Abstract
One of the aims of clinical genetics is to identify gene mutations or genomic rearrangements that may underlie complex presentations of phenotypic features, such as multiple congenital malformations and mental retardation. During the decade after publication of the first article on array-based comparative genome hybridization, this technique has supplemented karyotyping as the prime genome-wide screening method in patients with idiopathic multiple congenital malformations and mental retardation. The use of this novel, discovery-based, approach has dramatically increased the detection rate of genomic imbalances. Array-based comparative genome hybridization detects copy number changes in the genome of patients and healthy subjects, some of which may represent phenotypically neutral copy number variations. This prompts the need for properly distinguishing between those copy number changes that may contribute to the clinical phenotype amid a pool of neutral copy number variations. We briefly review the characteristics of copy number changes in relation to their clinical relevance. Second, we discuss several published workflow schemes to identify copy number changes putatively contributing to the phenotype, and third, we propose a three-step procedure aiming to rapidly evaluate copy number changes on a case-by-case basis as to their potential contribution to the phenotype of patients with idiopathic multiple congenital malformations and mental retardation. This workflow is gene-centered and should aid in identification of disease-related candidate genes and in estimating the recurrence risk for the disorder in the family.
Similar content being viewed by others
Main
Clinical genetics investigates mutations underlying complex associations of phenotypic features, such as multiple congenital malformations and mental retardation (MCAMR). These conditions can be caused by a mutation in a specific gene (e.g., Lesch Nyhan syndrome) or by a genome rearrangement. The latter, including aneuploidies, aneusomies, and chromosome rearrangements, existing either in all cells of an individual or in mosaic form, are classically analyzed by karyotyping.1,2 During the past decades, techniques such as Fluorescence In Situ Hybridization (FISH) allowed detection of small aneuploid segments of a single or several chromosomes, which seem to underlie clinically recognizable conditions such as Wolf-Hirschhorn syndrome, Williams Beuren syndrome, Smith Magenis syndrome, Charcot-Marie-Tooth syndrome, DiGeorge syndrome, which are collectively termed Genomic Disorders.3 Detection of microdeletions and microduplications by FISH or by Multiplex Ligation-mediated Probe Amplification proved to be a major contribution to clinical genetic diagnosis.4–6
Both FISH and Multiplex Ligation-mediated Probe Amplification are limited to investigating only a few loci at a time. Applying probes on a support (e.g., a glass slide) allowed interrogating larger numbers of genomic loci simultaneously.7 Thus, array-based comparative genome hybridization (array-CGH) has, because of a much higher resolution than classical karyotyping, produced both a dramatically increased detection rate and additional discoveries of much smaller aneuploid segments in the human genome. These subsequently allowed identification of specific genes for syndromes such as CHARGE (coloboma, heart anomalies, chonal atresia, retardation, genital and ear anomalies),8 Peters Plus,9 del(17)(q21),10,11 Pitt-Hopkins,12 recurrent 1q21.1 rearrangements,13 and thrombocytopenia-absent-radius (TAR) syndrome14 (for a review, see Ref. 15). In addition, a large number of “unique” cases of de novo segmental aneuploidy have been identified (for reviews, see Refs. 2 and 16). The use of karyotyping and array-based segmental aneuploidy profiling as two distinct and complementary approaches to analyzing genome rearrangements has considerably widened our understanding of the mutations underlying congenital malformations and psychomotor retardation. However, novel problems have emerged and a revision of our approach to dealing with genome rearrangements and imbalances in patients with these conditions seems warranted.
ASSESSMENT OF THE CLINICAL SIGNIFICANCE OF COPY NUMBER CHANGES IS NOT STRAIGHTFORWARD
As outlined in Table 1, several mechanistic relationships between segmental aneuploidies and phenotypic features may exist and may occur simultaneously. This complicates straightforward implementation of array-based aneuploidy profiling in clinical genetics. Similar to karyotyping, array-based aneuploidy profiling, uncovers, in addition to pathogenic imbalances, apparently phenotypically neutral polymorphisms, albeit at a much higher rate.17–22 As of February 1, 2010, a compilation of data from 35 publications in the Database of Human Genomic Variants (DGV) lists 49,988 entries, comprising 29,133 copy number variants (CNVs) representing 8,410 CNV loci.23 A CNV is defined as a DNA segment longer than 1 kb with a variable copy number compared with a reference genome.21 Because array-based aneuploidy profiling measures differences in fluorescence intensities after hybridization to an array of bacterial artificial chromosomes or oligonucleotide probes of a patient DNA sample in comparison with a reference sample, these assays actually measure DNA copy number changes (CNCs). “Variants” are generally being considered as alterations or uncommon forms of no clinical significance. To account for this, and to avoid confusion, the term CNC, rather than CNV has been proposed.24,25 Detection of significant numbers of CNCs in healthy individuals has presented us with the challenge of distinguishing phenotypically neutral CNCs from those contributing to the patient's phenotype.21,22
SIZE AND GENE CONTENT OF CNCs
Some features of these CNCs, such as size, the number of protein-encoding genes involved, and prevalence in the general population, have been proposed as tools to differentiate among these possible interpretations. Vermeesch et al.26 and de Vries et al.27 suggested that polymorphisms may be smaller in size than pathogenic rearrangements. Thienpont et al.28 suggested that not mere size but rather the number of genes contained within a CNC may determine its plausibility as contributing to the phenotype of the patient. However, a single, critical gene in a small CNC may be sufficient to cause a specific disorder. When analyzing 278 patients and 48 healthy parents with a 3,783 bacterial artificial chromosomes-based array,29 we found that, in 20 patients, segmental aneuploidies have arisen de novo, ranging in size from 0.11 up to 8.16 Mb, whereas familial polymorphisms ranged between 0.22 and 4.70 Mb.30 By classical karyotyping, an appreciable number of phenotypically neutral transmitted unbalanced chromosomal rearrangements and euchromatic variants has been detected (for a compilation see, Barber, The Chromosome Anomaly Collection, http://www.ngrl.org.uk/wessex/collection/index.htm).17 In a recent study of ∼2,500 healthy subjects, Itsara et al.22 uncovered phenotypically neutral variants >500 kb in 5-10% and variants >1 Mb in 1-2% of individuals. Given this high degree of overlap in size and high frequency of occurrence in healthy individuals, pathogenic segmental aneuploidies and transmitted benign, familial, CNCs cannot be distinguished based on their size alone.22
PREVALENCE AND DE NOVO FORMATION RATES OF CNCs
Recent studies of microdeletions flanked by low copy repeats (LCRs) have revealed significant prevalences in affected and in healthy individuals.31–34 As a result, the relative risk for a phenotypic feature (e.g., epilepsy) conveyed by such a microdeletion ranges well below the level needed for a dominant negative effect.35,36 This further complicates clinical interpretation in case such a microdeletion has been detected. In studies of families with such cases, transmission of a segmental aneuploidy from a healthy parent is taken as not conveying a phenotypic effect.37–39 Thus, only segmental aneuploidies arising de novo in a sporadic patient will be taken into account. In this way, it is generally assumed that de novo formation of CNCs is a rare event. To date, a single genome-wide estimate of the rate of de novo formation of CNCs has been published.40 By Representational Oligonucleotide Microarray Analysis (ROMA), Sebat et al.40 found two de novo CNCs in a population of 196 healthy individuals. This amounts to a de novo generation rate of CNCs of ∼0.01. A different estimate has been derived from the de novo deletion and duplications rates of the Duchenne Muscular Dystropy (DMD) gene, which extrapolated to the entire genome would amount to ∼0.14 insertion/deletion events per generation.41,42 This estimate is indirect and does not take into account the highly variable rates of de novo aneuploidy formation at different loci. Recent measurements of the de novo deletion/duplication rate at four different loci of genomic disorders were directly assayed in single sperm cells. Frequencies of de novo rearrangements ranging from ∼6 × 10−5 for the hereditary neuropathy with liability to pressure palsies (HNPP)/Charcot-Marie-Tooth disease type 1A (CMT1A) locus to 2 × 10−6 at the Smith-Magenis syndrome locus were found.43 For the recurrent 22q11.21 deletion associated with velocardiofacial syndrome, the mutation rate was estimated to be as high as 2.5 × 10−4.44 All of these loci are known to undergo recurrent rearrangements by nonallelic homologous recombination (NAHR). However, the total number of these loci is not known. In addition, no de novo rearrangement rates have yet been determined for those which may have arisen by microhomology mediated processes.45 In contrast, the average de novo mutation rate for single nucleotide polymorphisms in the human genome is estimated to be 2.5 × 10−8 per nucleotide per generation.46 Thus, de novo structural rearrangements may occur at a higher frequency within the general population.47 Accordingly, the per-locus mutation rate for genomic rearrangements has been estimated at a 100- to 10,000-fold the rate of point mutations.48
DELETIONS REVEALING RECESSIVE MUTATIONS
A ramification of such a high frequency of de novo CNC formation is that CNCs involving losses of one or several protein-encoding genes lead much more frequently to a hemizygous mutation than do heterozygous (gene) mutations. In case a heterozygous gene mutation coincides with a loss of the allele on the trans chromosome, the carrier experiences a state equivalent to a compound heterozygous mutation. As has been pointed out,3,49 hemizygous losses may ”unmask” autosomal recessive mutations. This is probably a frequent, albeit underreported, mechanism of action of recessive mutations.50 In a patient with a hemizygous loss of the 4p16 region, such “unmasking” of an autosomal recessive mutation (in the WFS1 gene) on the structurally normal chromosome 4 has been shown to contribute to the complex clinical phenotype of this patient with a combined Wolfram- and Wolf-Hirschhorn syndrome.51 This means that patients harboring a deletion may have inherited a mutation in a phenotypically relevant gene within the deletion from their other parent, thus provoking an autosomal recessive disorder. In this way, the Hardy-Weinberg equilibrium, assumed to underlie allele frequencies in the general population, may get distorted. A systematic investigation of such a distortion and the resultant lower frequency of pathogenic gene-mutations in comparison with a putatively higher frequency of CNCs conveying hemizygous gene losses has yet to be undertaken.
SEGMENTAL ANEUPLOIDIES OVERLAPPING WITH KNOWN CNVs
A further complication in the process of distinguishing pathogenic and phenotypically neutral CNCs results from the finding that probably pathogenic, segmental aneuploidies in patients frequently encompass, or at least overlap with, one or even several CNCs that have also been detected in healthy individuals. This suggests that haploinsufficiency for a gene contained within a CNC is not necessarily pathogenic in patients with losses involving these CNCs. Recent studies of recurrent losses and gains flanked by segmental duplications in regions such as 1q21, 15q11.2, and 15q13.3 in cohorts of patients with epilepsy,13,31,34 schizophrenia,33,52 mental retardation,13,38,39 and autism spectrum disorders32,37 have shown a wide range of phenotypes, including “normal,” in carriers of these segmental aneuploidies. Interestingly, in families with a patient carrying a deletion in 15q13.3, the number of healthy individuals exceeds the number of patients by almost 3 to 1.38 To evaluate a putative contribution to the patient's phenotype, if any, criteria such as cosegregation with the clinical phenotype in the patient's family, should be taken into account.21
Considering only CNCs that emerged de novo and cosegregate with the clinical disorder in the pedigree of the patient may provide a rigorous, albeit conservative, approach to identifying genes that may be causally related to the phenotype of the patient under study.3,21,53 This does not necessarily mean that haploinsufficiency or a 3-fold gene dosage resulting from the CNC is the only pathogenic mutational mechanism underlying the phenotype of the patient. For instance, analyses of pedigrees of families with multiple patients with autism spectrum disorder suggest that unaffected carrier mothers may transmit a mutation to their sons who then manifest several aspects of this complex disorder up to a full blown autism.54 Recent studies of large cohorts of subjects with microdeletions in the 15q13.3 region have demonstrated such a mechanism in roughly half of the families.31,32,34 Because such a parent-of-origin effect may be involved in families in which affected children inherit a CNC, such a mechanism should also be considered.
POSITION EFFECTS OF SEGMENTAL ANEUPLOIDY
In addition, loss or gain of a stretch of DNA devoid of protein-encoding genes should not necessarily be discarded as phenotypically irrelevant.55–57 As has been shown recently in patients with Williams-Beuren syndrome, deletions may influence expression levels of the adjacent, nondeleted genes.58 Recently, Castermans et al.59 showed that the AMYSIN gene located at 300 kb from a breakpoint of a ring 14 became silenced and this may account for the autistic phenotype of the patient. These examples indicate that other mechanisms than mere gene copy number alteration, such as gene silencing, may be involved in determining clinical phenotypes and therefore should be taken into account during interpretation of results from array-based aneuploidy profiling.
It is conceivable, in particular with large, gene-rich, CNCs that several of the aforementioned mechanisms (Table 1) may operate simultaneously. This complicates interpretation of CNCs in clinical practice even more.
DISTINGUISHING POLYMORPHIC AND PATHOGENIC CNCs
The interpretation of CNCs in clinical genetics aims to answer two basic questions. First, does the detected aneuploidy (e.g., CNC) contribute to explaining the clinical phenotype of the patient? Second, what is the risk of recurrence for this CNC and the resulting clinical phenotype?
The aforementioned characteristics of CNCs obviously complicate the process of clinical interpretation. In 2007, Lee et al.21 listed a number of characteristics associated with pathogenic CNCs. Thus, a CNC inherited from an affected parent, or similar to one found in an affected relative, which does not overlap with a CNV listed in the DGV, which is rich in protein-encoding genes and does contain OMIM genes is more likely to be pathogenic than a CNC inherited from a healthy parent or that is similar to one found in a healthy relative. Trying to translate these characteristics into a workflow would require full knowledge of the CNC complement of both parents and, if available, multiple relatives. As discussed before, although a full family examination may be preferable, mandatory array-CGH of all relevant family members before the interpretational process (as recommended by Lee et al.21) may be costly and time-consuming. Thus, a workflow based on extensive family investigation may, however justifiable from a clinical genetic point of view, be difficult to achieve in daily practice.
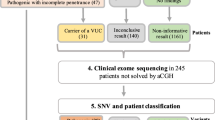
Recently, streamlined workflow schemes basing the evaluation of CNCs on their presence or absence in online databases have been proposed.60–65 These approaches all rely heavily on anonymous compilations of large population-based studies in which the investigated DNA donors were assumed to be healthy, although they have not been exhaustively phenotyped. Thus, these data sets may have been “contaminated” by CNCs that may have had a phenotypic impact. As discussed by Pinto et al.,66 compilations based on HapMap samples suffer from the limitation that no medical information has been obtained. In addition, the DNA for these samples has been isolated from Epstein Barr Virus-transformed lymphoblastoid cell lines, which may harbor transformed-induced genomic rearrangements.66,67 In addition, the boundaries of the CNVs in for instance the DGV have been mapped with array platforms with different levels of resolution.49 This makes it difficult to decide as to whether the CNC found in a given patient overlaps with a reported CNV. On the other hand, multiple appearances of the same CNV in multiple, unrelated healthy individuals decrease the likelihood that this CNV is associated with a significant deleterious phenotype. To avoid relying too heavily on such databases and to rapidly discern between putatively contributing CNCs and those that are likely to be phenotypically neutral, we propose a gene-centered approach (see also Fig. 1).
-
1
As a first step to evaluate each CNC, available databases (such as ECARUCA, DECIPHER65; for URLs, see Appendix) should be searched to determine whether the same CNC or one overlapping with the one under scrutiny has previously been found in a patient with a similar or overlapping phenotype. Although ECARUCA and DECIPHER appear the currently most useful databases, others (such as Genopedia/Phenopedia, KEGG, OMIM) may also be of help. Because genome-wide aneuploidy profiling is a relatively novel discovery-based approach a large number of “novel,” previously unreported CNCs are likely to continue to emerge in the near future. In addition, as compilation of data is an ongoing process, these databases are as yet not an exhaustive source of information. Another complication is the lack of consistency in descriptions of clinical phenotypes. Recently, a web-based tool has been developed aimed at a systematic and comprehensive phenotypic description.68 Therefore, an entirely “novel” (i.e., not yet described) CNC should not be discarded, but further interrogated as outlined below. For these cases, we need to proceed to step 2.
-
2
As a second step, previously not described CNCs should then be evaluated regarding their plausibility of contributing to the clinical phenotype. To do so, the functions and the tissue transcription patterns of any protein-encoding gene contained within a given CNC should be evaluated. In this step also, genes in the regions flanking the CNC that may be affected in their transcription levels should be considered.57 A systematic evaluation of strategies for gene-prioritization is currently not available yet within DECIPHER, a tool for prioritizing the likelihood that genes within the CNC may contribute to the phenotype has been incorporated.65 First priority should be given to genes that are known to be susceptible to haploinsufficiency and thus may explain (part) of the patient's phenotype. However, a gene involved in an autosomal recessive condition should not be discarded outright, because it may also explain the phenotype when the deletion unmasks a mutation on the other allele.51,69 Second, the temporal and spatial expression patterns of the gene should in principle be commensurate with the pathogenic process leading to the clinical phenotype. Thus, for a gene to contribute to a developmental disorder, it should be expressed during formation of the relevant tissue and organ. This criterion does not allow to discard or to prove a pathogenic role of a CNC, but merely evaluates its plausibility based on existing information on the genes within that CNC. In addition, CNCs containing genes that are not transcribed in tissues involved in the clinical phenotype (e.g., development of the heart in a patient with congenital heart disease) are less likely to contribute. However, such a CNC may contain an hitherto undiscovered microRNA that in the future be linked to a disease phenotype.70 For now, these CNCs are classified as “less likely to contribute” (see Fig. 1) and may be revisited at a later stage.
-
3
The third step investigates whether a CNC has either arisen de novo or cosegregates with an inherited phenotype in a family (van Daalen et al., unpublished data). In a sporadic patient, a de novo CNC is more likely to be causal than an inherited one. However, familial occurrence, i.e., inheritance from an apparently unaffected parent, does not necessarily imply that the CNC is unrelated to the phenotype.54 For instance, unmasking of a recessive mutation (see criterion 2, Refs. 51, 69), an alteration of the size of the imbalance,24 position effects,56–58 and subclinical, unrecognized manifestations in the carrier parent are still consistent with a pathogenic contribution of a given CNC. An additional complication is the recent description of highly variable phenotypes in subjects with microdeletions in regions such as 1q21.1 and 15q13.3.13,31 In case such CNCs were found in a patient and in healthy family members, we should continue searching for other potentially contributing CNCs or other mutations.53 In this way, recurrent microdeletions,31–39 which are currently viewed as CNVs of uncertain clinical significance, may in some cases be found to contribute to the patient's phenotype.
Sequential application of these criteria should, first, allow us to rapidly select CNCs that are most likely contributing to the clinical phenotype of the patient. Second, the number of costly and time-consuming follow-up studies, sometimes chasing familial polymorphisms,53 would be minimized. Third, the clinical interpretation for genetic counseling of the patient's aneuploidy profile can move ahead even before all relevant family members have been analyzed.
MECHANISMS OF ORIGIN OF CNCs AND DETERMINATION OF THEIR RECURRENCE RISK
Once the CNCs likely to contribute to the phenotype have been identified, an attempt can be made to answer the second core question in clinical genetics: what is the risk of recurrence for this mutation and the resulting clinical phenotype? The recurrence risk for a genomic rearrangement depends on whether the mutation has arisen de novo or is the result of a transmitted chromosomal structure predisposing to the formation of CNCs. A de novo interstitial or terminal loss or gain may arise through a homology driven nonallelic recombination event (NAHR), microhomology-driven processes such as FoSTeS, or nonhomologous end-joining (NHEJ) during repair of a double strand break.16,45,71,72 Rearrangements mediated by NAHR generally result in a recognizable syndrome such as the del(17)(q12) syndrome that presents with a combination of renal disease, diabetes, epilepsy, and characteristic facial dysmorphisms.73 This type of segmental aneuploidy represents the classical “genomic disorder” as defined by Lupksi et al.3,74
Most of the CNCs detected in genome screening studies seem to be more or less “unique” cases,2,16,30,75–76 with breakpoint regions that do not contain LCRs, clusters of olfactory receptor genes, or any of the other known chromosomal architectural features and thus have breakpoints not coinciding with other cases.2,30 Those most likely have arisen through nonhomologous end joining of spurious double strand breaks or through a microhomology-driven process.16,45,71 Therefore, their breakpoints cannot as clearly be defined as those underlying genomic disorders. The variability in size of the imbalances may explain why the phenotypes in these patients often are not readily clinically recognizable. In addition, given the incidental nature of spurious double strand breaks, their recurrence at exactly the same site is likely to be much lower than that of LCR-based Genomic Disorders. Patients with this kind of genomic rearrangements are often the most puzzling, because their phenotypes are highly complicated and variable.30,78,79
CNCs may also have resulted from a de novo or a familial reciprocal translocation giving rise to an unbalanced chromosome complement. The recurrence risk of the latter event can be determined unambiguously. A recent publication of such a familial translocation leading to a 5.0 Mb deletion of region 1q44 in a patient with a Dandy Walker variant and agenesis of the corpus callosum provides a case in point.80 The presence of this translocation indicates a recurrence risk for the same clinical constellation of 1 out of 4 and for carrying the translocation without clinical signs of 1 out of 2. Without knowledge of the translocation, the recurrence risk for segmental aneuploidy would have been estimated to be vanishingly small, whereas the second constellation would not even have been considered.80 A second point is that this particular translocation escaped routine karyotyping. It was detected only after array-CGH revealed a segmental loss of 1q44 that had prompted subsequent FISH probing. This, and many other, cases emphasize the continuing need for confirmation of the chromosomal structures underlying the imbalances by karyotyping or FISH.2,80
IMPACT OF A CNC DISCOVERY-BASED DIAGNOSTIC WORKUP ON GENETIC COUNSELLING
The majority of MCAMR patients carry genome imbalances that do not conform to the criteria for single gene-based syndromes or other well-circumscribed conditions such as recurrent Genomic Disorders resulting from the specific architecture of the human genome, which could have been detected by a targeted approach.3,11,81 Therefore, genome-wide aneuploidy profiling may be the most fruitful first step of genetic investigation in such patients.2,82 Precise breakpoint mapping using high-resolution array-based platforms and long range PCR will allow differentiating between cases with Genomic Disorders, which are flanked by LCRs, segmental duplications, or other known architectural features, and cases with “unique” breakpoints.45 Third, information regarding the structure of the genome rearrangement is essential because the recurrence risk for the rearrangement depends upon its structure as outlined above.2,80
The approach outlined above prompted us to invert the diagnostic workup of patients with idiopathic MCAMR. Instead of trying to subsume the patient's phenotype to a preconceived notion of either a (consistent) syndrome or a genome disorder, we should make use of high-resolution genome-wide screening techniques to detect novel CNCs, as the preferred initial genetic investigation in MCAMR patients.2,82 These CNCs need to be subsequently evaluated for their power to explain phenotypic features of the patients. Following the criteria outlined above may help to rapidly identify the CNCs contributing to the phenotype and thus allow us to fully garner the potential of this novel discovery-driven approach to clinical genetic diagnostics. These efforts need to be complemented by a systematic and comprehensive ontology of clinical phenotypes.68
The recent attempts to develop decision trees to determine the potential phenotypic contribution of novel CNCs,63–65 and the one outlined above, clearly indicate a need in the current era of genome-wide screening methods. This need may become even more pressing in view of the large amounts of data currently being generated in exome and whole genome sequencing efforts.83,84 The strategies adopted in the published decision trees are remarkably distinct. Although one may, on theoretical grounds, argue in favor or against certain approaches (see above), eventually only large, possibly multicenter studies, such as attempted by Buysse et al.,64 may allow to judge the relative merits and drawbacks of each individual approach. During such an evaluation, minimizing the number of CNCs or other mutations of “uncertain significance”64 may be a particularly important parameter for weighing the clinical utility of the respective strategies.
In summary, in the era of genome-wide aneuploidy screening, newly discovered CNCs may lead us to inverting the diagnostic approach to clinical genetic diagnosis in which novel classes of clinically relevant genomic imbalances will be detected and used to pinpoint genes more or less likely to contribute to the clinical phenotype. Integrating precise information regarding the breakpoint regions, and the structure of the rearrangement as determined by classical karyotyping and FISH investigation will allow us to determine the mutagenic mechanism. Subsequently, these data should be interpreted in terms of the patient's pathology and to eventually estimate its recurrence risk.
References
Hochstenbach R, Ploos van Amstel HK, Poot M . Microarray-based genome investigation: molecular karyotyping or segmental aneuploidy profiling?. Eur J Hum Genet 2006; 14: 262–265.
Hochstenbach R, van Binsbergen E, Engelen JJ, et al. Array analysis and karyotyping: workflow consequences based on a retrospective study of 36,325 patients with idiopathic developmental delay in the Netherlands. Eur J Med Genet 2009; 52: 161–169.
Lee JA, Lupski JR . Genomic rearrangements and gene copy-number alterations as a cause of nervous system disorders. Neuron 2006; 52: 103–121.
Ravnan JB, Tepperberg JH, Papenhausen P, et al. Subtelomere FISH analysis of 11,688 cases: an evaluation of the frequency and pattern of subtelomere rearrangements in individuals with developmental disabilities. J Med Genet 2006; 43: 478–489.
Rooms L, Reyniers E, Wuyts W, et al. Multiplex ligation-dependent probe amplification to detect subtelomeric rearrangements in routine diagnostics. Clin Genet 2006; 69: 58–64.
Kirchhoff M, Bisgaard AM, Bryndorf T, Gerdes T . MLPA analysis for a panel of syndromes with mental retardation reveals imbalances in 5.8% of patients with mental retardation and dysmorphic features, including duplications of the Sotos syndrome and Williams-Beuren syndrome regions. Eur J Med Genet 2007; 50: 33–42.
Pinkel D, Segraves R, Sudar D, et al. High resolution analysis of DNA copy number variation using comparative genomic hybridization to microarrays. Nat Genet 1998; 20: 207–211.
Vissers LE, van Ravenswaaij CM, Admiraal R, et al. Mutations in a new member of the chromodomain gene family cause CHARGE syndrome. Nat Genet 2004; 36: 955–957.
Lesnik Oberstein SA, Kriek M, White SJ, et al. Peters Plus syndrome is caused by mutations in B3GALTL, a putative glycosyltransferase. Am J Hum Genet 2006; 79: 562–566.
Koolen DA, Vissers LE, Pfundt R, et al. A new chromosome 17q21.31 microdeletion syndrome associated with a common inversion polymorphism. Nat Genet 2006; 38: 999–1001.
Sharp AJ, Hansen S, Selzer RR, et al. Discovery of previously unidentified genomic disorders from the duplication architecture of the human genome. Nat Genet 2006; 38: 1038–1042.
Zweier C, Peippo MM, Hoyer J, et al. Haploinsufficiency of TCF4 causes syndromal mental retardation with intermittent hyperventilation (Pitt-Hopkins syndrome). Am J Hum Genet 2007; 80: 994–1001.
Mefford HC, Sharp AJ, Baker C, et al. Recurrent rearrangements of chromosome 1q21.1 and variable pediatric phenotypes. N Engl J Med 2008; 359: 1685–1699.
Klopocki E, Schulze H, Strauss G, et al. Complex inheritance pattern resembling autosomal recessive inheritance involving a microdeletion in thrombocytopenia-absent radius syndrome. Am J Hum Genet 2007; 80: 232–240.
Slavotinek AM . Novel microdeletion syndromes detected by chromosome microarrays. Hum Genet 2008; 124: 1–17.
Stankiewicz P, Beaudet AL . Use of array CGH in the evaluation of dysmorphology, malformations, developmental delay, and idiopathic mental retardation. Curr Opin Genet Dev 2007; 17: 182–192.
Barber JC . Directly transmitted unbalanced chromosome abnormalities and euchromatic variants. J Med Genet 2005; 42: 609–629.
Eichler EE . Widening the spectrum of human genetic variation. Nat Genet 2006; 38: 9–11.
Freeman JL, Perry GH, Feuk L, et al. Copy number variation: new insights in genome diversity. Genome Res 2006; 16: 949–961.
McCarroll SA, Altshuler DM . Copy-number variation and association studies of human disease. Nat Genet 2007; 39: S37–S42.
Lee C, Iafrate AJ, Brothman AR . Copy number variations and clinical cytogenetic diagnosis of constitutional disorders. Nat Genet 2007; 39: S48–S54.
Itsara A, Cooper GM, Baker CS, et al. Population analysis of large copy number variants and hotspots of human genetic disease. Am J Hum Genet 2009; 84: 148–161.
Iafrate AJ, Feuk L, Rivera MN, et al. Detection of large-scale variation in the human genome. Nat Genet 2004; 36: 949–951.
South ST, Rope AF, Lamb AN, et al. Expansion in size of a terminal deletion: a paradigm shift for parental follow-up studies. J Med Genet 2008; 45: 391–395.
Curry CJ, Mao R, Aston E, et al. Homozygous deletions of a copy number change detected by array CGH: a new cause for mental retardation?. Am J Med Genet A 2008; 146A: 1903–1910.
Vermeesch JR, Fiegler H, de Leeuw N, et al. Guidelines for molecular karyotyping in constitutional genetic diagnosis. Eur J Hum Genet 2007; 15: 1105–1114.
de Vries BB, Pfundt R, Leisink M, et al. Diagnostic genome profiling in mental retardation. Am J Hum Genet 2005; 77: 606–616.
Thienpont B, Mertens L, de Ravel T, et al. Submicroscopic chromosomal imbalances detected by array-CGH are a frequent cause of congenital heart defects in selected patients. Eur Heart J 2007; 28: 2778–2784.
Vissers LE, de Vries BB, Osoegawa K, et al. Array-based comparative genomic hybridization for the genomewide detection of submicroscopic chromosomal abnormalities. Am J Hum Genet 2003; 73: 1261–1270.
Poot M, Eleveld MJ, van't Slot R, Ploos van Amstel HK, Hochstenbach R . Recurrent copy number changes in mentally retarded children harbor genes involved in cellular localization and the glutamate receptor complex. Eur J Hum Genet 2010; 18: 39–46.
Sharp AJ, Mefford HC, Li K, et al. A recurrent 15q13.3 microdeletion syndrome associated with mental retardation and seizures. Nat Genet 2008; 40: 322–328.
Miller DT, Shen Y, Weiss LA, et al. Microdeletion/duplication at 15q13.2q13.3 among individuals with features of autism and other neuropsychiatric disorders. J Med Genet 2009; 46: 242–248.
Stefansson H, Rujescu D, Cichon S, et al. Large recurrent microdeletions associated with schizophrenia. Nature 2008; 455: 232–236.
Helbig I, Mefford HC, Sharp AJ, et al. 15q13.3 microdeletions increase risk of idiopathic generalized epilepsy. Nat Genet 2009; 41: 160–162.
Dibbens LM, Mullen S, Helbig I, et al. Familial and sporadic 15q13.3 microdeletions in idiopathic generalized epilepsy: precedent for disorders with complex inheritance. Hum Mol Genet 2009; 18: 3626–3631.
de Kovel CG, Trucks H, Helbig I, et al. Recurrent microdeletions at 15q11.2 and 16p13.11 predispose to idiopathic generalized epilepsies. Brain 2010; 133: 23–32.
Depienne C, Moreno-De-Luca D, Heron D, et al. Screening for genomic rearrangements and methylation abnormalities of the 15q11-q13 region in autism spectrum disorders. Biol Psychiatry 2009; 66: 349–359.
van Bon BW, Mefford HC, Menten B, et al. Further delineation of the 15q13 microdeletion and duplication syndromes: a clinical spectrum varying from non-pathogenic to a severe outcome. J Med Genet 2009; 46: 511–523.
Doornbos M, Sikkema-Raddatz B, Ruijvenkamp CA, et al. Nine patients with a microdeletion 15q11.2 between breakpoints 1 and 2 of the Prader-Willi critical region, possibly associated with behavioural disturbances. Eur J Med Genet 2009; 52: 108–115.
Sebat J, Lakshmi B, Malhotra D, et al. Strong association of de novo copy number mutations with autism. Science 2007; 316: 445–449.
van Ommen GJ . Frequency of new copy number variation in humans. Nat Genet 2005; 37: 333–334.
White SJ, den Dunnen JT . Copy number variation in the genome; the human DMD gene as an example. Cytogenet Genome Res 2006; 115: 240–246.
Turner DJ, Miretti M, Rajan D, et al. Germline rates of de novo meiotic deletions and duplications causing several genomic disorders. Nat Genet 2008; 40: 90–95.
Wilson DI, Cross IE, Wren C, Scambler PJ, Burn J, Goodship J . Minimum prevalence of chromosome 22q11 deletions. Am J Hum Genet 1994; 55: A169.
Vissers LE, Bhatt SS, Janssen IM, et al. Rare pathogenic microdeletions and tandem duplications are microhomology-mediated and stimulated by local genomic architecture. Hum Mol Genet 2009; 18: 3579–3593.
Nachman MW, Crowell SL . Estimate of the mutation rate per nucleotide in humans. Genetics 2000; 156: 297–304.
Kidd JM, Cooper GM, Donahue WF, et al. Mapping and sequencing of structural variation from eight human genomes. Nature 2008; 453: 56–64.
Lupski JR . Genomic rearrangements and sporadic disease. Nat Genet 2007; 39: S43–S47.
Sharp AJ . Emerging themes and new challenges in defining the role of structural variation in human disease. Hum Mutat 2009; 30: 135–144.
Coman DJ, Gardner RJ . Deletions that reveal recessive genes. Eur J Hum Genet 2007; 15: 1103–1104.
Flipsen-Ten Berg K, van Hasselt PM, Eleveld MJ, et al. Unmasking of a hemizygous WFS1 gene mutation by a chromosome 4p deletion of 8.3 Mb in a patient with Wolf-Hirschhorn syndrome. Eur J Hum Genet 2007; 15: 1132–1138.
International Schizophrenia Consortium Rare chromosomal deletions and duplications increase risk of schizophrenia. Nature 2008; 455: 237–241.
Joziasse IC, van der Smagt JJ, Poot M, et al. A duplication including GATA4 does not co-segregate with congenital heart defects. Am J Med Genet A 2009; 149A: 1062–1066.
Zhao X, Leotta A, Kustanovich V, et al. A unified genetic theory for sporadic and inherited autism. Proc Natl Acad Sci USA 2007; 104: 12831–12836.
Beckmann JS, Estivill X, Antonarakis SE . Copy number variants and genetic traits: closer to the resolution of phenotypic to genotypic variability. Nat Rev Genet 2007; 8: 639–646.
Reymond A, Henrichsen CN, Harewood L, Merla G . Side effects of genome structural changes. Curr Opin Genet Dev 2007; 17: 381–386.
Henrichsen CN, Chaignat E, Reymond A . Copy number variants, diseases and gene expression. Hum Mol Genet 2009; 18: R1–R8.
Merla G, Howald C, Henrichsen CN, et al. Submicroscopic deletion in patients with Williams-Beuren syndrome influences expression levels of the nonhemizygous flanking genes. Am J Hum Genet 2006; 79: 332–341.
Castermans D, Thienpont B, Volders K, et al. Position effect leading to haploinsufficiency in a mosaic ring chromosome 14 in a boy with autism. Eur J Hum Genet 2008; 16: 1187–1192.
Edelmann L, Hirschhorn K . Clinical utility of array CGH for the detection of chromosomal imbalances associated with mental retardation and multiple congenital anomalies. Ann NY Acad Sci 2009; 1151: 157–166.
Gijsbers AC, Lew JY, Bosch CA, et al. A new diagnostic workflow for patients with mental retardation and/or multiple congenital abnormalities: test arrays first. Eur J Hum Genet 2009; 17: 1394–1402.
Koolen DA, Pfundt R, de Leeuw N, et al. Genomic microarrays in mental retardation: a practical workflow for diagnostic applications. Hum Mutat 2009; 30: 283–292.
Van Vooren S, Coessens B, Schoumans J, Robinson PN, de Ravel T . Cartagena satellite symposium: clinical application of copy number variation. Eur Cytogenetics Assoc Newsl 2009; 24: 30–33.
Buysse K, Delle Chiae B, Van Coster R, et al. Challenges for CNV interpretation in clinical molecular karyotyping: lessons learned from a 1001 sample experience. Eur J Med Genet 2009; 52: 398–403.
Firth HV, Richards SM, Bevan AP, et al. DECIPHER: database of chromosomal imbalance and phenotype in humans using ensembl resources. Am J Hum Genet 2009; 84: 524–533.
Pinto D, Marshall C, Feuk L, Scherer SW . Copy-number variation in control population cohorts. Hum Mol Genet 2007; 16: R168–R173.
Simon-Sanchez J, Scholz S, Fung HC, et al. Genome-wide SNP assay reveals structural genomic variation, extended homozygosity and cell-line induced alterations in normal individuals. Hum Mol Genet 2007; 16: 1–14.
Robinson PN, Köhler S, Bauer S, Seelow D, Horn D, Mundlos S . The human phenotype ontology: a tool for annotating and analyzing human hereditary disease. Am J Hum Genet 2008; 83: 610–615.
Breckpot J, Takiyama Y, Thienpont B, et al. A novel genomic disorder: a deletion of the SACS gene leading to spastic ataxia of Charlevoix-Saguenay. Eur J Hum Genet 2008; 16: 1050–1054.
Mencía A, Modamio-Høybjør S, Redshaw N, et al. Mutations in the seed region of human miR-96 are responsible for nonsyndromic progressive hearing loss. Nat Genet 2009; 41: 609–613.
Hastings PJ, Ira G, Lupski JR . A microhomology-mediated break-induced replication model for the origin of human copy number variation. PLoS Genet 2009; 5: e1000327.
Zhang F, Khajavi M, Connolly AM, Towne CF, Batish SD, Lupski JR . The DNA replication FoSTeS/MMBIR mechanism can generate genomic, genic and exonic complex rearrangements in humans. Nat Genet 2009; 41: 849–853.
Mefford HC, Clauin S, Sharp AJ, et al. Recurrent reciprocal genomic rearrangements of 17q12 are associated with renal disease, diabetes, and epilepsy. Am J Hum Genet 2007; 81: 1057–1069.
Murakami T, Garcia CA, Reiter LT, Lupski JR . Charcot-Marie-Tooth disease and related inherited neuropathies. Medicine (Baltimore) 1996; 75: 233–250.
Hoyer J, Dreweke A, Becker C, et al. Molecular karyotyping in patients with mental retardation using 100K single-nucleotide polymorphism arrays. J Med Genet 2007; 44: 629–636.
Fan YS, Jayakar P, Zhu H, et al. Detection of pathogenic gene copy number variations in patients with mental retardation by genomewide oligonucleotide array comparative genomic hybridization. Hum Mutat 2007; 28: 1124–1132.
Menten B, Maas N, Thienpont B, et al. Emerging patterns of cryptic chromosomal imbalance in patients with idiopathic mental retardation and multiple congenital anomalies: a new series of 140 patients and review of published reports. J Med Genet 2006; 43: 625–633.
Poot M, Beyer V, Schwaab I, et al. Disruption of CNTNAP2 and additional structural genome changes in a boy with speech delay and autism spectrum disorder. Neurogenetics 2010; 11: 81–89.
Poot M, van't Slot R, Leupert R, Beyer V, Passarge E, Haaf T . Three de novo losses and one insertion within a percentric inversion of chromosome 6 in a patient with complete absence of expressive speech and reduced pain perception. Eur J Med Genet 2009; 52: 27–30.
Poot M, Kroes HY, V D Wijst SE, et al. Dandy-Walker complex in a boy with a 5 Mb deletion of region 1q44 due to a paternal t(1;20)(q44;q13.33). Am J Med Genet A 2007; 143: 1038–1044.
Shaffer LG, Bejjani BA . Medical applications of array CGH and the transformation of clinical cytogenetics. Cytogenet Genome Res 2006; 115: 303–309.
Newman WG, Hamilton S, Ayres J, et al. Array comparative genomic hybridization for diagnosis of developmental delay: an exploratory cost-consequences analysis. Clin Genet 2007; 71: 254–259.
Choi M, Scholl UI, Ji W, et al. Genetic diagnosis by whole exome capture and massively parallel DNA sequencing. Proc Natl Acad Sci U S A 2009; 106: 19096–19101.
Ng SB, Buckingham KJ, Lee C, et al. Exome sequencing identifies the cause of a mendelian disorder. Nat Genet 2010; 42: 30–35.
Seidman JG, Seidman C . Transcription factor haploinsufficiency: when half a loaf is not enough. J Clin Invest 2002; 109: 451–455.
Yobb TM, Somerville MJ, Willatt L, et al. Microduplication and triplication of 22q11.2: a highly variable syndrome. Am J Hum Genet 2005; 76: 865–876.
Ullmann R, Turner G, Kirchhoff M, et al. Array CGH identifies reciprocal 16p13.1 duplications and deletions that predispose to autism and/or mental retardation. Hum Mutat 2007; 28: 674–682.
Horsthemke B, Wagstaff J . Mechanisms of imprinting of the Prader-Willi/Angelman region. Am J Med Genet A 2008; 146A: 2041–2052.
Antonarakis SE, Lyle R, Dermitzakis ET, Reymond A, Deutsch S . Chromosome 21 and down syndrome: from genomics to pathophysiology. Nat Rev Genet 2004; 5: 725–738.
Wolf NI, Sistermans EA, Cundall M, et al. Three or more copies of the proteolipid protein gene PLP1 cause severe Pelizaeus-Merzbacher disease. Brain 2005; 128( pt 4): 743–751.
Vandewalle J, Van Esch H, Govaerts K, et al. Dosage-dependent severity of the phenotype in patients with mental retardation due to a recurrent copy-number gain at Xq28 mediated by an unusual recombination. Am J Hum Genet 2009; 85: 809–822.
FitzPatrick DR . Transcriptional consequences of autosomal trisomy: primary gene dosage with complex downstream effects. Trends Genet 2005; 21: 249–253.
Regis S, Grossi S, Corsolini F, Biancheri R, Filocamo M . PLP1 gene duplication causes overexpression and alteration of the PLP/DM20 splicing balance in fibroblasts from Pelizaeus-Merzbacher disease patients. Biochim Biophys Acta 2009; 1792: 548–554.
Balikova I, Martens K, Melotte C et al. Autosomal-dominant microtia linked to five tandem copies of a copy-number-variable region at chromosome 4p16. Am J Hum Genet 2008 82: 181–187.
Acknowledgements
This research is in part supported by a grant from the Dutch Foundation for Brain Research (Hersenstichting) grant no. 2008(1) 34.
Author information
Authors and Affiliations
Corresponding author
Additional information
Disclosure: The authors declare no conflicts of interest.
APPENDIX: URLs OF THE DATABASES
APPENDIX: URLs OF THE DATABASES
The Chromosome Anomaly Collection, http://www.ngrl.org.uk/wessex/collection/index.htm; Database of Genomic Variants, http://projects.tcag.ca/variation/; DECIPHER, https://decipher.sanger.ac.uk/application/; ECARUCA, http://agserver01.azn.nl:8080/ecaruca/ecaruca.jsp; Genopedia, http://www.hugenavigator.net/HuGENavigator/startPagePedia.do; Human Phenotype Ontology, http://www.human-phenotype-ontology.org; KEGG (Kyoto Encyclopedia of Genes and Genomes), http://www.genome.jp/kegg/; OMIM (Online Mendelian Inheritance in Man), http://www.ncbi.nlm.nih.gov/omim; Phenopedia, http://www.hugenavigator.net/HuGENavigator/startPagePhenoPedia.do.
Rights and permissions
About this article
Cite this article
Poot, M., Hochstenbach, R. A three-step workflow procedure for the interpretation of array-based comparative genome hybridization results in patients with idiopathic mental retardation and congenital anomalies. Genet Med 12, 478–485 (2010). https://doi.org/10.1097/GIM.0b013e3181e3914a
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1097/GIM.0b013e3181e3914a