We sought to measure the diagnostic utility of WES for various categories of anomalies while attempting to identify candidate genes associated with these disorders.
Materials and methods
This study was approved by the CHU Sainte-Justine’s research ethics board. After obtaining informed consent from the parents, we prospectively recruited, over a 3-year period, 89 fetuses (from terminated pregnancies) and stillborns with: (i) at least two major malformations, (ii) severe ventriculomegaly (atria >15 mm bilaterally) and/or structural brain malformations, or (iii) an anomaly associated with a high risk of perinatal lethality. We recruited from three other centers (Mount Sinai Hospital, Toronto, Canada; Hôpitaux Universitaires de Strasbourg, France, and the Children’s Hospital of East Ontario, Ottawa, Canada) 12 additional cases that met these criteria for a total of 101 cases. These cases were unexplained and none of them was exposed to a known teratogen. Neither karyotyping (n = 58 cases) nor chromosomal microarray analysis (n = 84 cases) showed any candidate or pathogenic CNVs (Supplementary Table S1online). Autopsies were performed in 85 cases (Supplementary Table S1).
Genomic DNA was extracted from the umbilical cord, other fetal tissues, amniocytes or chorionic villi (Supplementary Table S1). For 83 cases WES was performed in trios, whereas for 18 cases it was performed in the affected cases but not in their parents, as indicated in Supplementary Table S1. The exome was captured using the Agilent SureSelect (V4 or V5) exome capture kit followed by 100 bp paired-end sequencing in a configuration of four exomes per lane on the Illumina HiSeq (2000 or 2500) system.
Exome sequence data processing, alignment (using a Burrows–Wheeler algorithm), and variant calling were performed according to the Broad Institute’s GATK (v4) best practices (http://www.broadinstitute.org/gatk/guide/topic?name=best-practices), and variant annotation was performed using Annovar. The median coverage of the target bases was 110 × with 95% of the target bases being covered ≥10 ×. We focused on variants affecting the exome and intronic bases up to positions −3 and +6 from the exon boundaries. Only variants whose positions were covered at ≥10 × and supported by at least four variant reads constituting ≥20% of the total reads for each called position were retained. This typically yielded an average of ~22,000 variants. This variant list was subsequently reduced to an average of ~400 rare variants by filtering out those that were present in ≥0.5% of in-house exome data sets (n = 600) from unrelated projects, as well as variants present in the 1000 Genomes Project or the Exome Variant Server (http://evs.gs.washington.edu/EVS/) with minor allele frequencies ≥0.5%. Putative DNMs (typically <10 per exome) were then extracted from the rare variant list by further excluding those that were present in the exomes of the parents. The sequencing reads carrying putative DNMs were inspected visually in each trio, using the Integrative Genomics Viewer, to exclude obvious false positives. Interpretation of sequence variants was done according to the standards of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology.5 Clinically significant results were returned to the parents.
Results
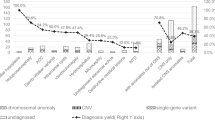
We studied 101 fetal cases with severe anomalies who presented mainly during the second trimester of pregnancy. We classified these cases into six categories based on their phenotypes: (i) isolated bilateral renal agenesis or dysgenesis (n = 11), (ii) VACTERL (vertebral defects, anal atresia, cardiac defects, tracheoesophageal fistula, renal anomalies, and limb abnormalities) association (n = 9), (iii) cerebral anomalies (n = 36), (iv) suspected ciliopathies (n = 5), (v) miscellaneous patterns of multiple malformations (n = 32), and (vi) fetal akinesia (n = 8) (Supplementary Table S1).
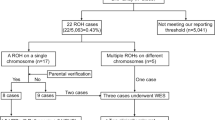
In 16 families with a suspected autosomal recessive disorder because of parental consanguinity, the presence of multiple affected siblings and/or the nature of the phenotype (e.g., anomalies suggestive of ciliopathies, which are mainly autosomal recessive), WES was performed in the affected individuals (n = 18) but not in their parents. The remaining cases (n = 83), all sporadic, were investigated by WES in trios. We first aimed to systematically identify DNMs in these trios to determine whether our series was enriched for such mutations. We observed a high number of putative DNMs (between 12 and 40 DNMs) in the three probands for which genomic DNA was extracted from cultured amniocytes or chorionic villi (Supplementary Table S1 online). Sanger validation of some of these variants using DNA extracted from another tissue suggested that the large majority of these putative DNMs are false positives, possibly related to culture artifacts. In DNA samples extracted from tissues, we identified and Sanger-validated a total of 116 DNMs, including 5 nonsense, 6 frameshift-causing indel, 77 missense and 2 canonical splice site variants (Supplementary Table S2). We documented the presence of at least one likely gene disrupting DNM (nonsense, frameshift or canonical splicing variant) in 11 of these cases (11%). The expected percentage of DNMs in a coding and splicing sequence that are synonymous is ~29%.6,7 Among the SNV and canonical splice site DNMs identified herein (n = 103), 19 were synonymous (18%) and 84 were functional (77 missenses, 5 stopgains and 2 splicing variants). This distribution indicates a significant enrichment in functional DNMs (P = 0.017, exact binomial test), thus suggesting that some of these are likely pathogenic.
For each category of anomalies, we next determined whether the WES data sets contained rare de novo or inherited variants in known disease genes associated or possibly associated with the reported phenotypes. We also sought to identify candidate genes based on their involvement in unrelated families or their known function in model organisms.
Renal a/dysgenesis
We studied eight sporadic cases with isolated bilateral renal agenesis (Supplementary Table S1 online). All these cases were males. Such a predominance of males has previously been documented in renal agenesis.8 In addition, we studied three cases (two females and one male) with isolated bilateral multicystic kidneys without any sign of lower urinary tract obstruction. In these cases, WES did not reveal any DNMs or rare potentially recessive variants in genes previously associated with congenital anomalies of the kidney and urinary tract (CAKUT).
Candidate gene
We identified DNMs in the GREB1L gene in two unrelated cases with bilateral renal agenesis (051-082-IEO and CONGE-070) (Supplementary Tables S1 and S3). Renal ultrasound was normal in the parents of these cases. One DNM is a predicted-damaging missense (NM_001142966; c.C2903T;p.A968V), whereas the other is a stopgain (NM_001142966; c.C293G:p.S98X) that truncates most of the protein (Supplementary Tables S1 and S3). Moreover, only one unvalidated loss-of-function variant in GREB1L is reported in the Exome Aggregation Consortium database, suggesting that this gene does not tolerate this type of variant. Using an established framework of gene-specific mutation rates,9 we observed an enrichment in functional DNMs in GREB1L in our series (P = 0.00035, two-tailed exact Poisson test), suggesting that damaging DNMs in this gene are pathogenic. GREB1L shares some identity with GREB1, a chromatin-bound estrogen receptor coactivator.10,11 The zebrafish homolog of GREB1L is induced by the Hedgehog pathway12 and a truncating mutation in this gene causes abnormal swim bladder inflation (http://zfin.org/ZDB-GENE-030131-4022), which can be a sign of abnormal Hedgehog signaling.13 Interestingly, germ-line mutations in Shh cause renal aplasia in mice.14
VACTERL association
We studied nine cases with at least two features of the VACTERL association (OMIM 192350), including at least one core feature (anorectal or tracheoesophageal defects) (Supplementary Table S1). WES revealed a de novo truncating mutation in CHD7 (OMIM 608892) (NM_017780; c.C5428T:p.R1810X) in a fetus (CONGE-79) with esophageal atresia and tetralogy of Fallot (Table 1 and Supplementary Tables S1 and S3). DNMs in CHD7 have been reported to cause these defects in the context of the CHARGE syndrome (coloboma of the eye, heart defects, atresia of the choanae, retardation of growth and/or development, genital and/or urinary abnormalities, and ear abnormalities and deafness), which overlaps with the VACTERL association.15
WES also showed the presence of a truncating DNM (NM_015375; c.C2125T:p.R709X) in DSTYK (OMIM 612666) in a fetus (051-141-VWD) with left heart hypoplasia and esophageal atresia (Table 1 and Supplementary Tables S1 and S3). This mutation is located in an upstream exon, which suggests that it is likely inducing nonsense-mediated decay of the transcript (Supplementary Tables S1 and S3). Dominant variants in DSTYK, including truncating variants, have been involved in CAKUT.16 Our case did not show any kidney anomalies, raising the possibility that the mutation is not penetrant, as previously observed in some families with CAKUT.16 Alternatively, mutations in this gene might cause a broader spectrum of congenital anomalies than initially suspected, explaining the phenotype of our fetal case. DSTYK functions as a modulator of firbroblast growth factor signaling.16 Knockdown of Dstyk in the zebrafish causes multiple defects of morphogenesis that are consistent with a disruption of firbroblast growth factor signaling.16 Interestingly, firbroblast growth factor signaling has been involved in esophageal development.17 Moreover, Dstyk is expressed in the developing heart and a decrease in its expression causes heart failure in zebrafish larvae.16
In the other cases, WES did not reveal any DNMs or rare biallelic variants in genes previously associated with monogenic forms of the VACTERL association.
Cerebral anomalies
We studied a total of 36 fetal cases with major structural brain anomalies, eight of which also had at least one major peripheral anomaly (Supplementary Table S1). In total, we identified eight mutations in seven genes (TUBA1A (OMIM 602529), TUBB3 (OMIM 602661), TUBB (OMIM 191130), PEX1 (OMIM 602136), PDHA1 (OMIM 300502), ACTB (OMIM 102630), and HIVEP2 (OMIM 616977)) that are pathogenic or likely pathogenic (Table 2). Interestingly, four of these mutations affect genes that code for tubulins (TUBA1A, TUBB3, and TUBB). Since the phenotypes observed in these eight fetuses were somewhat nonspecific, these mutations were not readily suspected by the clinicians. However, their phenotypes were consistent with what was reported for these genes with the exception of the cases with mutations in TUBB, ACTB, and HIVEP2, which are described hereafter.
Fetus CONGE-043 bears a missense DNM in TUBB (NM_001293212; c.C920T:p.P307L), which has not been reported in the Exome Aggregation Consortium database and which is predicted to be damaging by the tools used in this study (Table 2 and Supplementary Tables S1 and S3). DNMs in TUBB have been previously described in six children who displayed microcephaly and mild anomalies of the corpus callosum and basal ganglia.18,19 Some of these individuals also displayed circumferential skin creases.19 Fetus CONGE-043 showed a similar but much more severe phenotype with the presence of microlissencephaly, agenesis of the corpus callosum, dysmorphic basal ganglia, cerebellar hypoplasia, and circumferential skin creases (Table 2 and Supplementary Table S1). The cortex of our case was characterized by the presence of glomerular structures and a voluminous germinal area, which has been associated with tubulinopathies, further supporting the pathogenicity of the variant in TUBB.
Fetus 051-021-LIC bears a de novo missense in ACTB (NM_001101; c.G617A:p.R206Q), which has not been reported in the Exome Aggregation Consortium database and which is predicted to be damaging (Table 2 and Supplementary Tables S1 and S3). DNMs in ACTB cause Baraitser–Winter syndrome, which includes intellectual disability and neuronal migration defects of the cortex.20 The variant identified in fetus 051-021-LIC has been reported in ClinVar (SCV000322073) as pathogenic, but without any description of the associated phenotype. This variant clusters with several other variants that have been associated with Baraitser–Winter syndrome.20 Fetus 051-021-LIC presented with severe ventriculomegaly of the lateral and third ventricles with a narrow aqueduct, but there was no sign of neuronal migration defects on pathological examination at 35 weeks of gestation (Table 2 and Supplementary Table S1 online). Thus, this ACTB variant appears pathogenic but the associated phenotype is atypical.
Fetus 051-039-KUY presented at 20 weeks of gestation with severe triventriculomegaly and occipital encephalocele. WES showed that this fetus carried a truncating DNM in HIVEP2 (OMIM 143054) (NM_006734; c.2968_2971del:p.K990fs), which is represented by only one RefSeq isoform (Table 2 and Supplementary Tables S1 and S3). This variant is located in an upstream exon and is thus predicted to induce nonsense-mediated decay of the transcript. Nine children with DNMs in HIVEP2, including eight truncating mutations, have been reported.21,22 All of these children displayed intellectual disability with minimal or no structural changes on the magnetic resonance images. One possibility is that DNMs in HIVEP2 are associated with a more variable phenotype than previously suspected, explaining the structural defects observed in our case. Alternatively, this fetus might be affected by two different genetic disorders with the DNM in HIVEP2 causing a neurodevelopmental disorder that was not readily apparent in the fetus and mutation(s) in another gene causing her brain structural defects. We found that the fetus was a compound heterozygote for predicted-damaging variants in DNAH7, which has been associated with primary ciliary dyskinesia (Table 2 and Supplementary Tables S1 and S3).23 Interestingly, ciliary dysfunction can disrupt the ependymal flow through the aqueduct, causing its closure and subsequent triventricular hydrocephalus.24,25
Malformations suggestive of ciliopathies
We studied five fetuses with at least two of the following features of ciliopathies: occipital meningocele, large/hyperechogenic kidneys and vermis hypoplasia (Table 3). Four of these fetuses were compound heterozygous for pathogenic or likely pathogenic variants in known ciliopathy genes (CEP290 (OMIM 610142), TCTN1 (OMIM 609863), TMEM67 (OMIM 609884), and BBS10 (610148)), whereas the remaining one was a compound heterozygote for variants of unclear significance in TMEM67, which is also associated with a ciliopathy (Table 3 and Supplementary Tables S1 and S3).
Multiple malformations
We studied 32 cases with at least two major malformations (with patterns not corresponding to those included in the other categories) (Supplementary Table S1). We considered that six of them (19%) were likely explained by mutations in known disease genes, including FRAS1 (OMIM 607830), FLNB (OMIM 603381), FAM20C (OMIM 611061), TGFBR1 (OMIM 190181), EP300 (OMIM 602700), and MNX1 (OMIM 142994) (Table 4 and Supplementary Tables S1 and S3). The phenotype of these cases was consistent with that previously associated with these genes, except for fetus 051-008-TNK with a mutation in EP300 and fetus CONGE-005 with a mutation in MNX1. We describe these two fetuses in more detail hereafter.
Fetus 051-008-TNK presented during the second trimester with intrauterine growth restriction, spina bifida, and postaxial polydactyly. WES showed the presence of a truncating DNM in EP300 (NM_001429; c.102_105del:p.G34fs) (Table 4 and Supplementary Tables S1 and S3). Dominant mutations in EP300 cause Rubinstein–Taybi syndrome. The autopsy of fetus 051-008-TNK showed signs of Rubinstein–Taybi syndrome, including the typical facial features and broad halluces (Supplementary Table S1). However, postaxial polydactyly and neural tube defects are atypical features of EP300-related Rubinstein–Taybi syndrome, having been described only in single cases with this disorder.26,27 Interestingly, the loss of Ep300 disrupts neurulation in mice.28
Fetus CONGE-005 presented with akinesia and a short spine at 12 weeks of gestation. The fetus was delivered at term but died within minutes of birth with several anomalies including lumbosacral agenesis, anal imperforation, and scrotal agenesis (Supplementary Figure S1). The parents were consanguineous and previously had three newborns who died in the immediate neonatal period with a similar phenotype characterized by a short spine and the variable presence of sacrococcygial teratoma and anal imperforation. The conjunction of caudal agenesis with a presacral mass, such as a teratoma, is consistent with Currarino syndrome, which is caused by dominant mutations in MNX1 (OMIM 176450). Interestingly, we found that fetus CONGE-005 is homozygous and his parents heterozygous for the variant c.T2C/p.M1T in MNX1 (NM_005515) (Table 4 and Supplementary Tables S1 and S3). This gene has two isoforms—a longer isoform 1 of 401 amino acids (NP_005506.3) and a shorter isoform 2 of 189 amino acids (NP_001158727.1) that lacks the N-terminus present in isoform 1 where this variant is located. Specifically, p.M1T alters the initiation methionine of the longer isoform of the protein, likely impairing its production as the second in-frame methionine is far downstream at amino acid 221. Three cases, all with neonatal diabetes, have been reported with biallelic missense variants in MNX1.29,30 One of these cases also had Currarino syndrome 29. Our case has a much more severe phenotype than that found in any of the previously reported cases with mutations in MNX1. We postulate that the p.M1T variant is hypomorphic, being tolerated in the heterozygotes, but causing severe defects in the homozygotes.
Candidate genes
Fetus 051-043-BEV presented with scoliosis associated with vertebral segmentation defects. As this fetus was the product of a consanguineous union, we focused our attention on homozygous variants (Table 4 and Supplementary Table S1). Among these, we identified a rare, predicted-damaging missense variant in FKBP8 (OMIM 604840) (NM_012181; c.C572T:p.P191L) (Table 4 and Supplementary Tables S1 and S3). Interestingly, Fkpb8 mutant mice show extensive fusion of vertebrae, suggesting that the segmentation defects observed in the fetus are also caused by disruption of this gene.31,32
Fetus CONGE-074 presented during the second trimester with multiple anomalies, including phocomelia (Table 4 and Supplementary Table S1). This fetus carried a truncating DNM in VEGFA (OMIM 192240) (NM_003376; c.C789A:p.C263X), which is likely inducing nonsense-mediated decay of the transcript, as it is located in an upstream exon that is present in all RefSeq isoforms (Table 4 and Supplementary Table S3). VEGFA codes for a critical regulator of blood vessel development. Remarkably, the loss of a single Vegfa allele is lethal in mouse embryos between days 11 and 12.33,34 These embryos show impaired early vascular development and multiple structural anomalies, which remain poorly characterized because of the early lethality. At least some of the defects observed in our fetus might be explained by a defect of blood vessel development. For instance, it appears that the maternal administration of thalidomide during pregnancy causes fetal phocomelia by disrupting angiogenesis.35
Fetal akinesia
We studied eight akinetic fetuses from seven unrelated families (Supplementary Table S1). We identified pathogenic biallelic mutations in RYR1 (OMIM 180901) in two siblings from a consanguineous union (MALFO 1.13 and MALFO 2.13) and in an unrelated case (CONGE-060) (Table 5 and Supplementary Tables S1 and S3). Recessive mutations in RYR1 are mainly associated with myopathies presenting in childhood, but have previously been described in a few fetuses with akinesia.36 We did not find any rare variants in genes previously associated with akinesia in the remaining cases.
Discussion
The discovery of severe anomalies in a fetus raises the critical question of the prognosis and recurrence risk. Clinicians typically order prenatal chromosomal microarray analysis testing following the discovery of major fetal anomalies with the aim of identifying the cause of the disorder and establishing a precise prognosis. Although the diagnostic yield of chromosomal microarray analysis is relatively low, it currently represents the most powerful test for the investigation of fetal structural defects. The goal of this study was to evaluate the added value of WES for the investigation of these anomalies. By mainly recruiting cases from terminated pregnancies, we aimed to enrich our series with severe anomalies while complementing their phenotypic characterization with pathological data.
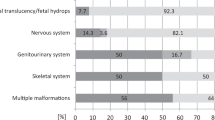
The diagnostic yield (19%) observed in our study is similar to those recently obtained in similar studies, with yields of 10% (3/35),37 20% (17/84)38, and 21% (5/24)39. Our yield was higher for ciliopathies (80%), cerebral anomalies (19%), multiple malformations (19%), and fetal akinesia (25%), but lower for renal a/dysgenesis (0%) and the VACTERL association (11%). The diagnosis was initially suspected by the clinicians in 6 of these families (those with mutations in FRAS1, TCTN1, CEP290, TMEM67, and BBS10) but not in the remaining 13 families (those with mutations in CHD7, TUBA1A, TUBB3, TUBB, PEX1, PDHA1, EP300, FLBN, MNX1, TGFBR1, and RYR1). In some of these latter cases, the phenotypes were not specific enough to invoke the diagnoses of interest. This lack of phenotypic specificity represents a major challenge of prenatal diagnosis and is explained by several factors, including the immaturity of the fetuses and our limited ability to phenotype them prenatally. For instance, single-gene disorders can be difficult to distinguish prenatally from common disorders that are genetically more complex, as illustrated by fetus CONGE-79, who presented with known features of CHD7-related disorders, such as esophageal atresia and tetralogy of Fallot, that can also be associated with the more common VACTERL association.
In other instances, the molecular diagnosis was not suspected because the phenotype was atypical and not corresponding to the phenotype usually found in children and adults. Some of these cases might represent the severe end of the phenotypic spectrum associated with known disease genes. These include the fetuses with mutations in TUBB, MNX1, and RYR1 who displayed a similar but much more severe phenotype than that previously described in individuals with mutations in these genes. Other cases might represent presentations of known disease genes that are substantially different from those previously documented. This might be the case of the fetus with a DNM in EP300 who presented with features such as spina bifida and postaxial polydactyly that are not usually found in EP300-related disorders. Similarly, DNMs in DSTYK (051-141-VWD), ACTB (051-021-LIC), and HIVEP2 (051-039-KUY) were found in fetuses with phenotypes that have not been previously associated with these genes. It remains unclear whether these DNMs explain the anomalies of these last three fetuses. This clinical heterogeneity associated with fetal anomalies and our limited ability to correlate genotypes and phenotypes in ongoing pregnancies represent challenges for the integration of WES in prenatal diagnosis.
The overall diagnostic yield of 19% is slightly lower than that obtained with the use of WES for the investigation of rare disorders in children or adults, which is ~25–31%.40 The lower yield of WES for some groups of anomalies could be explained by the involvement of epigenetic mechanisms or more complex genetic models (see, for example, ref.3). Alternatively, some of these severe anomalies could be caused by mutations in sets of genes other than those involved in disorders presenting after birth. For instance, we identified a novel candidate gene for renal agenesis, GREB1L, which is associated with a lethal phenotype that would not be ascertained beyond the immediate neonatal period. Very little is known about the genetic basis of the rare and severe disorders specifically encountered in fetuses. Systematic genomic studies of fetal anomalies will thus provide insight into the distinctive function of numerous genes acting during embryonic or fetal life while enhancing the diagnostic utility of genomic sequencing for prenatal diagnosis.
References
Lee CN, Lin SY, Lin CH, Shih JC, Lin TH, Su YN. Clinical utility of array comparative genomic hybridisation for prenatal diagnosis: a cohort study of 3171 pregnancies. BJOG 2012;119:614–625.
Lovrecic L, Remec ZI, Volk M, Rudolf G, Writzl K, Peterlin B. Clinical utility of array comparative genomic hybridisation in prenatal setting. BMC Med Genet 2016;17:81.
Sifrim A, Hitz MP, Wilsdon A et al. Distinct genetic architectures for syndromic and nonsyndromic congenital heart defects identified by exome sequencing. Nat Genet 2016;48:1060–1065.
Samuels ME. Saturation of the human phenome. Curr Genomics 2010;11:482–499.
Richards S, Aziz N, Bale S et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med 2015;17:405–424.
Kryukov GV, Pennacchio LA, Sunyaev SR. Most rare missense alleles are deleterious in humans: implications for complex disease and association studies. Am J Hum Genet 2007;80:727–739.
Iossifov I, O’Roak BJ, Sanders SJ et al. The contribution of de novo coding mutations to autism spectrum disorder. Nature 2014;515:216–221.
Harewood L, Liu M, Keeling J et al. Bilateral renal agenesis/hypoplasia/dysplasia (BRAHD): postmortem analysis of 45 cases with breakpoint mapping of two de novo translocations. PLoS ONE 2010;5:e12375.
Samocha KE, Robinson EB, Sanders SJ et al. A framework for the interpretation of de novo mutation in human disease. Nat Genet 2014;46:944–950.
Ghosh MG, Thompson DA, Weigel RJ. PDZK1 and GREB1 are estrogen-regulated genes expressed in hormone-responsive breast cancer. Cancer Res 2000;60:6367–6375.
Mohammed H, D’Santos C, Serandour AA et al. Endogenous purification reveals GREB1 as a key estrogen receptor regulatory factor. Cell Rep 2013;3:342–349.
Xu J, Srinivas BP, Tay SY et al. Genomewide expression profiling in the zebrafish embryo identifies target genes regulated by Hedgehog signaling during vertebrate development. Genetics 2006;174:735–752.
Winata CL, Korzh S, Kondrychyn I, Zheng W, Korzh V, Gong Z. Development of zebrafish swimbladder: the requirement of Hedgehog signaling in specification and organization of the three tissue layers. Dev Biol 2009;331:222–236.
Cain JE, Rosenblum ND. Control of mammalian kidney development by the Hedgehog signaling pathway. Pediatr Nephrol 2011;26:1365–1371.
Winberg J, Gustavsson P, Papadogiannakis N et al. Mutation screening and array comparative genomic hybridization using a 180K oligonucleotide array in VACTERL association. PLoS ONE 2014;9:e85313.
Sanna-Cherchi S, Sampogna RV, Papeta N et al. Mutations in DSTYK and dominant urinary tract malformations. N Engl J Med 2013;369:621–629.
Korzh S, Winata CL, Zheng W et al. The interaction of epithelial Ihha and mesenchymal Fgf10 in zebrafish esophageal and swimbladder development. Dev Biol 2011;359:262–276.
Breuss M, Heng JI, Poirier K et al. Mutations in the beta-tubulin gene TUBB5 cause microcephaly with structural brain abnormalities. Cell Rep 2012;2:1554–1562.
Isrie M, Breuss M, Tian G et al. Mutations in either TUBB or MAPRE2 cause circumferential skin creases kunze type. Am J Hum Genet 2015;97:790–800.
Verloes A, Di Donato N, Masliah-Planchon J et al. Baraitser–Winter cerebrofrontofacial syndrome: delineation of the spectrum in 42 cases. Eur J Hum Genet 2015;23:292–301.
Steinfeld H, Cho MT, Retterer K et al. Mutations in HIVEP2 are associated with developmental delay, intellectual disability, and dysmorphic features. Neurogenetics 2016;17:159–164.
Srivastava S, Engels H, Schanze I et al. Loss-of-function variants in HIVEP2 are a cause of intellectual disability. Eur J Hum Genet 2016;24:556–561.
Zhang YJ, O’Neal WK, Randell SH et al. Identification of dynein heavy chain 7 as an inner arm component of human cilia that is synthesized but not assembled in a case of primary ciliary dyskinesia. J Biol Chem 2002;277:17906–17915.
Ibanez-Tallon I, Pagenstecher A, Fliegauf M et al. Dysfunction of axonemal dynein heavy chain Mdnah5 inhibits ependymal flow and reveals a novel mechanism for hydrocephalus formation. Hum Mol Genet 2004;13:2133–2141.
Lee L. Riding the wave of ependymal cilia: genetic susceptibility to hydrocephalus in primary ciliary dyskinesia. J Neurosci Res 2013;91:1117–1132.
Negri G, Milani D, Colapietro P et al. Clinical and molecular characterization of Rubinstein–Taybi syndrome patients carrying distinct novel mutations of the EP300 gene. Clin Genet 2015;87:148–154.
Solomon BD, Bodian DL, Khromykh A et al. Expanding the phenotypic spectrum in EP300-related Rubinstein–Taybi syndrome. Am J Med Genet A 2015;167A:1111–1116.
Yao TP, Oh SP, Fuchs M et al. Gene dosage-dependent embryonic development and proliferation defects in mice lacking the transcriptional integrator p300. Cell 1998;93:361–372.
Flanagan SE, De Franco E, Lango Allen H et al. Analysis of transcription factors key for mouse pancreatic development establishes NKX2-2 and MNX1 mutations as causes of neonatal diabetes in man. Cell Metab 2014;19:146–154.
Bonnefond A, Vaillant E, Philippe J et al. Transcription factor gene MNX1 is a novel cause of permanent neonatal diabetes in a consanguineous family. Diabetes Metab 2013;39:276–280.
Shirane M, Ogawa M, Motoyama J, Nakayama KI. Regulation of apoptosis and neurite extension by FKBP38 is required for neural tube formation in the mouse. Genes Cells 2008;13:635–651.
Wong RL, Wlodarczyk BJ, Min KS et al. Mouse Fkbp8 activity is required to inhibit cell death and establish dorso-ventral patterning in the posterior neural tube. Hum Mol Genet 2008;17:587–601.
Carmeliet P, Ferreira V, Breier G et al. Abnormal blood vessel development and lethality in embryos lacking a single VEGF allele. Nature 1996;380:435–439.
Ferrara N, Carver-Moore K, Chen H et al. Heterozygous embryonic lethality induced by targeted inactivation of the VEGF gene. Nature 1996;380:439–442.
Vargesson N. Thalidomide-induced teratogenesis: history and mechanisms. Birth Defects Res C Embryo Today 2015;105:140–156.
Romero NB, Monnier N, Viollet L et al. Dominant and recessive central core disease associated with RYR1 mutations and fetal akinesia. Brain 2003;126:2341–2349.
Carss KJ, Hillman SC, Parthiban V et al. Exome sequencing improves genetic diagnosis of structural fetal abnormalities revealed by ultrasound. Hum Mol Genet 2014;23:3269–3277.
Yates CL, Monaghan KG, Copenheaver D et al. Whole-exome sequencing on deceased fetuses with ultrasound anomalies: expanding our knowledge of genetic disease during fetal development. Genet Med 2017.
Drury S, Williams H, Trump N et al. Exome sequencing for prenatal diagnosis of fetuses with sonographic abnormalities. Prenat Diagn 2015;35:1010–1017.
Sawyer SL, Hartley T, Dyment DA et al. Utility of whole-exome sequencing for those near the end of the diagnostic odyssey: time to address gaps in care. Clin Genet 2016;89:275–284.
Srour M, Shimokawa N, Hamdan FF et al. Dysfunction of the cerebral glucose transporter SLC45A1 in individualswith intellectual disability and epilepsy. Am J Hum Genet 2017;100:824–830.
Westerfield LE, Stover SR, Mathur VS et al. Reproductive genetic counseling challenges associated with diagnostic exomesequencing in a large academic private reproductive genetic counseling practice. Prenat Diagn 2015;35:1022–1029.
Acknowledgments
This work was supported by grants from the Canadian Institutes of Health Research (CRI 88413 and MOP-136913) and the Fonds de la Recherche du Québec-Santé, as part of the research programs of the Integrated Research Network in Perinatology of Quebec and Eastern Ontario and Réseau de Génétique Médicale Appliquée du Québec. We are grateful to the bioinformatic analysis team of the Réseau de Génétique Médicale Appliquée du Québec (Dan Spiegelman, Ousmane Diallo, Edouard Henrion, Alexandre Dionne-Laporte) for the primary analysis of the sequence data.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Disclosure
The authors declare no conflict of interest.
Electronic supplementary material
Rights and permissions
About this article
Cite this article
Boissel, S., Fallet-Bianco, C., Chitayat, D. et al. Genomic study of severe fetal anomalies and discovery of GREB1L mutations in renal agenesis. Genet Med 20, 745–753 (2018). https://doi.org/10.1038/gim.2017.173
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2017.173
This article is cited by
-
Insights on the Role of α- and β-Tubulin Isotypes in Early Brain Development
Molecular Neurobiology (2023)
-
A novel missense mutation in GREB1L identified in a three-generation family with renal hypodysplasia/aplasia-3
Orphanet Journal of Rare Diseases (2022)
-
A novel autosomal dominant GREB1L variant associated with non-syndromic hearing impairment in Ghana
BMC Medical Genomics (2022)
-
Update on Mayer—Rokitansky—Küster—Hauser syndrome
Frontiers of Medicine (2022)
-
Application of exome sequencing for prenatal diagnosis of fetal structural anomalies: clinical experience and lessons learned from a cohort of 1618 fetuses
Genome Medicine (2022)