Abstract
Purpose:
The purpose of this study was to determine the molecular consequences of the variant c.3700 A>G in the cystic fibrosis transmembrane conductance regulator (CFTR) gene, a variant that has been predicted to cause a missense mutation in the CFTR protein (p.Ile1234Val).
Methods:
Clinical assays of CFTR function were performed, and genomic DNA from patients homozygous for c.3700 A>G and their family members was sequenced. Total RNA was extracted from epithelial cells of the patients, transcribed into complementary DNA, and sequenced. CFTR complementary DNA clones containing the missense mutation p.Ile1234Val or a truncated exon 19 (p.Ile1234_Arg1239del) were constructed and heterologously expressed to test CFTR protein synthesis and processing.
Results:
In vivo functional measurements revealed that the individuals homozygous for the variant c.3700 A>G exhibited defective CFTR function. We show that this mutation in exon 19 activates a cryptic donor splice site 18 bp upstream of the original donor splice site, resulting in deletion of six amino acids (r.3700_3717del; p.Ile1234_Arg1239del). This deletion, similar to p.Phe508del, causes a primary defect in folding and processing. Importantly, Lumacaftor (VX-809), currently in clinical trial for cystic fibrosis patients with the major cystic fibrosis–causing mutation, p.Phe508del, partially ameliorated the processing defect caused by p.Ile1234_Arg1239del.
Conclusion:
These studies highlight the need to verify molecular and clinical consequences of CFTR variants to define possible therapeutic strategies.
Genet Med 16 8, 625–632.
Similar content being viewed by others
Main
Cystic fibrosis (CF) is a common genetic disease caused by mutations in the cystic fibrosis transmembrane conductance regulator (CFTR/ABCC7) gene (NM_000492.3) that leads to reduction of CFTR anion channel function on the surface of fluid-transporting epithelial tissues, such as the airways, gastrointestinal tract, and sweat ducts.1,2 The clinical phenotype is variable, but severely affected individuals typically suffer from airway obstruction with recurrent episodes of inflammation, infection, and pancreatic insufficiency.3,4 Approximately 2,000 different variants in the CFTR gene have been reported in the CF Mutation Database,5 yet the molecular consequences of less than 10% of these variants are understood.6 The need to overcome this gap in knowledge is urgent, particularly given the recent success in developing drugs that target the basic defects caused by CFTR mutation. VX-770 (or Ivacaftor) has been approved in North America and Europe as a drug for patients bearing the relatively rare mutation: p.Gly551Asp.7,8 The p.Gly551Asp mutation causes defective channel gating,9,10,11 and VX-770 is thought to be effective in partially restoring lung function in patients with p.Gly551Asp because it enhances the channel activity of this mutant.12,13,14,15 The most common mutation in Europe and North America, p.Phe508del, has been studied extensively and has been found to impair CFTR protein folding during synthesis.16,17,18,19 Knowledge regarding the molecular defects caused by p.Phe508del has driven the development of targeted, interventional compounds, such as VX-809 (or Lumacaftor). Although VX-809 has shown promise in partially ameliorating the protein folding defect caused by p.Phe508del in vitro, it is yet to be determined if it will have clinical efficacy in combination with VX-770 in a phase III clinical trial.20,21,22 The development of these drugs highlights the therapeutic relevance of understanding the basic defects caused by mutation. Unfortunately, a small percentage of the documented CFTR variations have been interrogated with respect to their consequences on CFTR protein folding and/or function. Coupled with global excitement about the therapeutic potential of small molecules, including VX-809 and VX-770, there is a growing demand to determine whether patients with rare CFTR variants will be effectively treated with these new drugs.
A recent, large-scale study showed that individuals in North America and Europe bearing the rare variant c.3700 A>G, predicted to cause the missense mutation p.Ile1234Val or alternative splicing, exhibited variable CF disease severity.6 This variant, although rare in North America (present in only 15 patients, http://cftr2.org), is relatively common in the Middle East. Statistics from the World Health Organization (http://www.who.int/genomics/publications) report p.Ile1234Val as the second most common CF-causing mutation in the Middle East (12.3% occurrence in patients from Jordan, Kuwait, Lebanon, Oman, Qatar, Saudi Arabia, and United Arab Emirates), with the exception of two countries, Bahrain and Israel, in which the occurrence is less than 3.8 and 0.06%, respectively. The mutation 1548delG is most common in these seven countries (17.2%), whereas 2043delG is most common in Bahrain (30.8%) and W1282X is most common in Israel (36.1%). Furthermore, the c.3700 A>G (p.Ile1234Val) mutation is specific to Middle Eastern individuals originating from Bedouin tribes, and although diagnostic tests are sensitive enough to identify this mutation at an early age, currently there is no effective treatment for patients with this CF-causing genotype.23
Interestingly, Sosnay et al.6 recently showed that there were no functional consequences of introducing the missense mutation (p.Ile1234Val) in CFTR complementary DNA (cDNA) with respect to CFTR protein synthesis, processing, and/or function. Because these cell biological findings are discordant with the clinical phenotype and defective CFTR function in in vivo measurements, we were prompted to test the effect of this variant on splicing.
In this study, we show that the c.3700 A>G variant caused aberrant splicing leading to the in-frame deletion of six amino acids (p.Ile1234_Arg1239del) in a conserved region of the CFTR protein and that this deletion leads to its misfolding during synthesis, as in the case of p.Phe508del. These insights led us to test the efficacy of an investigational compound in trials for p.Phe508del and we found that, in cell culture, VX-809 (Lumacaftor) partially ameliorated the folding defect of p.Ile1234_Arg1239del-CFTR. Hence, these studies support the rationale for defining the molecular consequences of rare CFTR mutations in guiding decisions regarding future therapeutic interventions.
Materials and Methods
Clinical studies
The pancreas status was determined by measuring fecal elastase concentration in stool. The sweat chloride test was performed using standardized protocol (Wescor, Logan, UT). Nasal potential difference measurements allowed assessment of CFTR function as change (Δ) in potential difference following chloride-free and isoproterenol perfusion (ΔCl-free+Iso) in the presence of amiloride. Sweat gland potential difference was measured between a topically placed electrode and a subcutaneously inserted needle following sweat stimulation by pilocarpine iontophoresis. Finally, the β-adrenergic sweat secretion rate was assessed using an evaporimeter (CyberDerm RG-1; Dasylab, Broomall, PA) and following a sweat stimulation sequence of (i) cholinergic secretion with carbachol (0.01 mg; Alcon, Mississauga, ON, Canada); (ii) inhibition of cholinergic secretion with atropine (8.8 mg; Sandoz; Boucherville, QC, Canada); and (iii) pure β-adrenergic secretion with a cocktail containing 8.8 mg atropine, 4.4 mg isoproterenol hydrochloride (Sandoz), and 0.93 mg aminophylline (Hospira, Saint-Laurent, QC, Canada), as described previously.24
Genomic mutation analysis
Genomic DNA from peripheral leukocytes from the patient was analyzed by direct sequence analysis. All exons and flanking intron sequences of CFTR (NM_000492.3) were sequenced both in forward and reverse directions. Sanger sequencing was performed according to standard protocols using BigDye terminator v1.1 (Life Technologies, Carlsbad, CA), and sequencing products were separated on an ABI model 3730 Capillary Sequencer (Life Technologies) and analyzed using SeqPilot software (JSI Medical Systems, Kippenheim, Germany). Exons are numbered based on traditional numbering. Sequence nomenclature is based on the recommended nomenclature of the Human Genome Variation Society.
Reverse transcriptase–polymerase chain reaction analysis
Total RNA was extracted from nasal epithelial cells of the patients homozygous for c.3700 A>G using PerfectPure RNA cultured cell kit (5Prime, Hamburg, Germany) according to manufacturer’s protocol. RNA was then transcribed into cDNA using SuperScript first-strand synthesis system for reverse transcriptase–polymerase chain reaction (PCR; Invitrogen, Carlsbad, CA) using manufacturer’s protocol. Primers amplifying the entire CFTR transcript as a series of overlapping fragments were designed. Standard PCR conditions were used. Primer sequences are available on request.
Generation of mutant CFTR constructs
p.Ile1234Val-CFTR was generated in human wild-type (WT)-CFTR cDNA (pcDNA3.1) by Norclone Biotech Laboratories (London, ON, Canada). p.Ile1234_Arg1239del-CFTR was generated in human WT-CFTR cDNA (pcDNA3.1), using the KAPA HiFi HotStart PCR Kit (KAPA Biosystems, Woburn, MA). The deletion was generated using the following primers: 5′-CAT ATT AGA GAA CAT TTC CTT CTC AGT GGG CCT CTT GGG AAG AAC TGG ATC-3′ (sense); and 5′-GAT CCA GTT CTT CCC AAG AGG CCC ACT GAG AAG GAA ATG TTC TCT AAT ATG-3′ (antisense). Plasmid DNA was prepared using the GenElute Plasmid Miniprep Kit (Sigma, St Louis, MO), and the presence of the deletion, as well as the integrity of CFTR cDNA, was confirmed by DNA sequencing (TCAG, Toronto, ON, Canada).
Studies of CFTR protein processing and function
Human embryonic kidney cells were transiently transfected with WT-CFTR or p.Ile1234Val-CFTR using PolyFect Transfection Reagent, according to the manufacturer’s protocol (Qiagen, Venlo, The Netherlands). Baby hamster kidney (BHK) cells were transiently transfected with WT-CFTR or p.Ile1234_Arg1239del-CFTR using GenJet, according to the manufacturer’s protocol (SignaGen Laboratories, Rockville, MD). Human embryonic kidney cells transiently expressing CFTR proteins were maintained in Dulbecco’s modified Eagle’s medium (Wisent, St-Bruno, Quebec, Canada) supplemented with nonessential amino acids (Life Technologies) and 10% fetal bovine serum (Wisent) at 37 °C with 5% CO2 (HEPA incubator; Thermo Electron Corporation, Waltham, MA); BHK cells transiently expressing CFTR proteins were maintained in Dulbecco’s modified Eagle’s medium/F12 (Wisent) supplemented with 10% fetal bovine serum at 37 °C with 5% CO2. p.Phe508del-CFTR was stably expressed in BHK cells (obtained from Dr Lukacs at McGill University, Montreal, QC) and maintained in Dulbecco’s modified Eagle’s medium/F12 supplemented with 10% fetal bovine serum and 200 µg/ml methotrexate (Sigma) at 37 °C with 5% CO2.
Human embryonic kidney cells expressing WT-CFTR or p.Ile1234Val-CFTR and BHK cells expressing WT-CFTR, p.Phe508del-CFTR, or p.Ile1234_Arg1239del-CFTR were grown at 37 °C for 24 h and subsequently lysed in modified radioimmunoprecipitation assay buffer (50 mmol/l Tris–HCl, 150 mmol/l NaCl, 1 mmol/l ethylenediaminetetraacetic acid (pH 7.4), 0.2% (v/v) sodium dodecyl sulfate, and 0.1% (v/v) Triton X-100) containing a protease inhibitor cocktail (Roche, Indianapolis, IN) for 10 min, and the soluble fractions were analyzed by sodium dodecyl sulfate–polyacrylamide gel electrophoresis on 6% gels. After electrophoresis, proteins were transferred to nitrocellulose membranes and incubated in 5% (w/v) milk. CFTR bands were detected using the human CFTR-NBD2-specific25 (amino acids 1204–1211) murine monoclonal antibody 596 (1:30,000, University of North Carolina at Chapel Hill, Chapel Hill, NC) and horseradish peroxidase–conjugated goat anti-mouse IgG secondary antibody (1:5,000) and subsequently exposed to film for 0.5–5 min as required. Calnexin was used as a loading control and was detected using a calnexin-specific rabbit antibody (1:5,000, Sigma) and horseradish peroxidase–conjugated goat anti-rabbit IgG secondary antibody (1:5,000) and subsequently exposed to film for 0.5–5 min as required. Relative levels of CFTR proteins were quantitated by densitometry of immunoblots using ImageJ 1.42 Q software (National Institutes of Health, Bethesda, MD), and reported values were normalized to calnexin expression levels.
To evaluate protein glycosylation status, BHK cells expressing WT-CFTR, p.Phe508del-CFTR, or p.Ile1234_Arg1239del-CFTR were grown at 37 °C for 24 h and subsequently lysed in modified radioimmunoprecipitation assay buffer as described above. Lysates were treated with either endoglycosidase H or peptide-N-glycosidase F (both from New England Biolabs, Ipswich, MA) according to the manufacturer’s protocol. Immunoblots were obtained using the human CFTR-NBD2-specific murine monoclonal antibody 596 as described above.
To test function, human embryonic kidney cells overexpressing WT-CFTR or p.Ile1234Val-CFTR were grown in 12-well plates, and on formation of a monolayer, the cells were incubated overnight with 10 mmol/l of the halide-sensitive fluorophore 6-methoxy-N-(3-sulfopropyl)quinolinium (SPQ; Invitrogen Molecular Probes, Carlsbad, CA), at 37 °C and 5% CO2. The next day, the cells were washed three times with phosphate-buffered saline to make the extracellular fluid free of SPQ. Cells were then kept in chloride-containing physiological solution, Hank’s buffered saline solution. Fluorescence measurements were made using Gemini EM Fluorescence microplate reader (Molecular Devices, Sunnyvale, CA). Intracellular SPQ was excited at 350-nm wavelength and the emission was measured at 450 nm, reporting chloride concentration. After reading baseline fluorescence in presence of physiological solution, a chloride gradient was established via exchange of the extracellular solution with a chloride-free, nitrate-containing buffer (NaNO3: 136 mmol/l, KNO3: 3 mmol/l, Ca(NO3)2: 2 mmol/l, glucose: 11 mmol/l, 4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid (HEPES): 20 mmol/l (pH 7.2 and osmolarity 300 mOsm)). CFTR was stimulated using the cyclic adenosine monophosphate (cAMP) agonist forskolin (10 μmol/l). After reaching equilibrium for chloride flux, the extracellular solution was replaced with physiological solution containing chloride, allowing SPQ to quench. The fluorescence measurements are expressed as the change in fluorescence relative to the fluorescence measurement just before CFTR stimulation. Data are summarized as initial rate of change in relative fluorescence units in the first 4 min of stimulation. Each condition was repeated with at least four technical replicates on the same plate, and a total of at least three biological replicates were conducted per condition.
Data analysis
All data are represented as mean ± SE. Prism 4.0 software (GraphPad Software, San Diego, CA) was used for statistical analysis. Nonpaired Student’s t-tests, one-way analysis of variance, and two-way analysis of variance were conducted as appropriate, and P values less than 0.05 were considered significant. Each experiment was replicated at least three times.
Results
Siblings in a Qatari family homozygous for the CFTR variant c.3700 A>G exhibit clinical features of CF and lack of CFTR function in in vivo measurements
A 27-year-old man and his 15-year-old sister, both from Qatar and homozygous for the mutation c.3700 A>G (p.Ile1234Val), presented for further CF diagnostic evaluation. The man, patient 1, was diagnosed at 4 months of age and had experienced recurrent lung infection over the years. He was hospitalized twice, at the ages of 13 and 15 years, for acute pulmonary exacerbation. In addition, he was recently admitted at the age of 27 years for mild hemoptysis. Sputum cultures grew Staphylococcus aureus and Haemophilus influenzae repeatedly. More recently, his sputum grew Mycobacterium abcessus. His lung function was always within normal ranges. His pancreas function had been progressively worsening, as indicated by decreased fecal elastase levels (122 μg EL/g stool). His sweat chloride concentration was 90 mmol/l (CF diagnostic cutoff >60 mmol/l).26
The sister, patient 2, was diagnosed at 2 months of age due to family history, persistent cough, and failure to thrive. She showed a similar disease course and remained pancreatic sufficient. She had experienced multiple chest infections and had been hospitalized several times for more severe pulmonary exacerbations. Her sputum cultures grew S. aureus, H. influenza, and Streptococcus pneumoniae repeatedly. Her lung function also was maintained in the normal range. Her sweat chloride concentration was 74 mmol/l, greater than the CF diagnostic cutoff.26
Due to the discrepant findings between clinical presentation and the biological evaluation of the CFTR mutant c.3700 A>G, we decided to perform additional functional tests. Nasal potential difference measurements demonstrated a ΔCl-free+Iso result of 1.8 mV in patient 1 and −7.4 in patient 2, compatible with ΔCl-free+Iso values seen in CF ( Figure 1a ). However, the ΔCl-free+Iso result of patient 2 was close to our institutional borderline cutoff of −7.5 mV, suggesting presence of some residual CFTR function. The potential difference measured in the sweat glands was hyperpolarized, as demonstrated for patients with CF. However, it was more negative in patient 2 (−47.2 mV) compared with patient 1 (−35.8 mV), for whom it was in a more borderline range. The β-adrenergic sweat secretory test demonstrated complete absence of β-adrenergic sweat secretion for patients 1 and 2 ( Figure 1b ).
Qatari siblings with cystic fibrosis transmembrane conductance regulator ( CFTR ) variant c.3700 A>G (predicted to cause the CFTR missense mutation: p.Ile1234Val) exhibit loss of CFTR function in nasal potential difference (NPD) and sweat secretion assays. (a) The original NPD tracing demonstrates absence of any CFTR-mediated response in patient 1 (ΔCl-free+Iso = 1.8 mV) and a relatively borderline CFTR-mediated response (ΔCl-free+Iso = −7.4 mV) in patient 2. Amiloride (Amil.) inhibited the epithelial sodium channel (ENaC), thereby limiting sodium reabsorption and allowing for measurement of the CFTR-mediated chloride response. (b) Evaporimeter tracings from healthy controls (β-adrenergic (β) peak = cholinergic (Chol.) peak), obligate heterozygote (β peak = ½ Chol. peak), and cystic fibrosis (no β peak) are presented to illustrate range of responses with this assay. Tracings from patients 1 and 2 do not show any β-adrenergic response. Short-lived responses following injection of β-adrenergic drugs represent experimental artifact caused by necessary movement of the evaporimeter.
Determining the consequences of p.Ile1234Val on CFTR folding, processing, and function
The CFTR variant c.3700 A>G has been predicted to create a missense mutation (p.Ile1234Val). We expressed a missense mutant construct in mammalian cells and found that CFTR protein processing was similar to that of WT-CFTR in that the protein was expressed as both the Golgi-modified, complex glycosylated, band C (migrating as broad band of 170 kDa) as well as the endoplasmic reticulum–modified, core glycosylated, band B (migrating as a sharper band of 150 kDa) ( Figure 2a ). Function of the missense mutation as a cAMP-activated chloride channel was compared with that of the WT-CFTR using a fluorescence-based assay. The fluorescence intensity of SPQ is sensitive to halide concentrations and can be loaded into cells to monitor transmembrane chloride ion fluxes in response to agonists of cAMP, namely forskolin, and the phosphodiesterase inhibitor 3-isobutyl-1-methylxanthine (IBMX). In the presence of an outwardly directed chloride gradient, the addition of the above agonists caused an increase in SPQ fluorescence in cells expressing WT-CFTR ( Figure 2b ). Therefore, this mutation caused no change in the CFTR protein folding, processing, or function, showing that it does not account for the disease phenotype.
In vitro characterization of p.Ile1234Val missense mutation introduced into cystic fibrosis transmembrane conductance regulator (CFTR) complementary DNA and expressed in heterologous expression system. (a) Immunoblots show the processing of wild-type (WT)-CFTR and p.Ile1234Val-CFTR in human embryonic kidney (HEK) cells after 24 h at 37 °C. WT-CFTR and p.Ile1234Val-CFTR were expressed as both the Golgi-modified, complex glycosylated (mature) band C form (broad 170-kDa band) as well as the endoplasmic reticulum–modified, core glycosylated (immature), band B form (sharp 150-kDa band). Maturation (expressed as percentage of band C/(band B + band C)) of WT-CFTR and p.Ile1234Val-CFTR was quantified for three independent trials, and there was no significant difference between the two (P > 0.05). (b) Fluorescence-based anion flux assay in HEK cells show WT-CFTR and p.Ile1234Val-CFTR function after stimulation using a cAMP agonist (gray bar, forskolin, 10 μmol/l) or vehicle (dimethyl sulfoxide) alone (empty bar). Flux responses in the first 4 min after cAMP stimulation (reported as an initial rate of change in relative fluorescence units (RFU)) for WT-CFTR and p.Ile1234Val-CFTR were quantified for three independent trials (four technical replicates each trial), and there was no significant difference between the two (P > 0.05). Ctl, control; SPQ, 6-methoxy-N-(3-sulfopropyl)quinolinium.
Routine diagnostic testing by direct sequence analysis of genomic DNA from peripheral leukocytes revealed a homozygous sequence alteration (c.3700 A>G) in the patients with no additional known disease-causing mutations. Although sequence change was predicted to cause an isoleucine to valine substitution at position 1234, in silico splicing analysis suggested that the mutation might affect splicing of exon 19 to exon 20, with deletion of 18 bp (5′-ATA AGT CCT GGC CAG AGG-3′), coding for residues 1234–1239. To evaluate whether this sequence variant had a functional impact on splicing, we performed analysis of mRNA from nasal epithelial cells of the patients.
Identification of alternative splicing leading to the deletion of six amino acids (p.Ile1234_Arg1239del) in NBD2 of CFTR
Reverse transcriptase–PCR was performed using primers that amplified all exons in overlapping fragments of the CFTR transcript. One of the fragments containing exons 18–21 showed a slightly smaller size compared with that of a normal control visualized on an agarose gel ( Figure 3a , b ). Direct sequence analysis of the smaller-sized fragment generated from the patient revealed a deletion of the last 18 bp of exon 19, r.3700_3717del, with little or no WT transcript ( Figure 3c ). This indicates that the c.3700 A>G substitution identified in the genomic DNA results in aberrant splicing of the CFTR transcript, by activating a cryptic donor splice site 18 bp upstream of the original donor splice site ( Figure 3d ). The 18-bp deletion removes six amino acids (p.Ile1234_Arg1239del) in the second nucleotide-binding domain (NBD2) of the CFTR protein.
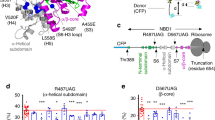
Aberrant splicing due to c.3700 A>G mutation in cystic fibrosis transmembrane conductance regulator ( CFTR ) predicts a deletion of six amino acids (p.Ile1234_Arg1239del). (a) Strategy for reverse transcriptase–polymerase chain reaction amplification of a region containing exon 19. (b) The amplified product visualized on an agarose gel. The expected size of 499 bp is seen in an unaffected control, whereas a slightly smaller band is seen in the patient. (c) Direct sequence analysis of the fragment in the patient revealed a deletion of the last 18 bp of exon 19. (d) A schematic overview of the genomic DNA with activation of a cryptic donor splice site in exon 19. The c.3700 A>G mutation in exon 19 activates a cryptic donor splice site 18 bp upstream of the original donor splice site. This results in a truncated exon 19 with a predicted deletion of six amino acids (r.3700_3717del; p.Ile1234_Arg1239del) in CFTR. mRNA, messenger RNA.
To determine the structural and functional consequences of this deletion, it was introduced in human WT-CFTR cDNA and expressed in mammalian cells. We found that similar to p.Phe508del, p.Ile1234_Arg1239del-CFTR is misprocessed and likely retained in the endoplasmic reticulum as it fails to express complex glycosylation ( Figure 4a ). However, the band C/(band B + band C) ratio, an indicator of CFTR processing, suggested that p.Ile1234_Arg1239del-CFTR is less severe than p.Phe508del because it had a significantly (P < 0.01) greater ratio ( Figure 4a ). Similar to p.Phe508del, the major form of p.Ile1234_Arg1239del-CFTR was found to be sensitive to endoglycosidase H, indicative of core glycosylation and retention in the endoplasmic reticulum ( Figure 4b ). However, similar to WT-CFTR, the heavier form of p.Ile1234_Arg1239del-CFTR was insensitive to endoglycosidase H yet sensitive to peptide-N-glycosidase F, indicative of complex glycosylation ( Figure 4b ). Further, we tested the effect of a therapeutic small molecule currently in clinical trials for p.Phe508del patients (a corrector compound, VX-809 or Lumacaftor) and found that it partially rescued the p.Ile1234_Arg1239del processing defect to a level comparable with that of p.Phe508del rescued by Lumacaftor (P < 0.01; Figure 4c ).
p.Ile1234_Arg1239del-CFTR is misprocessed, retained in the endoplasmic reticulum, and partially rescued by Lumacaftor (currently in clinical trials for p.Phe508del patients). (a) Immunoblots show steady-state levels of wild-type (WT)-CFTR, p.Phe508del-CFTR, and p.Ile1234_Arg1239del-CFTR in baby hamster kidney (BHK) cells after 24 h at 37 °C. WT-CFTR was expressed as both the Golgi-modified, complex glycosylated (mature) band C form (broad 170-kDa band) and the endoplasmic reticulum–modified, core glycosylated (immature), band B form (sharp 150-kDa band). p.Phe508del-CFTR and p.Ile1234_Arg1239del-CFTR were expressed primarily as the immature, band B form. Maturation (expressed as percentage of band C/(band B + band C)) of WT-CFTR, p.Phe508del-CFTR, and p.Ile1234_Arg1239del-CFTR was quantified for three independent trials, and a significant difference was found between the mutant proteins (P < 0.01). (b) To evaluate glycosylation status, immunoblots show the sensitivity of WT-CFTR, p.Phe508del-CFTR, and p.Ile1234_Arg1239del-CFTR to endoglycosidase H and peptide-N-glycosidase F. White arrowhead, complex glycosylated; black arrowhead, core glycosylated; gray arrowhead, deglycosylated. (c) Immunoblots show the processing of WT-CFTR, p.Phe508del-CFTR, and p.Ile1234_Arg1239del-CFTR in BHK cells in the absence (dimethyl sulfoxide) and presence of 3 µmol/l Lumacaftor (or VX-809) after 24 h at 37 °C. Equivalent sample loading was confirmed by immunoblots of calnexin protein expression. Maturation (expressed as percentage of band C/(band B + band C)) of p.Phe508del-CFTR and p.Ile1234_Arg1239del-CFTR was quantified for three independent trials, and there was a significant difference between untreated (dimethyl sulfoxide) and Lumacaftor (VX-809)-treated cultures, as indicated by asterisks (**P < 0.01). CFTR, cystic fibrosis transmembrane conductance regulator.
Discussion
This study highlights the importance of interrogating the consequences of CF-causing mutations using multiple research tools to develop rational therapeutic strategies. Adjunctive functional in vivo tests, i.e., nasal potential difference measurements and the novel β-adrenergic sweat secretion assay,24 confirmed that the variant c.3700 A>G led to reduced CFTR channel function on the apical membrane of these CF-affected tissues. This clinical phenotype is incompatible with the prediction that this variant caused the missense mutation, p.Ile1234Val. Cell-based studies of the predicted missense mutation introduced into CFTR cDNA showed that the predicted missense mutation, p.Ile1234Val, caused no apparent defects in protein folding, processing, or function. These findings prompted a detailed analysis of the entire CFTR gene and CFTR mRNA obtained from the nasal epithelium of a homozygous patient with the detection of aberrant splicing. We found that this variant caused aberrant splicing, with the deletion of six residues (p.Ile1234_Arg1239del) in a conserved region of the CFTR protein. Interestingly, deletion of this region of CFTR resulted in defective folding and processing, a protein-processing defect similar to that exhibited by p.Phe508del. This comparison motivated studies of Lumacaftor, a compound currently in clinical trial for patients with the p.Phe508del mutation and shown in cell culture to partially rescue the processing defect of p.Phe508del-CFTR. Importantly, a partial rescue was also induced by Lumacaftor of the p.Ile1234_Arg1239del-CFTR mutant in a heterologous expression system, and this finding motivates our future work to determine the extent of functional rescue that can be mediated by this compound in epithelial tissues derived from patients with this mutation.
The effect of p.Ile1234_Arg1239del on protein folding and processing is consistent with bioinformatics predictions
Examination of homology models and bioinformatics analyses predict that p.Ile1234_Arg1239del will cause significant conformational defects in the full-length CFTR protein. As shown in Supplementary Figure S1 online, residues 1234–1239 are modeled to constitute a β-turn between antiparallel β-strands (β2 and β3) in NBD2. Residues p.Ile1235_Gln1238 are integral to the putative β-turn structural motif as predicted by Chou and Fasman (based on 408 β-turns in 29 proteins).27 p.Ile1234_Arg1239 is N-terminal of NBD2 functional motifs (Walker A, residues 1244–1251; Q-loop, 1291–1297; signature C motif, 1346–1350; Walker B, 1366–1370; D-loop, 1374–1377; H-loop, 1401–1403), and therefore deletion of this sequence likely results in an altered and nonnative assembly of CFTR.
Interestingly, the CF Mutation Database5 contains several disease-associated mutations within the p.Ile1234_Arg1239 sequence and includes c.3705T>G (p.Ser1235Arg), c.3709G>A (p.Gly1237Ser), c.3713A>G (p.Gln1238Arg), c.3712C>T (p.Gln1238X), and c.3717G>C (p.Arg1239Ser). These mutations were each found in one or two individuals of European descent (i.e., Belgian, Spanish, French, and English), and the clinical presentation varied from a mild phenotype, in which the mutation was detected at 40 years of age (p.Gly1237Ser), with pancreatic sufficiency and forced expiratory volume in 1 s >70%, to a more severe CF phenotype (p.Gln1238X, in trans with p.Phe508del) that was diagnosed at birth and manifested with pancreatic insufficiency. These findings suggest that this region of NBD2 is mutation sensitive. Importantly, because the current studies show that the variant c.3700 A>G caused CF disease because it led to aberrant splicing and not the missense mutation predicted, we suggest that the molecular consequences for all of these variants should be interrogated in detail.
Future studies are required to understand the genotype–phenotype relationship
Both individuals exhibited pancreatic sufficiency at the time of the study, consistent with a relatively mild form of CF and possibly modest residual CFTR function. The capacity for patients bearing the mutation c.3700 A>G to exhibit residual CFTR function is yet to be rigorously tested. We tested the propensity for normal RNA splicing in one of the patients, as shown in Figure 3 . Only the aberrantly spliced CFTR mRNA was detected; hence, the mild pancreatic disease exhibited by the homozygous patients described in this study is unlikely to be conferred by modest expression of the normal transcript. This does not rule out the possibility that the propensity for normal RNA splicing may be different in pancreatic tissue or that other patients with this mutation may exhibit some disposition for normal RNA processing. In addition, modifier genes have been shown to be important contributors to lung and pancreatic disease.28,29 Polymorphisms in SLC26A9 have been shown to be associated with CF-related diabetes and meconium ileus.28,29 In our future work, it will also be important to determine the SLC26A9 genotype in a larger number of patients with the c.3700 A>G mutation in CFTR to examine the contribution of this modifier gene to exocrine pancreas function.
Currently, we are developing the methods necessary to measure the residual function in epithelial cultures from patients with this rare mutation. Whereas Ussing chamber studies of residual function conferred by p.Phe508del (or e.g. 3849+10kbC>T) in respiratory epithelia are possible using primary cultures derived from transplanted lungs, this is not possible for studies of the c.3700 A>G mutation because lung transplantation is not performed for CF patients in Qatar.30 There is optimism that recent advances in stem cell biology will enable the generation of epithelial tissues from patients with rare CFTR mutations and that such cultures will provide insight into consequences of rare CFTR mutation in epithelial transport function.31
In summary, these findings highlight the importance of experimentally verifying the clinical phenotype and the functional consequences of CF-causing variations because such studies will ultimately inform effective medical intervention. The discovery of aberrant splicing leading to deletion of six residues and the encouraging findings in cell culture studies that Lumacaftor (VX-809) partially rescues defective protein folding may have a significant impact on medical treatment of CF in the Middle East, where this mutation affects most patients. Furthermore, we predict that the anticipated consequences of other CF-causing mutations that have not yet been fully characterized in vitro may require investigation into the status of alternative splicing to more effectively treat CF disease in these patients.
Disclosure
The authors declare no conflicts of interest.
References
Riordan JR, Rommens JM, Kerem B, et al. Identification of the cystic fibrosis gene: cloning and characterization of complementary DNA. Science 1989;245:1066–1073.
Clarke LL, Grubb BR, Gabriel SE, Smithies O, Koller BH, Boucher RC . Defective epithelial chloride transport in a gene-targeted mouse model of cystic fibrosis. Science 1992;257:1125–1128.
Kerem E, Corey M, Kerem BS, et al. The relation between genotype and phenotype in cystic fibrosis–analysis of the most common mutation (delta F508). N Engl J Med 1990;323:1517–1522.
Ratjen F . Update in cystic fibrosis 2008. Am J Respir Crit Care Med 2009;179:445–448.
Cystic Fibrosis Mutation Database. http://www.genet.sickkids.on.ca. Accessed September 2013.
Sosnay PR, Siklosi KR, Van Goor F, et al. Defining the disease liability of variants in the cystic fibrosis transmembrane conductance regulator gene. Nat Genet 2013;45:1160–1167.
Davies JC, Wainwright CE, Canny GJ, et al.; VX08-770-103 (ENVISION) Study Group. Efficacy and safety of ivacaftor in patients aged 6 to 11 years with cystic fibrosis with a G551D mutation. Am J Respir Crit Care Med 2013;187:1219–1225.
Kotha K, Clancy JP . Ivacaftor treatment of cystic fibrosis patients with the G551D mutation: a review of the evidence. Ther Adv Respir Dis 2013;7:288–296.
Howell LD, Borchardt R, Cohn JA . ATP hydrolysis by a CFTR domain: pharmacology and effects of G551D mutation. Biochem Biophys Res Commun 2000;271:518–525.
Derand R, Bulteau-Pignoux L, Becq F . The cystic fibrosis mutation G551D alters the non-Michaelis-Menten behavior of the cystic fibrosis transmembrane conductance regulator (CFTR) channel and abolishes the inhibitory Genistein binding site. J Biol Chem 2002;277:35999–36004.
Bompadre SG, Sohma Y, Li M, Hwang TC . G551D and G1349D, two CF-associated mutations in the signature sequences of CFTR, exhibit distinct gating defects. J Gen Physiol 2007;129:285–298.
Serohijos AW, Hegedus T, Aleksandrov AA, et al. Phenylalanine-508 mediates a cytoplasmic-membrane domain contact in the CFTR 3D structure crucial to assembly and channel function. Proc Natl Acad Sci USA 2008;105:3256–3261.
Thibodeau PH, Richardson JM 3rd, Wang W, et al. The cystic fibrosis-causing mutation deltaF508 affects multiple steps in cystic fibrosis transmembrane conductance regulator biogenesis. J Biol Chem 2010;285:35825–35835.
Okiyoneda T, Veit G, Dekkers JF, et al. Mechanism-based corrector combination restores ?F508-CFTR folding and function. Nat Chem Biol 2013;9:444–454.
Van Goor F, Hadida S, Grootenhuis PD, et al. Correction of the F508del-CFTR protein processing defect in vitro by the investigational drug VX-809. Proc Natl Acad Sci USA 2011;108:18843–18848.
Ren HY, Grove DE, De La Rosa O, et al. VX-809 corrects folding defects in CFTR through action on membrane-spanning domain1 (MSD1). Mol Biol Cell 2013;24:3016–3024.
Clancy JP, Rowe SM, Accurso FJ, et al. Results of a phase IIa study of VX-809, an investigational CFTR corrector compound, in subjects with cystic fibrosis homozygous for the F508del-CFTR mutation. Thorax 2012;67:12–18.
Abdul Wahab A, Al Thani G, Dawod ST, Kambouris M, Al Hamed M . Heterogeneity of the cystic fibrosis phenotype in a large kindred family in Qatar with cystic fibrosis mutation (I1234V). J Trop Pediatr 2001;47:110–112.
Quinton P, Molyneux L, Ip W, et al. ß-adrenergic sweat secretion as a diagnostic test for cystic fibrosis. Am J Respir Crit Care Med 2012;186:732–739.
Cui L, Aleksandrov L, Chang XB, et al. Domain interdependence in the biosynthetic assembly of CFTR. J Mol Biol 2007;365:981–994.
Farrell PM, Rosenstein BJ, White TB, et al.; Cystic Fibrosis Foundation. Guidelines for diagnosis of cystic fibrosis in newborns through older adults: Cystic Fibrosis Foundation consensus report. J Pediatr 2008;153:S4–S14.
Chou PY, Fasman GD . Prediction of beta-turns. Biophys J 1979;26:367–383.
Blackman SM, Commander CW, Watson C, et al. Genetic modifiers of cystic fibrosis-related diabetes. Diabetes 2013;62:3627–3635.
Sun L, Rommens JM, Corvol H, et al. Multiple apical plasma membrane constituents are associated with susceptibility to meconium ileus in individuals with cystic fibrosis. Nat Genet 2012;44:562–569.
Chiba-Falek O, Parad RB, Kerem E, Kerem B . Variable levels of normal RNA in different fetal organs carrying a cystic fibrosis transmembrane conductance regulator splicing mutation. Am J Respir Crit Care Med 1999;159:1998–2002.
Wong AP, Bear CE, Chin S, et al. Directed differentiation of human pluripotent stem cells into mature airway epithelia expressing functional CFTR protein. Nat Biotechnol 2012;30:876–882.
Ramsey BW, Davies J, McElvaney NG, et al. A CFTR potentiator in patients with cystic fibrosis and the G551D mutation. N Engl J Med 2011;365:1663–1672.
Van Goor F, Hadida S, Grootenhuis PD, et al. Rescue of CF airway epithelial cell function in vitro by a CFTR potentiator, VX-770. Proc Natl Acad Sci USA 2009;106:18825–18830.
Eckford PD, Li C, Ramjeesingh M, Bear CE . Cystic fibrosis transmembrane conductance regulator (CFTR) potentiator VX-770 (ivacaftor) opens the defective channel gate of mutant CFTR in a phosphorylation-dependent but ATP-independent manner. J Biol Chem 2012;287:36639–36649.
Jih KY, Hwang TC . Vx-770 potentiates CFTR function by promoting decoupling between the gating cycle and ATP hydrolysis cycle. Proc Natl Acad Sci USA 2013;110:4404–4409.
Du K, Sharma M, Lukacs GL . The DeltaF508 cystic fibrosis mutation impairs domain-domain interactions and arrests post-translational folding of CFTR. Nat Struct Mol Biol 2005;12:17–25.
Acknowledgements
We thank the study participants for their contribution. The authors are also grateful to Jordan Dunitz (Division of Pulmonary, Allergy, Critical Care and Sleep Medicine, University of Minnesota, MN, USA) for his participation in procuring tissue. These studies were supported by operating grants to C.E.B. from the Canadian Institutes of Health Research (CIHR), Cystic Fibrosis Canada, and the Al Qamra Holding Group. T.G. was supported by a CIHR Investigator Award. P.N.R. was supported by an operating grant from Genome Canada. I.A.J. was supported by the Hamad Medical Corporation, Qatar National Research Fund, and the Al Qamra Holding Group. S.V.M. was supported by Peterborough K.M. Hunter and Ontario Graduate Studentships. The work concerning the CFTR protein and the mutant versions was partially funded by the Al Qamra Holding Group, located in Qatar, where this mutation is the major CF-causing mutation.
Author information
Authors and Affiliations
Corresponding author
Supplementary information
Supplementary Figure S1
(PDF 866 kb)
Rights and permissions
About this article
Cite this article
Molinski, S., Gonska, T., Huan, L. et al. Genetic, cell biological, and clinical interrogation of the CFTR mutation c.3700 A>G (p.Ile1234Val) informs strategies for future medical intervention. Genet Med 16, 625–632 (2014). https://doi.org/10.1038/gim.2014.4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2014.4
Keywords
This article is cited by
-
Approaching two decades of cystic fibrosis research in Qatar: a historical perspective and future directions
Multidisciplinary Respiratory Medicine (2019)
-
Nanomolar-potency ‘co-potentiator’ therapy for cystic fibrosis caused by a defined subset of minimal function CFTR mutants
Scientific Reports (2019)
-
An Overview of the Homozygous Cystic Fibrosis Transmembrane Conductance Regulator Mutation c.3700 A>G (p.Ile1234Val) in Qatar
Current Genetic Medicine Reports (2019)
-
Inhalational Anesthetics Induce Neuronal Protein Aggregation and Affect ER Trafficking
Scientific Reports (2018)