Abstract
The spatiotemporal regulation of immune cells in lymph nodes (LNs) is crucial for mounting protective T-cell responses, which are orchestrated by dendritic cells (DCs). However, it is unclear how the DC subsets are altered by the inflammatory milieu of LNs. Here, we show that the inflamed LNs of Listeria-infected mice are characterized by the clustering of neutrophils and monocytes and IFN-γ production. Significantly, the early inflammatory responses are coupled with the differentiation of not one, but two types of CD64+CD11c+MHCII+ inflammatory DCs. Through the assessment of chemokine receptor dependency, gene expression profiles, growth factor requirements and DC-specific lineage mapping, we herein unveil a novel inflammatory DC population (we termed ‘CD64+ cDCs’) that arises from conventional DCs (cDCs), distinguishable from CD64+ monocyte-derived DCs (moDCs) in inflamed LNs. We determined that Listeria-induced type I IFN is a critical inflammatory cue for the development of CD64+ cDCs but not CD64+ moDCs. Importantly, CD64+ cDCs displayed a higher potential to activate T cells than CD64+ moDCs, whereas the latter showed more robust expression of inflammatory genes. Although CD64+ and CD64− cDCs were able to cross-present soluble antigens at a high dose to CD8+ T cells, CD64+ cDCs concentrated and cross-presented a minute amount of soluble antigens delivered via CD64 (FcγRI) as immune complexes. These findings reveal the role of early inflammatory responses in driving the differentiation of two inflammatory DC subsets empowered with distinct competencies.
Similar content being viewed by others
Introduction
The lymph node (LN) is a well-designed organ that orchestrates the immune response upon challenge by an infection or antigen. Structurally, LNs comprise the cortex, the paracortex and the medulla.1, 2 B cells mainly reside in the cortex, forming subcapsular follicles, whereas T cells and dendritic cells (DCs) are positioned in the paracortex area. DCs recognize, take up and process pathogens and antigens, and present the degraded products to T cells via major histocompatibility complex (MHC) molecules, together with co-stimulatory molecules and cytokines. Static immunohistochemistry (IHC) and two-photon confocal live imaging have been used to examine the dynamic regulation of intranodal immune responses, which are tightly coupled with the microanatomy of LNs.3, 4, 5, 6, 7, 8, 9, 10 Discrete locations of DC subsets also have been reported.11, 12 The inflammatory milieu alters gene expression, dynamically remodels the architecture of the LN, inhibits the systemic spreading of pathogens and directs the subsequent adaptive immune responses.13, 14, 15, 16, 17, 18
Inflammatory DCs are a DC population induced by infection and inflammation settings. They are termed moDCs (monocyte-derived DCs) or TipDCs (TNFα- and iNOS-producing DCs).19 Since the original report that phagocytic monocytes can differentiate into DCs within LNs,20 inflammatory DCs have been identified by the expression of various surface markers (for example, Ly6C, F4/80, CCR2, DC-SIGN (CD209), CD206, FcɛRI and CD64 (FcγRI)) in many settings.21 However, their development remains to be determined.22 Recent studies revealed that CD64 can be used to distinguish moDCs or macrophages from conventional DCs (cDCs).23, 24, 25, 26, 27, 28 FcγR-mediated internalization of antigen–antibody immune complexes (ICs) induced DC maturation and MHC I-restricted cross-presentation in vitro.29 However, the identity of DC subsets that cross-present IC antigens has not been demonstrated.30 Although CD8α+-resident cDCs and CD103+ migratory cDCs have been shown to be superior in cross-presentation,31 CD11b+ cDCs and inflammatory DCs also cross-present antigens delivered via FcγR in certain settings.32, 33, 34 However, the role of FcγR expression, particularly that of CD64, in inflammatory DCs in cross-presentation and priming of T cells remains unclear.
We previously reported that systemic infection with Listeria monocytogenes (LM, a Gram-positive intracellular bacterium that induces the Th1 response) recruits neutrophils, monocytes and natural killer (NK) cells to the T-cell zone of the mouse spleen, where NK cells, positioned around neutrophil/monocyte clusters, are activated to produce IFN-γ, which in turn activates the neighboring monocytes to differentiate into inflammatory DCs.35 The dynamics were subsequently visualized by multi-photon intravital microscopy,36 and similar immune responses were observed in the peritoneum infected with another intracellular parasite, Toxoplasma gondii.37 No prior imaging study has examined the location and dynamics of inflammatory DCs in LNs, and thus understanding of how inflammatory DCs participate in the immune response requires more detailed analysis of inflamed LNs. Therefore, using Listeria as a model pathogen, we investigated myeloid cell subsets in inflamed, skin-draining LNs. We observed that CD64+CD11c+MHCII+ DC-like cells were increased in the LNs of infected mice subcutaneously infected with Listeria. We show that CD64+ cells are a heterogeneous population originated from both monocytes (CD64+ moDCs) and cDCs (CD64+ cDCs). Importantly, although CD64+ moDCs display robust inflammatory gene expression, CD64+ cDCs have a higher potential to stimulate T-cell proliferation and differentiation. Furthermore, CD64+ cDCs show efficient uptake and cross-presentation of a soluble antigen delivered in ICs. These findings delineate the early inflammatory responses that drive the differentiation of two types of inflammatory DCs, including a previously unrecognized population of CD64+ cDCs harboring the cross-presenting ability of ICs.
Materials and methods
Ethics statement
All animal experimental procedures were performed in accordance with the Animal Protection Act of the Ministry of Agriculture, Food and Rural Affairs and the Laboratory Animal Act of the Ministry of Health and Welfare/Ministry of Food and Drug Safety, and approved by the Institutional Animal Care and Use Committee (IACUC; KA2010-21) of the Korea Advanced Institute of Science and Technology (KAIST).
Mice and antibodies
All mice were in C57BL/6 background and housed under specific pathogen-free conditions at the KAIST animal facility. Gender and age-matched mice were used. Ccr7−/− (B6.129P2(C)-Ccr7tm1Rfor/J) and Ccr2−/− (B6.129S4-Ccr2tm1Ifc/J) mice were kindly provided by Dr RM Locksley (UCSF), while Batf3−/− (B6.129S(C)-Batf3tm1Kmm/J) and Ifnar−/− (B6.129S2-Ifnar1tm1Agt/Mmjax) mice were provided by Dr HK Lee (Korea Advanced Institute of Science and Technology). Flt3−/− (Flt3tm1Irl) mice were provided by Dr JH Choi (Hanyang University). The antibodies used are listed in Supplementary Table 1.
Infection and immunization
Live wild-type (WT) Listeria monocytogenes (104 or 2 × 103 CFU; strain, 10403S), 108 CFU of HKLM, 108 CFU of ΔhlyLM for infection and 20 μg of ovalbumin (OVA) or 1, 5 and 20 μg of Alexa Fluor 647-conjugated OVA protein (A647-OVA) emulsified in 40 μl complete Freund’s adjuvant (CFA, Sigma-Aldrich, St Louis, MO, USA) for immunization were subcutaneously injected into mouse footpads.
Cell isolation
Popliteal LNs were passed through a cell strainer (SPL, Pocheon-si, Gyeonggi-do, Korea) to generate single-cell suspensions. For DC isolation, LNs were digested with 1.6 mg ml−1 collagenase IV (Worthington Chemicals, Lakewood, NJ, USA) and 20 μg ml−1 DNase I (Roche, Basel, Switzerland) in Hanks’ Balanced Salt Solution supplemented with 10 mM HEPES, 1 mM MgCl2 and 1.8 mM CaCl2 for 25 min at 37 °C, incubated with 0.01 M EDTA (pH 8.0) for 5 min at room temperature, and processed into a single-cell suspension for antibody staining. Single-cell suspensions were prepared by mechanically disrupting the sample with a 19-Ga needle and then passing the sample through a 70 μm filter.
Flow cytometry and sorting
Cells isolated from LNs were blocked with the anti-CD16/32 antibody (clone 2.4G2) and then stained for surface molecules. For anti-CD64 staining, 5% mouse serum was added. For intracellular staining of IFN-γ and CCR7, the cells were pre-stained for surface molecules, fixed with 3.8% formaldehyde, permeabilized with 0.5% saponin, and then stained with the indicated antibodies. DAPI (4, 6-diamidino-2-phenylindole; Roche) staining or a LIVE/DEAD Fixable Violet Dead Cell Stain Kit (Invitrogen, Carlsbad, CA, USA) was used to exclude dead cells. Lineage (CD3, CD19, NK1.1)-positive cells were excluded from the subsequent analysis of myeloid cells in the LNs. Live cells were counted using counting beads (Invitrogen). The cells were sorted using an ARIA II or ARIA III (BD Biosciences, San Jose, CA, USA). Data were acquired using an LSRFortessa flow cytometer (BD Biosciences) and analyzed with FlowJo software (Treestar, Ashland, OR, USA).
Immunohistochemistry
For the staining of B220, CD11b, CD11c, IFN-γ Listeria, Ly6G, Lyve-1, NK1.1 and PNAd, LNs were harvested, fixed in 2% paraformaldehyde/phosphate-buffered saline for 2 h at 4 °C and embedded in optimal cutting temperature (OCT) compound (Sakura Finetek USA Inc., Torrance, CA, USA). Frozen LNs were cut into 6-μm sections with a Leica cryomicrotome (CM 1850, Leica, Wetzlar, Germany). For the staining of CD64 and CD169, LNs were embedded in OCT compound without fixation, cut into sections and dehydrated in acetone. The staining of 7/4 was performed in both fixed and unfixed sections. LN sections were incubated with 1% H2O2 and 0.1% sodium azide/phosphate-buffered saline for the quenching of endogenous peroxidase when tyramide amplification (Invitrogen) was used, and/or blocked with 5% normal goat serum containing 1% blocking solution (Invitrogen). B220, CD11b, CD169 and 7/4 were stained with fluorescein isothiocyanate (FITC)- or Alexa Fluor 647-conjugated antibodies. Listeria was detected with a rabbit anti-Listeria antibody and visualized with an Alexa Fluor 594-conjugated goat anti-rabbit secondary antibody. Ly6G, Lyve-1 and PNAd were stained with biotin-conjugated primary antibodies and visualized with Alexa Fluor 555-conjugated streptavidin (SA). To detect IFN-γ and NK1.1, we stained LN sections with biotin-conjugated primary antibodies followed by horseradish peroxidase-conjugated SA. The signal was further amplified using tyramide-biotin followed by Alexa Fluor 555-conjugated SA. For CD64 staining, LN sections were blocked with 5% normal goat serum and mouse serum containing a 1% blocking solution and stained with a phycoerythrin (PE)-conjugated anti-CD64 antibody. The signal was further amplified using an horseradish peroxidase-conjugated anti-phycoerythrin antibody, biotin-tyramide and Alexa Fluor 555-conjugated SA. CD11c was visualized with a biotin-conjugated anti-CD11c antibody, and the signal was amplified with horseradish peroxidase-conjugated SA and Alexa Fluor 555-conjugated tyramide. For detecting eYFP+ cells from Clec9a-cre::ROSA-eYFP mixed bone marrow (BM) chimera mice, LN sections were stained with a fluorescein isothiocyanate-conjugated anti-GFP antibody cross-reactive to eYFP (Abcam, Cambridge, UK). The stained sections were mounted with Vectashield (Vector Laboratories, Burlingame, CA, USA). Micrographs were acquired with a Nikon ECLIPSE 80i and analyzed with the NIS-Elements Br software (Nikon Instruments Inc., Tokyo, Japan).
Generation of mixed bone marrow chimeric mice
C57BL/6 (CD45.2) mice were exposed twice to 450 rad of γ- or X-ray irradiation at a 3 h interval and then subjected to tail-vein injection of a mixture of BM cells from WT (CD45.1) and Ccr2−/− (CD45.2) or Flt3−/− (CD45.2) or Clec9a-cre::ROSA-eYFP mice (5 × 105 cells for each, except 1 × 106 cells from Clec9a-cre::ROSA-eYFP BM cells). Mice were used for experiments 6–10 weeks after reconstitution. BM cells from Clec9a-cre::ROSA-eYFP mice were kindly provided by Dr Barbara Schraml (Klinikum der Universität München).
Titration of bacteria
Mice were infected with 2 × 103 CFU WT Listeria monocytogenes by injecting them subcutaneously into the footpads. At the indicated times following infection, draining popliteal LNs were collected and dissolved in 0.1% Triton X-100/phosphate-buffered saline. Cell lysates were serially diluted and plated on brain heart infusion (BHI) agar plates. Bacterial colonies were counted 24 h later.
IC formation
For the experiments of IC-mediated antigen uptake, OVA-rabbit IgG ICs were prepared by incubating A647-OVA (1 μg) and rabbit control (25 μg, Jackson ImmunoResearch Laboratories, West Grove, PA, USA) or anti-OVA (25 μg, Abcam) antibody at 37 °C for 30 min.
In vitro T-cell stimulation
To examine the T-cell-stimulating ability of DC subsets for soluble antigens, mice were immunized with OVA (20 μg)/CFA by subcutaneous injection to footpads, and then 3 days later, DC subsets were sorted from LN cells. OT-II or OT-I cells from uninfected mice were sorted by ARIA II. Sorted T cells were labeled with 5 μM CFDA SE (carboxyfluorescein diacetate, succinimidyl ester; Vybrant CFDA SE cell tracer kit; Invitrogen). Sorted DCs (5 × 103 cells each type) were co-cultured with 2.5 × 104 OT-I cells with or without OVA257–264 peptide (SIINFEKL, 1 μg ml−1) for 3 days or 104 OT-II cells with OVA323–339 peptide (ISQAVHAAHAEINEAGR, 100 ng ml−1) for 5 days in the presence of human IL-2 (50 Uml−1, Peprotech, Rocky Hill, NJ, USA). Proliferation was determined by flow cytometry based on the dilution of CFDA SE. Supernatants were collected, and IFN-γ production was determined by ELISA (BD Biosciences).
To compare cross-priming ability among DC subsets that took up antibody-complexed OVAs, mice were injected with CFA alone on the footpads and popliteal LNs were isolated. The single-cell suspensions of the LNs were prepared by following the procedure for DC isolation. LN cells were plated in tissue culture plates and pulsed with soluble A647-OVA or A647-OVA-IC (equivalent to 1 μg OVA per LN) for 1 h. DCs that took up OVA were assessed as A647+ by flow cytometry. OVA+ DCs (CD64+ moDCs and CD64+ cDCs, 4 × 103 cells) and OT-I cells (2 × 104 cells) were sorted and co-cultured for 4 days in the presence of human IL-2 (50 U ml−1). Proliferation was determined by flow cytometry, based on the dilution of CFDA SE. We observed similar results with OVA-rabbit IgG and OVA-mouse IgG ICs.
Statistical analysis
Data were analyzed using GraphPad PRISM 6 software. A two-tailed Student’s t-test was applied unless otherwise indicated. The results are expressed as the mean±s.e.m. Significance was presented at P-values⩽0.05 (*), ⩽0.01 (**) and ⩽0.001 (***), as indicated in the figures.
Results
Listeria infection induces the formation of intranodal inflammatory foci and the production of IFN-γ
To better understand how the various immune cell types in LNs are choreographed to mount protective immunity during local infection, we infected the footpads of mice with Listeria and analyzed cellular dynamics in skin-draining LNs by IHC and flow cytometry. Subcutaneous infection has been used to track immune responses in sync.9, 38, 39 In uninfected mice, a few CD11b+ myeloid cells are dispersed in the medullary region of the LN. Two days after infection, however, CD11b+ cell clusters were observed at three distinct locations: below the subcapsular sinus (SCS), in the interfollicular area (IFA) and in the paracortex (Figure 1a–d). These CD11b+ cell clusters lasted for 2 or 3 days and dissipated thereafter. Neutrophils (7/4+Ly6G+) were the major cell types detected in the clusters at these early time points, but they disappeared by day 4 (Supplementary Figure 1a and b). Monocytes (7/4+Ly6G−) were observed for a prolonged time in the clusters and spread into the deep paracortex thereafter. Interestingly, CD11c+ cells were observed primarily at the circumferences of the CD11b+ clusters (Supplementary Figure 1c and d). This result is similar to previous reports using subcutaneous infection with various pathogens, which triggers neutrophil swarming38 and the re-distribution of NK cells and IFN-γ production.13, 38, 39, 40, 41
Listeria infection-induced intranodal inflammation. (a–f) Mice were subjected to footpad infection with 2 × 103 wild-type Listeria. Popliteal lymph nodes were collected at the indicated times and sectioned. Fluorescence micrographs of myeloid cells (CD11b), natural killer (NK) cells (NK1.1), B cells (B220), Listeria and IFN-γ are color-matched and shown. The SCS, IFA and paracortex are denoted by stars (*), arrowheads and arrows, respectively. The original magnifications of a, c and e are ×40. Magnified images of the insets in a, c and e at day 2 and 3 are shown in b (×200) and d and f (×100). Scale bars, 200 μm. Data are representative of three independent experiments (N=2 mice per group).
Listeria was detected in LNs on day 1 post infection and cleared by day 4 (data not shown). The Listeria were located within the CD11b+ clusters on days 2 and 3 (Figure 1a and b), which might reflect that the pathogen is trapped in myeloid cell clusters to prevent its spread, as previously illustrated.13 IFN-γ production was detected on days 2–4 (Figure 1c and d). Recruited NK cells produced IFN-γ more robustly than T cells (Supplementary Figure 1e–g). IFN-γ production was tightly associated with CD11b+ cell clusters (Figure 1c and d), and IFN-γ seems to be produced from NK cell clusters in the SCS, IFA and cortical ridge (Figure 1e and f).
The CD11b+ clusters described here coincide with the locations where T cells were shown to be activated.6, 7, 8, 9, 10 Neither heat-killed (HKLM) nor listeriolysin O-deficient (ΔhlyLM) Listeria, which cannot invade the cytosol of the cells but can enter the draining LN, induced the clustering of CD11b+ or NK cells or the production of IFN-γ (Supplementary Figure 1h–j), indicating that intracellular invasion of Listeria is a pre-requisite for the induction of intranodal inflammation.
Collectively, these results show that Listeria infection induces a dynamic re-organization of various immune cell types in LNs, which may both confine Listeria and form core inflammatory foci that regulate the production of IFN-γ.
Time-course analysis of DC differentiation in the inflamed LN
To analyze the dynamic phenotypic changes of the myeloid cells that infiltrate the LNs of Listeria-infected mice, we first examined numerous cell surface markers, including CD64. We first excluded dead and lymphoid cells (T, B and NK cells) and then separated out the neutrophils (Ly6G+CD11b+; Figure 2a and b(i), purple) and inflammatory monocytes (7/4hiLy6Chi; Figure 2a and b(ii), yellow-green). In agreement with previous reports,23, 24, 25, 26, 27, 28 CD64 expression was observed among the inflammatory monocytes (Figure 2a and b(ii, iii), yellow-green). The expression of CD64 was detected as early as day 1 post infection and transiently increased during the course of infection, implying its regulation by the inflammatory milieu. Starting on day 2 post infection, the CD64+ monocytes gradually differentiated into CD11c+MHCII+ DCs, designated here as ‘CD64+ moDCs’ (Figure 2a and b(iv), green). We also observed a dramatic increase of distinct CD64+ cells among 7/4−CD11b+ cells, beginning on day 2 and persisting throughout the course of infection (Figure 2a and b(v), orange; and Supplementary Figure 2a). The majority of the 7/4−CD64+ cells expressed a low level of Ly6C and a high level of MHCII (Figure 2a and b(vi)), and thus resembled the Ly6CloMHCIIhi cells in the previously reported ‘Mo-waterfall’ plot that depicts the progressive differentiation of monocytes into macrophages.24 The CD64+Ly6CloMHCII+ cells were clearly distinct from CD64+ moDCs (Ly6ChiMHCIIint). Although 30–70% of the 7/4+CD64+ monocytes were CD11c+MHCII+ DCs (Figure 2a and b(iv)), ~80% of the CD64+Ly6CloMHCII+ cells expressed CD11c and MHCII (Figure 2a and b(vii)) at levels similar to those seen for CD64−CD11b+ cDCs (CD64− cDCs; Figure 2a and b(viii), blue). Of note, we excluded NK1.1+ cells before analyzing CD11b+ cells as some activated NK cells (B220+, CD11bint)42 also expressed CD64 (Supplementary Figure 2b).
Time-course analysis of myeloid cells that infiltrate Listeria-infected lymph nodes (LNs). (a) A gating scheme to delineate myeloid cells from the popliteal LNs of mice infected with 2 × 103 wild-type Listeria. Cells are color-coded: neutrophils, purple; 7/4hi inflammatory monocytes, yellow-green; CD64+ moDCs, green; 7/4−CD64+ cells, orange; CD64+Ly6CloMHCII+ cells, red; and CD64−CD11b+ cDCs, blue. The cells analyzed and presented in b are marked with Roman numerals (i–viii). (b) Flow cytometric analysis of cells separated and denoted in a. Cells were collected at the indicated days post infection. (c) Quantification of the results presented in b. Data are representative of three independent experiments (mean±s.e.m.; N=4 mice per group).
Quantifications of the LN cells in Listeria-infected mice revealed distinct dynamics of the myeloid cell types during the inflammatory response (Figure 2c and Supplementary Figure 2c). In accordance with the IHC data (Figure 1 and Supplementary Figure 1a–d), neutrophils displayed very transient infiltration of LNs, peaking at day 2 post infection, whereas 7/4+ inflammatory monocytes increased both in number and percentage beginning on day 1. The number and percentage of CD64+ moDCs followed the trend of monocytes with a one-day delay at the initiation. The 7/4−CD64+ and CD64+Ly6CloMHCII+ cells increased beginning on day 2 but subsided after day 3. Finally, CD64− cDCs transiently increased on days 1 and 2, preceding the emergence of CD64+Ly6CloMHCII+ cells. In sum, we found the CD64+Ly6CloMHCII+ DC-like cells that were present predominantly at inflamed LNs and expressed higher levels of MHCII compared to the CD64+ moDCs.
CCR7 is necessary for the recruitment of CD64+Ly6CloMHCII+ cells
Cells enter LNs via two routes: those from peripheral tissues (for example, skin) migrate via lymphatic vessels in a CCR7-dependent manner,43, 44 whereas blood-circulating cells enter LNs through high endothelial venules in a CD62L-dependent manner.45 Previous studies showed that skin-resident DCs migrate to LNs through afferent lymphatic vessels and home near high endothelial venules in the paracortex,46, 47, 48 where T cells are stimulated.6, 7, 8, 9, 10 Our IHC analysis showed that 7/4+CD64+ monocytes and 7/4−CD64+ cells, distinct from 7/4−CD64+CD169+ medullary MPs, were within the CD11b+ clusters in the paracortex (Supplementary Figure 3). CD11b+ cells clustered very near PNAd+ high endothelial venules and Lyve-1+ lymphatic vessels (Supplementary Figure 4a). Cluster-forming monocytes or 7/4−CD64+ cells did not infiltrate through high endothelial venules and their conversion into DC-like cells was CD62L-independent, as the administration of anti-CD62L antibody before Listeria infection decreased lymphocytes (Supplementary Figure 4b), but not monocytes and any of the CD64+ cells, in LNs (Supplementary Figure 4c).
Next, we tested whether CCR7 is required for the recruitment of CD64+ cells in our setting. First, we checked the LNs for resident and migratory DCs in WT and Ccr7-deficient (Ccr7−/−) mice after Listeria infection (Supplementary Figure 4d). Resident DCs were present in Ccr7−/− mice to a comparable level to WT mice. However, many fewer migratory DCs were found in Ccr7−/− mice compared to WT mice, as previously reported.43 On days 2 and 3 post infection, CD11b+ cell clusters and IFN-γ production were observed beneath the SCS and in the IFA in the LNs of Ccr7−/− mice, similar to the pattern seen in WT mice (Supplementary Figure 4e), indicating the dispensable role of CCR7 in CD11b+ cell cluster formation and IFN-γ production, particularly near the SCS. No difference in the infiltration of monocytes or their differentiation to CD64+ moDCs between WT and Ccr7−/− mice was observed (Figure 3a–d). However, we found significant reductions of 7/4−CD64+ cells and CD64+Ly6CloMHCII+ cells in Ccr7−/− mice (Figure 3e–h). The number of CD64− cDCs was also substantially reduced, affirming that they represent skin-originated migratory DCs (Figure 3i and j). The resemblance of CD64+Ly6CloMHCII+ cells to CD64− cDCs, in terms of their CCR7 dependency for migration to LNs, prompted us to investigate the origin of the CD64+Ly6CloMHCII+ cells in more detail.
CCR7 is required for the accumulation of CD64+Ly6CloMHCII+ cells in inflamed lymph nodes (LNs). Wild-type or Ccr7−/− mice were infected with 2 × 103 Listeria and popliteal LNs were isolated on the indicated days. Cells were separated and categorized as described in Figure 2. (a, c, e, g and i) Flow cytometric analysis of the indicated cells. Numbers indicate the percentages of the gated cells among the indicated cells. (b, d, f, h and j) Quantification of the indicated cells gated as described in a, c, e, g and i. Data are representative of three independent experiments (mean±s.e.m. of N=4 mice per group).
Intrinsic CCR2 is not required for the accumulation of CD64+Ly6CloMHCII+ cells in inflamed LNs
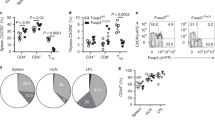
To check whether the CD64+Ly6CloMHCII+ cells originate from Ly6Chi monocytes, we used Ccr2-deficient (Ccr2−/−) mice, which lack Ly6Chi monocytes in blood circulation because their mobilization from BM depends on CCR2.49 As expected, few Ly6Chi monocytes were detected in the blood (data not shown) and LNs (Figure 4a and b) of Ccr2−/− mice. The number of 7/4−CD64+ cells and CD64+Ly6CloMHCII+ cells, but not CD64− cDCs, was also substantially decreased in the LNs (Figure 4c–h), but some remaining 7/4−CD64+ cells were Ly6CloMHCII+ (Figure 4e and f). These results suggest that CD64+Ly6CloMHCII+ cells include CCR2-independent cells, which do not originate from monocytes. To further substantiate this notion, we generated mixed-BM chimeric mice by transferring a 1:1 mixture of BM cells from WT (CD45.1) and Ccr2−/− (CD45.2) mice into irradiated WT mice. We analyzed the number of cells in the LNs 3 days after Listeria infection (Figure 4i). Although Ccr2−/− BM cells were severely compromised in their ability to re-constitute the CD64+ moDC population, the WT and Ccr2−/− cells almost equally re-populated CD64+Ly6CloMHCII+ cells to a level similar to that seen for CD64− cDCs. This result clearly suggests that a majority of CD64+Ly6CloMHCII+ cells originate from cells other than CCR2-dependent monocytes.
Intrinsic CCR2 is dispensable for the accumulation of CD64+Ly6CloMHCII+ cells in lymph nodes (LNs). (a–h) Wild-type (WT) or Ccr2−/− mice were infected with 2 × 103 Listeria and popliteal LNs were isolated on the indicated days. Cells were separated and categorized as described in Figure 2. (a, c, e and g) Flow cytometric analysis of indicated cells. Numbers indicate the percentages of gated cells among the indicated cells. (b, d, f and h) Quantification of the indicated cells gated as described in a, c, e and g. Data are representative of three independent experiments (mean±s.e.m. of N=4 mice per group). (i–k) Mice were reconstituted with a 1:1 mixture of BM cells from WT and Ccr2−/− mice (i) or WT and Flt3−/− (j) or a 1:2 mixture of BM cells from WT and Clec9a-cre::ROSA-eYFP mice (k). They were then infected with 2 × 103 Listeria and LN cells were analyzed 3 days later. (i and j) The ratios between the indicated cells from knockout and WT BM cells are shown. Chimerism was normalized to those of B cells (WT/Ccr2−/− chimera) or neutrophils (WT/Flt3−/− chimera). (k) Percentages of eYFP+ cells among DC subsets are shown. Data are pooled from three (i) or two (k) independent experiments. (j) Data are representative of two independent experiments. Each symbol represents an individual mouse; horizontal lines indicate the mean (±s.e.m. of N=3–6 mice per group per experiment). P-values are shown.
FLT3 dependence and Clec9a-cre-mediated fate mapping of CD64+Ly6CloMHCII+ cells
Next, we generated mixed-BM chimeric mice with WT and Flt3−/− BM cells to test whether the accumulation of CD64+Ly6CloMHCII+ cells in the LNs depends intrinsically on FLT3, a critical growth factor receptor for DC development and homeostasis50 (Figure 4j). We found that while CD64+ moDCs differentiated equally well from both WT and Flt3−/− BM cells, both CD64+Ly6CloMHCII+ cells and CD64− cDCs that originated from Flt3−/− BM cells were present in fewer numbers than those from WT BM cells. We further examined the origin of CD64+Ly6CloMHCII+ cells by generating the mixed-BM chimeric mice with WT and Clec9a-cre::ROSA-eYFP BM cells. In Clec9a-cre::ROSA-eYFP mice, cDC-originated cells, but not monocyte-derived cells, are genetically marked and visualized by the expression of eYFP.51 We found that while eYFP was expressed minimally in CD64+ moDCs, a substantial portion of CD64+Ly6CloMHCII+ cells and CD64− cDCs expressed eYFP similarly (Figure 4k). Our IHC analysis of the LNs of Clec9a-cre::ROSA-eYFP BM chimeric mice demonstrates the localization of CD64+eYFP(Clec9a)− moDCs and CD64+eYFP(Clec9a)+ cells that correspond to CD64+Ly6CloMHCII+ cells within CD64+ cell clusters (Supplementary Figure 5). We noted that CD64+eYFP(Clec9a)+ cells and CD64−eYFP(Clec9a)+ cDCs are positioned at the periphery of CD64+ cell clusters, which coincides with the location of CD11c+ cells (Supplementary Figure 1c and d). Thus, our results on the FLT3 requirement and fate mapping prove the cDC origin of CD64+Ly6CloMHCII+ cells.
Transcriptome analysis reveals the cDC lineage of the CD64+Ly6CloMHCII+ cells
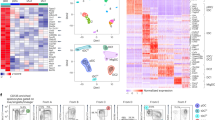
To confirm the cDC lineage of the CD64+Ly6CloMHCII+ cells based on gene expression profiles, we compared CD64+ moDCs, CD64+Ly6CloMHCII+ cells and CD64− cDCs by analyzing core cDC signature molecules and transcriptome analysis (Figure 5). Migratory DC subsets express core cDC signature genes including Flt3, Ccr7, CD26 (Dpp4) and Zbtb46,52 but not Mertk, a macrophage-specific gene.53 Our flow cytometric analysis revealed that CD64+Ly6CloMHCII+ cells and CD64− cDCs displayed almost identical high-level expression profiles of CD26 and FLT3, whereas these markers were expressed at a very low level on CD64+ moDCs (Figure 5a). We could detect intracellular CCR7, but not cell surface CCR7, probably reflecting the internalization of CCR7 during the migration of cells to LNs. CCR7 was detected in CD64+Ly6CloMHCII+ cells and CD64− cDCs, but not in CD64+ moDCs (Figure 5a). A previous study used CD88 to distinguish moDCs from cDCs.54 We found that CD88 is expressed only on the CD64+ moDCs, but not on CD64+Ly6CloMHCII+ cells and CD64− cDCs (Figure 5a). MERTK was detected on some CD64+ moDCs, but not on CD64+Ly6CloMHCII+ cells or CD64− cDCs (Figure 5b). Our findings on the differential expression of lineage marker proteins support the cDC lineage of the CD64+Ly6CloMHCII+ cells.
CD64+Ly6CloMHCII+ cells display gene expression profiles of cDC-lineage cells. (a) The expressions of CD26, FLT3, CCR7 and CD88 were analyzed in the indicated cells using flow cytometry at day 3 post infection. Gray-filled histograms, isotype controls. Data are representative of three independent experiments (N=2 or 3 mice per group). (b) Mice were infected with 2 × 103 Listeria and popliteal lymph nodes (LNs) were isolated on day 3. CD64+ moDCs, CD64+Ly6CloMHCIIhi cells and CD64− cDCs were analyzed for MERTK expression by flow cytometry. Numbers indicate the percentages of MERTK-expressing cells. (c–e) DCs from infected (CD64+ moDCs, CD64+Ly6CloMHCII+ cells and CD64− cDCs) and uninfected (steady-state (s.s.)) migratory CD11b+ cDCs. LNs were sorted and subjected to RNA sequencing (RNA-seq). Pearson correlation coefficients were calculated among the cDC signature gene expression profiles (c) and whole transcript expression profiles (d) of the DC subsets. Numbers indicate the correlation coefficients. (e) Principal component analysis of whole-gene expression profiles of the DC subsets. The percentage of the total variance of the genes representing each axis is shown in parentheses. Each dot represents an individual sample and is color-coded for each DC type.
To further establish the lineage relationships among CD64+ moDCs, CD64− cDCs and CD64+Ly6CloMHCII+ cells, we sorted them from Listeria-infected LNs at day 3 and CD11b+ migratory cDCs from uninfected mice. Then, we performed whole-transcriptome analysis through RNA sequencing of the sorted cells (Supplementary Figure 6). Correlation analysis with core cDC signature gene sets 52 and whole transcripts revealed that CD64+Ly6CloMHCII+ cells showed higher correlation coefficients with CD64− cDCs and steady-state CD11b+ cDCs than with CD64+ moDCs (Figure 5c and d). Principal component analysis also confirmed that the lineage relationships were closer among the cDC subsets than with the CD64+ moDCs (Figure 5e). Altogether, our findings establish the cDC lineage of the CD64+Ly6CloMHCII+ cells. Therefore, we hereinafter term them ‘CD64+ cDCs.’
Type I IFN receptor signaling contributes to CD64+ cDC generation
A scatter plot analysis revealed genes showing >2-fold differences in expression (Figure 6a and Supplementary Table 2). There were only ~800 differentially expressed genes between CD64+ cDCs and CD64− cDCs, indicating their high degree of relatedness. A heat map analysis of differentially expressed genes with ⩾3-fold differences in expression yielded six distinct gene clusters (Figure 6b). The genes in cluster 1, which were more highly expressed in all cDC populations compared to the CD64+ moDCs, included core cDC signature genes, such as Ccr7 and Zbtb46, and transcription factors that are necessary for DC development (for example, Relb, Id2, Irf4 and Batf3). Cluster 4, and particularly cluster 5, included genes related to the inflammatory immune responses, innate immune signaling and cell death (for example, Tlrs, Ddx58, Dhx58, Nlrp3 and Bid), suggesting that the CD64+ moDCs play major roles in these responses.
Type I IFN receptor signaling enables CD64+ cDC generation. (a) Scatter plots of whole-gene expression the DC subsets. The average gene expression of duplicates of each DC subset is shown. The numbers of genes showing <1.5-fold difference in expression between subsets are shown in black. The genes showing ⩾2-fold difference in expression are numbered and color-matched with the relevant DC subsets. RNAs were analyzed from biological duplicates of each DC subset. There were more genes showing ⩾2-fold differences in expression when we compared each cDC subset to the CD64+ moDCs, versus comparing the cDC subsets to one another. (b) Heat map of differentially expressed genes (those whose expression levels showed a ⩾3-fold change between cell types and a <3-fold difference between duplicates). Genes were grouped into six clusters and are listed along with their gene ontologies in Supplementary Table 2. (c and d) Wild-type (WT) or Batf3−/− mice were infected with 2 × 103 Listeria and popliteal lymph nodes (LNs) were collected at day 3 (c) or day 1 and 2 (d) after infection. (c) Cells were analyzed by flow cytometry. (d) Viable bacteria were quantified. Each symbol represents an individual mouse; horizontal lines indicate the mean (±s.e.m. of N=3 mice per group). (e–g) WT or Ifnar−/− mice were infected with 2 × 103 Listeria and popliteal LNs were collected at day 2 (e and f) or day 1 and 2 (g) after infection. (e and f) Cells were analyzed by flow cytometry. (e) The cell numbers of CD64+ moDCs and 7/4−CD64+ cells within the LNs are shown in the graphs. (f) The sum of CD64− cDCs (blue bar) and CD64+ cDCs (red bar) cell numbers is shown in the graphs. Mean±s.e.m. of N=4 mice per group. (g) Viable bacteria were quantified. Data are representative of two independent experiments. Each symbol represents an individual mouse; horizontal lines indicate the mean (±s.e.m. of N=3 mice per group).
BATF3 is a major transcription factor required for the development of CD8α+ and its related CD103+ cDCs but dispensable for that of CD11b+ cDCs, although it is expressed in both CD8α+ and CD11b+ cDCs.55, 56 We found that all three CD11b+ cDC populations, but not CD64+ moDCs, express BATF3 (Figure 6b) and are present in LNs of Batf3−/− mice to a comparable level to WT mice (Figure 6c). Interestingly, unlike the systemic infection of Listeria which enters the spleen via CD8α+ DCs,55, 57 we found no decrease of Listeria entry to LNs in Batf3−/− mice, indicating that CD8α+ or CD103+ cDCs do not serve as a portal for Listeria to enter LNs (Figure 6d).
Genes involved in type I IFN receptor signaling (for example, Stat1, Stat2 and Irf9) were upregulated in all DC subsets under inflammatory conditions (Figure 6b; cluster 4). As previous studies showed that Listeria infection induced type I IFNs,58, 59, 60 we tested the role of type I IFN receptor signaling in DC subset differentiation in WT and Ifnar−/− mice at 2 days after infection with Listeria (Figure 6e–g). Type I IFN receptor signaling was dispensable for the development of CD64+ moDCs, but there were fewer CD64+ cDCs in Ifnar−/− mice than in WT mice. Notably, the number of CD64− cDCs was correspondingly increased, such that there was no change in the sum of CD64+ and CD64− cDCs (Figure 6f), suggesting the possibility that CD64− cDCs acquire CD64 in response to type I IFN receptor signaling. This decrease of CD64+ cDC differentiation was not due to fewer Listeria delivered to the LNs, because we found no difference in the bacterial load in the LNs between WT and Ifnar−/− mice (Figure 6g). Our study suggests that inflammatory signals drive CD11b+ cDCs to differentiate into CD64+ cDCs that induce distinct gene expression from monocytes or moDCs.
CD64+ cDCs are potent antigen presenting cells that are highly efficient in cross-presenting soluble and IC antigens
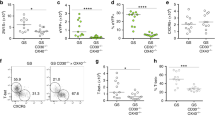
We examined whether CD64+ cDCs can be generated by a common immunization protocol using CFA. We immunized mice on the footpad with OVA protein admixed with CFA and analyzed the popliteal LNs 3 days later. We detected three DC subsets, CD64+ moDCs, CD64+ cDCs and CD64− cDCs (Figure 7a), that displayed CD26, FLT3 and CCR7 expression profiles similar to those observed in Listeria-infected LNs (Figure 7b).
CD64+ cDCs are potent antigen-presenting cells capable of cross-priming CD8+ T cells. (a) Mice were subjected to footpad immunization with OVA (20 μg) in CFA, popliteal lymph node (LN) cells were obtained on day 3 and DC subsets were analyzed by flow cytometry. Cells were analyzed by flow cytometry and designated as in Figure 2. (b) Expressions of the cDC signature molecules (CD26, FLT3 and CCR7) among the indicated cells. Gray-filled histograms, isotype controls. Data are representative of three independent experiments (N=2 or 3 mice per group). (c and d) CD64+ moDCs, CD64+ cDCs and CD64− cDCs (5 × 103) were sorted from LNs of mice immunized as described in (a) and co-cultured with CFDA SE-labeled OT-I cells (2.5 × 104) for 3 days with or without 1 μg ml−1 OVA257–264 (SIINFEKL) peptides (c) or OT-II cells (104) for 5 days with 100 ng ml−1 OVA323–339 peptides (d). (c) Proliferation of OT-I cells assessed by CFDA SE dilution. Gray-filled histograms, OT-I cells only. Numbers indicate the percentage of proliferating cells, indicated by CFDA SE dilution. (d) Upper panels: proliferation of OT-II cells was assessed by CFDA SE dilution. DC subsets are shown and color-coded; gray-filled histograms, OT-II cells only. Lower graph: the amount of IFN-γ in the culture supernatants is measured by ELISA. Data are representative of three independent experiments. (e) Mice were subjected to footpad injection with CFA. Popliteal LN cells were obtained on day 3 and pulsed with A647-OVA alone (1 μg, white) or OVA-ICs prepared by incubating A647-OVA (1 μg) and rabbit control (25 μg, gray) or anti-OVA antibody (25 μg, black) for 3 h in vitro. Percentages of A647-OVA+ cells among DC subsets are shown. Data are representative of three independent experiments with biological triplicates (mean±s.e.m.). (f) LN cells were prepared and treated as described in e, but with a 1 h pulse of OVA-IC. A647-OVA+ DCs (4 × 103) were sorted and co-cultured with CFDA SE-labeled OT-I cells (2 × 104) for 4 days. Proliferation of OT-I cells was assessed by CFDA SE dilution. DC subsets are shown and color-coded; gray-filled histograms, OT-I cells only. Data are representative of three independent experiments.
Next, we assessed the T-cell-stimulating capacity of the DC subsets. Three days after mice were immunized with OVA/CFA, we sorted CD64+ moDCs, CD64+ cDCs and CD64− cDCs from the total LN cells and co-cultured the sorted cells with OVA-specific transgenic CD8+ (OT-I) or CD4+ (OT-II) T cells. Remarkably, CD64+ cDCs and CD64− cDCs, but not CD64+ moDCs, cross-primed OT-I cells without exogenously added SIINFEKL antigenic peptides, although all cell types were capable of inducing T-cell proliferation in the presence of SIINFEKL peptides (Figure 7c). Similarly, CD64+ cDCs and CD64− cDCs induced better OT-II cell proliferation and IFN-γ production compared to CD64+ moDCs (Figure 7d). Next, we examined antigen uptake by the DC subsets (Supplementary Figure 7a–c). We traced cells from inflamed LNs after pulse with fluorophore-conjugated OVA (A647-OVA) by flow cytometry. MoDCs were most efficient in taking up OVA, and CD64− cDCs were the poorest (Supplementary Figure 7a). However, CD64+ cDCs were the largest population among OVA+ DC subsets in cell numbers (Supplementary Figure 7b), reflecting their large occupancy in DC populations (Supplementary Figure 7c). The expression of MHCII and a co-stimulatory molecule, CD86, at higher levels on CD64+ and CD64− cDCs than that on moDCs, as observed both in Listeria infection and OVA/CFA immunization, may contribute to the more potent T cell-stimulation by cDCs (Supplementary Figure 7d).
CD64 is an Fcγ receptor that mediates the internalization of antigen–antibody ICs. To compare the ability of the three DC subsets to take up ICs, whole LN cells were isolated from mice injected with CFA alone and incubated in vitro with an IC of A647-OVA and control or anti-OVA antibody. OVA-ICs, but not OVA plus control IgG, increased dramatically the uptake of OVA by CD64+ moDCs and CD64+ cDCs, but not by CD64− cDCs, compared to OVA alone (Figure 7e). Next, we examined the capacity of CD64+ DCs to cross-present antibody-complexed antigens. CFA-inflamed LN cells were pulsed with OVA-IC in vitro and then OVA+CD64+ DCs were sorted and co-cultured with OT-I cells (Figure 7f). Both CD64+ moDCs and CD64+ cDCs induced the proliferation of OT-I cells; however, CD64+ cDCs were superior in cross-priming antibody-complexed antigens than CD64+ moDCs, although the latter cells were better at IC uptake. Overall, these findings suggest that CD64+ cDCs induced by infection or CFA immunization serve as potent antigen-presenting cells that are highly capable of cross-priming both soluble and antibody-complexed antigens.
Discussion
This report describes the highly coordinated regulation of inflammatory responses in bacterially infected LNs. We show that local Listeria infection through the skin induced the formation of CD11b+ myeloid cell clusters composed of neutrophils and inflammatory monocytes in draining LNs, and triggered IFN-γ production by NK cells surrounding these clusters. This inflammatory response in LNs appears to be correlated to Th1 differentiation (data not shown). Importantly, we show that inflammatory conditions generate two types of CD64+ DCs: CD64+ moDCs and CD64+ cDCs. Flow cytometry showed that the expression of the core cDC signature molecules was observed on CD64+ cDCs but not on CD64+ moDCs. Conversely, the macrophage signature protein, MERTK, was detected on CD64+ moDCs but not on CD64+ cDCs. For recruitment to LNs, CCR7 is required by CD64+ cDCs but not by CD64+ moDCs, whereas intrinsic CCR2 is absolutely required by CD64+ moDCs but not by CD64+ cDCs. CD64+ cDCs, but not CD64+ moDCs, depend on FLT3 for LN accumulation and can be marked by the cDC lineage-tracer gene, Clec9a. Our transcriptome analysis further corroborated that CD64+ cDCs are more closely related to CD64− cDCs than to CD64+ moDCs. Ifnar−/− mice were characterized by a decrease in CD64+ cDCs and a corresponding increase in CD64− cDCs, but no change in CD64+ moDCs. Finally, CD64+ cDCs were capable of stimulating T-cell proliferation and differentiation, and cross-priming CD8+ T cells, whereas CD64+ moDCs were much less effective in this regard. This result is in line with a previous study reporting the different T-cell stimulating ability between moDCs and cDCs.54 Altogether, for the first time, our comprehensive analysis of the various CD64+ cells provides convincing evidence for bona fide cDC-originated CD64+Ly6CloMHCII+ cells that act as very potent T-cell stimulators (Supplementary Table 3).
Previous studies used Clec9a (DNGR-1) and Zbtb46 to genetically trace cDC-lineage cells.51, 61, 62 In Clec9a-cre::ROSA-eYFP mice, some CD64+CD11c+MHCII+ cells in the kidney were shown to be eYFP-labeled and derived from common DC precursors.51 We also verified the cDC origin of CD64+ cDCs by using mixed-BM chimera generated with BM cells of WT and Clec9a-cre::ROSA-eYFP or WT and Flt3−/−mice. It was reported that inflammation in the colon recruits two types of CX3CR1-GFPint cells to the intestine in CX3CR1-GFP mice:61 Ly6CloMHCIIhi and Ly6ChiMHCIIint cells. Microarray analysis showed that the Ly6Clo cells exhibited gene expression profiles that were quite distinct from those of Ly6Chi cells. Moreover, Ly6Clo cells displayed the expression of cDC signature genes (Ccr7, Flt3 and Zbtb46) and stimulated naive T cells better than Ly6Chi cells. Thus, we postulate that the inflammation-induced intestinal Ly6CloCX3CR1-GFPint cells (CD11chiMHCIIhi) and Ly6ChiCX3CR1-GFPint cells (CD11clo/intMHCIIint) would be equivalent to the CD64+ cDCs and CD64+ moDCs described herein, respectively. Unfortunately, our numerous attempts at adoptive transfer to show that CD64+ cDCs were derived from DC precursors but not monocytes failed, probably because inflammatory conditions yield an environment that is unfavorable for the survival or differentiation of transferred cells. Recently, GM-CSF-cultured BM cells were found to be composed of cDCs (GM-DCs) and MPs (GM-Macs).63 These GM-DCs resembled our CD64+ cDCs in terms of their surface expression of CD64, core cDC signature genes and T-cell-stimulating ability. Interestingly, our mixed-BM chimeric mice of WT and Ccr2−/− cells exhibited lower (although not significantly different) chimerism of the CD64+Ly6CloMHCII+ cells compared to CD64− cDCs (Figure 4i), probably reflecting the heterogeneity of the CD64+ cells. Furthermore, we found that some, but not all, CD64+ moDCs express MERTK. As the co-expression of CD64 and MERTK has been used to define macrophages, a portion of CD64+ moDCs may contain cells that can be classified as macrophages. In the future, questions on the ontogenic relationships between the aforementioned DC and macrophage-like cells should be answered through the development of more sophisticated mouse tools.
We show that CD64+ cDCs express higher levels of MHCII and constitute the largest DC population in inflamed LNs; it is likely that they serve as major antigen-presenting cells for CD4+ T cells. Importantly, we demonstrate that CD64+ cDCs are capable of cross-presenting to CD8+ T cells, in particular, low dose antigens complexed with antibodies, indicating that they are specialized to concentrate minute antigens through FcγRs. Although CD8α+ or CD103+ cDCs got attention from past studies for their efficient cross-presentation, CD11b+ cDCs have also been shown to be capable of cross-presenting antigens which are complexed with cognate antibodies.64, 65, 66 As a part of CD11b+ cDCs, the function of CD64+ cDCs described in our study is in line with previous studies. As we showed that Ab-complexed antigens could be taken up more efficiently by CD64-expressing cells, we speculate that CD64+ cDCs might play a role in a secondary response to infection by utilizing antibodies that are generated against the infectious agent in primary infection. Pre-existing antibodies could form ICs and facilitate the uptake of pathogens by DCs via Fc receptors, which will lead to the activation of T cells at a lower concentration of antigens. In contrast, CD64+ moDCs, which showed a more robust inflammatory response program in our transcriptome analysis, may serve to eliminate pathogens and provide the inflammatory milieu. In concert, our findings shed new light on the inflammatory responses in LNs that can drive the differentiation of multiple types of DCs. Further investigations are needed to examine how the core competencies of the different DC subsets cooperate to orchestrate the innate and adaptive immune responses.
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Change history
13 April 2018
This article has been corrected since Online Publication and a correction has also been published.
References
Matsuno K, Ueta H, Shu Z, Xue-Dong X, Sawanobori Y, Kitazawa Y et al. The microstructure of secondary lymphoid organs that support immune cell trafficking. Arch Histol Cytol 2010; 73: 1–21.
Willard-Mack CL . Normal structure, function, and histology of lymph nodes. Toxicol Pathol 2006; 34: 409–424.
Qi H, Kastenmuller W, Germain RN . Spatiotemporal basis of innate and adaptive immunity in secondary lymphoid tissue. Annu Rev Cell Dev Biol 2014; 30: 141–167.
Cahalan MD, Parker I . Choreography of cell motility and interaction dynamics imaged by two-photon microscopy in lymphoid organs. Annu Rev Immunol 2008; 26: 585–626.
Bousso P . T-cell activation by dendritic cells in the lymph node: lessons from the movies. Nat Rev Immunol 2008; 8: 675–684.
Hickman HD, Takeda K, Skon CN, Murray FR, Hensley SE, Loomis J et al. Direct priming of antiviral CD8+ T cells in the peripheral interfollicular region of lymph nodes. Nat Immunol 2008; 9: 155–165.
Kastenmuller W, Brandes M, Wang Z, Herz J, Egen JG, Germain RN . Peripheral prepositioning and local CXCL9 chemokine-mediated guidance orchestrate rapid memory CD8+ T cell responses in the lymph node. Immunity 2013; 38: 502–513.
Groom JR, Richmond J, Murooka TT, Sorensen EW, Sung JH, Bankert K et al. CXCR3 chemokine receptor-ligand interactions in the lymph node optimize CD4+ T helper 1 cell differentiation. Immunity 2012; 37: 1091–1103.
Chtanova T, Han SJ, Schaeffer M, van Dooren GG, Herzmark P, Striepen B et al. Dynamics of T cell, antigen-presenting cell, and pathogen interactions during recall responses in the lymph node. Immunity 2009; 31: 342–355.
Sung JH, Zhang H, Moseman EA, Alvarez D, Iannacone M, Henrickson SE et al. Chemokine guidance of central memory T cells is critical for antiviral recall responses in lymph nodes. Cell 2012; 150: 1249–1263.
Gerner MY, Kastenmuller W, Ifrim I, Kabat J, Germain RN . Histo-cytometry: a method for highly multiplex quantitative tissue imaging analysis applied to dendritic cell subset microanatomy in lymph nodes. Immunity 2012; 37: 364–376.
Kissenpfennig A, Henri S, Dubois B, Laplace-Builhe C, Perrin P, Romani N et al. Dynamics and function of Langerhans cells in vivo: dermal dendritic cells colonize lymph node areas distinct from slower migrating Langerhans cells. Immunity 2005; 22: 643–654.
Kastenmuller W, Torabi-Parizi P, Subramanian N, Lammermann T, Germain RN . A spatially-organized multicellular innate immune response in lymph nodes limits systemic pathogen spread. Cell 2012; 150: 1235–1248.
Turley SJ, Fletcher AL, Elpek KG . The stromal and haematopoietic antigen-presenting cells that reside in secondary lymphoid organs. Nat Rev Immunol 2010; 10: 813–825.
Liao S, Ruddle NH . Synchrony of high endothelial venules and lymphatic vessels revealed by immunization. J Immunol 2006; 177: 3369–3379.
Malhotra D, Fletcher AL, Turley SJ . Stromal and hematopoietic cells in secondary lymphoid organs: partners in immunity. Immunol Rev 2013; 251: 160–176.
Soderberg KA, Payne GW, Sato A, Medzhitov R, Segal SS, Iwasaki A . Innate control of adaptive immunity via remodeling of lymph node feed arteriole. Proc Natl Acad Sci USA 2005; 102: 16315–16320.
Webster B, Ekland EH, Agle LM, Chyou S, Ruggieri R, Lu TT . Regulation of lymph node vascular growth by dendritic cells. J Exp Med 2006; 203: 1903–1913.
Serbina NV, Salazar-Mather TP, Biron CA, Kuziel WA, Pamer EG . TNF/iNOS-producing dendritic cells mediate innate immune defense against bacterial infection. Immunity 2003; 19: 59–70.
Randolph GJ, Inaba K, Robbiani DF, Steinman RM, Muller WA . Differentiation of phagocytic monocytes into lymph node dendritic cells in vivo. Immunity 1999; 11: 753–761.
Segura E, Amigorena S . Inflammatory dendritic cells in mice and humans. Trends Immunol 2013; 34: 440–445.
Merad M, Sathe P, Helft J, Miller J, Mortha A . The dendritic cell lineage: ontogeny and function of dendritic cells and their subsets in the steady state and the inflamed setting. Annu Rev Immunol 2013; 31: 563–604.
Langlet C, Tamoutounour S, Henri S, Luche H, Ardouin L, Gregoire C et al. CD64 expression distinguishes monocyte-derived and conventional dendritic cells and reveals their distinct role during intramuscular immunization. J Immunol 2012; 188: 1751–1760.
Tamoutounour S, Henri S, Lelouard H, de Bovis B, de Haar C, van der Woude CJ et al. CD64 distinguishes macrophages from dendritic cells in the gut and reveals the Th1-inducing role of mesenteric lymph node macrophages during colitis. Eur J Immunol 2012; 42: 3150–3166.
Tamoutounour S, Guilliams M, Montanana Sanchis F, Liu H, Terhorst D, Malosse C et al. Origins and functional specialization of macrophages and of conventional and monocyte-derived dendritic cells in mouse skin. Immunity 2013; 39: 925–938.
Bain CC, Scott CL, Uronen-Hansson H, Gudjonsson S, Jansson O, Grip O et al. Resident and pro-inflammatory macrophages in the colon represent alternative context-dependent fates of the same Ly6Chi monocyte precursors. Mucosal Immunol 2013; 6: 498–510.
Schlitzer A, McGovern N, Teo P, Zelante T, Atarashi K, Low D et al. IRF4 transcription factor-dependent CD11b+ dendritic cells in human and mouse control mucosal IL-17 cytokine responses. Immunity 2013; 38: 970–983.
Plantinga M, Guilliams M, Vanheerswynghels M, Deswarte K, Branco-Madeira F, Toussaint W et al. Conventional and monocyte-derived CD11b(+) dendritic cells initiate and maintain T helper 2 cell-mediated immunity to house dust mite allergen. Immunity 2013; 38: 322–335.
Regnault A, Lankar D, Lacabanne V, Rodriguez A, Thery C, Rescigno M et al. Fcgamma receptor-mediated induction of dendritic cell maturation and major histocompatibility complex class I-restricted antigen presentation after immune complex internalization. J Exp Med 1999; 189: 371–380.
Platzer B, Stout M, Fiebiger E . Antigen cross-presentation of immune complexes. Front Immunol 2014; 5: 140.
Nierkens S, Tel J, Janssen E, Adema GJ . Antigen cross-presentation by dendritic cell subsets: one general or all sergeants? Trends Immunol 2013; 34: 361–370.
Platt CD, Ma JK, Chalouni C, Ebersold M, Bou-Reslan H, Carano RA et al. Mature dendritic cells use endocytic receptors to capture and present antigens. Proc Natl Acad Sci USA 2010; 107: 4287–4292.
den Haan JM, Bevan MJ . Constitutive versus activation-dependent cross-presentation of immune complexes by CD8(+) and CD8(−) dendritic cells in vivo. J Exp Med 2002; 196: 817–827.
Baker K, Qiao SW, Kuo TT, Aveson VG, Platzer B, Andersen JT et al. Neonatal Fc receptor for IgG (FcRn) regulates cross-presentation of IgG immune complexes by CD8-CD11b+ dendritic cells. Proc Natl Acad Sci USA 2011; 108: 9927–9932.
Kang SJ, Liang HE, Reizis B, Locksley RM . Regulation of hierarchical clustering and activation of innate immune cells by dendritic cells. Immunity 2008; 29: 819–833.
Waite JC, Leiner I, Lauer P, Rae CS, Barbet G, Zheng H et al. Dynamic imaging of the effector immune response to listeria infection in vivo. PLoS Pathog 2011; 7: e1001326.
Goldszmid RS, Caspar P, Rivollier A, White S, Dzutsev A, Hieny S et al. NK cell-derived interferon-gamma orchestrates cellular dynamics and the differentiation of monocytes into dendritic cells at the site of infection. Immunity 2012; 36: 1047–1059.
Chtanova T, Schaeffer M, Han SJ, van Dooren GG, Nollmann M, Herzmark P et al. Dynamics of neutrophil migration in lymph nodes during infection. Immunity 2008; 29: 487–496.
Coombes JL, Han SJ, van Rooijen N, Raulet DH, Robey EA . Infection-induced regulation of natural killer cells by macrophages and collagen at the lymph node subcapsular sinus. Cell Rep 2012; 2: 124–135.
Bajenoff M, Breart B, Huang AY, Qi H, Cazareth J, Braud VM et al. Natural killer cell behavior in lymph nodes revealed by static and real-time imaging. J Exp Med 2006; 203: 619–631.
Garcia Z, Lemaitre F, van Rooijen N, Albert ML, Levy Y, Schwartz O et al. Subcapsular sinus macrophages promote NK cell accumulation and activation in response to lymph-borne viral particles. Blood 2012; 120: 4744–4750.
Blasius AL, Barchet W, Cella M, Colonna M . Development and function of murine B220+CD11c+NK1.1+ cells identify them as a subset of NK cells. J Exp Med 2007; 204: 2561–2568.
Ohl L, Mohaupt M, Czeloth N, Hintzen G, Kiafard Z, Zwirner J et al. CCR7 governs skin dendritic cell migration under inflammatory and steady-state conditions. Immunity 2004; 21: 279–288.
Forster R, Schubel A, Breitfeld D, Kremmer E, Renner-Muller I, Wolf E et al. CCR7 coordinates the primary immune response by establishing functional microenvironments in secondary lymphoid organs. Cell 1999; 99: 23–33.
Girard JP, Moussion C, Forster R . HEVs, lymphatics and homeostatic immune cell trafficking in lymph nodes. Nat Rev Immunol 2012; 12: 762–773.
Sixt M, Kanazawa N, Selg M, Samson T, Roos G, Reinhardt DP et al. The conduit system transports soluble antigens from the afferent lymph to resident dendritic cells in the T cell area of the lymph node. Immunity 2005; 22: 19–29.
Bajenoff M, Granjeaud S, Guerder S . The strategy of T cell antigen-presenting cell encounter in antigen-draining lymph nodes revealed by imaging of initial T cell activation. J Exp Med 2003; 198: 715–724.
Lindquist RL, Shakhar G, Dudziak D, Wardemann H, Eisenreich T, Dustin ML et al. Visualizing dendritic cell networks in vivo. Nat Immunol 2004; 5: 1243–1250.
Serbina NV, Pamer EG . Monocyte emigration from bone marrow during bacterial infection requires signals mediated by chemokine receptor CCR2. Nat Immunol 2006; 7: 311–317.
Waskow C, Liu K, Darrasse-Jeze G, Guermonprez P, Ginhoux F, Merad M et al. The receptor tyrosine kinase Flt3 is required for dendritic cell development in peripheral lymphoid tissues. Nat Immunol 2008; 9: 676–683.
Schraml BU, van Blijswijk J, Zelenay S, Whitney PG, Filby A, Acton SE et al. Genetic tracing via DNGR-1 expression history defines dendritic cells as a hematopoietic lineage. Cell 2013; 154: 843–858.
Miller JC, Brown BD, Shay T, Gautier EL, Jojic V, Cohain A et al. Deciphering the transcriptional network of the dendritic cell lineage. Nat Immunol 2012; 13: 888–899.
Gautier EL, Shay T, Miller J, Greter M, Jakubzick C, Ivanov S et al. Gene-expression profiles and transcriptional regulatory pathways that underlie the identity and diversity of mouse tissue macrophages. Nat Immunol 2012; 13: 1118–1128.
Nakano H, Moran TP, Nakano K, Gerrish KE, Bortner CD, Cook DN . Complement receptor C5aR1/CD88 and dipeptidyl peptidase-4/CD26 define distinct hematopoietic lineages of dendritic cells. J Immunol 2015; 194: 3808–3819.
Edelson BT, Kc W, Juang R, Kohyama M, Benoit LA, Klekotka PA et al. Peripheral CD103+ dendritic cells form a unified subset developmentally related to CD8alpha+ conventional dendritic cells. J Exp Med 2010; 207: 823–836.
Hildner K, Edelson BT, Purtha WE, Diamond M, Matsushita H, Kohyama M et al. Batf3 deficiency reveals a critical role for CD8alpha+ dendritic cells in cytotoxic T cell immunity. Science 2008; 322: 1097–1100.
Neuenhahn M, Kerksiek KM, Nauerth M, Suhre MH, Schiemann M, Gebhardt FE et al. CD8alpha+ dendritic cells are required for efficient entry of Listeria monocytogenes into the spleen. Immunity 2006; 25: 619–630.
O'Connell RM, Saha SK, Vaidya SA, Bruhn KW, Miranda GA, Zarnegar B et al. Type I interferon production enhances susceptibility to Listeria monocytogenes infection. J Exp Med 2004; 200: 437–445.
Auerbuch V, Brockstedt DG, Meyer-Morse N, O'Riordan M, Portnoy DA . Mice lacking the type I interferon receptor are resistant to Listeria monocytogenes. J Exp Med 2004; 200: 527–533.
Carrero JA, Calderon B, Unanue ER . Type I interferon sensitizes lymphocytes to apoptosis and reduces resistance to Listeria infection. J Exp Med 2004; 200: 535–540.
Zigmond E, Varol C, Farache J, Elmaliah E, Satpathy AT, Friedlander G et al. Ly6C hi monocytes in the inflamed colon give rise to proinflammatory effector cells and migratory antigen-presenting cells. Immunity 2012; 37: 1076–1090.
Meredith MM, Liu K, Darrasse-Jeze G, Kamphorst AO, Schreiber HA, Guermonprez P et al. Expression of the zinc finger transcription factor zDC (Zbtb46, Btbd4) defines the classical dendritic cell lineage. J Exp Med 2012; 209: 1153–1165.
Helft J, Bottcher J, Chakravarty P, Zelenay S, Huotari J, Schraml BU et al. GM-CSF mouse bone marrow cultures comprise a heterogeneous population of CD11c(+)MHCII(+) macrophages and dendritic cells. Immunity 2015; 42: 1197–1211.
Baker K, Rath T, Lencer WI, Fiebiger E, Blumberg RS . Cross-presentation of IgG-containing immune complexes. Cell Mol Life Sci 2013; 70: 1319–1334.
Schuurhuis DH, van Montfoort N, Ioan-Facsinay A, Jiawan R, Camps M, Nouta J et al. Immune complex-loaded dendritic cells are superior to soluble immune complexes as antitumor vaccine. J Immunol 2006; 176: 4573–4580.
de Jong JM, Schuurhuis DH, Ioan-Facsinay A, van der Voort EI, Huizinga TW, Ossendorp F et al. Murine Fc receptors for IgG are redundant in facilitating presentation of immune complex derived antigen to CD8+ T cells in vivo. Mol Immunol 2006; 43: 2045–2050.
Acknowledgements
This study was supported by grants from the National Research Foundation of Korea (NRF-2012M3A9B4027955, NRF-2017R1A2B4007817).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Experimental & Molecular Medicine website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Min, J., Yang, D., Kim, M. et al. Inflammation induces two types of inflammatory dendritic cells in inflamed lymph nodes. Exp Mol Med 50, e458 (2018). https://doi.org/10.1038/emm.2017.292
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/emm.2017.292
This article is cited by
-
An Organotypic Human Lymph Node Model Reveals the Importance of Fibroblastic Reticular Cells for Dendritic Cell Function
Tissue Engineering and Regenerative Medicine (2024)
-
A synthetic bioactive peptide of the C-terminal fragment of adhesion protein Fibulin7 attenuates the inflammatory functions of innate immune cells in LPS-induced systemic inflammation
Inflammation Research (2024)
-
Lipid A analog CRX-527 conjugated to synthetic peptides enhances vaccination efficacy and tumor control
npj Vaccines (2022)
-
Antigen presentation by dendritic cells and their instruction of CD4+ T helper cell responses
Cellular & Molecular Immunology (2020)