Abstract
Purpose
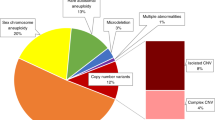
Noninvasive prenatal screening (NIPS) using cell-free DNA in maternal blood is highly sensitive for detecting fetal trisomies 21, 18, and 13. Using a genome-wide approach, other chromosome anomalies can also be detected. We report on the origin, frequency, and clinical significance of these other chromosome aberrations found in pregnancies at risk for trisomy 21, 18, or 13.
Methods
Whole-genome shallow massively parallel sequencing was used and all autosomes were analyzed.
Results
In 78 of 2,527 cases (3.1%) NIPS was indicative of trisomy 21, 18, or 13, and in 41 (1.6%) of other chromosome aberrations. The latter were of fetal (n = 10), placental (n = 22), maternal (n = 1) or unknown (n = 7). One case lacked cytogenetic follow-up. Nine of the 10 fetal cases were associated with an abnormal phenotype. Thirteen of the 22 (59%) placental aberrations were associated with fetal congenital anomalies and/or poor fetal growth (<p10), which was severe (<p2.3) in six cases.
Conclusion
Genome-wide NIPS in pregnancies at risk for trisomy 21, 18, or 13, reveals a chromosomal aberration other than trisomy 21, 18 or 13 in about one-third of the abnormal cases. The majority involves a fetal or placental chromosome aberration with clinical relevance for pregnancy management.
Similar content being viewed by others
Introduction
Noninvasive prenatal screening (NIPS) is a test based on the analysis of cell-free DNA in blood of pregnant women. It is a safe and reliable method to screen for trisomies 21, 18, and 13,1 both in high- and low-risk pregnancies.2 NIPS may also reveal other chromosome aberrations.3 Both the origin of the cell-free DNA as well as the combination of sequencing technique and bioinformatical analysis will determine the types and size of detectable chromosome aberrations.4 When whole-genome sequencing and analysis are performed, other fetal chromosome aberrations than the common aneuploidies can be detected.5, 6 Since the majority of cell-free DNA is of maternal origin, maternal copy-number variants (CNVs) and chromosome anomalies due to a maternal malignancy can also be detected, possibly leading to false-positive NIPS results.3, 7, 8, 9 Moreover, NIPS can reveal chromosomal anomalies limited to the placenta (not present in the fetus), because the so-called cell-free DNA fetal fraction, about 10% at 10 weeks of gestation,10 is derived from the placental cytotrophoblast.11, 12, 13 Therefore, to confirm that a chromosomal anomaly detected with NIPS is fetal, diagnostic testing is necessary. In rare cases NIPS will not detect a trisomy, because it is present in the fetus but not in the placenta.14
In the Netherlands, screening for untreatable disorders is subject to a Ministerial License. As from 1 April 2014, the eight Dutch University Medical Centers received a license to offer NIPS as an alternative option to invasive testing in patients at elevated risk for trisomy 21, 18, or 13. The project was called the TRIDENT study (Trial by Dutch laboratories for Evaluation of Non-Invasive Prenatal Testing), and overall data for the first 5 months have been published.15, 16 Here we present the chromosome aberrations other than trisomy 21, 18 and 13 that were detected during the first year of TRIDENT. Clinical outcomes as well as cytogenetic follow-up studies during and after pregnancy are available for almost all these pregnancies.
Materials and methods
Study population and sample collection
Between 1 April 2014 and 1 April 2015, 3,306 high-risk pregnant women chose NIPS as opposed to diagnostic (invasive) testing or no testing. Women had access to these choices when their risk for a fetus with trisomy 21, 18, or 13 was ≥1:200 based on first trimester combined test results or because of medical history (previous child with a trisomy or parental carriership of a Robertsonian translocation involving chromosome 13 or 21). Exclusion criteria included vanishing twins and are described in more detail in Oepkes et al.15 All women signed an informed consent. A license for the study was granted by the Minister of Health (11016-118701-PG). The license did not allow analysis of the sex chromosomes. The study was approved by the local University Medical Center Ethics Committees.
The cohort in the present study partially overlaps with the cohort previously described, which reported on the common trisomies.15
Massively parallel shotgun sequencing and analysis
Whole-genome shallow massively parallel shotgun sequencing was performed in all cases as described previously.15 A minimum of 5 million mapped reads was set, but most cases were sequenced at a depth of 10–12 million reads. In 753 of 3,306 cases (data from two centers) only the z-scores of chromosomes 21, 18, and 13 were determined. In all other cases (2,553/3,306), WISECONDOR was used for analyzing the whole genome with a resolution of approximately 10–15 Mb at the sequencing depth used.15, 17 The z-score cut-off that was employed for calling trisomies and subchromosomal aberrations based on the sliding window approach was 3. For calling trisomies an additional threshold was used that at least 50% of all bins on a chromosome should be called. For each sample a WISECONDOR output plot was produced and visually inspected as has been described.17 The results of these 2,553 cases are presented here.
Follow-up pre- and postnatal cytogenetic studies
In case of an abnormal NIPS result, parents were counseled by a clinical geneticist and informed consent was obtained for follow-up testing. Depending on the type of chromosomal aberration, maternal DNA was tested with array (Infinium_CytoSNP_850K genotyping array, Illumina Inc., San Diego, CA, USA, CytoScan HD, Affymetrix, Inc, Santa Clara, CA, USA or 180k oligo-array, Agilent, Santa Clara, CA, USA) and/or invasive testing (chorionic villus sampling (CVS) or amniocentesis) was performed. Uniparental disomy studies were performed by single-nucleotide polymorphism array (Illumina or Affymetrix; see above) or by polymorphic microsatellite repeat analysis, in cases of trisomy 6, 7, 11, 14, 15, 16, and 20.
Routine methods were used for prenatal cytogenetic investigations. Chorionic villi were investigated with array using DNA isolated from cytotrophoblast and mesenchymal core separately.18 Uncultured amniotic fluid cells (AF) cells were investigated with fluorescent in situ hybridization or array, and cultured cells with conventional karyotyping, fluorescent in situ hybridization, or array. Chromosomal mosaicism was investigated according to the European Cytogenetic Guidelines.19
In cases of suspected confined placental mosaicism (CPM) postnatal cytogenetic studies of the placenta were performed. One to five chorionic villi biopsies from different quadrants of the placenta were analyzed by karyotyping, fluorescent in situ hybridization, quantitative fluorescence polymerase chain reaction (Devyser compact v3 QF-PCR, Devyser, Hägersten, Sweden; Aneufast v3.0, Nijmegen, Genomed Ltd, Kent, UK; or QSTRplus v2, Elucigene, Manchester, UK), or array. Both cytotrophoblast and mesenchymal core were investigated in most cases.18 If parents refrained from invasive testing, newborn cord blood and/or a skin biopsy was available in most cases.
Classification of abnormal findings
Abnormal findings other than trisomy 21, 18, or 13 were classified into four categories:
-
1
Fetal aberrations: chromosome aberration confirmed in AF, cord blood, or skin.
-
2
Placental aberrations: chromosome aberrations confirmed in first trimester chorionic villi or postnatal placenta but absent in AF/cord blood/postnatal tissue, or chromosome anomalies typically involved in CPM, but with no or insufficient placental tissues available for cytogenetic confirmation. This was based on large chorionic villi cytogenetic studies showing that trisomies such as trisomy 3, 7, 8, 9, 16, and 20 are rarely confirmed in the fetus.12, 20, 21, 22, 23 If in such a case only 1 or 2 placenta biopsies were available and these were normal, a placental aberration was considered not to be excluded due to placental site variation.24
-
3
Maternal CNV: a CNV that is confirmed with array on maternal DNA. A maternal CNV can be inferred from the WISECONDOR results when a small gain or loss with a respectively very high or low z-score is restricted to a small part of a chromosome.
-
4
Unresolved: classification not possible despite pre- and postnatal cytogenetic follow-up investigations.
Follow-up clinical studies
In most cases, a fetal anatomy scan was performed around 20 weeks of gestation as part of standard screening, and when CPM was suspected, scans were repeated in the second and third trimester to monitor fetal growth. Postnatally, clinical follow-up was collected. Intrauterine growth restriction (IUGR) was defined as a birthweight-for-gestational age below the 2.3th percentile (<p2.3) and small for gestational age (SGA) as birthweight below the 10th percentile (<p10).25
Results
In 2,527 of 2,553 (99%) cases, results of genome-wide analysis were reported. In 78 (3.1%) cases a common trisomy was found (trisomy 21 (n = 61), trisomy 18 (n = 11), and trisomy 13 (n = 6)). In 41 (1.6%) cases another chromosome aberration was involved. In one case of trisomy 8, the pregnancy was terminated in a private clinic before follow-up testing could be performed. In the other 40 cases, follow-up was available. A fetal origin of the chromosome aberration was identified in 10 cases (25%), a placental origin in 22 cases (55%), and a maternal origin in 1 case (2%). In 7 cases (18%) the origin remained unknown.
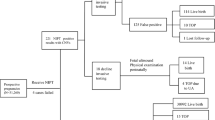
An overview of all 41 cases with a chromosome aberration other than trisomy 21, 18, and 13, their origin, and pregnancy outcome is provided in Figure 1.
The origin of chromosome aberrations other than the common trisomies and pregnancy outcome. AGA, appropriate growth for gestational age; CNV, copy-number variation; DD, developmental delay; FU, follow-up investigations; IUD, intrauterine death; IUGR, intrauterine growth restriction (<p2.3); MCA, multiple congenital anomalies; NIPS, noninvasive prenatal screening; SGA, small for gestational age (<p10); TOP, termination of pregnancy.
In 10/40 (25%) cases the chromosome aberration was shown to be fetal (Table 1 and Supplementary Table S1 online). After confirmation, five pregnancies were terminated and one case ended in an intrauterine death (case 1.5, Table 1 and Supplementary Table S1). Four pregnancies resulted in live births, but three of the infants had multiple congenital anomalies (MCA). The fourth case (case 1.3, Table 1 and Supplementary Table S1) had trisomy 15 mosaicism in uncultured but not in cultured amniotic fluid cells. This child developed normally up to the last examination at 11 months of age. For details about cytogenetic and clinical follow-up, see Supplementary Table S1.
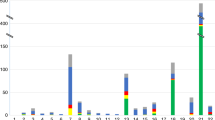
In 22/40 (55%) cases CPM was most likely (Table 2 and Supplementary Table S2 online). Trisomy 7 and trisomy 16 were most commonly involved. Table 2 summarizes the clinical outcome per chromosome aberration. In all but one case of trisomy 2, the pregnancy ended with live birth. MCA were detected in 5/22 (23%) cases (cases 2.1, 2.8, 2.15, 2.16, and 2.19, Supplementary Table S2). Despite normal cytogenetic results in amniotic fluid and/or cord blood, an association with the trisomy and/or uniparental disomy cannot be excluded in four of five cases. In case 2.15, hypomethylation of the H19 differentially methylated region allowed to make the diagnosis of Silver–Russell syndrome. IUGR (<p2.3) was found in 6/22 (27%) cases: two cases with MCA (2.1 and 2.16, Supplementary Table S2) and four cases with isolated IUGR (2.2, 2.6, 2.11, 2.14, Supplementary Table S2). In 7/22 (32%) other cases, SGA (birth weight between p2.3 and p10) was found: three cases with MCA (2.8, 2.15, and 2.19, Supplementary Table S2) and four cases with SGA only (2.13, 2.17, 2.20, and 2.21, Supplementary Table S2). In only 9/22 (41%) cases, the pregnancy was uneventful. Details about the cytogenetic and clinical follow-up studies are shown in Supplementary Table S2.
Maternal CNVs were commonly detected, but most were of benign nature or at the most of unknown clinical significance. Eight such CNVs were reported in the initial phase of the study but with current experience would not be reported anymore, and therefore are not included in the analysis. Only one, a 1.4-Mb duplication on chromosome 17p, associated with Charcot–Marie–Tooth (CMT1A), was disclosed because of its clinical relevance. The mother was already known to be clinically affected and did not opt for invasive testing.
Finally, in 7/40 (18%) cases, the origin of the abnormal NIPS result could not be determined despite clinical and cytogenetic follow-up investigations (Supplementary Table S3 online).
Discussion
Our data show that when genome-wide NIPS is used in pregnant women tested because of an increased risk for one of the common aneuploidies, one-third of the chromosome aberrations are different from trisomy 21, 18, or 13. Genetic follow-up testing confirmed a fetal, placental, or maternal origin of the chromosome aberration in respectively 25%, 55%, and 2% of the abnormal cases. The present study is unique in that near complete (98%) clinical and cytogenetic follow-up was achieved. Especially placental analyses, which usually are not performed,26 allowed us to confirm the large contribution of CPM to abnormal NIPS findings other than trisomy 21, 18, and 13. The frequency of pathogenic maternal CNVs in our series (0.04%, 1/2,553) is comparable to that reported by Brison et al.8 (0.05%, 5/9,882). It is to be expected that in a general obstetric population the relative proportions of the different categories of chromosome aberrations, as shown in this paper, will be different with an expected decrease of common trisomies3 as well as of CPM, which is associated with abnormal first trimester combined test results.
All women had received pretest counseling on possible findings with genome-wide NIPS. When an aberration other than the common trisomies was detected this result was disclosed. In 25% of the cases, diagnostic follow-up testing showed the chromosome aberration to be fetal. All these fetal chromosome anomalies were known to be associated with developmental delay and/or MCA. One case with trisomy 15 mosaicism developed normally up to the age of 11 months. In 55% of the cases CPM was involved, and our data on postnatal outcome confirm the association with an increased risk for fetal developmental and growth problems.27, 28, 29 It is to be expected that the adverse pregnancy outcome in several cases of suspected CPM is caused by occult fetal mosaicism.30 Therefore, it is important that clinicians are aware that the term CPM can be misleading since strict confinement to the placenta can never be proven. Other possible explanations for fetal pathology in case of a placental trisomy are uniparental disomy of an imprinted chromosome, or uniparental disomy leading to homozygosity for a recessive disease allele.31 In one case a maternal pathogenic CNV was found of which the fetus potentially was a carrier. Finally, in 18% of the cases follow-up cytogenetic studies did not reveal the chromosome aberration and therefore their origin remains unknown. An unnoticed vanishing twin, low-grade chromosomal mosaicism, or other biological factors such as sample quality can be responsible for these false-positive results. Because these data show a significant risk for obstetric pathology and fetal anomalies, we advise close obstetric surveillance in all cases of abnormal NIPS results other than the common trisomies, even if the fetal karyotype as determined by diagnostic testing is normal.
We have shown that in an elevated risk population, genome-wide NIPS may detect a range of clinically relevant chromosome anomalies. Questionnaire studies showed that most Dutch pregnant women and health-care providers support testing for a wider range of severe disorders.32, 33 Recently, the American College of Medical Genetics and Genomics did not recommend to screen for autosomal aneuploidies other than those involving chromosomes 21, 18 and 13 nor for genome-wide CNVs.34 It is argued that the former chromosome aberrations are rare, that they may lead to unnecessary diagnostic procedures and even unnecessary pregnancy terminations, and that counseling is challenging.34 As to the frequency, in the present study the rare fetal chromosome anomalies as a group were more frequent than trisomy 13 and almost as frequent as trisomy 18, confirming previous studies.35 All fetal chromosome aberrations but one were severe and relevant for the parents to know. Five couples chose to terminate the pregnancy; in one case the fetus died in utero. Genome-wide NIPS indeed led to 29 (1.1%) extra invasive procedures but also to an increased prenatal detection of true fetal chromosome aberrations that otherwise potentially would have remained undetected: in more than half of the cases with a fetal chromosome aberration in which ultrasound investigations were performed, the 20-week anatomy scan was normal and therefore falsely reassuring. Moreover, with a targeted analysis not all invasive tests would have been avoided since unusual placental aneuploidies2, 36, 37 as well as benign maternal CNVs, present in ~0.1% of pregnancies (45/51,000),38 may lead to test failures or false-positive results for chromosomes 21, 18, or 13, requiring invasive testing as well for a definitive diagnosis.
Pregnancy termination without confirmatory testing occurred once. In most cases this can be prevented by adequate pre- and posttest counseling emphasizing the importance of confirmatory testing. NIPS posttest counseling is challenging but, due to the origin of the cell-free DNA fetal fraction, the NIPS result in fact resembles the cytogenetic result of direct or short-term culture villi. Therefore, for counseling about the clinical significance of an abnormal NIPS result and about the most appropriate diagnostic test, the clinician can use data from a large number of studies on cytogenetic analysis in short- and long-term cultured villi.21, 22, 23, 28, 39, 40 Apart from factors such as gestational age and patient preferences, the choice for CVS or amniocentesis will largely depend on the type of chromosome aberration involved.21 We realize that access to appropriately trained genetic counselors and clinical geneticists differs worldwide and therefore national recommendations will be needed in addition to society guidelines.
In conclusion, about one-third of all chromosome aberrations found with NIPS in pregnant women at increased risk for the common aneuploidies is different from trisomy 21, 18, and 13. This study revealed their origin to be fetal or placental in ~80% of the cases. Fetal as well as placental chromosome anomalies were associated with MCA in the fetus/child and/or impaired fetal growth. It can therefore be concluded that findings other than the common trisomies are clinically relevant for pregnancy management, and that there is potential clinical utility in performing genome-wide NIPS.
References
Lo YM, Corbetta N, Chamberlain PF et al. Presence of fetal DNA in maternal plasma and serum. Lancet. 1997;350:485–487.
Taylor-Phillips S, Freeman K, Geppert J et al. Accuracy of non-invasive prenatal testing using cell-free DNA for detection of Down, Edwards and Patau syndromes: a systematic review and meta-analysis. BMJ Open 2016;6:e010002.
Brady P, Brison N, Van Den Bogaert K et al. Clinical implementation of NIPT—technical and biological challenges. Clin Genet. 2016;89:523–530.
Vermeesch JR, Voet T, Devriendt K . Prenatal and pre-implantation genetic diagnosis. Nat Rev Genet 2016;17:643–656.
Pescia G, Guex N, Iseli C et al. Cell-free DNA testing of an extended range of chromosomal anomalies: clinical experience with 6,388 consecutive cases. Genet Med. 2017;19:169–175.
Fiorentino F, Bono S, Pizzuti F et al. The clinical utility of genome-wide non invasive prenatal screening. Prenat Diagn. 2017;37:593–601.
Snyder MW, Simmons LE, Kitzman JO et al. Copy-number variation and false positive prenatal aneuploidy screening results. N Engl J Med 2015;372:1639–1645.
Brison N, Van Den Bogaert K, Dehaspe L et al. Accuracy and clinical value of maternal incidental findings during noninvasive prenatal testing for fetal aneuploidies. Genet Med. 2017;19:306–313.
Bianchi DW, Chudova D, Sehnert AJ et al. Noninvasive prenatal testing and incidental detection of occult maternal malignancies. JAMA. 2015;314:162–169.
Wang E, Batey A, Struble C, Musci T, Song K, Oliphant A . Gestational age and maternal weight effects on fetal cell-free DNA in maternal plasma. Prenat Diagn. 2013;33:662–666.
Flori E, Doray B, Gautier E et al. Circulating cell-free fetal DNA in maternal serum appears to originate from cyto- and syncytio-trophoblastic cells. Case report. Hum Reprod. 2004;19:723–724.
Grati FR, Malvestiti F, Ferreira JC et al. Fetoplacental mosaicism: potential implications for false-positive and false-negative noninvasive prenatal screening results. Genet Med. 2014;16:620–624.
Srebniak MI, Diderich KE, Noomen P, Dijkman A, de Vries FA, van Opstal D . Abnormal non-invasive prenatal test results concordant with karyotype of cytotrophoblast but not reflecting abnormal fetal karyotype. Ultrasound Obstet Gynecol 2014;44:109–111.
Van Opstal D, Srebniak MI & Polak J et al. False negative NIPT results: risk figures for chromosomes 13, 18 and 21 based on chorionic villi results in 5967 cases and literature review. PLoS One. 2016;11:e0146794.
Oepkes D, Page-Christiaens LC, Bax CJ et al. Trial by Dutch laboratories for evaluation of non-invasive prenatal testing. Part I—clinical impact. Prenat Diagn. 2016;36:1083–1090.
van Schendel RV, Page-Christiaens L, Beulen L et al. Trial by Dutch laboratories for evaluation of non-invasive prenatal testing. Part II— women’s perspectives. Prenat Diagn. 2016;36:1091–1098.
Straver R, Sistermans EA, Holstege H, Visser A, Oudejans CB, Reinders MJ . WISECONDOR: detection of fetal aberrations from shallow sequencing maternal plasma based on a within-sample comparison scheme. Nucleic Acids Res 2014;42:e31.
Smidt-Jensen S, Christensen B, Lind AM . Chorionic villus culture for prenatal diagnosis of chromosome defects: reduction of the long-term cultivation time. Prenat Diagn. 1989;9:309–319.
Hastings R, Howell P, Bricarelli FD, Kristoffersson U, Cavani S. Specific Constitutional Cytogenetic Guidelines. Accessed on 2012. http://www.e-c-a.eu/en/GUIDELINES.html.
Wolstenholme J . Confined placental mosaicism for trisomies 2, 3, 7, 8, 9, 16, and 22: their incidence, likely origins, and mechanisms for cell lineage compartmentalization. Prenat Diagn. 1996;16:511–524.
Van Opstal D, Srebniak MI . Cytogenetic confirmation of a positive NIPT result: evidence-based choice between chorionic villus sampling and amniocentesis depending on chromosome aberration. Expert Rev Mol Diagn 2016;16:513–520.
Malvestiti F, Agrati C, Grimi B et al. Interpreting mosaicism in chorionic villi: results of a monocentric series of 1001 mosaics in chorionic villi with follow-up amniocentesis. Prenat Diagn. 2015;35:1117–1127.
Hahnemann JM, Vejerslev LO . European collaborative research on mosaicism in CVS (EUCROMIC)—fetal and extrafetal cell lineages in 192 gestations with CVS mosaicism involving single autosomal trisomy. Am J Med Genet 1997;70:179–187.
Schuring-Blom GH, Keijzer M, Jakobs ME et al. Molecular cytogenetic analysis of term placentae suspected of mosaicism using fluorescence in situ hybridization. Prenat Diagn. 1993;13:671–679.
Visser GH, Eilers PH, Elferink-Stinkens PM, Merkus HM, Wit JM . New Dutch reference curves for birthweight by gestational age. Early Hum Dev 2009;85:737–744.
Lebo RV, Novak RW, Wolfe K, Michelson M, Robinson H, Mancuso MS . Discordant circulating fetal DNA and subsequent cytogenetics reveal false negative, placental mosaic, and fetal mosaic cfDNA genotypes. J Transl Med 2015;13:260.
Lestou VS, Kalousek DK . Confined placental mosaicism and intrauterine fetal growth. Arch Dis Child Fetal Neonatal Ed 1998;79:F223–226.
Toutain J, Labeau-Gauzere C, Barnetche T, Horovitz J, Saura R . Confined placental mosaicism and pregnancy outcome: a distinction needs to be made between types 2 and 3. Prenat Diagn. 2010;30:1155–1164.
Kalousek DK, Howard-Peebles PN, Olson SB et al. Confirmation of CVS mosaicism in term placentae and high frequency of intrauterine growth retardation association with confined placental mosaicism. Prenat Diagn. 1991;11:743–750.
Benn P . Trisomy 16 and trisomy 16 mosaicism: a review. Am J Med Genet 1998;79:121–133.
Eggermann T, Soellner L, Buiting K, Kotzot D . Mosaicism and uniparental disomy in prenatal diagnosis. Trends Mol Med 2015;21:77–87.
van Schendel RV, Dondorp WJ, Timmermans DR et al. NIPT-based screening for Down syndrome and beyond: what do pregnant women think? Prenat Diagn. 2015;35:598–604.
Tamminga S, van Schendel RV, Rommers W et al. Changing to NIPT as a first-tier screening test and future perspectives: opinions of health professionals. Prenat Diagn. 2015;35:1316–1323.
Gregg AR, Skotko BG, Benkendorf JL et al. Noninvasive prenatal screening for fetal aneuploidy, 2016 update: a position statement of the American College of Medical Genetics and Genomics. Genet Med. 2016;18:1056–1065.
Van Opstal D, de Vries F, Govaerts L et al. Benefits and burdens of using a SNP array in pregnancies at increased risk for the common aneuploidies. Hum Mutat. 2015;36:319–326.
Snyder HL, Curnow KJ, Bhatt S, Bianchi DW . Follow-up of multiple aneuploidies and single monosomies detected by noninvasive prenatal testing: implications for management and counseling. Prenat Diagn. 2016;36:203–209.
Sistermans E, Straver R, Faas BH . Maternal malignancies detected with noninvasive prenatal testing. JAMA. 2015;314:2192.
van den Boom D, Ehrich M, Kim SK . Copy-number variation and false positive results of prenatal screening. N Engl J Med 2015;373:2584.
Wolstenholme J, Emslie JB, Connors S . Association of Clinical Cytogeneticists chorion villus sampling database 1987-2000. Prenat Diagn. 2006;26:420–427.
Battaglia P, Baroncini A, Mattarozzi A et al. Cytogenetic follow-up of chromosomal mosaicism detected in first-trimester prenatal diagnosis. Prenat Diagn. 2014;34:739–747.
Acknowledgements
This work was supported by a grant from the Netherlands Organisation for Health Research and Development (ZonMw, 200340002). The authors thank all members of the Dutch NIPT Consortium for their contribution to this study.
Author information
Authors and Affiliations
Consortia
Corresponding author
Ethics declarations
Competing interests
GCP-C reports being a full-time employee of Illumina since 18 January 2016. The other authors declare no conflict of interest.
Additional information
Supplementary material is linked to the online version of the paper at
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Van Opstal, D., van Maarle, M., Lichtenbelt, K. et al. Origin and clinical relevance of chromosomal aberrations other than the common trisomies detected by genome-wide NIPS: results of the TRIDENT study. Genet Med 20, 480–485 (2018). https://doi.org/10.1038/gim.2017.132
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2017.132
Keywords
This article is cited by
-
Clinical outcomes of screen-positive genome-wide cfDNA cases for trisomy 20: results from the global expanded NIPT Consortium
Molecular Cytogenetics (2024)
-
Psychological impact of additional findings detected by genome-wide Non-Invasive Prenatal Testing (NIPT): TRIDENT-2 study
European Journal of Human Genetics (2024)
-
Population screening for 15q11-q13 duplications: corroboration of the difference in impact between maternally and paternally inherited alleles
European Journal of Human Genetics (2024)
-
Laboratory performance of genome-wide cfDNA for copy number variants as compared to prenatal microarray
Molecular Cytogenetics (2023)
-
Phenotypic findings and pregnancy outcomes of fetal rare autosomal aneuploidies detected using chromosomal microarray analysis
Human Genomics (2022)