Abstract
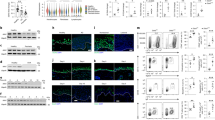
Two heterodimeric receptors consisting of interleukin (IL)-20R2 are shared by three of the IL-20 family of cytokines, IL-19, IL-20 and IL-24. Along with IL-22, these cytokines are downstream effectors of IL-23 and have been implicated in keratinocyte functions and the pathogenesis of psoriasis. Surprisingly, whereas knocking out either the IL-23 or IL-22 gene abolished imiquimod-induced psoriatic phenotypes in mice, similar attempt for IL-20R2 had little effect. Here, we report that the apparent disparity may result from a new IL-20R2 isoform encoded by an alternatively spliced transcript which survived the previous attempt for IL-20R2 gene knockout via the exon I deletion.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 6 digital issues and online access to articles
$119.00 per year
only $19.83 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Perera GK, Di Meglio P, Nestle FO . Psoriasis. Annu Rev Pathol 2012; 7: 385–422.
Di Cesare A, Di Meglio P, Nestle FO . The IL-23/Th17 axis in the immunopathogenesis of psoriasis. J Invest Dermatol 2009; 129: 1339–1350.
Ettehadi P, Greaves MW, Wallach D, Aderka D, Camp RD . Elevated tumour necrosis factor-alpha (TNF-alpha) biological activity in psoriatic skin lesions. Clin Exp Immunol 1994; 96: 146–151.
Zheng Y, Danilenko DM, Valdez P, Kasman I, Eastham-Anderson J, Wu J et al. Interleukin-22, a T(H)17 cytokine, mediates IL-23-induced dermal inflammation and acanthosis. Nature 2007; 445: 648–651.
van der Fits L, Mourits S, Voerman JS, Kant M, Boon L, Laman JD et al. Imiquimod-induced psoriasis-like skin inflammation in mice is mediated via the IL-23/IL-17 axis. J Immunol 2009; 182: 5836–5845.
Chan JR, Blumenschein W, Murphy E, Diveu C, Wiekowski M, Abbondanzo S et al. IL-23 stimulates epidermal hyperplasia via TNF and IL-20R2-dependent mechanisms with implications for psoriasis pathogenesis. J Exp Med 2006; 203: 2577–2587.
He M, Liang P . IL-24 transgenic mice: in vivo evidence of overlapping functions for IL-20, IL-22, and IL-24 in the epidermis. J Immunol 2010; 184: 1793–1798.
Sa SM, Valdez PA, Wu J, Jung K, Zhong F, Hall L et al. The effects of IL-20 subfamily cytokines on reconstituted human epidermis suggest potential roles in cutaneous innate defense and pathogenic adaptive immunity in psoriasis. J Immunol 2007; 178: 2229–2240.
Rutz S, Wang X, Ouyang W . The IL-20 subfamily of cytokines—from host defence to tissue homeostasis. Nat Rev Immunol 2014; 14: 783–795.
Blumberg H, Conklin D, Xu WF, Grossmann A, Brender T, Carollo S et al. Interleukin 20: discovery, receptor identification, and role in epidermal function. Cell 2001; 104: 9–19.
Wang M, Tan Z, Zhang R, Kotenko SV, Liang P . Interleukin 24 (MDA-7/MOB-5) signals through two heterodimeric receptors, IL-22R1/IL-20R2 and IL-20R1/IL-20R2. J Biol Chem 2002; 277: 7341–7347.
Wang M, Liang P . Interleukin-24 and its receptors. Immunology 2005; 114: 166–170.
Kotenko SV, Izotova LS, Mirochnitchenko OV, Esterova E, Dickensheets H, Donnelly RP et al. Identification of the functional interleukin-22 (IL-22) receptor complex: the IL-10R2 chain (IL-10Rbeta) is a common chain of both the IL-10 and IL-22 (IL-10-related T cell-derived inducible factor, IL-TIF) receptor complexes. J Biol Chem 2001; 276: 2725–2732.
Sheikh F, Baurin VV, Lewis-Antes A, Shah NK, Smirnov SV, Anantha S et al. Cutting edge: IL-26 signals through a novel receptor complex composed of IL-20 receptor 1 and IL-10 receptor 2. J Immunol 2004; 172: 2006–2010.
Ouyang W, Rutz S, Crellin NK, Valdez PA, Hymowitz SG . Regulation and functions of the IL-10 family of cytokines in inflammation and disease. Annu Rev Immunol 2011; 29: 71–109.
Yang XO, Chang SH, Park H, Nurieva R, Shah B, Acero L et al. Regulation of inflammatory responses by IL-17 F. J Exp Med 2008; 205: 1063–1075.
Wiekowski MT, Leach MW, Evans EW, Sullivan L, Chen SC, Vassileva G et al. Ubiquitous transgenic expression of the IL-23 subunit p19 induces multiorgan inflammation, runting, infertility, and premature death. J Immunol 2001; 166: 7563–7570.
Wolk K, Haugen HS, Xu W, Witte E, Waggie K, Anderson M et al. IL-22 and IL-20 are key mediators of the epidermal alterations in psoriasis while IL-17 and IFN-gamma are not. J Mol Med 2009; 87: 523–536.
Zhang R, Tan Z, Liang P . Identification of a novel ligand-receptor pair constitutively activated by ras oncogenes. J Biol Chem 2000; 275: 24436–24443.
Van Belle AB, de Heusch M, Lemaire MM, Hendrickx E, Warnier G, Dunussi-Joannopoulos K et al. IL-22 is required for imiquimod-induced psoriasiform skin inflammation in mice. J Immunol 2012; 188: 462–469.
Wahl C, Muller W, Leithauser F, Adler G, Oswald F, Reimann J et al. IL-20 receptor 2 signaling down-regulates antigen-specific T cell responses. J Immunol 2009; 182: 802–810.
Ueyama A, Yamamoto M, Tsujii K, Furue Y, Imura C, Shichijo M et al. Mechanism of pathogenesis of imiquimod-induced skin inflammation in the mouse: a role for interferon-alpha in dendritic cell activation by imiquimod. J Dermatol 2014; 41: 135–143.
Cai Y, Fleming C, Yan J . New insights of T cells in the pathogenesis of psoriasis. Cell Mol Immunol 2012; 9: 302–309.
Gottlieb AB, Krueger JG, Sandberg Lundblad M, Gothberg M, Skolnick BE . First-in-human, phase 1, randomized, dose-escalation trial with recombinant anti-IL-20 monoclonal antibody in patients with psoriasis. PloS One 2015; 10: e0134703.
Wang M, Tan Z, Thomas EK, Liang P . Conservation of the genomic structure and receptor-mediated signaling between human and rat IL-24. Genes Immun 2004; 5: 363–370.
Thomas EK, Nakamura M, Wienke D, Isacke CM, Pozzi A, Liang P . Endo180 binds to the C-terminal region of type I collagen. J Biol Chem 2005; 280: 22596–22605.
Ma Z, Luo D, Huang A, Xu Y, Wang Y, Wei Y et al. pKILLIN: a versatile positive-selection cloning vector based on the toxicity of Killin in Escherichia coli. Gene 2014; 544: 228–235.
Liang P, Meade JD, Pardee AB . A protocol for differential display of mRNA expression using either fluorescent or radioactive labeling. Nat Protoc 2007; 2: 457–470.
Acknowledgements
We would like to thank Dr Ursula Maria Wegenka for making the IL-20R2 knockout mice available and her kind advice for this study. This work was supported in part by a 863 grant (2012AA02A305) and grants (2012ZX09103301; 2011ZX09401005) from the Chinese Ministry of Science and Technology (PL), a 973 grant (2012CB910700) from the Chinese Ministry of Education (PL), and a grant (81171955) from the Chinese Natural Science Foundation (PL). We thank Jamie Walden and Jonathan Meade from GenHunter Corporation for proofreading the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Zhou, H., Liu, X., Yu, R. et al. Alternative splicing directs two IL-20R2 isoforms and is responsible for the incomplete gene knockout via the exon I ablation. Genes Immun 17, 220–227 (2016). https://doi.org/10.1038/gene.2016.13
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gene.2016.13
This article is cited by
-
Alternative splicing and cancer: a systematic review
Signal Transduction and Targeted Therapy (2021)
-
Alternative splicing isoforms in health and disease
Pflügers Archiv - European Journal of Physiology (2018)