Abstract
Purpose:
Diagnostic exome sequencing was immediately successful in diagnosing patients in whom traditional technologies were uninformative. Herein, we provide the results from the first 500 probands referred to a clinical laboratory for diagnostic exome sequencing.
Methods:
Family-based exome sequencing included whole-exome sequencing followed by family inheritance−based model filtering, comprehensive medical review, familial cosegregation analysis, and analysis of novel genes.
Results:
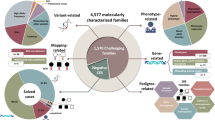
A positive or likely positive result in a characterized gene was identified in 30% of patients (152/500). A novel gene finding was identified in 7.5% of patients (31/416). The highest diagnostic rates were observed among patients with ataxia, multiple congenital anomalies, and epilepsy (44, 36, and 35%, respectively). Twenty-three percent of positive findings were within genes characterized within the past 2 years. The diagnostic rate was significantly higher among families undergoing a trio (37%) as compared with a singleton (21%) whole-exome testing strategy.
Conclusion:
Overall, we present results from the largest clinical cohort of diagnostic exome sequencing cases to date. These data demonstrate the utility of family-based exome sequencing and analysis to obtain the highest reported detection rate in an unselected clinical cohort, illustrating the utility of diagnostic exome sequencing as a transformative technology for the molecular diagnosis of genetic disease.
Genet Med 17 7, 578–586.
Similar content being viewed by others
Introduction
To date, ~7,000 rare diseases are known (National Institutes of Health Office of Rare Diseases Research), and each specific genetic disease is considered to be rare. Collectively, 25–30 million people in the United States are estimated to be affected by a rare genetic disorder and up to 25% of pediatric inpatient admissions are attributable to these diseases.1 As a whole, rare genetic disorders affect a significant number of individuals and have a great impact on the health system.2
Elucidation of the genetic basis of a rare Mendelian disease is quite challenging and the majority of patients remain undiagnosed despite extensive workup.3 The conventional diagnostic process used by most medical geneticists starts with recognition of specific phenotypic features and is generally paired with sequential laboratory testing, contingent on previous tests being uninformative. A recent retrospective analysis demonstrated that after being referred to a medical geneticist for diagnostic testing, less than half (46%) of patients with clinical and family histories consistent with a genetic etiology received a genetic diagnosis.3
Whole-exome sequencing (WES) provides a one-step simultaneous interrogation of virtually all exonic and adjacent intronic sequences and has been remarkably successful both in a diagnostic setting (diagnostic exome sequencing (DES)) and as a discovery tool (research exome sequencing),4 especially for disorders characterized by significant genetic heterogeneity.5 The diagnostic rate for clinical DES in unselected patients, who generally underwent exhaustive single-gene or gene-panel tests before DES, is reported to be ~25%.6,7
In this report, we analyzed 500 unselected consecutive cases that were referred to our laboratory for DES. The results demonstrate that DES is an integral tool for genetic diagnosis, especially for elucidating the molecular etiology of genetic diseases.
Materials and Methods
Terminology
Characterized gene. A gene known to be associated with a clinical phenotype based on the Human Gene Mutation Database (HGMD) or OMIM-Morbid database or the medical literature.
Novel gene. A gene that is not currently known to underlie a Mendelian genetic condition.
Trio. Three individuals from a family, consisting of the proband plus two first-degree relatives, most commonly the biological mother and father.
Whole-exome sequencing. Library capture and sequencing of virtually all the coding exons within the genome. WES includes the collective of research and DES.
Diagnostic exome sequencing. WES performed in the clinical setting in a Clinical Laboratory Improvement Amendments–certified diagnostic laboratory for single-patient diagnostic purposes.
Research exome sequencing. WES in the research setting. Research exome sequencing often entails a study design that includes multiple families with a similar phenotype and uses bioinformatics to narrow down a common disease gene.
Patients/study population
Patients were ascertained sequentially through clinical samples sent to Ambry Genetics Laboratory for DES beginning in September 2011. Clinicians were encouraged to refer all first-degree and affected second- or third-degree family members along with the proband for testing. Patient identifiers were removed. Solutions’ Institutional Review Board determined the study to be exempt from the Office for Human Research Protections Regulations for the Protection of Human Subjects (45 CFR 46) under category 4. Retrospective data analysis of anonymized data exempted the study from the requirement to receive consent from patients.
The clinical and test histories, along with a full pedigree provided by the referring physician(s), for each case were carefully summarized and tabulated by a molecular geneticist or a genetic counselor. The following data points were included: age of the proband at date of sample receipt; gender; consanguinity; previous clinical and/or differential diagnoses; a short clinical synopsis; family history; an 18-category organ system(s) involvement categorization; and whether the proband had intellectual disability, a positive brain magnetic resonance imaging result, multiple congenital anomalies, seizures/epilepsy, ataxia, autism spectrum disorder, or psychiatric disease ( Table 1 ).
Whole-exome sequencing
Genomic deoxyribonucleic acid was isolated from whole blood from all the probands and accompanying relatives. For many families, the whole exomes of the trio samples were prepared. Enrichment was performed using the SureSelect Target Enrichment System (Agilent Technologies, Santa Clara, CA) or SeqCap EZ VCRome 2.0 (Roche NimbleGen, Madison, WI).8
Characterized and novel gene databases
Internal gene databases were created based on two main classes of genes: characterized and novel (Supplementary Figure S1 online). The characterized and disease-causing (ChAD) gene database included genes that are associated with syndromes listed in the HGMD9 and the OMIM database. Modifier genes, disease association risk alleles (such as genes in which the only reported variants are common disease alleles that modify risk by less than twofold), and genes in which only somatic alterations are reported were excluded from the ChAD database. Novel genes were defined as those not known to underlie a Mendelian condition at the time of data analysis and were not included in the analysis of 84 cases based on specifications of the clinical order. Any RefSeq gene not included in the ChAD database was included in the novel gene database. The databases featured a dynamic design for shuffling genes between the ChAD and novel gene database in real time and were curated on a weekly basis to incorporate the latest discoveries.
Bioinformatics annotation, filtering of variants, and FIND
The sequence data were aligned to the reference human genome (GRCh37), and variant calls were generated using CASAVA (Consensus Assessment of Sequence And Variation, Illumina) and Pindel.10 The HGMD,9 the Single Nucleotide Polymorphism database,11 the 1000 Genomes Project,12 HapMap data,13 and online search engines (e.g., PubMed) were used to search for previously described gene mutations and polymorphisms. Data were annotated with the Ambry Variant Analyzer tool, including nucleotide and amino acid conservation, biochemical nature of amino acid substitutions, population frequency (Exome Variant Server (National Heart, Lung, and Blood Institute Grand Opportunity Exome Sequencing Project: http://evs.gs.washington.edu/EVS/) and the 1000 Genomes Project),12 and predicted functional impact (including PolyPhen14 and SIFT15 in silico prediction tools). Sequence alignments of the reads were viewed using IGV (Integrative Genomics Viewer) software.16
Stepwise filtering included the removal of common SNPs, intergenic and 3′/5′ UTR variants, non–splice related intronic variants, and synonymous variants (except those at the first and last nucleotide position of an exon) ( Figure 1 ). The filtering pipeline protected all variants annotated within the HGMD and/or the OMIM databases. Variants were then filtered further based on family history and possible inheritance models using the “Family History Inheritance-Based Detection” (FIND) bioinformatics program. FIND uses the affected status of each family member whose exome was sequenced and compares the proband’s genotype of each detected alteration to the genotype of the family members (most commonly parents). Alterations survive the filtering if they are consistent with Mendelian inheritance models and the affected status of each family member. A minimum of four inheritance models were executed for each family (autosomal dominant (AD), autosomal recessive (AR), X-linked recessive, X-linked dominant, and Y-linked in male probands). For increased sensitivity of potential phenocopies and genes associated with reduced penetrance, autosomal and X-linked “proband-only” models were also generated for each proband that captured heterozygous, homozygous, compound heterozygous, and hemizygous alterations irrespective of cosegregation based on family members’ genotypes.
Bioinformatics filtering and medical review of genes and alterations produced by exome sequencing. Bioinformatics and analysis variant classification schema. ACMG, American College of Medical Genetics and Genomics; AD, autosomal dominant; AR, autosomal recessive; ESP, NHLBI GO Exome Sequencing Project; HGMD, Human Gene Mutation Database; MAF, minor allele frequency; XLD, X-linked dominant; XLR, X-linked recessive; YL, Y-linked; 1000 Genomes, 1000 Genomes Project.
PRECISE and potentially causal alterations
Each candidate mutation was assessed by a molecular geneticist to identify the most likely causative mutation(s) using the “Personalized Medical Review with Enhanced and Comprehensive Assessment” (PRECISE) analysis method ( Figure 1 ). To further increase sensitivity despite potential phenocopies or genes associated with reduced penetrance, and for families in which there is uncertainty regarding the affected status among members of the trio, additional FIND models were run using alternative combinations of affected and unaffected designations. Stringent criteria were imposed to identify potential causative alleles. Each alteration was evaluated for its pathogenicity (alteration classification). The first level of PRECISE filtering removed alterations classified as polymorphisms or artifacts using population frequency in the Exome Variant Server and the 1000 Genomes Project,12 evolutionary conservation, and review of the medical literature. Classification of gene alterations followed Ambry’s clinical variant classification scheme (http://www.ambrygen.com/sites/default/files/Reclassification%20Chart.pdf). For rare missense changes, information regarding conservation of the corresponding amino acid and predictions of PolyPhen and SIFT15 were also used to aid in, but not determine, the interpretation of the variants. Each gene was then assessed for the level of phenotypic overlap with the proband and reported patients. Significant gene overlap with previously reported patients and consistent inheritance pattern and disease mechanism (gain versus loss of function) are required to classify alterations in known gene(s) as candidates. Data from animal models,17 protein function, gene family, corresponding pathways, and location in a known microdeletion syndrome locus were assessed to prioritize an alterations in a novel gene as a likely or possible molecular etiology. Characterized genes with significant or uncertain phenotypic overlap were considered “candidate genes.” At least one American Board of Medical Genetics–certified molecular geneticist and one genetic counselor and/or a second molecular geneticist performed independent reviews for each individual case.
Variant confirmation, cosegregation analysis, and parental confirmation
Most candidate alterations were confirmed using automated fluorescence dideoxy (“Sanger”) sequencing. For selected single-nucleotide substitutions detected from trio exome sequencing with high Q-score and read depth above laboratory-established confidence thresholds (coverage >40×, CASAVA Q score >80, and mutant ratio >35% for heterozygous calls), Sanger confirmation was not performed.18 Amplification primers were designed using PrimerZ.19 Sequencing was performed on an ABI3730 (Life Technologies, Carlsbad, CA) using standard procedures. Cosegregation Sanger analysis was performed when additional family members were available and segregation data would assist with interpretation. In addition, for apparent de novo alterations, short tandem repeat testing was performed to confirm parentage if trio exome data were not available and confirmatory of parentage.
Reporting of primary results
The primary report focused on deleterious mutations or variants of unknown significance in genes related to the current major clinical concern (Supplementary Figure S2 online). For characterized gene findings, we classified the overall DES results into four categories based on the combined assessment of the deleterious nature of the alteration and the level of phenotypic overlap among the proband and reported patients with alterations in that candidate gene: (i) positive: relevant alteration(s) detected; (ii) likely positive: relevant alteration(s) detected; (iii) uncertain: alteration(s) of uncertain clinical relevance detected; (iv) negative: no relevant alterations detected (Supplementary Figure S2 online). If either the phenotypic overlap or the deleterious nature of the alterations was classified as uncertain, then the overall DES result category was uncertain. Significant findings in clinically novel genes were interpreted as (i) likely positive: relevant alteration(s) detected or (ii) possibly positive: alteration(s) of uncertain clinical significance detected. In addition, we included “notable findings” in the primary report. These comprised gene alterations with at least minimal clinical overlap but that were unlikely to contribute to the proband’s indicated major clinical concern. Notable alterations in characterized genes were classified as such based on one or more of the following criteria: (i) patient’s overall phenotypic spectrum inconsistent with gene association; (ii) failure to cosegregate with affected status; (iii) one allele in an AR condition; and (iv) alteration is most likely benign based on frequency, conservation, and in silico predictions. Alterations in novel genes excluded after cosegregation analysis were also listed as “notable findings.”
Results
Demographics and major reasons for referral
Consistent with previous reports,6 the most frequent referral indication for DES was for pediatric-related neurological conditions (65%; Table 1 ). Sixty-four percent of probands had intellectual disability and/or developmental delay, 34% had a positive brain magnetic resonance imaging result, 28% had multiple congenital anomalies, and 24% had seizures and/or epilepsy.
Exome sequencing strategy
DES was performed on the proband only (141 patients), proband plus one first-degree relative (21 patients), or trio families (338 patients, including 288 parents–proband trios) (Supplementary Table S1 online). Mean coverage of the captured region was 107× per sample, with ~92% covered with at least 10×, an average of ~88% with base call quality of Q30 or greater, and an overall average mean quality score of approximately Q35 (Supplementary Table S2 online).
Rates of diagnosis
Overall, 30.4% (152/500) of patients undergoing DES had a positive gene finding in a characterized gene ( Table 2 ). Approximately one-quarter (130/500, 26.0%) received a definitive molecular diagnosis (Supplementary Table S3 online) and 4.2% (22/500) received a likely positive result with relevant alteration(s) detected in characterized genes (Supplementary Table S4 online). Among the 416 patients who underwent novel gene analysis, 31 (7.5%) were positive for a novel gene finding (data not shown). The overall positive rate among all gene types was 38.5% (160/416). Uncertain findings in characterized genes were found in 8.8% of probands (44/500). Approximately half of all patients had no relevant gene findings (215/416, 51.7%).
Inheritance patterns among positive findings
Among the 152 positive/likely positive cases, a total of 163 gene molecular defects were identified, including 86 AD (52.8%), 51 AR (31.3%), and 26 X-linked (16.0%) conditions ( Table 3 ). De novo alterations accounted for approximately half of all the identified gene defects (80/163; 49.1%). Among positive cases for which both parents’ specimens were available, de novo alterations in AD, X-linked recessive, and X-linked dominant diseases accounted for 90.7, 40.9, and 75.0% of the molecular defects, respectively. Among the 86 AD mutations, 60 arose de novo in the germ line, 5 probands inherited the mutation from a symptomatic parent, and 2 carried a mutation with indisputable pathogenicity in genes known to be associated with reduced penetrance and variable expressivity. Among the 26 probands with X-linked findings, 13 boys were hemizygous for a maternally inherited mutation and 1 female patient inherited the mutation from an affected mother.
Dual diagnoses
Among the 152 probands with a positive or likely positive finding in a characterized or novel gene, 23 (15.1%) had multiple gene findings (data not shown). Among these, 11 patients received a dual molecular diagnosis (Supplementary Table S5 online), in which the two significant findings are associated with nonoverlapping clinical presentations (patients 2, 4, 5, 6, 8, 9, and 10) or possibly both contribute to the major phenotypes (patients 1, 3, 7, and 11).
Mutation types among the 211 identified mutant alleles
The 152 positive/likely positive cases are associated with 214 mutant alleles (Supplementary Table S6 online), including 165 (77.1%) single-nucleotide substitutions (111 missense, 37 nonsense, and 17 affecting splicing), 45 (21.3%) small (<20 nt) deletion/insertion/indels (42 frameshift and 3 in-frame deletions), and 4 (1.9%) large deletion/indels (ranging in size from 41 nt to 102 nt, confirmed by Sanger sequencing). Approximately half (46.7%) of the 214 mutant alleles were truncating (nonsense, splicing, frameshift, or large deletions) and 53.3% were nontruncating (missense or in-frame deletions).
Genes with mutant findings within multiple patients
Mutations were identified in 141 different genes, 23 (16.3%) of which were found in two or more unrelated patients (i.e., “recurrent genes”; Table 4 ). Mutations in 16 genes were each identified in two patients. Mutations in five genes (ACTG2, FOXG1, IQSEC2, SMC1A, and DYNC1H1) were each observed in three patients, and mutations in two genes (ANKRD11 and KMT2A) were each observed in four patients. Some patients had recurrent clinical phenotypes with molecular defects in different genes. Seven patients presenting with hereditary spastic paraplegia and/or ataxia had mutations in six genes: ALS2, CYP7B1, FA2H, ITPR1 (de novo mutations detected in two patients), KCNC3, and SPAST. Similar to a previous report,6 mutations in Coffin–Siris syndrome (two for ARID1B and one for ARID1A) and Cornelia de Lange syndrome (three for SMC1A and one for SMC3) genes were also seen in multiple patients.
Recently characterized genes
Interestingly, 35 probands harbored mutations in genes discovered since January 2012 (23.2% of positives) (Supplementary Table S7 online). These included eight (5.3%) cases in which the gene was not yet listed in HGMD/OMIM Morbid at the time of analysis but was included in our internal ChAD gene database. An amended report was issued to reflect new gene discovery for four probands (2.6%).
Trio WES
Most patients opted for the reflexive testing option, which entailed exome sequencing of probands and analysis of characterized genes and, if negative, reflex to trio exome sequencing and analysis of all genes. This strategy posed an inherent bias toward positive findings among proband-only cases. However, this bias is removed when comparing trios versus proband-only test strategies among patients opting for nonreflexive testing. The positive rate observed among patients undergoing the trio test strategy was 37.3% (82/220), whereas the positive rate among the singleton test strategy was 20.6% (14/68), a statistically significant difference (P < 0.01) (Supplementary Table S8 online).
Discussion
Recent publications report a DES diagnostic rate plateau of ~25%.6,7 Herein, we report a diagnostic rate of 30.4% among relevant characterized genes. In addition, alterations in novel genes may provide a likely or possible explanation for the molecular etiology in 12% of patients with no significant findings in characterized genes. This rate is ~20% higher among characterized gene findings and 60% higher when considering novel gene findings as compared with those reported in similar clinical cohorts.6,7 The diagnostic rate for WES can depend on several factors, including overall gene coverage, bioinformatics filtering, and/or level of manual medical review. The difference in our detection rates could be attributed to several factors.
The use of family trio exome sequencing has been recommended to reduce the rate of uncertain findings.20 Our data demonstrate the increased diagnostic utility of a trio WES strategy. In addition, trio sequencing reduces analytic cost, highly prioritizes de novo changes (in sporadic cases), obviates the need for numerous low-throughput Sanger cosegregation analyses, and reduces overall turnaround time. Trio sequencing also adds to the clinical sensitivity with regard to the interpretation of clinically novel genes. Further, the diagnostic rate is improved by comprehensive, tailored, manual medical review that does not rely on preset phenotype-driven gene lists but rather on a carefully curated and frequently updated characterized gene/phenotype database and cross-checking by at least one certified molecular geneticist in addition to at least two reviews by molecular geneticists and/or genetic counselors. The resultant well-defined gene/alteration list after trio filtering based on family history enables the possibility of a thorough literature search for each gene prioritized, further minimizing the chance of missing a de novo mutation and/or positive finding in a newly discovered locus.
The utility of a “medical exome” that specifically targets ~4,600 medically relevant genes has recently been suggested as a possible alternative to the capture of all ~20,000 genes.20 The rate of new gene characterization is accelerating rapidly; in the past 6 years, the number of OMIM phenotypes for which the molecular basis is understood has doubled.4 That approximately one-quarter of our relevant gene findings were located within these newly characterized genes argues against the utility of gene panels or a “medical exome” with a predetermined list of genes. Further, the high ratio of positive findings in newly characterized genes underscores the importance of translating the most up-to-date research discoveries into clinical exome analysis to improve diagnostic sensitivity.
In addition to cost savings and time savings, the superiority of exome sequencing over single-gene tests and/or gene panels is exemplified by three main findings: (i) the observation of oligogenic findings and/or dual diagnoses in 7% of positive cases; (ii) novel gene findings in ~8% of patients; and (iii) a high ratio of positive findings among newly characterized genes.
The 11 patients with likely dual molecular diagnoses (7% positive cases), the 35 patients with alterations in recently characterized genes (23% of positive cases), and the 31 patients with novel gene findings (14% of positive cases) highlight the importance of DES in the context of complex phenotypes resulting from genes not yet characterized or from the combined/synergistic effect of two or more underlying pathogenic genetic alterations. In such cases, traditional models of single-gene testing would probably have proven unsuccessful in providing an accurate, comprehensive diagnosis.
The recurrent gene findings were most often observed among newly characterized genes. For example, there was only one report published in 2012 regarding the identification of de novo mutations in KMT2A (MLL) in five patients with Wiedemann–Steiner syndrome.21 In our first 500 unselected exome families, we detected mutations in this gene in four probands ( Table 4 ). Similar examples included recurrent positive findings in our cohort for the newly published genes PACS1 (identical mutation hot spot),22 TUBB4A,23 ACTG2,24 UBE3B,25,26 and GNAO127 ( Table 4 and Supplementary Table S7 online). The predominance of newly characterized genes among the recurrent gene findings is probably due to the lack of available clinical tests, either by single-gene analysis or by a panel test, for most of these new disease loci. Therefore, DES is the sole possible clinical approach for a definitive molecular diagnosis in these families. The reanalysis of previously reported DES cases based on new publications can also increase clinical sensitivity of the DES test. For example, we made a diagnosis for GNAO1-related epileptic encephalopathy on the same day the corresponding article was e-published in the American Journal of Human Genetics,27 and we then identified another de novo mutation in this gene in another patient through retrospective data mining.
Interestingly, the results also uncovered some unexpected modes of inheritance. The initial assumption for a rare genetic syndrome in a consanguineous family is that the disease is caused by a homozygous variant inherited through both parents. However, such an assumption may mislead the molecular diagnostic efforts and mask the real underlying inheritance pattern. In our cohort, a de novo missense mutation in TRPS1 was identified in a patient with a family history of consanguinity presenting with atypical trichorhinophalangeal syndrome.28 A de novo frameshift mutation in CHD7 was detected in a proband whose parents are first cousins, and the gene finding is consistent with his clinical presentation. In another family, the presence of two siblings of opposite genders, both with megacystis and echogenic bowel, led to a suspicion of AR inheritance of the disorder. DES performed on the proband−parents trio was unrevealing based on the AR model; however, the AD filter prioritized an apparently de novo novel change, c.770G>A (p.R257H), in the ACTG2 gene. Cosegregation analysis by targeted Sanger sequencing confirmed the DES findings and demonstrated that the affected sister also carried this heterozygous alteration. These results indicate both pathogenicity and a possible gonadal mosaic origin of this mutation (Tuzovic et al., personal communication). This broad spectrum of results illustrates how DES analysis based on all applicable inheritance patterns provides an unbiased scheme to pin down the real causative mutations.
Laboratories face a difficult dilemma when reporting DES diagnostic rates. Striving to increase the sensitivity of exome sequencing could be misconstrued as “overinterpretation.” In fact, a recent report shows that up to one-third of positive cases may be attributable to overinterpretation.20
Following a systematic and comprehensive variant classification scheme reduces this risk. However, the analysis of the phenotypic overlap among the patient and previously reported patients is more complicated. Despite this, it is difficult to argue against some level of contribution of a bona fide mutation in a well-characterized gene. The uncertainty arises with the more poorly described genes. However, our data add to the observations emerging from exome sequencing that the current understanding about the phenotypic spectrum of even well-described syndromes is expanding. This observation has implications for the interpretation of exome results and may argue against the notion of overinterpretation. Overall, a careful evaluation of both the alteration (“analytic validity”)29 and the overlap of the patient’s clinical features with those associated with the gene (“clinical validity”)29 both during DES analysis and after reporting is essential.
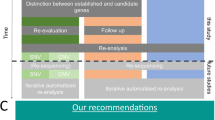
The rapid pace of new disease gene discovery, the significant ratio of positive findings in new disease genes, and the recurrence of many of the newly characterized loci illustrated in our cohort accentuate the importance of curating an up-to-date disease gene database. Our observations also highlight the utility of family-based exome sequencing, including trio exome sequencing, familial cosegregation analysis, inheritance-based model filtering, and comprehensive medical review. Furthermore, the data demonstrate the clinical sensitivity of DES over gene panels, especially with respect to conditions of genetic heterogeneity. Ongoing efforts to enhance the gene coverage of current DES platforms may further justify DES as the first-tier test.
Finally, the high diagnostic rate of DES clearly highlights the potential medical–economic savings to patients and insurance companies paying for testing. Almost 40% of patients analyzed in this cohort had a positive finding, more than doubling the rate of diagnosis achieved following a traditional molecular testing approach.3 Reaching a diagnosis results in an end to the expensive, time-consuming, and potentially invasive diagnostic odyssey that poses a heavy burden both for families and the health-care system.30,31,32 The high diagnostic rate of DES, the implication for patient care after a diagnosis, and the clear cost savings make exome sequencing well suited to become the standard of care in diagnostic medicine.
Disclosure
All of the authors are employed by and receive a salary from Ambry Genetics. Exome sequencing is among its commercially available tests.
References
Costa T, Scriver CR, Childs B . The effect of Mendelian disease on human health: a measurement. Am J Med Genet 1985;21:231–242.
Giampietro PF, Greenlee RT, McPherson E, Benetti LL, Berg RL, Wagner SF . Acute health events in adult patients with genetic disorders: the Marshfield Epidemiologic Study Area. Genet Med 2006;8:474–490.
Shashi V, McConkie-Rosell A, Rosell B, et al. The utility of the traditional medical genetics diagnostic evaluation in the context of next-generation sequencing for undiagnosed genetic disorders. Genet Med 2014;16:176–182.
Boycott KM, Vanstone MR, Bulman DE, MacKenzie AE . Rare-disease genetics in the era of next-generation sequencing: discovery to translation. Nat Rev Genet 2013;14:681–691.
Ku CS, Cooper DN, Polychronakos C, Naidoo N, Wu M, Soong R . Exome sequencing: dual role as a discovery and diagnostic tool. Ann Neurol 2012;71:5–14.
Yang Y, Muzny DM, Reid JG, et al. Clinical whole-exome sequencing for the diagnosis of mendelian disorders. N Engl J Med 2013;369:1502–1511.
Gahl WA, Markello TC, Toro C, et al. The National Institutes of Health Undiagnosed Diseases Program: insights into rare diseases. Genet Med 2012;14:51–59.
Gnirke A, Melnikov A, Maguire J, et al. Solution hybrid selection with ultra-long oligonucleotides for massively parallel targeted sequencing. Nat Biotechnol 2009;27:182–189.
Stenson PD, Mort M, Ball EV, et al. The Human Gene Mutation Database: 2008 update. Genome Med 2009;1:13.
Ye K, Schulz MH, Long Q, Apweiler R, Ning Z . Pindel: a pattern growth approach to detect break points of large deletions and medium sized insertions from paired-end short reads. Bioinformatics 2009;25:2865–2871.
Sherry ST, Ward MH, Kholodov M, et al. dbSNP: the NCBI database of genetic variation. Nucleic Acids Res 2001;29:308–311.
Abecasis GR, Altshuler D, Auton A, et al. A map of human genome variation from population-scale sequencing. Nature 2010;467:1061–1073.
International HapMap Consortium. The International HapMap Project. Nature 2003;426:789–796.
Adzhubei IA, Schmidt S, Peshkin L, et al. A method and server for predicting damaging missense mutations. Nat Methods 2010;7:248–249.
Kumar P, Henikoff S, Ng PC . Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat Protoc 2009;4:1073–1081.
Robinson JT, Thorvaldsdóttir H, Winckler W, et al. Integrative genomics viewer. Nat Biotechnol 2011;29:24–26.
Robinson PN, Kohler S, Oellrich A, et al. Improved exome prioritization of disease genes through cross-species phenotype comparison. Genome Res 2014;24:340–348.
Strom SP, Lee H, Das K, et al. Assessing the necessity of confirmatory testing for exome-sequencing results in a clinical molecular diagnostic laboratory. Genet Med 2014;16:510–515.
Tsai MF, Lin YJ, Cheng YC, et al. PrimerZ: streamlined primer design for promoters, exons and human SNPs. Nucleic Acids Res 2007;35(Web Server issue):W63–W65.
Atwal PS, Brennan ML, Cox R, et al. Clinical whole-exome sequencing: are we there yet? Genet Med 2014;16:717–719.
Jones WD, Dafou D, McEntagart M, et al. De novo mutations in MLL cause Wiedemann-Steiner syndrome. Am J Hum Genet 2012;91:358–364.
Schuurs-Hoeijmakers JH, Oh EC, Vissers LE, et al. Recurrent de novo mutations in PACS1 cause defective cranial-neural-crest migration and define a recognizable intellectual-disability syndrome. Am J Hum Genet 2012;91:1122–1127.
Simons C, Wolf NI, McNeil N, et al. A de novo mutation in the ß-tubulin gene TUBB4A results in the leukoencephalopathy hypomyelination with atrophy of the basal ganglia and cerebellum. Am J Hum Genet 2013;92:767–773.
Thorson W, Diaz-Horta O, Foster J 2nd, et al. De novo ACTG2 mutations cause congenital distended bladder, microcolon, and intestinal hypoperistalsis. Hum Genet 2014;133:737–742.
Basel-Vanagaite L, Dallapiccola B, Ramirez-Solis R, et al. Deficiency for the ubiquitin ligase UBE3B in a blepharophimosis-ptosis-intellectual-disability syndrome. Am J Hum Genet 2012;91:998–1010.
Basel-Vanagaite L, Yilmaz R, Tang S, et al. Expanding the clinical and mutational spectrum of Kaufman oculocerebrofacial syndrome with biallelic UBE3B mutations. Hum Genet 2014;133:939–949.
Nakamura K, Kodera H, Akita T, et al. De Novo mutations in GNAO1, encoding a Gao subunit of heterotrimeric G proteins, cause epileptic encephalopathy. Am J Hum Genet 2013;93:496–505.
Casci I, Accousti W, Lacassie Y . Unexpected exome sequencing result: De novo TRPS1 mutation in an infant with infantile scoliosis, mild developmental delay, and history of consanguinity. Am J Med Genet A 2014;164A:1334–1337.
Biesecker LG, Green RC . Diagnostic clinical genome and exome sequencing. N Engl J Med 2014;370:2418–2425.
Iglesias A, Anyane-Yeboa K, Wynn J, et al. The usefulness of whole-exome sequencing in routine clinical practice. Genet Med 2014; 16: 922–931.
Srivastava S, Cohen JS, Vernon H, et al. Clinical whole exome sequencing in child neurology practice. Ann Neurol 2014;76:473–483.
Beaulieu CL, Majewski J, Schwartzentruber J, et al.; FORGE Canada Consortium. FORGE Canada Consortium: outcomes of a 2-year national rare-disease gene-discovery project. Am J Hum Genet 2014;94:809–817.
The Reference Sequence (RefSeq) Project. RefSeq: The NCBI handbook [Internet]. National Library of Medicine, National Center for Biotechnology Information: Bethesda, MD. http://www.ncbi.nlm.nih.gov/refseq/.
Acknowledgements
We are grateful to the patients and their families for their participation, and to their physicians and genetic counselors for providing samples and clinical histories.
Author information
Authors and Affiliations
Corresponding authors
Supplementary information
Supplementary Information
(ZIP 171 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 3.0 Unported License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/3.0/
About this article
Cite this article
Farwell, K., Shahmirzadi, L., El-Khechen, D. et al. Enhanced utility of family-centered diagnostic exome sequencing with inheritance model–based analysis: results from 500 unselected families with undiagnosed genetic conditions. Genet Med 17, 578–586 (2015). https://doi.org/10.1038/gim.2014.154
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gim.2014.154