Abstract
Purpose:
Predicting the phenotypic effects of mutations has become an important application in clinical genetic diagnostics. Computational tools evaluate the behavior of the variant over evolutionary time and assume that variations seen during the course of evolution are probably benign in humans. However, current tools do not take into account orthologous/paralogous relationships. Paralogs have dramatically different roles in Mendelian diseases. For example, whereas inactivating mutations in the NPC1 gene cause the neurodegenerative disorder Niemann-Pick C, inactivating mutations in its paralog NPC1L1 are not disease-causing and, moreover, are implicated in protection from coronary heart disease.
Methods:
We identified major events in NPC1 evolution and revealed and compared orthologs and paralogs of the human NPC1 gene through phylogenetic and protein sequence analyses. We predicted whether an amino acid substitution affects protein function by reducing the organism’s fitness.
Results:
Removing the paralogs and distant homologs improved the overall performance of categorizing disease-causing and benign amino acid substitutions.
Conclusion:
The results show that a thorough evolutionary analysis followed by identification of orthologs improves the accuracy in predicting disease-causing missense mutations. We anticipate that this approach will be used as a reference in the interpretation of variants in other genetic diseases as well.
Genet Med 18 10, 1029–1036.
Similar content being viewed by others
Introduction
With the revolutionary developments in sequencing technologies,1 molecular testing is now widely used to support clinical diagnosis and to identify unknown causes of genetic disorders.1,2,3 There are several approaches to evaluating the effect of a variant: (i) evidence-based, (ii) frequency-based, (iii) functional (variants with obviously drastic consequences such as nonsense and frameshift mutations), and (iv) predictive.4 The first three approaches are successful in determining the effects of variants but are limited when it comes to the variants of unknown significance.1 For novel variants, which comprise the majority of coding variation,5 in silico prediction is a quick way is to estimate potential consequences. Computational tools, such as PolyPhen6 and SIFT,7 are frequently used to evaluate genetic variations; however, they are not yet at the level of desired performance in terms of sensitivity and specificity, even for well-studied monogenic Mendelian diseases.4,8,9 These tools take into account the following key parameters: sequence conservation, structural constraints, and physiochemical properties of amino acids and known annotations, such as functionally important sites. Risk estimation is largely dependent on the molecular conservation, which is inferred from comparative sequence analysis,10 and is based on the fact that most disease-causing mutations cause a reduction of evolutionary fitness; therefore, they are not selected for and are not observed in homologs in other organisms.9 To identify homologous sequences in other organisms, current tools use automated sequence similarity searches followed by multiple sequence alignment (MSA) and clustering. Consequently, sets of similar sequences that are used in the downstream analysis usually include both orthologs and paralogs.6 This approach is based on the argument that disease-causing substitutions far more often affect protein structure than function,11 and although paralogous proteins may have a slightly different function, their structure is fully conserved.
However, the roles of paralogous genes in disease and health are different. In most cases of Mendelian diseases, only one of the paralogous genes is associated with the disease.12 In 87% of the gene pairs, only one pair is associated with disease, and this trend is observed in gene families with more than two members. Once a gene is duplicated, purifying selection pressure on one or both of the copies is relaxed and they become more prone to accumulating mutations. This divergence can lead to sub-functionalization or neofunctionalization,13 often resulting in different roles of paralogs in disease.
This pattern is observed in Niemann-Pick disease type C (NP-C), which is a neurovisceral lysosomal lipid storage disease.14,15,16 NP-C is inherited in an autosomal recessive pattern and is caused by mutations in either NPC1 or NPC2 genes.17 NPC1 and NPC2 proteins work in concert to transport cholesterol from the endosomal/lysosomal compartment.16,18 Homozygous loss of function in either protein perturbs lipid homeostasis, which results in pathogenicity. Ninety-five percent of affected individuals carry pathogenic mutations in the NPC1 gene,15 which recently attracted attention because of its role in the entry of the Ebola virus.19 By contrast, the NPC1 paralog, NPC1L1, is not associated with the disease. On the contrary, inactivating mutations in NPC1L1 reduce the risk of coronary heart disease.20 NPC1 deletion in mice causes hearing loss,21 defects in retina,22 and deficiency in cerebellum development,23 whereas NPC1L1 deficiency protects ApoE-/- mice against atherosclerosis.24
Diagnosis of NP-C is challenging because of the heterogeneity in symptoms and clinical presentation.16 Until recently, the diagnostic standard was filipin staining of unesterified cholesterol in fibroblasts obtained by skin biopsy.25,26 This test, however, is definitive in only approximately two-thirds of cases. NP-C diagnostics has been significantly improved through the discovery of cholesterol oxidation products (“oxysterols”) that are elevated in the plasma of NP-C subjects.27 The plasma oxysterol assay detects >97% of cases with 100% sensitivity.28 DNA sequencing offers another tool for NP-C diagnostics, but in practice detects only ~85% of NP-C cases due to the large number of private and noncoding sequence mutations.29 For novel missense mutations, in silico tools are indispensable for predicting potential NP-C. However, these tools use different algorithms and data sets to build MSAs to assess the variant effect, resulting in substantial inconsistencies.30 Researchers usually rely on agreement between several tools, which has the effect of increasing specificity while decreasing the sensitivity.31 Moreover, the tools that use conservation information do not discriminate between orthologous and paralogous proteins6 and thus include NPC1 paralogs, such as NPC1L1, in their analysis. Although including paralogs in risk-estimating data sets is convenient (this eliminates computationally demanding and often nontrivial steps to separate orthologs and paralogs), such simplification confounds the function-specific signal.
NP-C disease caused by NPC1 mutations is an ideal case study to understand the effects of paralogs in predicting disease-causing mutations because of a dramatic consequence of the duplication event that yielded NPC1L1. Moreover, many experimentally validated disease-causing mutations as well as alleles with high frequencies that are likely to be benign are known for this gene (Supplementary Table S1 online).
In this study, we established the precise evolutionary history of the NPC1 gene and identified evolutionary events that most likely affected its function. We used this information to build a computational approach that showed improved accuracy in categorizing damaging and benign single amino acid substitutions in NPC1.
Materials and Methods
Databases, multiple sequence alignments, and phylogenetic trees
Human NPC1 protein (NM_000271.4) was queried through BLASTP32 against the human genome to reveal the related sequences. Each hit was blasted individually against the RefSeq database. For each job, the full sequences were compiled and aligned using MAFFT default algorithm.33 Neighbor joining tree was built with the phylip package.34 From the tree, the NPC1 homologs clade was isolated. With the retrieved homologs, MAFFT version v7.154b E-INS-i algorithm was used to realign the full-length sequences. The phylogenetic trees were built using the maximum likelihood approach with PHYML software version 2014092935 and applying the JTT substitution model and the remaining parameters as default. The outgroups that were not considered to be NPC1 homologs based on RefSeq annotations and domain architectures were discarded from the MSA, NPC1 homologs were realigned, and the final phylogenetic tree was built.
Orthology assignment
Orthologs and paralogs were distinguished using the maximum likelihood phylogenetic tree. In case of major duplication events, a consistently more divergent duplicated clade was categorized as paralogs that are less likely to retain the original NPC1 function. The reference point for evolutionary distance was determined as the full-length NPC1 node. In the cases in which no divergence consistency between clades was observed (e.g., not all species in clade A were more diverged than those in clade B, or incomplete species set in both clades), the orthology assignment was deemed inconclusive. For the species-level duplications, the sequence, which was significantly diverged from the closest node of NPC1 orthologs, was categorized as paralogous.
Scoring the effect of single amino acid variants
PubMed 1997–2014 database was searched to identify relevant studies and case series. The initial search resulted in 312 articles. General review articles on NP-C disease, articles lacking genetic testing, and experimental findings not connected with clinical data were excluded. As a result, we identified 56 articles referencing a total of 572 mutations in the NPC1 gene. After excluding repetitive reports, insertion/deletion, frameshift and nonsense mutations, and benign SNPs, the final list of the most likely pathogenic nsSNVs comprised 166 variants that were referred to as “damaging” variants in this study. To retrieve the set of “benign” mutations, we used frequencies in human populations reported by Wassif et al.31 The variants found in humans with higher frequency than the most common deleterious variant, I1063T, were categorized as benign. However, we removed N222S, N961S, S1200G, and A521S from this list due to the reports suggesting that they might be damaging.
In MSA, misaligned amino acids are masked based on the queried position. First, residues adjacent to ambiguous amino acids (represented by X) are masked. Second, if a sequence has insertion or deletion in close proximity to the position of interest, then that residue is masked. Selection of the representative isoform also depends on the queried position. Because of the variations in splice sites, a single representative isoform may not align well with the rest of the proteins for each position. Therefore, selection of the representative isoform based on the queried position yields better alignment quality. In the algorithm, the “moderately variable” category was defined as a position having more than five different substitutions in a given set. Position was categorized as “hypervariable” if there were more than nine different substitutions (see the source code and related input files such as MSA and phylogenetic tree at http://genomics.utk.edu/saver/source.rar).
Statistical analyses
The performance of the algorithm is described by the following parameters: sensitivity; specificity; false discovery rate; accuracy; F1 score; and Matthews correlation coefficient. In the equations given here, TP, TN, FP, and FN refer to the number of true positives, true negatives, false positives, and false negatives, respectively.

Domain architecture prediction and risk map generation
To build domain architectures, we used the CDvist Web server with HMMER3 against Pfam 27.0 and HHsearch against Protein Data Bank options.36,37 Protein Data Bank HHsearch probability cutoff was adjusted to 98%.
We implemented the SAVER algorithm in a PYTHON3 script and ran it on all theoretical human NPC1 amino acid variants. For each position, we counted the allowed (benign) amino acids. The range was between 0 (no substitution allowed) and 19 (any substitution allowed). For secondary structure information, X-ray crystal structure (protein data bank ID: 3GKH) was used for N-terminal domain and Psipred prediction was used for the rest. The Protter Web application was used to generate the NPC1 membrane topology figure.38
Results
Distinct clusters of NPC1 homologs suggest different functions
NPC1 protein has 13 transmembrane (TM) regions with three luminal domains. The crystal structure of the N-terminal domain has been solved with bound cholesterol, indicating its role in cholesterol binding and transport.39 The pentahelical sterol-sensing domain, which resides between TM3 and TM8, is required for cholesterol egress from the lysosome. There are nine human genes that share similarity through their sterol-sensing domains and are identifiable by BLASTP initiated with NPC1: NPC1, NPC1-L1, PTCH1, PTCH2, PTCHD2, PTCHD3, PTCHD4, SCAB SREBF, and DISP. These related proteins also share the “Patched” domain, which has a role in cholesterol-dependent processes. Domain architectures of these proteins show significant differences, with only NPC1 and NPC1-L1 containing the N-terminal cholesterol-binding domain ( Figure 1a ). A phylogenetic tree constructed from the MSA of all Patched domain proteins shows distinct clades, where the NPC1-NPC1L1 clade is clearly separated from the rest ( Figure 1b ). These findings strongly suggest that other Patched-containing sequences should not be taken into account when examining function-specific characteristics of NPC1. In contrast, automated tools often include such functionally unrelated sequences in their data sets (Supplementary Figure S1 online).
Relationships between Patched domain–containing proteins. (a) The domain architectures of human Patched domain–containing proteins were retrieved using the CDvist Web server. Boxes with a white background represent PFAM domains. Cholesterol-binding domains (in blue) were retrieved using a PDB database profile. The cholesterol-binding domain was found exclusively in NPC1 and NPC1L1. (b) Some pairs such as PTCH1-PTCH2, NPC1-NPC1L1, and PTCHD3-PTCHD4 have a relatively recent common ancestor, whereas the other proteins are related to each other more distantly, as they are represented as single clades on the phylogenetic tree. According to the phylogenetic tree, the NPC1-NPC1L1 clade is clearly separated from other Patched domain–containing sequences. PDB, protein data bank.
Major events in NPC1 evolution
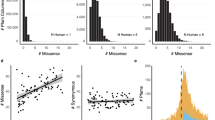
NPC1 is found in four of the five eukaryotic supergroups—unikonta, plants, chromalveolata, and excavates—and is missing from rhizaria. Phylogenetic analysis of NPC1 protein shows that the NPC1 gene followed vertical evolution. Thus, it is likely that NPC1 was present in the last eukaryotic common ancestor. Multiple gene duplication events are observed in taxonomic ranks from superorder to species level; among 397 species with NPC1, 195 (49%) have more than one copy (Supplementary Figure S2 online).
In the common ancestor of gnathostomata (jawed vertebrates), the NPC1 gene was duplicated, giving rise to the “NPC1-like” protein, which is present in most jawed vertebrates including humans (named NPC1L1). The NPC1L1 clade is greatly diverged from the root when compared to the gnathostomatan NPC1 clade ( Figure 2 ). NPC1 is present in each organism that has NPC1-L1; however, the opposite is not true. Moreover, the NPC1L1 clade has a longer average branch length from its root, indicating a greater divergence (Supplementary Figure S3 online). The NPC1L1 divergence and dispensability strongly suggest that its function is different from that of NPC1, which is consistent with the observation that mutations in NPC1-L1 do not cause the disease. We observed a very similar pattern of NPC1 duplication in neoptera ( Figure 2 and Supplementary Figure S3 online).
Maximum likelihood phylogenetic tree of NPC1 proteins and described sets. The star is placed at the root of full-length NPC1. On the left side, the black markers represent the closest NPC1 to the root for each organism. Green markers (Sets 1 and 2) show the orthologs, whereas red markers point to paralogs. Blue markers represent sequences that are ambiguous in terms of orthology. The gray-shaded clade contains a short version of NPC1. Set 1, which contains the HsNPC1 orthologs after the most recent duplication, is a subset of Set 2.
In fungi and amoebozoa, several duplications occurred, but only at the species and genus level, suggesting there was no major duplication event in these kingdoms.
In plants, there was NPC1 duplication in the common ancestor of flowering plants. More than one paralog is observed in Pentapetalae. However, the distances of two clades from the root are comparable ( Figure 2 ). Furthermore, some organisms have only one version of the gene from either clade, which suggests that one paralog is sufficient and neither copy is indispensable. Internal diversity was comparable in two clades (Supplementary Figure S3 online). Therefore, the paralogs may not have gained significantly different functions. Thus, the Homo sapiens NPC1 (HsNPC1) orthology assignment cannot be precisely performed in plants.
Unikonts (metazoa, fungi, and amoebozoa) and plants have the full-length NPC1 protein with 13 TM regions, except for Dictyostelium (Supplementary Figure S4 online). They all accommodate a lumenal N-terminal domain that binds to cholesterol. However, in Naegleria gruberi (excavate) and in most chromalveolates, this domain is missing, resulting in a shorter protein with 12 TM regions (Supplementary Figure S4 online). We found that all organisms that lack the NPC1 N-terminal domain have a separate protein (~300 amino acids) encoded in their genomes, which is homologous (~30% identity, ~50% similarity) to the N-terminal domain of the full-length HsNPC1. Oomycetes have both “full” and “short” versions of NPC1. In the phylogenetic tree, these two versions are distinctly separated. Except for Nannochloropsis gaditana (which has an atypical NPC1 with no sterol-sensing domain), all organisms with the short version of the NPC1 protein also have the separate cholesterol-binding protein. Moreover, the separate cholesterol-binding protein is found exclusively in the organisms that have the short NPC1. The separate cholesterol-binding protein is predicted to have a signal peptide at the N-terminus and a TM region at the C-terminus. Thus, concatenation of the separate cholesterol-binding protein and the short version of NPC1 substantially resembles HsNPC1. Exclusive coexistence of these two proteins suggests that they interact and function similarly to the full version of NPC1. The existence of both “full” and “short” versions in oomycetes and the vertical evolutionary patterns suggest that both versions could have been present in the last eukaryotic common ancestor, where either fusion or dissociation could have occurred; then, only one version was kept in all organisms, except for oomycetes, in which both were kept.
In addition to major duplication events, in each kingdom there were also species and/or genus level duplications. In such cases, one copy evolves slowly to keep the original function and the extra copies, which are not prone to the same levels of purifying selective pressure, diverge faster. We used the distance measurements from the common ancestor node in the phylogenetic tree to determine the slowest evolving gene (the clade with a shorter branch distance to the common ancestor), which in turn enabled us to find the functional orthologs. Notably, NPC1 was lost in many parasites including whole clades, such as microsporidia (fungi) and apicomplexa (chromalveolata). Except for N. gruberi, all species sequenced in the Excavata supergroup are parasitic (Trypanosomatidae family, Trichomonas vaginalis, and Giardia intestinalis) and contain no NPC1 in their genomes.
Defining HsNPC1 functional orthologs
Products of orthologous genes are very likely to perform the same function. Therefore, distinguishing HsNPC1 orthologs from other homologous proteins is critical for identifying potentially pathogenic variants specifically affecting HsNPC1 function. Detailed analysis of the phylogenetic tree of all NPC1 homologs guided HsNPC1 orthology assignment. The clades retaining the original NPC1 function were determined based on the agreement of three lines of evidence. First, we compared the distances of duplicated clades to the full-length NPC1 root ( Figure 2 ) to identify which one is less diverged. Second, we compared the organism content of the clades. If a clade is a subset of another, then the superset clade was considered the “original” one representing HsNPC1 orthologs. Finally, diversity within the clades was assessed; the less diverged clade is more likely to be ancestral (Supplementary Figure S3 online). When all three criteria agree, HsNPC1 orthologs can be identified with confidence. However, in some cases, the sequence divergence information was inconclusive. In those cases, none of the clades was a subset of another. Moreover, the diversity within the clades was comparable. Consequently, these sequences were not included in the set of HsNPC1 orthologs.
Evaluating missense mutations in HsNPC1: the scoring algorithm
Our master MSA included all homologs. We divided the master MSA into three sets grouped by the orthology relationships (see Figure 2 for details). The phylogenetic clade containing HsNPC1 after the most recent major evolutionary event, which is the birth of NPC1L1 in gnathostomata, was considered the core alignment. This alignment, referred to as “Set 1,” was given the highest importance in the evaluation algorithm. Set 2 includes Set 1 and also other unambiguous HsNPC1 orthologs. Finally, Set 3 contained all other HsNPC1 homologs, including paralogs, except for the short versions of NPC1.
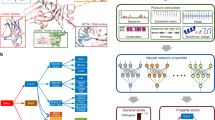
To predict the effect of missense mutations on HsNPC1 function, we propose an algorithm (SAVER: Single Amino Acid Variant Evaluator) that provides binary output from the MSA analysis of Sets 1 and 2 ( Figure 3 ). In the scoring part, Set 1 is given the highest weight because it contains HsNPC1 and its orthologs that evolved after the most recent duplication and the birth of many Mendelian diseases correlates with the time of most recent duplications.12 However, using only Set 1, which is limited in our case to bilaterian genomes, would not be sufficient for collecting the entire ancestral information. For this reason, Set 2 was used to compensate for the lack of evolutionary depth in Set 1. Because Set 2 was carefully constructed from sequences that are likely to conserve the ancestral function of NPC1, the amount of false signal it introduces is limited. Furthermore, the possibility of false signals in Set 2 was addressed by lowering its priority. Because sufficient evolutionary depth was reached with Set 2, specificity was not affected drastically by excluding sequences only in Set 3.
Sequencing and aligning errors are key factors causing misinterpretation. Thus, working with the cleanest possible data set, a nearly perfect alignment and well-constructed phylogenetic trees is critical for assessing the mutations. Ab initio elimination of sequences that have misaligned regions is not an optimum solution; therefore, we apply positional masking of misaligned regions so that only well-aligned positions are taken into account. Another challenge in eukaryotic sequence comparison is dealing with isoforms that can redundantly dominate the signal and cause artificial conserved positions. We resolve this issue by choosing a representative isoform for each gene that depends on the queried position.
For a single amino acid substitution from AA0 to AA1, scoring algorithms usually use the abundance of the AA1 in MSA. However, instead of counting the number of sequences with substitutions, we propose counting how many times a given replacement has occurred independently so that a single evolutionary event would not be counted multiple times. Multiple independent substitutions occurring in different clades suggest that a position tolerates mutations, whereas a single substitution compensated by a suppressor mutation can be in a potentially “irreplaceable” position.
Improved success in distinguishing between damaging and benign nsSNVs
We scanned literature to retrieve known NPC1 variants. Only single amino acid substitutions were taken into account. Only biochemically validated NP-C–causing mutations were considered as “damaging” variants. Frequencies of HsNPC1 variants from several exome sequencing data sets31 were used to define the benign mutation data set. We selected the common variants that have never been shown as pathogenic in any study and that have frequency greater than 0.028%, which is the frequency of the most commonly reported pathogenic variant, I1061T. Our compiled control set contained 166 damaging and 21 benign nsSNVs (Supplementary Table S1 online).
We tested our approach in comparison with automated tools PolyPhen-2, SIFT, and PROVEAN.6,7,40 The results indicate that our approach outperforms other tools in terms of sensitivity (~10% improvement), while causing a relatively low cost in specificity, and (ii) in terms of the overall quality, as measured by the Matthews correlation coefficient ( Table 1 ). The drastic improvement in sensitivity can be explained by the fact that our method eliminates the false evolutionary signals introduced by functionally diverged sequences that are included in the analysis by other tools (Supplementary Table S1 online).
We also applied our method to all theoretical amino acid substitutions in NPC1; 24282 (1278 positions in NPC1 sequence X 19 amino acid substitutions) theoretical single amino acid variants were evaluated in comparison with the automated methods described above (Supplementary Table S1 online). Ultimately, our method predicts 81% of the variants as damaging, whereas PolyPhen-2, PROVEAN, and SIFT predict 60, 70 and 66% as damaging, respectively. Because we suspected that our approach overpredicts damaging variants, we adjusted the cutoffs of other tools to fix the damaging rate at 81%. After the adjustment, the performance of two methods (PolyPhen-2 and PROVEAN) was improved; however, none of them reached the quality of our approach, as measured by the Matthews correlation coefficient value. Comparison between receiver-operating characteristics of the tools and our “sensitivity false-positive rate” datum shows a clear distinction of our result from the general trend of the others (Supplementary Figure S5 online).
An example of how inclusion of paralogous sequences might negatively affect the prediction is shown in Figure 4 . Known pathogenic mutations, N968S, G986S, G993A, and M995R (see Supplementary Table S1 online), are predicted as benign by all three automated tools, probably because the same substitutions are found in NPC1L1 paralogs that are included in their MSA sets ( Figure 4 ). We generated a risk map for NPC1-caused NP-C disease, where the topology of the human NPC1 is shown with the positions colored based on the number of allowed substitutions (Supplementary Figure S6 online). This risk map provides clues about the functionally critical regions of HsNPC1 (Supplementary Text S1 online), and the full list of potentially damaging and benign substitutions in this protein is provided as Supplementary Table S2 online. We have built a Web-based application for querying single amino acid variants in NPC1, which can serve as a reference for clinicians. It is freely available at http://genomics.utk.edu/saver/npc1.html.
An alignment window illustrating false effects of paralogs in predicting damaging mutations. Blue-shaded sequences are HsNPC1 orthologs and the rest are paralogs. For each tool, the red marker represents “predicted as damaging” and the green marker represents “predicted as benign.” Residues highlighted in red are the potential causes of predicting pathogenic variants as benign.
Discussion
In this work, we showed that it is possible to get closer to the desired level of predicting the effects of missense mutations by carefully analyzing the evolutionary history of a gene. A clear improvement is accomplished by taking into consideration only function-specific orthologous protein sequences. Remote homologs and paralogs that are likely to be functionally diverged should be removed from the analysis. In selecting functional counterparts, specific criteria based on a thorough phylogenetic analysis must be used.
The proposed approach depends greatly on manual work (constructing high-quality data sets, alignments, trees, and defining orthologs and paralogs) as well as reasoning, which depends on the output of a particular computational step. Thus, at this time, this approach cannot be fully automated and will not replace any of the available automated tools. However, revealing common trends and problems in identifying functional orthologs and testing this approach on other well-defined monogenic Mendelian diseases should lead to the development of the next generation of predictive automated methods directly applicable in clinical practice.
Disclosure
The authors declare no conflict of interest.
References
Katsanis SH, Katsanis N. Molecular genetic testing and the future of clinical genomics. Nat Rev Genet 2013;14:415–426.
Chang F, Li MM. Clinical application of amplicon-based next-generation sequencing in cancer. Cancer Genet 2013;206:413–419.
Ng SB, Buckingham KJ, Lee C, et al. Exome sequencing identifies the cause of a mendelian disorder. Nat Genet 2010;42:30–35.
Oliver GR, Hart SN, Klee EW. Bioinformatics for clinical next generation sequencing. Clin Chem 2015;61:124–135.
Tennessen JA, Bigham AW, O’Connor TD, et al.; Broad GO; Seattle GO; NHLBI Exome Sequencing Project. Evolution and functional impact of rare coding variation from deep sequencing of human exomes. Science 2012;337:64–69.
Adzhubei IA, Schmidt S, Peshkin L, et al. A method and server for predicting damaging missense mutations. Nat Methods 2010;7:248–249.
Ng PC, Henikoff S. SIFT: predicting amino acid changes that affect protein function. Nucleic Acids Res 2003;31:3812–3814.
Sunyaev SR. Inferring causality and functional significance of human coding DNA variants. Hum Mol Genet 2012;21:R10–R17.
Jordan DM, Ramensky VE, Sunyaev SR. Human allelic variation: perspective from protein function, structure, and evolution. Curr Opin Struct Biol 2010;20:342–350.
Ng PC, Henikoff S. Predicting the effects of amino acid substitutions on protein function. Annu Rev Genomics Hum Genet 2006;7:61–80.
Wang Z, Moult J. SNPs, protein structure, and disease. Hum Mutat 2001;17:263–270.
Dickerson JE, Robertson DL. On the origins of Mendelian disease genes in man: the impact of gene duplication. Mol Biol Evol 2012;29:61–69.
Lynch M, Conery JS. The evolutionary fate and consequences of duplicate genes. Science 2000;290:1151–1155.
Jahnova H, Dvorakova L, Vlaskova H, et al. Observational, retrospective study of a large cohort of patients with Niemann-Pick disease type C in the Czech Republic: a surprisingly stable diagnostic rate spanning almost 40 years. Orphanet J Rare Dis 2014;9:140.
Patterson MC, Hendriksz CJ, Walterfang M, Sedel F, Vanier MT, Wijburg F ; NP-C Guidelines Working Group. Recommendations for the diagnosis and management of Niemann-Pick disease type C: an update. Mol Genet Metab 2012;106:330–344.
Vanier MT. Niemann-Pick disease type C. Orphanet J Rare Dis 2010;5:16.
Vanier MT. Complex lipid trafficking in Niemann-Pick disease type C. J Inherit Metab Dis 2015;38:187–199.
Sleat DE, Wiseman JA, El-Banna M, et al. Genetic evidence for nonredundant functional cooperativity between NPC1 and NPC2 in lipid transport. Proc Natl Acad Sci USA 2004;101:5886–5891.
White JM, Schornberg KL. A new player in the puzzle of filovirus entry. Nat Rev Microbiol 2012;10:317–322.
Stitziel NO, Won HH, Morrison AC, et al.; Myocardial Infarction Genetics Consortium Investigators. Inactivating mutations in NPC1L1 and protection from coronary heart disease. N Engl J Med 2014;371:2072–2082.
King KA, Gordon-Salant S, Pawlowski KS, et al. Hearing loss is an early consequence of Npc1 gene deletion in the mouse model of Niemann-Pick disease, type C. J Assoc Res Otolaryngol 2014;15:529–541.
Yan X, Ma L, Hovakimyan M, et al. Defects in the retina of Niemann-pick type C 1 mutant mice. BMC Neurosci 2014;15:126.
Nusca S, Canterini S, Palladino G, et al. A marked paucity of granule cells in the developing cerebellum of the Npc1(-/-) mouse is corrected by a single injection of hydroxypropyl-ß-cyclodextrin. Neurobiol Dis 2014;70:117–126.
Davis HR Jr, Hoos LM, Tetzloff G, et al. Deficiency of Niemann-Pick C1 Like 1 prevents atherosclerosis in ApoE-/- mice. Arterioscler Thromb Vasc Biol 2007;27:841–849.
Börnig H, Geyer G. Staining of cholesterol with the fluorescent antibiotic “filipin”. Acta Histochem 1974;50:110–115.
Vanier MT, Latour P. Laboratory diagnosis of Niemann-Pick disease type C: the filipin staining test. Methods Cell Biol 2015;126:357–375.
Porter FD, Scherrer DE, Lanier MH, et al. Cholesterol oxidation products are sensitive and specific blood-based biomarkers for Niemann-Pick C1 disease. Sci Transl Med 2010;2:56ra81.
Jiang X, Sidhu R, Porter FD, et al. A sensitive and specific LC-MS/MS method for rapid diagnosis of Niemann-Pick C1 disease from human plasma. J Lipid Res 2011;52:1435–1445.
Stampfer M, Theiss S, Amraoui Y, et al. Niemann-Pick disease type C clinical database: cognitive and coordination deficits are early disease indicators. Orphanet J Rare Dis. 2013;8:35.
Castellana S, Mazza T. Congruency in the prediction of pathogenic missense mutations: state-of-the-art web-based tools. Brief Bioinform 2013;14:448–459.
Wassif CA, Cross JL, Iben J, et al. High incidence of unrecognized visceral/neurological late-onset Niemann-Pick disease, type C1, predicted by analysis of massively parallel sequencing data sets. Genet Med 2016;18:41–48.
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ. Basic local alignment search tool. J Mol Biol 1990;215:403–410.
Katoh K, Standley DM. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 2013;30:772–780.
Retief JD. Phylogenetic analysis using PHYLIP. Methods Mol Biol 2000;132:243–258.
Guindon S, Delsuc F, Dufayard JF, Gascuel O. Estimating maximum likelihood phylogenies with PhyML. Methods Mol Biol. 2009;537:113–37.
Finn RD, Bateman A, Clements J, et al. Pfam: the protein families database. Nucleic Acids Res 2014;42:D222–D230.
Adebali O, Ortega DR, Zhulin IB. CDvist: a webserver for identification and visualization of conserved domains in protein sequences. Bioinformatics 2015;31:1475–1477.
Omasits U, Ahrens CH, Müller S, Wollscheid B. Protter: interactive protein feature visualization and integration with experimental proteomic data. Bioinformatics 2014;30:884–886.
Kwon HJ, Abi-Mosleh L, Wang ML, et al. Structure of N-terminal domain of NPC1 reveals distinct subdomains for binding and transfer of cholesterol. Cell 2009;137:1213–1224.
Choi Y, Chan AP. PROVEAN web server: a tool to predict the functional effect of amino acid substitutions and indels. Bioinformatics 2015;31:2745–2747.
Acknowledgements
This work was supported in part by National Institutes of Health grant GM072295 (to I.B.Z.).
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Inforamtion
(ZIP 3636 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Adebali, O., Reznik, A., Ory, D. et al. Establishing the precise evolutionary history of a gene improves prediction of disease-causing missense mutations. Genet Med 18, 1029–1036 (2016). https://doi.org/10.1038/gim.2015.208
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2015.208