Abstract
Multiple sclerosis (MS) is an inflammatory, demyelinating disorder of the central nervous system that develops in genetically susceptible individuals. The majority of the MS-associated gene variants are located in genetic regions with importance for T-cell differentiation. Vitamin D is a potent immunomodulator, and vitamin D deficiency has been suggested to be associated with increased MS disease susceptibility and activity. In CD4+ T cells, we have analyzed in vitro vitamin D responsiveness of genes that contain an MS-associated single-nucleotide polymorphism (SNP) and with one or more vitamin D response elements in their regulatory regions. We identify IL2RA and TAGAP as novel vitamin D target genes. The vitamin D response is observed in samples from both MS patients and controls, and is not dependent on the genotype of MS-associated SNPs in the respective genes.
Similar content being viewed by others
Introduction
Multiple sclerosis (MS) is an inflammatory, demyelinating disorder of the central nervous system leading to various degrees of physical and cognitive disability. The prevalence is 0.5–2.0 cases per 1000 inhabitants, with the highest prevalence reported in Scandinavia.1, 2 MS typically appears in young adults and affects females more than twice as often as males.3 The leading hypothesis is that MS is caused by a complex interaction between multiple genes and environmental factors, which leads to central nervous system inflammation causing demyelination and axonal degeneration. To date, the best supported environmental risk factors in MS are low vitamin D status, Epstein–Barr virus infection and smoking.4
It is now well established that the HLA class II-DRB1 locus causes the primary genetic association in MS, with the HLA-DRB1*15:01 allele as the major genetic risk factor in most populations (odds ratio=3.1).5, 6 In 2011, the International Multiple Sclerosis Genetics Consortium and the Wellcome Trust Case Control Consortium 2 completed a genome-wide association study including more than 9000 MS cases and found evidence for 52 non-HLA genetic loci that are associated with MS. A strong presence of immunologically relevant genes, especially genes known to regulate T cell-mediated immunity, is evident within these loci. Apart from the HLA-DRB1*15:01 allele, all other genetic MS-risk loci each exert a relatively small effect (odds ratio=1.1–1.3).6 Interestingly, these genome-wide association studies implicated the relevance also for vitamin D-processing enzymes (that is, CYP27B1 and CYP24A1) in MS6 and one study shows that HLA-DRB1*15:01 is regulated by vitamin D in lymphoblastoid cell lines.7
These studies were followed by genome-wide analyses of immune-related loci using the ImmunoChip, which enhanced the catalog of MS-risk variants to 110.8 The vast majority of single-nucleotide polymorphisms (SNPs) identified as MS-risk loci are located in non-coding regions of the genome.6, 8 Interestingly, the MS-associated SNPs are enriched in DNase I hypersensitive sites, indicating a functional role in regulation of gene transcription.9
Sun exposure and diet are the major sources of vitamin D. It exists as a biologically inert compound, vitamin D, which requires hydroxylation in the liver to 25-hydroxy vitamin D (25(OH)D). This is the major circulating form of vitamin D that has to be further hydroxylated to become the active metabolite 1,25-dihydroxy vitamin D (1,25(OH)2D). This bioactive form of vitamin D was originally described as an essential hormone for mineral and bone homeostasis, and its biological effects are mediated by the nuclear receptor vitamin D receptor (VDR).10, 11 The discovery of VDR expression in monocytes and later in antigen-presenting cells suggested a potential role for 1,25(OH)2D also in immune cells.12, 13, 14, 15 More recently it has become clear that vitamin D is indeed an important regulator of innate as well as adaptive immunity.16, 17, 18, 19, 20 Naive T cells have a low expression of VDR, which is enhanced upon T-cell activation,13, 21, 22 whereby 1,25(OH)2D mediates a shift towards a more anti-inflammatory cytokine profile.23, 24, 25 T cells express 1α-hydroxylase (encoded by CYP27B1), the enzyme responsible for the hydroxylation of 25(OH)D to 1,25(OH)2D and can themselves metabolize circulating 25(OH)D to active 1,25(OH)2D.22, 26 Interestingly, 1,25(OH)2D acts directly on the CD4+ T-lymphocyte VDR to inhibit experimental autoimmune encephalomyelitis (EAE), an animal model for MS,27 whereas CD8+ T cells are dispensable for vitamin D-mediated experimental autoimmune encephalomyelitis protection.24 Furthermore, data from Correale and colleagues suggest that 1,25(OH)2D has an important role for T-cell homeostasis in MS, as 1,25(OH)2D acts directly on CD4+ T cells to modulate cytokine secretion and reduce proliferation, and induce regulatory T cells (Treg).28, 22 Taken together, these studies highlight the importance for 1,25(OH)2D on T cells, especially CD4+ T cells, in relation with MS.
VDR is a nuclear receptor and a DNA-binding transcription factor that heterodimerizes with the retinoid X receptor (RXR). Upon 1,25(OH)2D ligation, the VDR–RXR heterodimer recognizes vitamin D response elements (VDREs) within regulatory regions of its target genes, thereby affecting gene transcription.11 VDREs are composed of tandem motifs with the consensus PuG(G/T)TCA, which are often arranged as direct repeats separated by 3 bp (DR3). VDR–RXR can also recognize everted repeats of hexameric motifs spaced by 6 bp (ER6).29, 30, 31 A gene that is directly activated by a transcription factor such as VDR generally harbors a binding site for the transcription factor. Moreover, gene expression is normally rapidly induced upon activation of that particular transcription factor.
In the current study, we measured vitamin D response in human CD4+ T cells on expression of MS-associated genes,6, 8 with more than one VDREs in their regulatory region or with one VDRE and an MS-associated SNP in the same regulatory region. Exogenous addition of 1,25(OH)2D3 to CD4+ T cells in vitro induced the expression of IL2RA, whereas it repressed TAGAP expression. This change in gene expression correlated with increased cell-surface expression of the IL2RA encoded CD25 and reduced TAGAP protein levels in activated CD4+ T cells. The vitamin D response was independent of MS disease and MS-risk genotype in the two genes. However, in MS patients there was a significant correlation between IL2RA expression in CD4+ T cells and serum levels of 25(OH)D.
Results
VDR and CYP24A1 expression in CD4+ T cells
It has previously been reported that VDR expression is low in naive T cells, and that it is upregulated upon T-cell activation.12, 19, 22, 32, 33 In agreement with this, we observed low VDR expression both at the mRNA and protein levels in freshly purified CD4+ T cells. Upon stimulation with αCD3/CD28-coated beads, VDR expression increased (Figures 1a and b) and reached its maximum protein expression after 24–40 h stimulation (Figures 1b and c). To verify the ability of 1,25(OH)2D3 to induce activation of VDR signaling pathways in αCD3/CD28-stimulated human CD4+ T cells, we measured the expression levels of the well-documented VDR-target gene CYP24A1 encoding 24-hydroxylase.34 It has previously been shown that VDR is efficiently triggered only if 1,25(OH)2D3 is added to T lymphocytes, with high levels of VDR expression at the time of 1,25(OH)2D3 addition.19 Therefore, CD4+ T cells were stimulated with αCD3/CD28-coated beads for 40 h before the addition of 1, 10 or 100 nm 1,25(OH)2D3. After 24 h, CYP24A1 was efficiently induced, reaching maximum induction with 10 nm 1,25(OH)2D3 (Figure 1d).
VDR and CYP24A1 expression in CD4+ T cells. (a–c) Human CD4+ T cells were left unstimulated (0) or stimulated for indicated hours with αCD3/CD28 beads before cell harvesting. (a) mRNA expression levels of VDR were measured relative to RNaseP, and expression in the unstimulated cells was set to 1. The graph represents the median with error range from four independent experiments. (b) Whole-cell lysates from untreated or stimulated CD4+ T cells were immunoblotted with indicated antibodies. The immunoblot shows one representative experiment out of four. (c) Bands (as shown for one representative in b) were quantified and normalized as described. The graph shows the median level with error range of VDR from four independent experiments. (d) Human CD4+ T cells were stimulated with αCD3/CD28 beads for 40 h before the addition of indicated amounts of 1,25(OH)2D3 or vehicle control for 24 h. CYP24A1 mRNA expression was measured by quantitative real-time PCR using RNaseP as reference gene. The graph represents CYP24A1 expression in 1,25(OH)2D3-treated cells relative to its expression in vehicle-treated cells, with a maximum of 100% assigned for the group with maximum 1,25(OH)2D3-induced CYP24A1 expression, that is, 10 nm 1,25(OH)2D3. Each line represents measurements from CD4+ T cells from one healthy donor.
Selection of MS-risk genes
In a recent genome-wide association study including samples from 9772 MS cases from 14 different countries, 23 previously suggested associations were replicated and 29 novel susceptibility loci were identified.6 Combining these data with analysis of 14 498 MS cases on an ImmunoChip custom genotyping array, additional MS susceptibility variants were identified. The 110 established MS-risk variants represent 103 discrete loci outside the major histocompatibility complex.8 Using data from previous in silico analyses,35 we found that 80% of the 103 non-HLA MS-associated genes have one or more DR3-type VDREs in their gene promoter or regulatory regions (−10 to +5 kb). Of these genes, 23 genes contained two or more VDREs (19 genes with two VDREs and 2 genes each with three or four VDREs; see Supplementary Table 1). We chose to study genes that had either (i) more than one VDRE in their regulatory region (20 genes, see Supplementary Table 1) or (ii) that contained one VDRE and an MS-risk SNP in the regulatory region of the gene (3 genes; IL12B, SCO2 and IL2RA). These MS-risk genes were analyzed in silico for expression in CD4+ T cells. We further limited the number of genes by selecting those that showed high expression in the https://genome-euro.ucsc.edu/ genome browser (labeled ‘red’ in Supplementary Table 1) and that displayed gene expression higher than the average at http://biogps.org/ (that is, CD5, IL2RA, MALT1, RGS14, SP140, STAT3, STAT4, TAGAP, TCF7, TYK2, BATF and MYC); see Table 1. In addition, CD6 was analyzed, as the MS-associated SNPs in this region are located between the two VDRE-containing CD5 and CD6 genes (Table 1).
Calcitriol affects the expression of IL2RA, TAGAP, BATF and MYC in CD4+ T cells
A total of 13 genes (Table 1) were analyzed for 1,25(OH)2D3 responsiveness in CD4+ T cells isolated from six healthy donors. The cells were activated for 40 h with αCD3/CD28 to induce VDR expression before the addition of 10 nm 1,25(OH)2D3 or vehicle control (ethanol). Cells were collected 3, 6 and 24 h after addition of 1,25(OH)2D3 or vehicle, and gene expression was measured by quantitative real-time PCR relative to the three reference genes 18S rRNA, TBP and GAPDH as described in the Materials and Methods section. Genes that displayed pair-wise significant differences in expression in 1,25(OH)2D3-treated samples compared with vehicle-treated samples (paired Student’s t-test for each time point) for the three different reference genes were considered as 1,25(OH)2D3-responsive. CYP24A1 has previously been shown to be regulated by VDR.34 As expected, its expression was induced upon addition of 1,25(OH)2D3 compared with vehicle control. Of the tested MS-associated VDRE-containing genes (Table 1), IL2RA expression was induced by 1,25(OH)2D3 treatment, whereas TAGAP, MYC and BATF expression was downregulated (Figure 2). No significant 1,25(OH)2D3 response was observed for the other genes tested (data not shown).
Calcitriol response of MS susceptibility genes in CD4+ T cells. Human CD4+ T cells from six donors were treated as described in Figure 1d, with 10 nm 1,25(OH)2D3 before quantitative real-time PCR analysis of the MS-associated genes listed in Table 1. The graphs show median expression relative to TBP of indicated genes with error ranges in 1,25(OH)2D3- and vehicle-treated cells. Expression at the time of 1,25(OH)2D3 or vehicle addition is set to 1. Significant P-values, P<0.05, are given (Student’s paired t-test for each single time point). Only data from genes with significantly increased expression after 1,25(OH)2D3 addition are shown.
Using a paired Student’s t-test, we were able to compare 1,25(OH)2D3-induced versus vehicle-induced gene expression at each individual time point. To determine whether there is a change in differential expression over time for all the genes listed in Table 1, we performed a functional analysis of variance test. After Benjamini–Hochberg correction, only three of the genes from Figure 2 showed a significant P-value for all three reference genes, that is, CYP24A1, IL2RA and TAGAP (corrected P-values for gene expression relative to TBP; 1.9 e-13, 2.6 e-6 and 4.7 e-7, respectively). For further analyses, we chose to focus on genes that displayed significantly altered expression when normalized to each of the three reference genes in both statistical tests, that is, IL2RA and TAGAP. As before, CYP24A1 was used as a positive control.
Calcitriol inhibits TAGAP protein expression and induces cell-surface expression of CD25 in activated CD4+ T cells
To assess whether vitamin D has an impact on the protein level of TAGAP and CD25, encoded by TAGAP and IL2RA, respectively, CD4+ T cells were purified from healthy donors, activated for 40 h with αCD3/CD28 before addition of 10 nm 1,25(OH)2D3 or vehicle control (ethanol). Cells were collected 24 and 48 h after addition of 1,25(OH)2D3 or vehicle, and protein expression was measured by flow cytometry. We observed that TAGAP protein expression was induced upon T-cell activation as measured by flow cytometry and western blotting (Figures 3a and b). Addition of calcitriol resulted in a small but significant reduction of total TAGAP expression after 48 h of incubation (Figure 3b), whereas cell-surface expression of CD25 was slightly induced (Figure 3c). No significant changes in protein levels of either of the proteins were observed after 24 h with calcitriol (data not shown).
Calcitriol inhibits TAGAP protein expression and induces cell-surface expression of CD25 in CD4+ T cells. (a) Human CD4+ T cells were left unstimulated (0) or stimulated for 40 h with αCD3/CD28 beads before intracellular staining of TAGAP and flow cytometry. Median fluorescent intensity (MFI) before (0) and after (40) stimulation is indicated for each donor (N=4). Paired Student’s t-test, P=0.0040. The histogram plot shows one representative experiment (black line, isotype control unstimulated cells (0 h); gray line, isotype control stimulated cells (40 h); orange line, TAGAP-stained unstimulated cells (0 h); green line, TAGAP-stained stimulated cells (40 h). (b) Whole-cell lysates from unstimulated or stimulated CD4+ T cells were immunoblotted with indicated antibodies. Human CD4+ T cells were treated as described in Figure 1d, with 10 nM 1,25(OH)2D3 for 48 h before flow cytometry to measure (c) TAGAP (N=7) and (d) CD25 (N=5) expression. The graphs show the MFI of TAGAP and CD25 in vehicle- and calcitriol-treated cells. Paired Student’s t-test P=0.030 (TAGAP) and P=0.012 (CD25). Histogram plots show one representative experiment (black line, isotype control vehicle-treated cells; gray line, isotype control calcitriol-treated cells; red line, TAGAP (c) or CD25 (d) vehicle-treated cells, blue line, TAGAP- (c) or CD25 (d)-stained calcitriol-treated cells).
Expression of TAGAP and IL2RA is not influenced by MS or MS-risk genotype
To evaluate whether there is a correlation between the presence of an MS-risk variant in TAGAP or IL2RA, and their vitamin D response in CD4+ T cells in vitro, we analyzed gene expression in CD4+ T cells from 18 individuals (8 relapsing-remitting MS (RRMS) patients and 10 age- and sex-matched healthy controls) that were genotyped for rs1738074 (ImmunoChip top-hit8 in TAGAP 5′ untranslated region) and rs7090512 (IL2RA, a promoter SNP that appeared as secondary signal of association in the genome-wide association study6). Cells were treated with 10 nm 1,25(OH)2D3 or vehicle in the presence of αCD3/CD28 beads, collected after 6, 24 and 48 h, and relative gene expression of CYP24A1, TAGAP and IL2RA was measured by quantitative real-time PCR. CYP24A1 expression was induced in the cultivated cells upon 1,25(OH)2D3 treatment (data not shown). As in the initial screening in CD4+ T cells from healthy donors (Figure 2), we observed an increase in IL2RA and a decrease in TAGAP expression upon 1,25(OH)2D3 addition. No difference was observed between MS patients and healthy controls (Figure 4a). As the presence or absence of MS did not give rise to any difference in vitamin D response, all samples were combined and grouped according to genotype of MS-associated SNPs in IL2RA and TAGAP. No significant genotype-dependent difference in vitamin D response was observed for either TAGAP or IL2RA (Figure 4b).
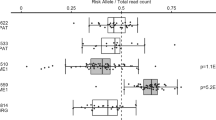
Calcitriol response of TAGAP and IL2RA in CD4+ T cells is not influenced by MS or MS-risk genotype. CD4+ T cells from 8 RRMS patients (MS) and 10 healthy controls (HC) genotyped for MS-risk SNPs in TAGAP (rs1738074) and IL2RA (rs7090512) were cultivated for 7–8 days with αCD3/CD28 and IL-2, treated as described in Materials and Methods before harvesting after 6, 24 and 48 h, with 1,25(OH)2D3 or vehicle control. The box plots with whiskers defining minimum and maximum show the log2 values of relative expression of indicted genes (relative to TBP) in 1,25(OH)2D3-treated cells divided by relative expression in vehicle-treated cells, that is, log2 of fold induction by 1,25(OH)2D3 at each time point for (a) MS patients compared with HCs and (b) the same samples sorted on IL2RA and TAGAP genotype, that is, samples carrying the minor allele (IL2RA: N=10, MAF (G)=0.3480; TAGAP: N=11, MAF (G)=0.46563) compared with samples homozygous for major allele (N=8 (IL2RA); N=7 (TAGAP)). The horizontal lines within the boxes represent the median of the groups, and Mann–Whitney U-test was performed to compare the groups (MS vs HC, AA vs AG/GG). The horizontal line at 0 indicates no induction by 1,25(OH)2D3.
IL2RA expression in CD4+ T cells correlates with serum levels of 25(OH)D
Our in vitro experiments indicate that TAGAP and IL2RA gene expression is regulated by 1,25(OH)2D3. To investigate whether there is a correlation between the expression of these vitamin D-responsive genes in immune cells and serum 25(OH)D levels, we measured the expression of IL2RA and TAGAP in CD4+ T cells purified from RRMS patients with known vitamin D status.36 Within this cohort, 14 patients had used vitamin D supplements within 6 months before sampling.37 Accordingly, the range of serum 25(OH)D levels was relatively large (52–260 nm).36 In these samples, we observed a positive correlation between relative IL2RA expression and 25(OH)D serum levels in MS patients (Figure 5). This was most likely not due to differential VDR expression levels, as VDR expression did not correlate with the serum 25(OH)D levels in the MS patients,36 neither did VDR expression correlate with IL2RA expression (data not shown). We did not observe any correlation between serum 25(OH)D levels and TAGAP expression (Figure 5).
IL2RA expression correlates with serum levels of 25(OH)D in CD4+ T cells from MS patients. CD4+ T cells were purified from a Dutch cohort of RRMS patients with known serum 25(OH)D levels as described in Materials and Methods. The graphs display linear regression analyses of serum 25(OH)D levels with the log2 of the expression of IL2RA and TAGAP relative to TBP in CD4+ T cells. r2 represents the coefficient of determination and P the uncorrected P-values.
Discussion
Low levels of vitamin D is a risk factor in MS, and active vitamin D can act directly on immune cells, as these cells express VDR. In the present study, we demonstrate that two MS-associated genes, IL2RA and TAGAP, that both contain VDREs, are regulated by vitamin D in CD4+ T cells in vitro. The change in gene expression correlated with change also at the protein level for CD25 and TAGAP. The vitamin D response of these genes was comparable in T cells from MS cases and age- and sex-matched healthy controls, and was independent of the genotype of their corresponding MS-associated SNPs within the IL2RA and TAGAP genes, respectively. Finally, there was a positive correlation between IL2RA expression in CD4+ T cells and the serum level of 25(OH)D in MS cases, indicating that vitamin D might have an impact on IL2RA expression in vivo.
For TAGAP and IL2RA, a twofold change in gene expression was observed after 1,25(OH)2D3 treatment. This rather modest response to vitamin D is comparable to what has been detected in other studies when analyzing IL-2 expression in the Jurkat T cell line38 or IFN-γ and IL-10 expression in human primary T cells.19 Furthermore, in a transcriptome-wide study performed in THP-1 cells, only 67 (16%) of 408 vitamin D-upregulated genes were induced more than 1.5-fold after 4 h of 1,25(OH)2D3 treatment.39 As 1,25(OH)2D3 triggers VDR signaling in T lymphocytes efficiently only in the presence of high VDR levels,19 T cells were activated with αCD3/CD28-coated beads to induce VDR expression before 1,25(OH)2D3 addition, and these beads were kept in the media during the 1,25(OH)2D3 treatment period. In the current work, we used 10 nm 1,25(OH)2D3 throughout the study. This is a concentration that is frequently used in equivalent studies19, 22 and is considered to be a pharmacological dose.22 This level of 1,25(OH)2D3 is ~100-fold higher than that of the circulating levels of 1,25(OH)2D3 (⩽100 pm) in healthy individuals.40 However, it is likely that the local concentration of 1,25(OH)2D3 in tissues and within cells expressing the 1-α-hydroxylase (from the CYP27B1 gene) may be considerably higher than the levels in the circulation.19, 22, 41 The initial screening (Figure 2) was performed in CD4+ T cells from six healthy donors that were not genotyped. The minor allele frequency of the MS-associated genes tested for vitamin D responsiveness differs between 0.010 (rs34536443 in TYK2) and 0.47 (rs1738074 in TAGAP). The genotype distribution will vary from gene to gene, and we cannot rule out the possibility that for some genes, especially those with low minor allele frequency, only one genotype is present, which could influence the vitamin D responsiveness of the gene. However, for IL2RA and TAGAP, the genotype of the MS-associated SNPs did not affect vitamin D responsiveness in the corresponding genes (Figure 4).The primary selection criterion for the current analyses of vitamin D responsiveness of MS-associated genes was that the genes to study should contain at least one VDRE. There are examples of genes that are regulated by VDR in a VDRE-independent manner where VDR inhibits gene expression by antagonizing certain transcription factors.42, 43, 44 Thus, our analyses are not comprehensive. However, when analyzing genome-wide chromatin immunoprecipitation (ChIP) data of VDR binding focusing on the top VDR sites (based on fold enrichment scoring), the majority of these genes contain a DR3–VDRE sequence.39, 45, 46, 47 Genome-wide VDR ChIP-sequencing studies have been performed in five different cell lines (GM10855 and GM10861,45 THP-1 cells,39, 48 LS180 colorectal cancer cells49 and LX3 hepatic stellate cells50) and one study was performed in primary human CD4+ T cells.51 Taken together, these data indicate that there are far more genomic VDR-binding sites (1000–10 000) per cell than target genes (100–500 per tissue). With a total of 21 776 non-overlapping VDR-binding sites, only 54 are common within the six data sets and 67% are unique for one of the analyzed data sets47 (summarized for the MS susceptibility genes in Supplementary Table 2). This indicates that VDR displays cell-specific binding patterns, and functional follow-up of each potential target gene in relevant tissue is therefore of great importance.
Among the 13 VDRE-containing genes tested in CD4+ T cells, all genes were occupied by VDR in T cells, although in some genes more than 10 000 bp upstream of the coding sequence (that is, IL2RA and TYK2;51 summarized in Supplementary Table 2), but only two of them were regulated by vitamin D in vitro. Whereas VDR-mediated gene repression occurs through a variety of mechanisms (reviewed in ref. 11), transcriptional activation by VDR is mediated through binding to its cognate VDREs. Surprisingly, the VDRE-containing TAGAP gene was downregulated upon 1,25(OH)2D3 addition, and a response was observed after only 3 h. Owing to this rapid response it is likely that TAGAP is regulated by VDR itself and not indirectly by another factor that is transcriptionally induced by VDR. Interestingly, in ChIP experiments from CD4+ T cells, one of the VDREs in TAGAP (+329 VDRE in Table 1) lies within the VDR ChIP-sequence-binding interval.51 Combined with our expression data, this suggests that VDR can bind TAGAP and thereby can regulate gene expression in human CD4+ T cells. Our data further indicated that the genotype of the MS-associated rs1738074, only 121 nucleotides away from the VDRE (+329), does not have any impact on vitamin D-mediated repression of TAGAP in CD4+ T cells. As TAGAP contains a VDRE in its 5′ region, it is likely that the vitamin D-activated VDR binds to the TAGAP promoter, as shown in CD4+ T cells,51 thus reducing the transcriptional activity of the gene. This might occur by recruiting corepressors to the site, which is the case for instance of the VDR-regulated MYC gene in human prostate cells.52 Alternatively, data from the encode database indicate that the VDRE in the 5′ region of TAGAP overlaps with a RELA ChIP-sequencing signal.53 This suggests that VDR might compete with RELA inhibiting TAGAP transcription in a mechanism similar to VDR-mediated IL-2 repression, where VDR blocks the binding of the transcription factor nuclear factor of activated T cells-1 to the IL-2 promoter.42
On the other hand, IL2RA was consistently induced after 1,25(OH)2D3 treatment. This was observed at a much later time point, that is, 24 h and could indicate that IL2RA induction was not directly mediated by VDR. A significant correlation between IL2RA expression in CD4+ T cells and serum levels of vitamin D in MS patients further indicates that vitamin D, although not necessarily directly, might modulate IL2RA expression. VDR expression can be induced by calcitriol,54 however, as there was no significant correlation between VDR expression and 25(OH)D levels36 or IL2RA expression (data not shown), the positive correlation between IL2RA expression and vitamin D status is not due to different VDR expression levels. In the current analyses we could not find any correlation between IL2RA response and the genotype of rs7090512. Furthermore, the MS-associated SNPs in IL2RA were not pulled down in the VDR ChIP-sequencing experiment in CD4+ T cells.51 The MS-associated SNPs in IL2RA are not within a predicted VDRE. The VDRE in IL2RA is situated almost 10 kb upstream of the IL2RA transcriptional start site (TSS), but VDR loci in a distance even more than 1 Mb from the gene’s transcriptional start site are accepted as regulatory sites.47 It remains to be shown whether this VDRE in IL2RA is a functional site for VDR in these cells.
TAGAP encodes the T-cell activation Rho-GTPase-activating protein (TAGAP). It has previously been shown that TAGAP mRNA expression is induced during T-cell activation,55 and we here show that TAGAP is induced also at the protein level upon T-cell activation (Figures 3a and b). The function of TAGAP as such is currently unknown. However, as GAPs are involved in several key immunological processes in T cells,56, 57 TAGAP might have a potential regulatory role in T-cell activation. In the current study, we show that 1,25 (OH)2D3 down-modulated TAGAP expression in activated CD4+ T cells. Polymorphisms in TAGAP have shown association to other autoimmune diseases such as type-1 diabetes, rheumatoid arthritis and Crohn’s disease,58, 59, 60, 61 and studies of colonic tissues in Crohn’s disease patients show elevated TAGAP expression in samples from patients with a more severe phenotype.62 It is not known whether TAGAP expression correlates with MS severity. Thus, whether vitamin D-mediated reduction in TAGAP expression in activated CD4+ T cells is beneficial for MS remains to be studied.
IL2RA expresses CD25, the alpha chain of the high-affinity IL-2 receptor, and increased cell-surface expression of CD25 observed in 1,25(OH)2D3-treated CD4+ T cells might indicate an increase in interleukin (IL)-2 responsiveness. Interestingly, there is a correlation between 25(OH)D levels and Treg cell function.63 Moreover, 1,25(OH)2D3 induces the development of Treg cells, with an enhanced suppressive activity in vitro.22 Even though IL-2 acts as trophic factor for anti-inflammatory Treg cells, it is also aiding the expansion of pro-inflammatory effector T cells.64 In the samples we have analyzed, both effector CD4+ T cells as well as Treg cells are likely to be present. An increase in IL-2 response of Treg cells would probably be beneficial for autoimmunity, whereas the opposite would be the case for effector CD4+ T cells.
Low vitamin D status has been suggested to be a risk factor in several diseases, and VDR ChIP sequencing of human primary CD4+ T cells indicates enrichment of VDR binding within regions associated with autoimmunity.51 In the current analyses we find that 80% of the MS-associated genes have one or more VDRE in their regulatory region; however, only 2 out of 13 selected VDRE-containing genes display vitamin D responsiveness in CD4+ T cells in vitro. This indicates that the presence of VDRE and binding of VDR is not sufficient to induce gene expression in vitro, and it remains to be analyzed whether this is different in an in vivo setting. Altogether our data show that the MS susceptibility genes TAGAP and IL2RA are regulated by vitamin D in CD4+ T cells. The mechanism behind this regulation remains to be elucidated; however, neither presence nor absence of MS nor the genotype of the MS-associated risk variants do seem to affect the vitamin D responsiveness of these genes.
Materials and methods
Reagents and antibodies
Calcitriol (1,25(OH)2D3; Sigma-Aldrich Corp., St Louis, MO, USA) was solubilized in absolute ethanol to 1 mm and further diluted in X-VIVO 15 medium (04–418, Lonza, Basel, Switzerland) before addition to the cell culture. The antibodies used were fluorescein isothiocyanate-conjugated mouse anti-human CD4 (clone RTF-4 g, Southern Biotech, Birmingham, AL, USA), fluorescein isothiocyanate-conjugated mouse IgG1 isotype control (15H6, Southern Biotech), allophycocyanin-conjugated anti-human CD25 (IgG1, Immunotools, Friesoythe, Germany), allophycocyanin-conjugated mouse isotype IgG1 (Immunotools), mouse anti-VDR (D-6, sc-13133, Santa Cruz Biotechnology, Dallas, TX, USA), mouse anti-GAPDH (6C5, sc-32233, Santa Cruz Biotechnology), rabbit anti-TAGAP (EPR15593, Abcam, Cambridge, UK), rabbit anti-Actin (A2066, Sigma-Aldrich Corp.), rabbit monoclonal IgG (EPR25A, Abcam), horseradish peroxidase-conjugated goat anti-mouse IgG and goat anti-rabbit IgG (Jackson ImmunoResearch Laboratories Europe Ltd., Suffolk, UK) and Alexa Fluor 647-conjugated goat anti-rabbit IgG (A-21245, Molecular Probes, Life Technologies, Carlsbad, CA, USA).
Isolation of human CD4+ T cells and cell culture
Peripheral blood mononuclear cells were isolated from whole blood by Lymphoprep (Axis Shield, Dundee, Scotland), before negative selection of CD4+ T cells using autoMACS Pro Separator (Miltenyi Biotec, Bergisch Gladbach, Germany) or Dynabeads Untouched Human CD4 T Cell Isolation kit (Life Technologies) to more than 95% purity as measured by flow cytometry (FacsCalibur, BD Biosciences, San Jose, CA, USA or Attune Acoustic Focusing Flow Cytometer, Life Technologies). Freshly purified CD4+ T cells were either frozen on liquid nitrogen for later use or resuspended in X-VIVO medium (Lonza) with human IL-2 (R&D Systems, Minneapolis, MS, USA) and αCD3/CD28 beads (1:1) (Human T-activator CD3/CD28 Dynabeads for cell expansion and activation, Life Technologies) for activation and 1,25(OH)2D3 treatment. If the cell density reached 2.5 million cells per ml, the wells were split. Cells from genotyped individuals were stored on liquid nitrogen (two million cells per tube). After thawing, the cells were expanded to achieve sufficient cell numbers for the analyses by growing them as the freshly isolated CD4+ T cells for 7-8 days before removal of aCD3/CD28 beads 24 h prior to addition of 1,25(OH)2D3 or vehicle control together with aCD3/CD28 beads. At the time intervals indicated in the text, cells were counted and collected by centrifugation. Before cell lysis for western blot analyses, magnetic beads were removed before cell harvesting.
Cell lysis, western blotting and flow cytometry
The cell pellet was resuspended in reducing SDS-loading buffer, sonicated and heated to 95 °C for 5 min. Proteins were separated by SDS-polyacrylamide gel electrophoresis using pre-made Criterion gels (BioRad, Hercules, CA, USA) and transferred to polyvinylidene fluoride membrane (BioRad) using a Hoefer Semi-Phor Seri-Dry transfer unit (Amersham Biosciences, Buckinghamshire, UK). The membrane was blocked in 3% skimmed milk in Tris-buffered saline (TBS, pH 7.4) containing 0.1% Tween-20 (Sigma-Aldrich) (TBS-T) before incubation with antibodies in TBS-T with 3% skimmed milk. Bound antibodies were visualized by incubation with secondary horseradish peroxidase-conjugated antibodies and Super Signal West Pico stable peroxide solution (Pierce Biotechnology, Rockford, IL, USA). Densitometry of the western blots was analyzed by the ImageJ software.65 Cell-surface protein expression, that is, CD4 and CD25, was analyzed with flow cytometry (FacsCalibur, BD Biosciences; or Attune Acoustic Focusing Cytometer, Life Technologies) and FlowJo software (Tree Star, Ashland, OR, USA) or Attune Cytometric Software (Life Technologies). For intracellular staining of TAGAP, cells were fixed in 3% paraformaldehyde before permeabilization with 0.05% saponin (S7900, Sigma). In all steps proceeding permeabilization, 0.05% saponin was present in all staining and washing solutions. Samples were analyzed with Attune Acoustic Focusing Flow Cytometer (Life Technologies) and FlowJo software (Tree Star).
Genotyping
TAGAP and IL2RA were genotyped for rs1738074 and rs7090512 using pre-made TaqMan genotyping assays (assay ID 2966098 and 1841420, respectively, Applied Biosystems, Life Technologies) or genotypes were extracted from genome-wide SNP data assessed by the Human Omni Express Bead Chip (Illumina, San Diego, CA, USA).
MS patients and controls
The Norwegian sample collection consisted of CD4+ T cells that were isolated from whole blood66 from untreated Norwegian females of Nordic origin with RRMS (n=8, mean age 42 years, s.d.=11) and female controls of Nordic origin (n=10, mean age 36 years, s.d.=6.4). The RRMS patients had not experienced a relapse or received steroids in the 3 months before enrollment. The Dutch collection consisted of cDNA from CD4+ T cells from RRMS patients (n=36, mean age 40, s.d.=10, 70% females) collected in the Netherlands.36 The patients used either interferon-β or no immunomodulation drugs, and they had not had any relapse for the last 6 weeks. Serum levels of 25(OH)D in the Dutch cohort were measured as described previously.36 All MS patients fulfilled the updated McDonald MS criteria.67 The Regional Committee for Medical and Health Research Ethics South East, Norway, and the ‘Atrium-Orbis-Zuyd’ (Heerlen, The Netherlands) approved the study. Written informed consent was obtained from all study participants.
RNA isolation, cDNA conversion and real-time quantitative PCR
RNA and cDNA conversion from the Dutch cohort has previously been described.36 For all other experiments, cells were resuspended in 350 μl RNA Protect Cell Reagent (Qiagen, Limburg, The Netherlands) and kept at −80 °C before RNA extraction using QIAshredder (Qiagen) for homogenization of the cells, and RNAeasy Plus Mini Kit (Qiagen) for isolation of RNA. RNA concentration, quality and integrity were verified by Nanodrop ND-1000 Spectrophotometer (Thermo Fisher Scientific Inc., Madison, WI, USA) and Agilent 2100 Bioanalyzer (Agilent Technologies, Santa Clara, CA, USA). Total RNA (560 ng) was reverse transcribed in a 20-μl reaction using Maxima First Strand cDNA Synthesis Kit for RT-quantitative real-time PCR (Thermo Scientific, Waltham, MA, USA) and the GeneAmp PCR system 9700 thermo cycler (Applied Biosystems) for a one-step PCR reaction (25 °C for 10 min, 50 °C for 30 min and 85 °C or 5 min). The cDNA was stored at −80 °C. Real-time quantitative PCR was performed on 7900HT Fast Real-Time PCR system or Viia7 Real-Time PCR system (both from Applied Biosystems, Life Technologies) with the standard curve method using TaqMan pre-made gene expression assays. A standard curve was generated for each RNA collection using a mixture of RNA for the specific collection. The samples were run in triplicates or duplicates, with 1–4 ng cDNA as input, and primers and probes (indicated below, all from Applied Biosystems, Life Technologies) were added to each reaction, including in addition 5 μl TaqMan Gene Expression Mastermix (Applied Biosystems, Life Technologies), 0,5 μl of 20 × Primer Probe mix (Applied Biosystems, Life Technologies) and 4 μl RNase-free water (Qiagen). The samples were run on a MicroAmp Optical 384-well reaction plate (Applied Biosystems, Life Technologies), with the following PCR conditions: 50 °C for 2 min, 95 °C for 10 min followed by 40 cycles of 95 °C for 15 s and 60 °C for 1 min. Data were analyzed by the sequence detection systems v. 2.3 (Applied Biosystems, Life Technologies). The following TaqMan gene expression assays were used (all from Applied Biosystems, Life Technologies): BATF, Hs00232390; CD5, Hs00204397; CD6, Hs00198752; CYP24A1, Hs00167999; IQCB1, Hs00274384; KIF21B, Hs01118430; MALT1, Hs01120052; MYC, Hs00153408; RGS14, Hs00374626; SP140, Hs00610654; STAT3, Hs01047580; STAT4, HS01028017; TAGAP, Hs00299284; TCF7, Hs00175273; TYK2, Hs00177464; and VDR, Hs00172113. All data were normalized to TBP and/or GAPDH and/or 18S rRNA, also TaqMan probes from Applied Biosystems, Life Technologies: 18S rRNA, 4319413E; GAPDH, Hs03929097; and TBP, 4326322E. Outliers from the triplets were excluded. In samples where only doublets were analyzed, samples showing a CT s.d. more than 0.5 were re-run. A negative control without cDNA and a no-reverse transcriptase control were included in each experiment. Relative expression was calculated as the ratio between the target gene and RNaseP or TBP, GAPDH and 18S rRNA reference genes. PCR specificity was verified by a single band after gel electrophoresis.
Statistics
Statistically significant differences between gene or protein expressions at single time points were assessed by a two-sided paired Student’s t-test and gene expression between two groups (MS versus controls, groups with different genotypes) were assessed by two-sided Mann–Whitney U-test. Linear regression was used to asses correlation between gene expression and serum levels of 25(OH)D (all Graph Pad Prism Software, Inc., San Diego, CA, USA). P-values below 0.05 were considered significant. To assess the statistical significance for a change in gene expression over time, a functional analysis of variance was used. For each gene, a natural cubic spline with three degrees of freedom was fitted to the difference between the control samples and the vitamin D-treated samples and tested with analysis of variance against the null model. To account for the correlation between the series, we used a linear mixed model68 with random effect for each series. As several independent genes were tested, Benjamini–Hochberg correction69 was performed on the P-values using R.70 Corrected P-values below 0.05 were considered significant.
References
Gourraud PA, Harbo HF, Hauser SL, Baranzini SE . The genetics of multiple sclerosis: an up-to-date review. Immunol Rev 2012; 248: 87–103.
Berg-Hansen P, Moen S, Harbo H, Celius E . High prevalence and no latitude gradient of multiple sclerosis in Norway. Mult Scler 2014; 20: 1780–1782.
Alonso A, Hernan MA . Temporal trends in the incidence of multiple sclerosis: a systematic review. Neurology 2008; 71: 129–135.
O’Gorman C, Lucas R, Taylor B . Environmental risk factors for multiple sclerosis: a review with a focus on molecular mechanisms. Int J Mol Sci 2012; 13: 11718–11752.
Olerup O, Zetterquist H . HLA-DRB1*01 subtyping by allele-specific PCR amplification: a sensitive, specific and rapid technique. Tissue Antigens 1991; 37: 197–204.
International Multiple Sclerosis Genetics C, Wellcome Trust Case Control C, Sawcer S Hellenthal G Pirinen M Spencer CC et al. Genetic risk and a primary role for cell-mediated immune mechanisms in multiple sclerosis. Nature 2011; 476: 214–219.
Ramagopalan SV, Maugeri NJ, Handunnetthi L, Lincoln MR, Orton SM, Dyment DA et al. Expression of the multiple sclerosis-associated MHC class II Allele HLA-DRB1*1501 is regulated by vitamin D. PLoS Genet 2009; 5: e1000369.
International Multiple Sclerosis Genetics C Beecham AH Patsopoulos NA Xifara DK Davis MF Kemppinen A et al. Analysis of immune-related loci identifies 48 new susceptibility variants for multiple sclerosis. Nat Genet 2013; 45: 1353–1360.
Maurano MT, Humbert R, Rynes E, Thurman RE, Haugen E, Wang H et al. Systematic localization of common disease-associated variation in regulatory DNA. Science 2012; 337: 1190–1195.
DeLuca HF, Zierold C . Mechanisms and functions of vitamin D. Nutr Rev 1998; 56: S4–S10 ; discussion S54–S75.
Haussler MR, Jurutka PW, Mizwicki M, Norman AW . Vitamin D receptor (VDR)-mediated actions of 1alpha,25(OH)(2)vitamin D(3): genomic and non-genomic mechanisms. Best Pract Res Clin Endocrinol Metab 2011; 25: 543–559.
Provvedini DM, Tsoukas CD, Deftos LJ, Manolagas SC . 1,25-dihydroxyvitamin D3 receptors in human leukocytes. Science 1983; 221: 1181–1183.
Veldman CM, Cantorna MT, DeLuca HF . Expression of 1,25-dihydroxyvitamin D(3) receptor in the immune system. Arch Biochem Biophys 2000; 374: 334–338.
Overbergh L, Decallonne B, Waer M, Rutgeerts O, Valckx D, Casteels KM et al. 1alpha,25-dihydroxyvitamin D3 induces an autoantigen-specific T-helper 1/T-helper 2 immune shift in NOD mice immunized with GAD65 (p524-543). Diabetes 2000; 49: 1301–1307.
Chen S, Sims GP, Chen XX, Gu YY, Chen S, Lipsky PE . Modulatory effects of 1,25-dihydroxyvitamin D3 on human B cell differentiation. J Immunol 2007; 179: 1634–1647.
Baeke F, van Etten E, Gysemans C, Overbergh L, Mathieu C . Vitamin D signaling in immune-mediated disorders: evolving insights and therapeutic opportunities. Mol Aspects Med 2008; 29: 376–387.
Bikle DD . Vitamin D and immune function: understanding common pathways. Curr Osteoporos Rep 2009; 7: 58–63.
Adams JS, Hewison M . Unexpected actions of vitamin D: new perspectives on the regulation of innate and adaptive immunity. Nat Clin Pract Endocrinol Metab 2008; 4: 80–90.
Baeke F, Korf H, Overbergh L, van Etten E, Verstuyf A, Gysemans C et al. Human T lymphocytes are direct targets of 1,25-dihydroxyvitamin D3 in the immune system. J Steroid Biochem Mol Biol 2010; 121: 221–227.
Peelen E, Knippenberg S, Muris AH, Thewissen M, Smolders J, Tervaert JW et al. Effects of vitamin D on the peripheral adaptive immune system: a review. Autoimmun Rev 2011; 10: 733–743.
Norman AW . Minireview: vitamin D receptor: new assignments for an already busy receptor. Endocrinology 2006; 147: 5542–5548.
Correale J, Ysrraelit MC, Gaitan MI . Immunomodulatory effects of vitamin D in multiple sclerosis. Brain 2009; 132 (Pt 5): 1146–1160.
Adorini L . Immunomodulatory effects of vitamin D receptor ligands in autoimmune diseases. Int Immunopharmacol 2002; 2: 1017–1028.
Meehan TF, DeLuca HF . The vitamin D receptor is necessary for 1alpha,25-dihydroxyvitamin D(3) to suppress experimental autoimmune encephalomyelitis in mice. Arch Biochem Biophys 2002; 408: 200–204.
Smolders J, Damoiseaux J, Menheere P, Hupperts R . Vitamin D as an immune modulator in multiple sclerosis, a review. J Neuroimmunol 2008; 194: 7–17.
Kongsbak M, von Essen MR, Levring TB, Schjerling P, Woetmann A, Odum N et al. Vitamin D-binding protein controls T cell responses to vitamin D. BMC Immunol 2014; 15: 35.
Mayne CG, Spanier JA, Relland LM, Williams CB, Hayes CE . 1,25-Dihydroxyvitamin D3 acts directly on the T lymphocyte vitamin D receptor to inhibit experimental autoimmune encephalomyelitis. Eur J Immunol 2011; 41: 822–832.
Tsoukas CD, Provvedini DM, Manolagas SC . 1,25-dihydroxyvitamin D3: a novel immunoregulatory hormone. Science 1984; 224: 1438–1440.
Thummel KE, Brimer C, Yasuda K, Thottassery J, Senn T, Lin Y et al. Transcriptional control of intestinal cytochrome P-4503A by 1alpha,25-dihydroxy vitamin D3. Mol Pharmacol 2001; 60: 1399–1406.
Drocourt L, Ourlin JC, Pascussi JM, Maurel P, Vilarem MJ . Expression of CYP3A4, CYP2B6, and CYP2C9 is regulated by the vitamin D receptor pathway in primary human hepatocytes. J Biol Chem 2002; 277: 25125–25132.
Thompson PD, Jurutka PW, Whitfield GK, Myskowski SM, Eichhorst KR, Dominguez CE et al. Liganded VDR induces CYP3A4 in small intestinal and colon cancer cells via DR3 and ER6 vitamin D responsive elements. Biochem Biophys Res Commun 2002; 299: 730–738.
Bhalla AK, Amento EP, Clemens TL, Holick MF, Krane SM . Specific high-affinity receptors for 1,25-dihydroxyvitamin D3 in human peripheral blood mononuclear cells: presence in monocytes and induction in T lymphocytes following activation. J Clin Endocrinol Metab 1983; 57: 1308–1310.
von Essen MR, Kongsbak M, Schjerling P, Olgaard K, Odum N, Geisler C . Vitamin D controls T cell antigen receptor signaling and activation of human T cells. Nat Immunol 2010; 11: 344–349.
Prosser DE, Jones G . Enzymes involved in the activation and inactivation of vitamin D. Trends Biochem Sci 2004; 29: 664–673.
Wang TT, Tavera-Mendoza LE, Laperriere D, Libby E, MacLeod NB, Nagai Y et al. Large-scale in silico and microarray-based identification of direct 1,25-dihydroxyvitamin D3 target genes. Mol Endocrinol 2005; 19: 2685–2695.
Smolders J, Thewissen M, Theunissen R, Peelen E, Knippenberg S, Menheere P et al. Vitamin D-related gene expression profiles in immune cells of patients with relapsing remitting multiple sclerosis. J Neuroimmunol 2011; 235: 91–97.
Smolders J, Peelen E, Thewissen M, Cohen Tervaert JW, Menheere P, Hupperts R et al. Safety and T cell modulating effects of high dose vitamin D3 supplementation in multiple sclerosis. PloS One 2010; 5: e15235.
Matilainen JM, Rasanen A, Gynther P, Vaisanen S . The genes encoding cytokines IL-2, IL-10 and IL-12B are primary 1alpha,25(OH)2D3 target genes. J Steroid Biochem Mol Biol 2010; 121: 142–145.
Heikkinen S, Vaisanen S, Pehkonen P, Seuter S, Benes V, Carlberg C . Nuclear hormone 1alpha,25-dihydroxyvitamin D3 elicits a genome-wide shift in the locations of VDR chromatin occupancy. Nucleic Acids Res 2011; 39: 9181–9193.
Holmoy T, Kampman MT, Smolders J . Vitamin D in multiple sclerosis: implications for assessment and treatment. Expert Rev Neurother 2012; 12: 1101–1112.
Jeffery LE, Burke F, Mura M, Zheng Y, Qureshi OS, Hewison M et al. 1,25-Dihydroxyvitamin D3 and IL-2 combine to inhibit T cell production of inflammatory cytokines and promote development of regulatory T cells expressing CTLA-4 and FoxP3. J Immunol 2009; 183: 5458–5467.
Alroy I, Towers TL, Freedman LP . Transcriptional repression of the interleukin-2 gene by vitamin D3: direct inhibition of NFATp/AP-1 complex formation by a nuclear hormone receptor. Mol Cell Biol 1995; 15: 5789–5799.
Takeuchi A, Reddy GS, Kobayashi T, Okano T, Park J, Sharma S . Nuclear factor of activated T cells (NFAT) as a molecular target for 1alpha,25-dihydroxyvitamin D3-mediated effects. J Immunol 1998; 160: 209–218.
Towers TL, Freedman LP . Granulocyte-macrophage colony-stimulating factor gene transcription is directly repressed by the vitamin D3 receptor. Implications for allosteric influences on nuclear receptor structure and function by a DNA element. J Biol Chem 1998; 273: 10338–10348.
Ramagopalan SV, Heger A, Berlanga AJ, Maugeri NJ, Lincoln MR, Burrell A et al. A ChIP-seq defined genome-wide map of vitamin D receptor binding: associations with disease and evolution. Genome Res 2010; 20: 1352–1360.
Seuter S, Neme A, Carlberg C . Characterization of genomic vitamin D receptor binding sites through chromatin looping and opening. PloS One 2014; 9: e96184.
Carlberg C . Genome-wide (over)view on the actions of vitamin D. Front Physiol 2014; 5: 167.
Tuoresmaki P, Vaisanen S, Neme A, Heikkinen S, Carlberg C . Patterns of genome-wide VDR locations. PloS One 2014; 9: e96105.
Meyer MB, Goetsch PD, Pike JW . VDR/RXR and TCF4/beta-catenin cistromes in colonic cells of colorectal tumor origin: impact on c-FOS and c-MYC gene expression. Mol Endocrinol 2012; 26: 37–51.
Ding N, Yu RT, Subramaniam N, Sherman MH, Wilson C, Rao R et al. A vitamin D receptor/SMAD genomic circuit gates hepatic fibrotic response. Cell 2013; 153: 601–613.
Handel AE, Sandve GK, Disanto G, Berlanga-Taylor AJ, Gallone G, Hanwell H et al. Vitamin D receptor ChIP-seq in primary CD4+ cells: relationship to serum 25-hydroxyvitamin D levels and autoimmune disease. BMC Med 2013; 11: 163.
Toropainen S, Vaisanen S, Heikkinen S, Carlberg C . The down-regulation of the human MYC gene by the nuclear hormone 1alpha,25-dihydroxyvitamin D3 is associated with cycling of corepressors and histone deacetylases. J Mol Biol 2010; 400: 284–294.
Consortium EP. An integrated encyclopedia of DNA elements in the human genome. Nature 2012; 489: 57–74.
Zella LA, Kim S, Shevde NK, Pike JW . Enhancers located within two introns of the vitamin D receptor gene mediate transcriptional autoregulation by 1,25-dihydroxyvitamin D3. Mol Endocrinol 2006; 20: 1231–1247.
Mao M, Biery MC, Kobayashi SV, Ward T, Schimmack G, Burchard J et al. T lymphocyte activation gene identification by coregulated expression on DNA microarrays. Genomics 2004; 83: 989–999.
Saoudi A, Kassem S, Dejean A, Gaud G . Rho-GTPases as key regulators of T lymphocyte biology. Small GTPases 2014; e-pub ahead of print 8 May 2014; doi:10.4161/sgtp.28208.
Pernis AB . Rho GTPase-mediated pathways in mature CD4+ T cells. Autoimmun Rev 2009; 8: 199–203.
Eyre S, Hinks A, Bowes J, Flynn E, Martin P, Wilson AG et al. Overlapping genetic susceptibility variants between three autoimmune disorders: rheumatoid arthritis, type 1 diabetes and coeliac disease. Arthritis Res Ther 2010; 12: R175.
Franke A, Balschun T, Sina C, Ellinghaus D, Hasler R, Mayr G et al. Genome-wide association study for ulcerative colitis identifies risk loci at 7q22 and 22q13 (IL17REL). Nat Genet 2010; 42: 292–294.
Festen EA, Goyette P, Green T, Boucher G, Beauchamp C, Trynka G et al. A meta-analysis of genome-wide association scans identifies IL18RAP, PTPN2, TAGAP, and PUS10 as shared risk loci for Crohn’s disease and celiac disease. PLoS Genet 2011; 7: e1001283.
Chen R, Stahl EA, Kurreeman FA, Gregersen PK, Siminovitch KA, Worthington J et al. Fine mapping the TAGAP risk locus in rheumatoid arthritis. Genes Immun 2011; 12: 314–318.
Connelly TM, Berg AS, Harris LR 3rd, Hegarty JP, Ruggiero FM, Deiling SM et al. T-cell activation Rho GTPase-activating protein expression varies with inflammation location and severity in Crohn’s disease. J Surg Res 2014; 190: 457–464.
Smolders J, Thewissen M, Peelen E, Menheere P, Tervaert JW, Damoiseaux J et al. Vitamin D status is positively correlated with regulatory T cell function in patients with multiple sclerosis. PloS One 2009; 4: e6635.
Boyman O, Sprent J . The role of interleukin-2 during homeostasis and activation of the immune system. Nat Rev Immunol 2012; 12: 180–190.
Abràmoff MD, Magalhães PJ, Ram SJ . Image processing with ImageJ. Biophoton Int 2004; 11: 36–43.
Bos SD, Page CM, Andreassen BK, Elboudwarej E, Gustavsen MW, Briggs F et al. Genome-wide DNA methylation profiles indicate CD8+ T cell hypermethylation in multiple sclerosis. PloS One 2015; 10: e0117403.
Polman CH, Reingold SC, Banwell B, Clanet M, Cohen JA, Filippi M et al. Diagnostic criteria for multiple sclerosis: 2010 revisions to the McDonald criteria. Ann Neurol 2011; 69: 292–302.
Pinheiro J, Bornkamp B, Glimm E, Bretz F . Model-based dose finding under model uncertainty using general parametric models. Stat Med 2014; 33: 1646–1661.
Benjamini Y, Hochberg Y . Controlling the false discovery rate: a practical and powerful approach to multiple testing. J R Stat Soc Series B (Methodol) 1995; 57: 289–300.
R Development Core Team. A language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria, 2013. Available at http://www.R-project.org/.
Acknowledgements
We thank all patients and controls for participation. The doctors and nurses at the Department of Neurology, Oslo University Hospital are thanked for their help with the collection of samples. The study was funded by grants from The South-Eastern Norway Regional Health Authority, Ullevål University Hospital Scientific Advisory Council (VIRUUS), the Odd Fellow society, Halvor Høies fund, Biogen Idec Norway AS, Novartis Norway AS and Merck Serono.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
TB has received an unrestricted grant for running expenses in this project from Biogen Idec Norway AS, ISL has received unrestricted grant for salary expenses from Novartis Norway AS and TH has received research grants from Merck Serono.
Additional information
Supplementary Information accompanies this paper on Genes and Immunity website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Berge, T., Leikfoss, I., Brorson, I. et al. The multiple sclerosis susceptibility genes TAGAP and IL2RA are regulated by vitamin D in CD4+ T cells. Genes Immun 17, 118–127 (2016). https://doi.org/10.1038/gene.2015.61
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/gene.2015.61
This article is cited by
-
Replication study of GWAS risk loci in Greek multiple sclerosis patients
Neurological Sciences (2019)
-
T‐cell activation RhoGTPase‐activating protein plays an important role in TH17‐cell differentiation
Immunology & Cell Biology (2017)