Abstract
Purpose: Usher syndrome is a major cause of genetic deafness and blindness. The hearing loss is usually congenital and the retinitis pigmentosa is progressive and first noticed in early childhood to the middle teenage years. Its frequency may be underestimated. Newly developed molecular technologies can detect the underlying gene mutation of this disorder early in life providing estimation of its prevalence in at risk pediatric populations and laying a foundation for its incorporation as an adjunct to newborn hearing screening programs.
Methods: A total of 133 children from two deaf and hard of hearing pediatric populations were genotyped first for GJB2/6 and, if negative, then for Usher syndrome. Children were scored as positive if the test revealed ≥1 pathogenic mutations in any Usher gene.
Results: Fifteen children carried pathogenic mutations in one of the Usher genes; the number of deaf and hard of hearing children carrying Usher syndrome mutations was 15/133 (11.3%). The population prevalence was estimated to be 1/6000.
Conclusion: Usher syndrome is more prevalent than has been reported before the genome project era. Early diagnosis of Usher syndrome has important positive implications for childhood safety, educational planning, genetic counseling, and treatment. The results demonstrate that DNA testing for Usher syndrome is feasible and may be a useful addition to newborn hearing screening programs.
Similar content being viewed by others
Main
Usher syndrome refers to a group of recessive disorders characterized by hearing loss (HL), retinitis pigmentosa (RP), and vestibular dysfunction. The three clinical classes are type 1 with congenital deafness and vestibular areflexia; type 2 with a congenital, moderate HL, and no vestibular areflexia; and type 3 with a progressive hearing and vestibular loss. All have RP with variable ages of diagnosis. Nine causative genes have so far been identified: MYO7A, USH1C, CDH23, PCDH15, and SANS (Usher type 1); USH2A, GPR98, and WHRN (Usher type 2); and USH3A (Usher type 3). Mutations in these genes show clinical variability that can range from nonsyndromic HL1–3 to isolated RP.4 The frequency has been reported to be ∼1/25,000 in the United States and Scandinavia.5–7 Studies of Schools for the Deaf have reached a consensus that ∼5% of such students have Usher syndrome.5,8–10 However, all the earlier studies sampled mostly severe and profoundly deaf children and would have overlooked those with milder HLs. Early surveys also were based on a clinical phenotype that can be difficult to diagnose in childhood. It has been generally accepted that deaf and hard of hearing (D/HOH) children had a high risk for RP.
MATERIALS AND METHODS
Subjects
High school students (ages 14–20 years) in special education classes for D/HOH children throughout Oregon were given kits that contained a questionnaire, consent forms, information on Usher syndrome, and a saliva collection kit. Nineteen schools were involved, including the Oregon school for the Deaf, from which 14 students were participants. Willing families provided written informed consent as well as assent. Samples, questionnaires, and signed consent forms were mailed to the Boys Town Hospital Gene Marker laboratory in Omaha, Nebraska, where the DNA was extracted. Students were encouraged to participate regardless of severity of loss, age at onset, laterality, or type of loss. Two hundred kits were distributed by the school teachers and nurses to children who took the kits to their families. Eighty-one were returned but three were discarded, two because the sample was insufficient, and one because a valid consent could not be obtained. Of the 78 individuals remaining, 37 reported a severe to profound HL, 18 reported a moderate to severe HL, three reported a mild HL, one reported a unilateral HL, and 19 did not specify the degree of HL. There were 41 men and 37 women. When compared with the Gallaudet report,11 this sample has an over representation of more severe losses. One hundred fifteen children receiving cochlear implants (CI) at the University of Iowa Implant Program were also involved. These children were all previously tested for GJB2/6 mutations.12 Forty-one (35.7%) were positive for GJB2/6 mutations. Of the remaining 74 individuals, DNA was available for 55. Human research institutional review boards at Boys Town National Research Hospital and the University of Iowa approved all the procedures.
All families and subjects received a written report of their findings. Families with children positive for the Usher screen were contacted by telephone and requested to come to the Casey Eye clinic for counseling and eye examination. Genetic counseling was available for all participants on request.
DNA extraction
The ORAGENE saliva sample collection kit (DNA Genotek, Inc., Ontario, Canada) was used to collect DNA in the Oregon study. DNA was extracted according to the manufacturer's instructions. For the CI children, 10 milliliters of whole blood were obtained by venipuncture and DNA was extracted using a Gentra PureGene DNA Extraction Kit (Qiagen, Valencia, CA) according to the manufacturer's instructions.
PCR and sequencing
GJB2 sequencing of exon 2 plus adjacent noncoding DNA including 5′ and 3′ splice sites was done on all subjects. Sequencing of the specific Usher exons was done to validate all pathogenic variants observed by microarray analysis. Gene specific sequencing was used to search for the second mutation in children where only one mutation had been detected by microarray analysis. Coding regions and splice sites of the GJB2, CDH23, MYO7A, USH1C, and USH2A genes were amplified using gene-specific primers. Deletions affecting GJB6 were screened as described previously.12 All amplification reactions were cycled using a standard protocol on a GeneMate Genius thermocycler (ISC BioExpress, Kaysville, UT). Sequencing was completed with a BigDye™ v3.1 Terminator Cycle Sequencing kit (Applied Biosystems, Foster City, CA), according to the manufacturer's instructions. Sequencing products were read using an ABI 3730 sequencer (Perkin Elmer, Waltham, MA). All sequencing chromatograms were compared to published cDNA sequence.
Microarray analysis
DNA samples negative for GJB2/6 mutations were subject to Usher syndrome testing (http://www.asperophthalmics.com) using Usher Chip v4.0 platform.13 All pathologic mutations were confirmed by standard Sanger sequencing. WHRN was the only Usher gene not represented on this version of the ASPER chip.
RESULTS
Six of 78 (7.7%) D/HOH children from Oregon were found to harbor mutations in the GJB2/6 gene. This is lower than that reported in the CI cohort reflecting the fact that the Oregon cohort had a wider range of HL severity. This was a surprising finding because we had felt that the presence of a pathologic genotype for GJB2/6 would significantly lower the likelihood of Usher syndrome in a high percentage of D/HOH children allowing us to avoid the cost of the Usher testing for them that at the time was more expensive.
Children negative for GJB2/6 HL (72 D/HOH and 55 CI) were then screened for known Usher mutations using the ASPER chip.13 The number of children carrying ≥1 Usher mutation among the D/HOH and CI children, negative for GJB2/6, was 8/72 (11.1%) and 7/55 (12.7%), respectively (Table 1). After the DNA testing was completed, it was revealed that only one Oregon student had been previously diagnosed with Usher syndrome. Frequency data from the two populations were not found to vary significantly from each other with regard to the frequency of Usher syndrome and were pooled with the combined frequencies being considered to extrapolate a frequency of Usher genotypes in the general population.
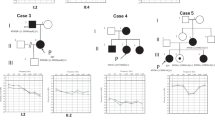
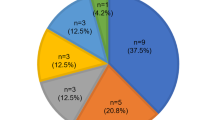
Mutations observed—mutations observed in this study and the genotypes of the cases detected are presented in Table 2. USH1D was observed to be the most frequent gene involved (6/15, 40%). Interestingly, one USH2A mutation (c.4338delCT) was detected in the CI cohort. Two children in the CI cohort were observed to be homozygous for USH1B: c. 448C>T, p.R150X but, because they were sibs, only one was counted in the prevalence estimate.
Frequency estimation—the frequency estimate needs to be adjusted for false negatives because of an imperfect test sensitivity and reduced specificity because of normal carriers. The sensitivity of the ASPER microarray was estimated by us in a separate sample of 77 United States individuals, each with a clinically established Usher phenotype and who had never been tested for Usher mutations before their ASPER testing. Of these, 30 were found to have ≥1 mutation in an Usher gene, giving a sensitivity of 30/77 = 0.39. The sensitivity of the ASPER chip has been observed to be between 35% and 55% in other studies13 (also L. Tranebjaerg, personal communication). To be conservative and thereby not overestimate the prevalence, our calculations assume a sensitivity of 0.5. The combined carrier rate of all Usher genetic types combined is assumed to be no >5% (14 and data not shown).
The adjusted prevalence of Usher syndrome in the D/HOH population can be calculated using the formula y = (1 − c) × x/s where x is the observed frequency, s is the sensitivity, and c is the carrier rate. The frequency for the combined groups, weighted by sample size, was calculated to be 9.2%. Thus, the frequency within the D/HOH childhood population is estimated at (1.0-0.05) × 0.092/0.5 = 0.175 (standard error ∼0.029; and the 95%tile lower limit is 0.117). Because a conservative estimate of the frequency of childhood deafness is ∼1/1000,15–17 one can extrapolate that the frequency of Usher syndrome in the general American population is 0.001 × 0.175, rounded up to ∼1/6000.
DISCUSSION
Prevalence
The estimate of the prevalence of Usher syndrome presented here is supported by other evidence. The frequency of detection observed here compares favorably with the results of two other studies of CI children where 7.0% and 10.4% were found to have RP by electroretinogram.18,19 Also, it is remarkable that, although of limited size, two different populations, one of children in a school system in Oregon with a wide range of HL severity and the other of children receiving CI in Iowa all of whom have a severe to profound HL, showed similar frequencies for the genetic types of Usher syndrome.
One must consider the possibility of selection bias, especially for the Oregon group, because parents chose to participate, a decision that may have been predicated on a perception of symptoms noted by the parents. There is less likelihood of such a self selection bias with the CI cohort.
The prevalence we have observed is nearly five times greater than what has been estimated from previous surveys. It rivals the frequencies seen for many other better known orphan diseases such as Duchenne muscular dystrophy(1/4,608),20 Huntington disease (∼1/15,000),21 and Lou Gehrig disease (1/40,000).22
Where are the extra Usher syndrome cases predicted by this study? Part of the answer is in the fact that not all persons with two pathogenic mutations develop RP, at least not as youngsters. There are reports of persons with mutations in MYO7A, CDH23, PCDH15, and USH1C who have been observed to have nonsyndromic HL.1–3,23,24 CDH23 is perhaps the largest category with about 1/3 reported as having nonsyndromic HL; however, many of these patients appeared to have mild retinal manifestations raising the question about a later onset type of RP.1 Another possible explanation for why Usher syndrome is underrepresented might be that people with Usher syndrome are not coming to the attention of physicians. Deaf, and especially deaf-blind, individuals face severe communication difficulties when visiting a doctor and might avoid doing so. Also, there is the possibility of under-diagnosis because of a lack of awareness by professionals of the high risk of Usher syndrome to D/HOH persons. The costs of genetic evaluations and of the sophisticated ophthalmologic examinations may deter some families from seeking a diagnosis. Finally, some Usher types may be associated with undiscovered morbidity that shortens lifespan.
These considerations point to a likelihood that the deaf-blind population is underserved in the United States. If the diagnoses are not being recorded, it implies that they are missed and that the patients are therefore not receiving the medical and social support they need. Other larger and better controlled genetic epidemiologic studies are needed to validate our results and, if confirmed, to search for answers regarding why the prevalence is much greater than previously believed.
The impact of new molecular technology
The molecular technology used to survey the D/HOH children in this study is at an early stage in its translation into the clinical realm but should be discussed in terms of the eventual implementation of this and similar technologies into newborn screening programs. ASPER (http://www.asperophthalmics.com) uses one type of platform that is widely used to test for Usher syndrome and has been applied to other genetic disorders as well.25 There are now competing technologies offered by SNPlex (https://products.appliedbiosystems.com) and Fluidigm (http://fluidigm.com/products) that have similar properties to ASPER. The cost is about the same for all, ranging from $150 to $500 per test. All function basically the same in that they detect known mutations but not novel DNA variants. For any particular gene, the maximum sensitivity of all the known-mutation based platforms should be about the same, 50–70%. Regardless of platform, the cost is attractively low and one can envision screening programs of high risk populations at a reasonable cost and with a reasonable pickup rate. It is anticipated that DNA-based diagnostics will gradually become an integral part of early childhood screening programs including those for newborn HL.26 Indeed, molecular diagnostics are being integrated into newborn screening for cystic fibrosis in much the same manner that it is proposed that it be used for childhood HL.27–29
DNA testing as an adjunct to newborn hearing screening
The sensitivity of any mutation-based screening test depends on the number of mutations being tested: the more mutations, the higher the sensitivity, but at a certain level, the addition of a new rare mutation adds little to sensitivity. Because each mutation adds some to the cost of a screening test, the balance of cost versus benefit needs to be considered as new mutations are added. Sensitivity can be greatly increased if the Usher genes can be sequenced using a probability-driven sequencing, resequencing, or one of the Next Generation Sequencing approaches.30–32 At the present time, these are fairly expensive and it is not practical to use them on a screening basis.
False positives are an issue with DNA testing of single gene recessive disorders. As a screening test, it detects children at a high risk for developing RP but follow-up is needed to confirm positive findings. First, if only one mutation is found, the second mutation must be sought to confirm the molecular diagnosis because there is a possibility that the child is just a carrier. The likelihood of an Usher carrier in the general population is about 5% and this can be taken as an approximate estimate of the false positive rate for the Usher screening test. It is unlikely, although not impossible, that any additional false positives would occur because of technical error because all results are confirmed by Sanger sequencing and many will have the family studied to confirm segregation. The possibility that the child may have a clinical variant that produces just an HL must be considered. In this sense, Usher screening is similar to other newborn screening in that it detects children at high risk for a serious symptom but follow-up confirmation is required. Consequently, the child must be examined by the ophthalmologist for RP. If the eye examination is negative, then yearly ophthalmologic follow-up would be advised. One should also be sure that the child is fully integrated into a pediatric otolaryngology service and that the family is aware of the availability of cochlear implantation. Genetic counseling is strongly recommended to aid in avoiding misinterpretation of the genetic findings.
In the context of most newborn hearing screening programs, the appropriate time for DNA screening is at the time of follow-up verification of the HL, which occurs between 3 and 12 months of age. At that time, a sample of blood from a heel stick can be taken and tested for genetic causes of HL. We propose a tiered approach as an adjunct to newborn hearing screening programs where DNA screening would be used to identify upward of 50% of the cases (that would include GJB2/6 and Usher gene mutations) with a simple and inexpensive DNA test. Those that are negative could go on to the more expensive second tier sequencing strategies, if funds are available. The first tier of this approach is diagrammed in Figure 1. As other genes are incorporated, the overall sensitivity will increase. A second tier would involve using probability-driven sequencing or Next Generation Sequencing technologies (as those become available and affordable), which could potentially bring the sensitivity up to close to 100% for all genetic types of childhood HL.
Proposed paradigm for screening D/HOH children for Usher syndrome. Testing for GJB2/6 will rule out Usher syndrome; if negative, testing continues for the most commonly occurring Usher mutations. The “Future Tests” box refers to the probable incorporation of tests for other genetic types of HL such as for the genes SLC26A4, OTOF, etc. The “confirm by sequencing” step should include a search for the second mutation when needed.
Benefit of early detection of Usher syndrome
For Usher syndrome, early diagnosis has many immediate and several long-term advantages to patients and their families. First, the election of cochlear implantation may be conditioned on the diagnosis of risk for future vision loss because combined sensory deficits are especially difficult for the patient, and the maintenance of hearing would be of great benefit if vision were lost. Not all profoundly deaf children below the age of 12 months receive CI in the United States.33,34 An Usher diagnosis would allow limited services to focus on children with potential Usher syndrome to insure that the parents have the opportunity to choose implantation at an early age. The choice of bilateral implants may be influenced by foreknowledge of a developing RP to enhance the ability for sound localization. Also, the late motor milestones that are observed in Usher 1 children with vestibular areflexia often leads to extensive tests (in addition to parental anxiety) to evaluate the reasons for the motor delay, and an early diagnosis of Usher syndrome provides a reason for such delay. Early habilitation and training programs need to take into account that Usher 1 children will need appropriate lighting. Knowledge that a child has Usher syndrome is so important to educators, that most schools for the deaf have at one time or another screened their student body for Usher syndrome. Genetic counseling is improved once a diagnosis of the etiology of the HL is understood, and parents may view a combined sensory deficit as having a different burden and such information may influence their family planning.
The greatest benefit of early identification lies in the potential for future treatment to delay the progression of the RP. Diet and lifestyle may be important determinants of the rate of progression of the retinal disease in Usher as was shown for adult onset macular degeneration.35 Animal studies raise possibility of using sunglasses in children suspected of having Usher syndrome to reduce the progression of the RP.36 There are also new advances in the area of neuroprotectives for the eye.37 Gene therapy for the eye has been effective for another retinal dystrophy, Leber's congenital Amaurosis38,39 and holds promise for the Usher syndromes.40,41 The principle behind early diagnosis is that early intervention would be the most effective. Many of the therapeutic possibilities, such as changes in the diet or the use of blue and UV light filtering glasses, are well suited to children and may prove beneficial in slowing the progression of the visual loss. If proven effective, such a simple therapy would give time for more effective gene-based therapies to be developed and to be proven safe for children. Although the progressive RP results in deafness and blindness in the adult, Usher syndrome is none-the-less a pediatric disorder since early diagnose and intervention would be predicted to be most effective in slowing the RP. If we wish to reduce the burden to individuals, families, and society, then we must focus our attention onto children and early diagnosis becomes paramount.
The purpose of this study was to determine the prevalence of Usher syndrome among D/HOH children and to demonstrate that DNA screening of D/HOH children is feasible. The results support the hypothesis that Usher syndrome is much more common than previously thought, although the reasons for this are unclear. Early diagnosis of Usher syndrome has important positive implications for childhood safety, educational attainment, and genetic counseling. Early diagnosis is now possible through DNA testing at a reasonable cost enabling large scale screening. Despite its importance as a public health issue, little attention has been paid to this disorder. Because it is so highly prevalent in the D/HOH child, we recommend that molecular testing should be part of the diagnostic workup of any D/HOH child as well as an adjunct to newborn hearing screening. The high frequency of Usher syndrome underscores the need for enhanced recognition of this disease as the most common cause of deafness and blindness in the United States.
References
Astuto LM, Bork JM, Weston MD, et al. CDH23 mutation and phenotype heterogeneity: a profile of 107 diverse families with Usher syndrome and nonsyndromic deafness. Am J Hum Genet 2002; 71: 262–275.
Ahmed ZM, Riazuddin S, Aye S, et al. Gene structure and mutant alleles of PCDH15: nonsyndromic deafness DFNB23 and type 1 Usher syndrome. Hum Genet 2008; 124: 215–223.
Ahmed ZM, Smith TN, Riazuddin S, et al. Nonsyndromic recessive deafness DFNB18 and Usher syndrome type IC are allelic mutations of USHIC. Hum Genet 2002; 110: 527–531.
Keats BJ, Savas S . Genetic heterogeneity in Usher syndrome. Am J Med Genet A 2004; 130: 13–16.
Boughman JA, Vernon M, Shaver KA . Usher syndrome: definition and estimate of prevalence from two high risk populations. J Chronic Dis 1983; 36: 595–603.
Sadeghi M, Kimberling WJ, Tranebjaerg L, Moller C . The prevalence of Usher syndrome in Sweden: a nationwide epidemiologic and clinical survey. Audiol Med 2004; 2: 220–228.
Nuutila A . Dystrophia retinae pigmentosa-dysacusis syndrome (DRD): a study of the Usher or Hallgren syndrome. J Genet Hum 1970; 18: 57–58.
Fraser GR . Epidemiology of profound childhood deafness. Audiology 1974; 13: 335–341.
Day CW . Current screening procedures for the Usher syndrome at residential schools for the deaf. Am Ann Deaf 1982; 127: 45–48.
Tamayo ML, Bernal JE, Tamayo G, Frias JL . A study of the etiology of deafness in an institutionalized population in Colombia (1992). Am J Med Genet 44: 405–408.
Gallaudet Research Institute Regional and National Summary Report of Data from the 2006-2007 Annual Survey of Deaf and Hard of Hearing Children and Youth. Washington, DC, Gallaudet University, 2007.
Snoeckx RL, Huygen PL, Feldmann D, et al. GJB2 mutations and degree of hearing loss: a multicenter study. Am J Hum Genet 2005; 77: 945–957.
Cremers FP, Kimberling WJ, Kulm M, et al. Development of a genotyping microarray for Usher syndrome. J Med Genet 2007; 44: 153–160.
Kimberling WJ . Estimation of the frequency of occult mutations for an autosomal recessive disease in the presence of genetic heterogeneity: application to genetic hearing loss disorders. Hum Mutat 2005; 26: 462–470.
Fortnum HM, Summerfield AQ, Marshall DH, Davis AC, Bamford JM . Prevalence of permanent childhood hearing impairment in the United Kingdom and implications for universal neonatal hearing screening: questionnaire based ascertainment study. BMJ 2001; 323: 536–540.
Van NK, Decoufle P, Caldwell K . Prevalence and characteristics of children with serious hearing impairment in metropolitan Atlanta, 1991-1993. Pediatrics 1999; 103: 570–575.
Maki-Torkko EM, Lindholm PK, Vayrynen MR, Leisti JT, Sorri MJ . Epidemiology of moderate to profound childhood hearing impairments in northern Finland. Any changes in ten years?. Scand Audiol 1998; 27: 95–103.
Mets MB, Young NM, Pass A, Lasky JB . Early diagnosis of Usher syndrome in children. Trans Am Ophthalmol Soc 2000; 98: 237–242.
Loundon N, Marlin S, Busquet D, et al. Usher syndrome and cochlear implantation. Otol Neurotol 2003; 24: 216–221.
Nigro G, Comi LI, Limongelli FM, et al. Prospective study of X-linked progressive muscular dystrophy in Campania. Muscle Nerve 1983; 6: 253–262.
Morrison PJ, Johnston WP, Nevin NC . The epidemiology of Huntington's disease in Northern Ireland. J Med Genet 1995; 32: 524–530.
Piemonte and Valle d'Aosta Register for Amyotrophic Lateral Sclerosis (PARALS) Incidence of ALS in Italy: evidence for a uniform frequency in Western countries. Neurology 2001; 56: 239–244.
Riazuddin S, Nazli S, Ahmed ZM, et al. Mutation spectrum of MYO7A and evaluation of a novel nonsyndromic deafness DFNB2 allele with residual function. Hum Mutat 2008; 29: 502–511.
Pennings RJ, Topsakal V, Astuto L, et al. Variable clinical features in patients with CDH23 mutations (USH1D-DFNB12). Otol Neurotol 2004; 25: 699–706.
Koenekoop RK, Lopez I, den Hollander AI, Allikmets R, Cremers FP . Genetic testing for retinal dystrophies and dysfunctions: benefits, dilemmas and solutions. Clin Experiment Ophthalmol 2007; 35: 473–485.
Levy HL, Albers S . Genetic screening of newborns. Annu Rev Genomics Hum Genet 2000; 1: 139–177.
Sontag MK, Hammond KB, Zielenski J, Wagener JS, Accurso FJ . Two-tiered immunoreactive trypsinogen-based newborn screening for cystic fibrosis in Colorado: screening efficacy and diagnostic outcomes. J Pediatr 2005; 147: S83–S88.
Lindau-Shepard BA, Pass KA . Newborn screening for cystic fibrosis by use of a multiplex immunoassay. Clin Chem 2010; 56: 445–450.
Wilcken B . Newborn screening for cystic fibrosis: techniques and strategies. J Inherit Metab Dis 2007; 30: 537–543.
Stone EM . Leber congenital amaurosis—a model for efficient genetic testing of heterogeneous disorders: LXIV Edward Jackson Memorial Lecture. Am J Ophthalmol 2007; 144: 791–811.
Voelkerding KV, Dames SA, Durtschi JD . Next-generation sequencing: from basic research to diagnostics. Clin Chem 2009; 55: 641–658.
Tucker T, Marra M, Friedman JM . Massively parallel sequencing: the next big thing in genetic medicine. Am J Hum Genet 2009; 85: 142–154.
Bradham T, Jones J . Cochlear implant candidacy in the United States: prevalence in children 12 months to 6 years of age. Int J Pediatr Otorhinolaryngol 2008; 72: 1023–1028.
Belzner KA, Seal BC . Children with cochlear implants: a review of demographics and communication outcomes. Am Ann Deaf 2009; 154: 311–333.
Chiu CJ, Klein R, Milton RC, Gensler G, Taylor A . Does eating particular diets alter the risk of age-related macular degeneration in users of the Age-Related Eye Disease Study supplements?. Br J Ophthalmol 2009; 93: 1241–1246.
Cideciyan AV, Jacobson SG, Aleman TS, et al. In vivo dynamics of retinal injury and repair in the rhodopsin mutant dog model of human retinitis pigmentosa. Proc Natl Acad Sci U S A 2005; 102: 5233–5238.
Sieving PA, Caruso RC, Tao W, et al. Ciliary neurotrophic factor (CNTF) for human retinal degeneration: phase I trial of CNTF delivered by encapsulated cell intraocular implants. Proc Natl Acad Sci U S A 2006; 103: 3896–3901.
Hauswirth WW, Aleman TS, Kaushal S, et al. Treatment of leber congenital amaurosis due to RPE65 mutations by ocular subretinal injection of adeno-associated virus gene vector: short-term results of a phase I trial. Hum Gene Ther 2008; 19: 979–990.
Cai X, Conley SM, Naash MI . RPE65: role in the visual cycle, human retinal disease, and gene therapy. Ophthalmic Genet 2009; 30: 57–62.
Williams DS . Usher syndrome: animal models, retinal function of Usher proteins, and prospects for gene therapy. Vision Res 2008; 48: 433–441.
Saihan Z, Webster AR, Luxon L, Bitner-Glindzicz M . Update on Usher syndrome. Curr Opin Neurol 2009; 22: 19–27.
Acknowledgements
Supported by grants from the Foundation Fighting Blindness, Hear See Hope Foundation, Howard Hughes Medical Institute, NIDCD Grant RO1 DCOO2842 and unrestricted Funds from Research to Prevent Blindness. Supported by a National Health and Medical Research Council (NHMRC) Postdoctoral Training Fellowship (to M.S.H.). Supported by a Doris Duke Clinical Research Fellowship (to A.E.S.).
Author information
Authors and Affiliations
Corresponding author
Additional information
Disclosure: The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Kimberling, W., Hildebrand, M., Shearer, A. et al. Frequency of Usher syndrome in two pediatric populations: Implications for genetic screening of deaf and hard of hearing children. Genet Med 12, 512–516 (2010). https://doi.org/10.1097/GIM.0b013e3181e5afb8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1097/GIM.0b013e3181e5afb8
Keywords
This article is cited by
-
Genetic insights into PHARC syndrome: identification of a novel frameshift mutation in ABHD12
BMC Medical Genomics (2023)
-
Adenylyl cyclase 6 plays a minor role in the mouse inner ear and retina
Scientific Reports (2023)
-
Hearing Loss with Vision Impairment: Usher Syndrome. A Case of the East Democratic Republic of Congo
Indian Journal of Otolaryngology and Head & Neck Surgery (2023)
-
Hereditäre Schwerhörigkeit
HNO (2023)
-
Novel mutations of the USH2A gene cause Usher syndrome in five Chinese families
BMC Ophthalmology (2022)