Abstract
In several human tumors, signal transducer and activator of transcription 3 (STAT3) and nuclear factor κB (NFκB) are activated and interact; how these STAT3–NFκB complexes are transported to the nucleus is not fully understood. In this study, we found that Rac1 was activated in starved cancer cells and that activated Rac1 coexisted with STAT3 and NFκB. Rac1 knockdown and overexpression of the dominant-negative mutant Rac1N19 inhibited the degradation of IκBα, an inhibitor of NFκB. MG132, an inhibitor of the ubiquitin proteasome pathway, increased the amount of non-phosphorylated IκBα, but not serine-phosphorylated IκBα, indicating that IκBα degradation by Rac1 in starved cancer cells is independent of IκBα serine phosphorylation by IKK. Rac1 knockdown also inhibited the nuclear translocation of STAT3–NFκB complexes, indicating that this translocation requires activated Rac1. We also demonstrated that the mutant STAT3 Y705F could form complexes with NFκB, and these unphosphorylated STAT3–NFκB complexes translocated into the nucleus and upregulated the activity of NFκB in starved cancer cells, suggesting that phosphorylation of STAT3 is not essential for its translocation. To our knowledge, this is the first study demonstrating the crucial role of Rac1 in the function of STAT3–NFκB complexes in starved cancer cells and implies that targeting Rac1 may have future therapeutic significance in cancer therapy.
Similar content being viewed by others
Introduction
Rac1 belongs to the Rho family of small GTPases, which participates in numerous pathways, including cytoskeleton reorganization, gene transcription, cell proliferation and survival.1, 2 Rac1 binds to signal transducer and activator of transcription 3 (STAT3) at the cell membrane, as well as inside the nucleus in COS-1 and smooth muscle cells treated with growth factors, and it appears to regulate the phosphorylation of tyrosine and serine residues.3, 4 It functions in the nuclear translocation of phosphorylated STATs (p-STATs) and β-catenin, and accumulates in the nucleus during the G2-phase, promoting cell division.5, 6
STAT3, one of seven STAT family members, is activated in response to interleukin-6 (IL6). Many cytokines use the common gp130 receptor to activate the phosphorylation of STAT3 on tyrosine residue 705, leading to the formation of STAT3 dimers through a reciprocal phosphotyrosine–SH2 domain interaction. STAT3 dimers move to the nucleus, a translocation that requires Rac1 and GTPase-activating protein MgcRacGAP,5 and facilitate various transcriptional activities. STAT3 is constitutively activated in nearly all human cancers.7 Target proteins transcribed by activated STAT3 are implicated in the fundamental events of tumor development, including proliferation, survival, invasion and angiogenesis.8
Nuclear factor-κB (NFκB) is a transcription factor associated with cell survival and proliferation, as well as immune and inflammatory responses, and can be activated by both canonical and non-canonical pathways. NFκB activation is triggered by growth factors and cytokines, such as tumor necrosis factor alpha, LTβ, IL1β and lipopolysaccharide, and is closely linked to human tumorigenesis. The genes expressed by NFκB suppress tumor cell death, promote tumor growth and provide tumors with an inflammatory microenvironment. NFκB consists of five members, Rel (c-Rel), RelA (p65), RelB, NFκB 1 (p50 and its precursor p105) and NFκB 2 (p52 and its precursor p100). They form both homo- and heterodimers, of which the best characterized is the p50 and p65 heterodimer. In resting cells, this heterodimer is sequestered in the cytoplasm by its association with the inhibitory subunit IκBα.9 Degradation of IκBα is the main regulatory step of the canonical NFκB pathway, and Rac GTPase regulates IκBα degradation by conveying the SCF (Skp1/Cul-1/F-box protein complex) complex and IκBα to membrane ruffles.10
STAT3 and NFκB interact.11, 12, 13 In our previous study, we found that STAT3 and NFκB are activated simultaneously in cancer cells by an intrinsic mechanism under stressful conditions and that they cooperatively induce various survival factors.12 Moreover, we showed that STAT3 and NFκB exist as identical nuclear complexes on proximal IL6 promoters and that STAT3 has a critical role in not only binding to IL6 promoters but also in retaining NFκB in the nucleus. However, where the STAT3–NFκB complex is formed, whether in the cytoplasm or nucleus, has not been elucidated, and if the STAT3–NFκB complex is formed in the cytoplasm, how is it transported to the nucleus? In this study, we found that activated Rac1 is required for IκBα degradation and the transport of the STAT3–NFκB complex to the nucleus, indicating a novel function of Rac1 GTPase.
Materials and methods
Cells, antibodies and other reagents
HeLa cells (human cervical cancer) were cultured in minimum essential medium supplemented with 10% heat-inactivated fetal bovine serum, 2 mM L-glutamine, 100 U ml−1 penicillin and 100 μg ml−1 streptomycin. Anti-STAT3 Y705, anti-STAT3, anti-IκBα, anti-phospho-IκBα and anti-p50 antibodies (Abs) were purchased from Cell Signaling Biotechnology (Danvers, MA, USA). Anti-p65, anti-HA, and anti-histone H3 Abs, and MG132 were from Santa Cruz Biotechnology (Dallas, TX, USA). The anti-V5 Ab and Hank’s balanced salt solution (HBSS) were from Invitrogen (Grand Island, NY, USA). The anti-Rac1 Ab was from Millipore (Darmstadt, Germany). The anti-tubulin Ab was from Calbiochem (Darmstadt, Germany).
Expression constructs and lentiviral vector transfections
Lentiviral constructs expressing STAT3 short hairpin RNA (shRNA), p65 shRNA and Rac1 shRNA were purchased from Sigma-Aldrich (St. Louis, MO, USA). The wild-type Rac1, Rac1Q61L, Rac1N17, wild-type STAT3, STAT3 Y705F, IKK2 and IKK2 K44M cDNAs were from Addgene (Cambridge, MA, USA). All the cDNA constructs were subcloned into pCDH-EF1-MCS-T2A-Puro, a lentiviral vector for cDNA expression (System Biosciences, Mountain View, CA, USA). Six tandem repeats of the STAT3-responsive element (and NFκB-responsive element were cloned into pGF1-mCMV, a lentiviral reporter vector (System Biosciences). All the lentiviral vectors were transfected into 293TN cells (System Biosciences) with Lipofectamine 2000 transfection reagent (Invitrogen). Particles were collected 2 days after the transfection of the lentiviral plasmids and were used to infect HeLa cells. Lentivirus-infected HeLa cells were puromycin selected for 1 week.
Rac1 GTPase activation assay
Cells were processed and assayed for Rac1 activity using a kit from Cell Biolabs (San Diego, CA, USA). Briefly, cell lysates were incubated with agarose beads coupled to the p21-binding domain of a p21-activated protein kinase. Bound Rac1 was analyzed by western blot using an anti-Rac1 Ab.
Real-time PCR
Cells were collected after treatment at the indicated time points. Total RNA was isolated using an RNeasy kit (Qiagen, Venlo, Netherlands). The PrimeScript RT kit (Takara, Shiga, Japan) was used to reverse transcribe the messenger RNA (mRNA) into cDNA. PCR was carried out using an ABI PRISM 7000 machine (Applied Biosystems, Carlsbad, CA, USA) with SYBR Premix Ex Taq II (Takara). The sequences of primers are as follows: hIL6: For, 5′-AAGCCAGAGCTGTGCAGATGAGTA-3′ and Rev, 5′-TGTCCTGCAGCCACTGGTTC-3′; β-actin: For, 5′-TGGCACCCAGCACAATGAA-3′ and Rev, 5′-CTAAGTCATAGTCCGCCTAGAAGCA-3′; and GFP: For, 5′-CACCCGCATCGAGAAGTACG-3′ and Rev, 5′-GGTGCCACCACCTTGAAGT-3′. Analysis of each sample was performed at least three times for each experiment, and the data in the figures are reported as relative quantitation: average values of 2−ΔΔCT±s.d.
Preparation of cell lysates, western blots and immunoprecipitation
The nuclear and cytoplasmic fractions were prepared using the NE-PER nuclear and cytoplasmic extraction reagents (Pierce, Rockford, IL, USA). For immunoprecipitation, 200 μg of proteins were incubated overnight with anti-p65 Ab or anti-STAT3 Ab, then with protein A agarose beads (Santa Cruz Biotechnology) for 1 h. The beads were washed five times, and the immunoprecipitate was extracted with 2 × SDS sample buffer. For preparing whole-cell lysates, cells were lysed in high-salt lysis buffer (20 mM Tris-HCl (pH 8.0), 1% Triton X-100, 2 mM EDTA and 1 mM phenylmethylsulfonyl fluoride), incubated on ice for 20 min and centrifuged for 20 min to remove cell debris. A total of 20 μg of whole-cell lysate was used in SDS–polyacrylamide gel electrophoresis. The proteins were then electrotransferred to a polyvinylidene difluoride membrane and incubated overnight with Abs at 4 °C. Then, the membranes were incubated with peroxidase-conjugated secondary Abs (Pierce) for 1 h at room temperature, and the signal was detected using an enhanced chemiluminescence detection kit (Amersham Biosciences, Piscataway, NJ, USA).
Confocal microscopy
HeLa cells were grown on Lab-Tek four-well glass chamber slides (NUNC, Rockford, IL, USA) and were incubated in HBSS for the indicated time periods. Cells were fixed and permeabilized with cold methanol for 5 min. They were then washed with phosphate-buffered saline, and incubated with primary Ab and secondary Ab conjugates. Images were collected on a laser scanning confocal microscope LSM710 (Carl Zeiss, Oberkochen, Germany) equipped with argon (488 nm) and krypton (568 nm) lasers, using a × 40 water immersion objective lens. Images were processed with ZEN 2009 light edition (Carl Zeiss).
Results
Rac1 is activated and colocalizes with STAT3, p65 and IκBα in starved cancer cells
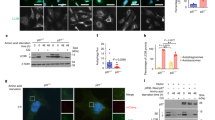
In a previous study, we showed that STAT3 and NFκB are activated in starved cancer cells and that activated STAT3 and NFκB cooperatively induce IL6 expression.12 We also found that STAT3 has critical roles in the nuclear retention of NFκB in starved cancer cells and that NADPH oxidases, activated by autophagic processes, are associated with STAT3 activation.14 p22phox-dependent NADPH oxidase function is dependent on the small GTPase Rac1.15 Rac1 and protein kinase C mediate the activation of STAT3 in endothelial cells following hypoxia–reoxygenation.16 The binding of STAT3 with Rac1 occurs predominantly at the cell membrane, but also inside the nucleus, and occurs through the binding of the coiled-coil domain of STAT3 to the 54 NH2-terminal residues of Rac1. In this study, we found that, in control cells, STAT3 and NFκB were dispersed in the cytoplasm, but in cells exposed to HBSS, STAT3, NFκB and IκBα localized to the membrane ruffles, where IκBα was degraded before STAT3 and NFκB translocated into the nucleus in a time-dependent manner, as demonstrated by confocal microscopy (Figures 1a and b). The small GTPase Rac1 is a regulator of actin assembly, which is essential for membrane ruffles. Taken together, this suggests that the small GTPase Rac1 functions in the activation of STAT3–NFκB complexes in starved cancer cells.
STAT3, p65 and IκBα relocated to the membrane ruffles in starved cancer cells. (a, b) HeLa cells were incubated in HBSS for the indicated time periods. Confocal microscopy analysis showing STAT3, p65 and IκBα (a), and a merged image of STAT3 and p65 (b). Arrows indicate membrane ruffles. DAPI, 4′,6-diamidino-2-phenylindole; HBSS, Hank’s balanced salt solution; STAT3, signal transducer and activator of transcription 3.
To investigate the connection between Rac1 and STAT3–NFκB complexes, we used the previous discovery that activated Rac1 assists NFκB activation by conveying the SCF complex and IκBα to membrane ruffles.10 To verify this finding, we performed a pull-down experiment using the Rac1 effector PAK linked to GST protein. HBSS induced activation of Rac1 in a time-dependent manner (Figure 2a). The next step was to view the colocalization of STAT3, NFκB and Rac1 after HBSS treatment in HeLa cells using confocal microscopy. We found that Rac1, p65, STAT3 and IκBα were colocalized at the membrane ruffles after a 4-h treatment with HBSS. In control cells, STAT3, p65 and Rac1 were dispersed throughout the cytoplasm (Figure 2b). Consistent with this, Rac1 and STAT3 bound to each other in the nuclei of starved cancer cells, evidenced by immunoprecipitation of the nuclear and cytoplasmic fractions (Figure 2c). These results suggest that Rac1 is required for activation of the STAT3–NFκB complexes in starved cancer cells.
Rac1 is activated and colocalized with STAT3 or p65 in starved cancer cells. (a) HeLa cells were incubated in HBSS for the indicated time periods, and total cell lysates were incubated with PAK PBD agarose beads. Bound Rac1 was detected with western blotting using a Rac1 Ab (upper panel). The amount of total Rac1 was also measured with western blotting (lower panel). PC, positive control (GTPγS), NC, negative control (GDP). (b) HeLa cells incubated in HBSS for 4 h were fluorescence-stained with anti-Rac1 Ab, anti-STAT3 Ab, anti-p65 Ab and DAPI. (c) HeLa cells expressing vector or Rac1 wild-type HA (Rac1WT-HA) and STAT3 wild-type V5 (ST3WT-V5) were western blotted with V5 Ab, HA Ab and tubulin Ab (left). HeLa cells expressing Rac1WT-HA and ST3WT-V5 were incubated with HBSS for 0 or 8 h. Nuclear and cytoplasmic fractions were prepared, immunoprecipitated with anti-STAT3 Ab and western blotted with anti-HA Ab and anti-V5 Ab (right). DAPI, 4′,6-diamidino-2-phenylindole; HBSS, Hank’s balanced salt solution; STAT3, signal transducer and activator of transcription 3.
Degradation of IκBα by Rac1 in starved cancer cells is independent of IκBα serine phosphorylation by IKK
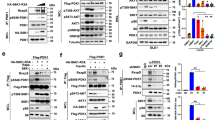
To investigate the effect of Rac1 on IκBα degradation, we examined the IκBα protein level in HeLa cells expressing non-target (NT) or Rac1 shRNA after HBSS treatment. As shown in Figure 3a, the protein level of IκBα decreased in a time-dependent manner when treated with HBSS in control cells, but not in Rac1 knockdown cells. As viewed by confocal microscopy, IκBα was diffusely spread throughout the cytoplasm of control cells, whereas upon exposure to HBSS, it disappeared. However, in Rac1 knockdown cells, IκBα was not degraded completely (Figure 3b). Because these results suggest a relationship between activated Rac1 and IκBα degradation, we next verified whether Rac1 activation depended on IκBα degradation.
Degradation of IκB kinase by Rac1 in starved cancer cells is independent of IκBα serine phosphorylation by IKK. (a) HeLa cells expressing NT shRNA or Rac1 shRNA were western blotted with Rac1 Ab and tubulin Ab (left). HeLa cells expressing NT shRNA or Rac1 shRNA were incubated in HBSS for the indicated time periods and western blotted with IκBα Ab and tubulin Ab (right). (b) HeLa cells expressing NT or Rac1 shRNA incubated in HBSS for the indicated time periods were fluorescence-stained with anti-Rac1 Ab, anti-IκBα and DAPI. Arrow indicate p105 (upper) and p50 (lower), respectively. (c) HeLa cells expressing control vector, Rac1WT-HA, Rac1Q61L-HA or Rac1N17-HA were western blotted with HA Ab and tubulin Ab (left). HeLa cells expressing control vector, Rac1WT-HA, Rac1Q61L-HA or Rac1N17-HA were incubated in HBSS for the indicated time periods and western blotted with IκBα Ab and tubulin Ab (right). (d) HeLa cells expressing NT shRNA, STAT3 shRNA or Rac1 shRNA were western blotted with STAT3 Ab, Rac1 Ab and tubulin Ab (left). HeLa cells expressing NT shRNA, STAT3 shRNA or Rac1 shRNA were incubated in HBSS for 6 h with and without MG132 (25 μM), and western blotted to detect phospho-serine IκBα, IκBα or tubulin (right). Ab, antibody; DAPI, 4′,6-diamidino-2-phenylindole; HBSS, Hank’s balanced salt solution; NT, non-target; short hairpin RNA; STAT3, signal transducer and activator of transcription 3.
We made HeLa cells that overexpressed Rac1WT-HA, the dominant-positive mutant Rac1Q61L-HA, or the dominant-negative mutant Rac1N17-HA and used a western blot to measure the amount of IκBα in HBSS-treated cells. As shown in Figure 3c, IκBα depletion had already occurred after 8 h in HeLa cells overexpressing vector, Rac1WT, and Rac1Q61, and IκBα was degraded in a time-dependent manner during HBSS treatment, whereas in HeLa cells overexpressing Rac1N17, IκBα was not completely degraded by HBSS. These data suggest that Rac1 is necessary for the degradation of IκBα.
To investigate serine-phosphorylated IκBα, MG132, an inhibitor of the ubiquitin proteasome pathway, was applied to HBSS-treated STAT3 or Rac1 knockdown HeLa cells. We found that MG132 treatment increased the level of non-phosphorylated IκBα, but not serine-phosphorylated IκBα, in starved cancer cells (Figure 3d). In addition, HeLa cells overexpressing IKK K44M, the kinase-dead mutant, had a similar NFκB activity as the control cells after HBSS treatment (Supplementary Figure 1a), as determined by the reporter assay with NFκB-responsive elements (Supplementary Figure 1b). These results indicate that Rac1 is required for IκBα degradation in starved cancer cells. This process is independent of the IκBα ubiquitination-mediated degradation via serine phosphorylation by IKK, but it is still dependent on degradation of non-phosphorylated IκBα.12 Furthermore, we recently reported that GCN–eIF2α pathway was found to be associated with NFκB activation, possibly through modified translational regulation, that is, decrease of IκBα translation along with global transloational attenuation and selective translational increase of specific genes including NFκB p65 subunit [14].
Rac1 activation is required for the nuclear translocation and activity of STAT3 and NFκB
Because STAT3 and NFκB were activated in starved cancer cells,12 we investigated whether the HBSS-induced Rac1 activation could lead to the nuclear translocation and activation of STAT3 and NFκB. First, using confocal microscopy, we analyzed the ability of STAT3 and NFκB to translocate into the nucleus using Abs against STAT3 and p65. In HeLa cells expressing NT shRNA, STAT3 and the p65 subunit translocated into the nucleus during HBSS treatment in a time-dependent manner, but not in Rac1 knockdown cells (Figure 4a). HeLa cells expressing the dominant-positive mutant Rac1Q61L-HA or the dominant-negative mutant Rac1N17-HA demonstrated the specific involvement of Rac1 in this translocation (Supplementary Figure 2). To verify that the translocation of STAT3 and NFκB to the nucleus was dependent on Rac1, the nuclear and cytoplasmic fractions were extracted from HeLa cells expressing NT shRNA or Rac1 shRNA after HBSS treatment, and western blotting was performed. Nuclear translocation was considerably inhibited during starvation in Rac1 knockdown cells, indicating that activated Rac1 was required for nuclear translocation of STAT3 and the p65 subunit (Figure 4b). Translocation of the p50 subunit into the nucleus was independent of Rac1. Using the transfected HeLa cells expressing NT shRNA or Rac1 shRNA with reporter vectors containing STAT3-responsive element or NFκB-responsive element, we found that translocation of STAT3 and NFκB correlated with their transactivation activity. STAT3 and NFκB activation was partially blocked in the HeLa cell lines expressing Rac1 shRNA, as compared with the NT control cell lines (Figure 4c). These results indicate that Rac1 induces STAT3 and NFκB activation, supporting the master role of this GTPase in the positioning and function of STAT3–NFκB complexes.
Rac1 activation induces the nuclear translocation and activity of the STAT3 and NFκB complex. (a) HeLa cells expressing NT shRNA or Rac1 shRNA were incubated in HBSS for the indicated time periods and fluorescence-stained with anti-p65 Ab, anti-STAT3 Ab and DAPI. (b) The nuclear and cytoplasmic fractions were prepared from NT-HeLa cells and Rac1 shRNA-HeLa cells incubated in HBSS for the indicated time periods, and western blotted to detect pY705–STAT3, STAT3, p65, p50, histone H3 and tubulin. (c) HeLa cells stably expressing NT shRNA or Rac1 shRNA were incubated in HBSS for the indicated time periods, and STAT3 reporter activity and NFκB reporter activity were measured by real-time PCR. Data are presented as relative quantitation (RQ)±s.d. *P<0.05 and **P<0.01 compared with the respective control. Ab, antibody; DAPI, 4′,6-diamidino-2-phenylindole; HBSS, Hank’s balanced salt solution; NT, non-target; shRNA, short hairpin RNA; NFκB, nuclear factor κB; STAT3, signal transducer and activator of transcription 3.
Unphosphorylated STAT3 binds to NFκB in starved cancer cells
In the JAK–STAT3 pathway, the most understood cytokine signaling pathway, JAK tyrosine kinases are activated by recruitment to the cytokine receptors and phosphorylate STAT3. Phosphorylated STAT3 proteins form dimers and move to the nucleus, where they are associated with various transcriptional activities. Rac1 has an important role in STAT3 tyrosine phosphorylation and nuclear translocation. Rac1 and MgcRacGAP can bind STAT3, and this Rac1–MgcRacGAP–STAT3 association is required for the STAT3 Y705 phosphorylation following cytokine stimulation.5 On the other hand, unphosphorylated STAT3, which activates gene expression by a mechanism distinct from that used by STAT3 dimers, is likely to be an important transcription factor both in cancer and in the cytokine response.17 Unphosphorylated STAT3 forms complexes with dimers of NFκB subunits and contributes to the second wave of induction of κB-responsive genes after IL6 stimulation.13
We demonstrated that Rac1 is activated in starved cancer cells (Figure 2a) and that the level of phosphorylated STAT3 is decreased after HBSS exposure in HeLa cells with Rac1 knockdown (Figure 4b). To verify whether phosphorylation of STAT3 is an essential step in activating STAT3–NFκB complexes in starved cancer cells, we measured the nuclear translocation efficiency of STAT3 in starved HeLa cells overexpressing wild-type STAT3 or the STAT3 Y705F mutant, which cannot be phosphorylated on residue 705. STAT3 translocated to the nucleus in STAT3 Y705F–HeLa cells, as well as in STAT3 WT–HeLa cells (Figure 5a), with both cell types having similar levels of the reporter with the NFκB-responsive elements as compared with the control cells (Figure 5b). Furthermore, unphosphorylated STAT3 interacted with NFκB in the nuclei, evidenced by immunoprecipitation of the nuclear and cytoplasmic fractions in starved HeLa cells expressing STAT3 WT and p65 or STAT3 Y705F and p65 (Figure 5c). Taken together, these results demonstrate that phosphorylation of STAT3 is not an essential step for the nuclear translocation or activity of the STAT3–NFκB complex in starved cancer cells.
Unphosphorylated STAT3 binds to NFκB in starved cancer cells. (a) Nuclear and cytoplasmic fractions were prepared from HeLa cells expressing STAT3WT (ST3W)-V5 or STAT3 Y705F (ST3F)-V5 incubated in HBSS for the indicated time periods and western blotted with pY705–STAT3 Ab, V5 Ab, p65 Ab, p50 Ab, histone H3 Ab and tubulin Ab. Arrows indicate pY705-STAT3 (left), p105 (right, upper) and p50 (right, lower), respectively. (b) HeLa cells stably expressing control vector, ST3W-V5 or ST3F-V5 were incubated in HBSS for the indicated time periods, and NFκB reporter activity was measured by real-time PCR. Data are presented as relative quantitation (RQ)±s.d. (c) HeLa cells expressing ST3WT-V5 and p65-HA, or ST3F-V5 and p65-HA, were western blotted with V5 Ab, HA Ab and tubulin Ab (left). The nuclear and cytoplasmic fraction were prepared from HBSS-incubated HeLa cells expressing ST3WT-V5 and p65-HA or ST3F-V5 and p65-HA, immunoprecipitated with anti-p65 Ab, and western blotted with STAT3 Ab and p65 Ab (right). *P<0.05 and **P<0.01 compared with the respective control. Ab, antibody; HBSS, Hank’s balanced salt solution; IP, immunoprecipitate; NFκB, nuclear factor κB; STAT3, signal transducer and activator of transcription 3.
Rac1 is required for translocation of the STAT3–NFκB complex in starved cancer cells
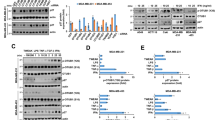
In our previous study, we found that STAT3 forms complexes with NFκB and that these complexes exist as identical nuclear complexes on the IL6 promoter in starved cancer cells.12 To investigate the role of Rac1 in the starvation-induced STAT3–NFκB interaction, HeLa cell lines stably expressing cDNA constructs (STAT3 WT and p65, with or without Rac1 shRNA) were prepared (Figure 6a). STAT3 and NFκB bound in the nuclei, evidenced by immunoprecipitation of the nuclear and cytoplasmic fractions in starved HeLa cells expressing STAT3 WT and p65. In Rac1 knockdown cells expressing STAT3 WT and p65, STAT3 and NFκB bound each other in the cytoplasm, but could not translocate to the nucleus during starvation (Figure 6b). Furthermore, cells knocked down for Rac1 by infection of Rac1 shRNA had lower levels of IL6 mRNA, similar to the knockdown of STAT3 or p65 in HeLa cells (Figure 6c). Collectively, the above data demonstrate that Rac1 activation during starvation is required for IκBα degradation and nuclear translocation of STAT3–NFκB complexes, thus contributing to IL6 induction through the formation of STAT3-NFκB complexes on the IL6 promoter.
Rac1 is required for nuclear translocation of the STAT3–NFκB complex in starved cancer cells. (a) Parental HeLa cells and HeLa cells expressing STAT3WT-V5 and p65-HA, and STAT3WT-V5, p65-HA and Rac1 shRNA were western blotted with V5 Ab, HA Ab, Rac1 Ab and tubulin Ab. (b) HeLa cells expressing ST3WT-V5 and p65-HA, and ST3WT-V5, p65-HA and Rac1 shRNA were incubated in HBSS for 8 h and had their nuclear and cytoplasmic fraction extracted and immunoprecipitated with anti-p65 Ab, followed by western blotting with STAT3 Ab and p65 Ab. (c) HeLa cells expressing NT shRNA, STAT3 shRNA, p65 shRNA or Rac1 shRNA were incubated in HBSS for 16 h, and the amounts of IL6 mRNA were measured by real-time PCR. Data are presented as relative quantitation (RQ)±s.d. Ab, antibody; C.F., cytoplasmic fraction; HBSS, Hank’s balanced salt solution; IP, immunoprecipitate; mRNA, messenger RNA; NT, non-target; N.F., nuclear fraction; NFκB, nuclear factor κB; shRNA, short hairpin RNA; STAT3, signal transducer and activator of transcription 3.
Discussion
In addition to confirming the translocation of GTP-bound Rac1 to the membrane ruffles, we also found that Rac1 activation promoted the recruitment of molecules crucial for activating STAT3–NFκB complexes, including the NFκB inhibitor, IκBα, to the membrane ruffles. More importantly, this occurred only if Rac1 was activated by treatment with HBSS (Figure 3). Supporting this, it was reported that Rac1 GTPase affects NFκB activation by conveying the SCF complex and IκBα to the membrane ruffle.10 In our study, during amino acid and serum deprivation of the HeLa cells, Rac1 was activated (Figure 2a), and the activated Rac1 regulated the degradation of IκBα (Figure 3). Moreover, phospho-serine IκBα was not detected, although the level of non-phosphorylated IκBα increased when starved cancer cells were treated with MG132, a proteasomal inhibitor (Figure 3d), indicating that the IκBα not phosphorylated by IKK is degraded by activated Rac1 in starved cancer cells to release free NFκB.
The Rho family of small GTPases has key roles in a variety of cellular functions, including regulation of the cell cycle, transcription and transformation.18 Rac1 regulates cell migration, membrane ruffling, production of superoxide and phagocytosis.18, 19, 20 It has been reported that Rac1 is involved in the assembly and activation of NADPH oxidase (Nox).21, 22 In our previous study, we found that Jak2/STAT3 was activated in amino-acid- and serum-deprivated cancer cells, as well as rapamycin-treated cancer cells.14 The activation of JAK2/STAT3 was totally dependent on reactive oxygen species produced by NADPH oxidase enzymes that were activated during early autophagic processes, because STAT3 phosphorylation was blocked completely by treatment with DPI, a specific inhibitor of Nox and partially by p22 phox knockdown.14 Supporting this, it was recently reported that reactive oxygen species induced by activated Rac1 stimulated STAT3 phosphorylation at Tyr705 and Ser727.23
The C-terminal region of Rac1 contains a functional NLS, suggesting a role for Rac1 in the nucleus. Consistent with this, GTP-bound Rac1 and MgcRacGAP function as a nuclear transport chaperone for activated STATs.5 In this study, using Rac1 knockdown HeLa cells, we demonstrated that Rac1 was required for the nuclear translocation of STAT3–NFκB complexes (Figures 4 and 6). Further studies on the composition and inter-relationship of STAT3–NFκB complexes during nuclear translocation in starved cancer cells are needed.
There are several reports that STAT3 and NFκB interact with each other. For example, Hagihara et al. demonstrated that STAT3 forms a complex with the p65 subunit of NFκB following stimulation of cells with IL1 and IL6, and that the bound STAT3 interacts with non-consensus sequences near the κB element of the SAA promoter. Moreover, they demonstrated that a complex including STAT3, p65 and p300 is essential for the synergistic induction of the SAA gene, which does not have a typical STAT3 response element in its promoter, by IL1 and IL6.24 Furthermore, a recent study found that IR treatment increased phosphorylation of p65 and STAT3, both of which eventually translocate into the nucleus in glioma. In the nucleus, CBP/p300 acetyltransferases bind to the p-p65, transfer acetyl groups and acetylate p65. p-STAT3 directly interacts with p-p65 at its transactivation domain and forms a p-p65–p-STAT3 complex in the nucleus. That study found that IR elevates p-p65/p-STAT3 complex formation and subsequent binding to the adjacent NFκB (+399) and STAT3 (+479) consensus sequences in the ICAM-1 intron 1.11 On the other hand, Yang et al. found that unphosphorylated STAT3 forms complexes with dimers of NFκB subunits to contribute to the second wave of induction of κB-responsive genes after IL6 stimulation. The intracellular concentration of unphosphorylated STAT3 increases when the STAT3 gene is activated in response to gp130-linked cytokines, allowing unphosphorylated STAT3 to compete more effectively with IκBα for unphosphorylated NFκB with which to form a novel transcription factor that induces RANTES expression by binding to the proximal κB site of the promoter.13 This suggests that only phosphorylated STAT3 can interact with p300/phosphorylated p65 efficiently, leading to p65 acetylation.
In our previous work,12 we found that the level of phosphorylated STAT3 is increased during starvation, that the activated STAT3 has an essential role in the nuclear retention of NFκB and that STAT3–NFκB exist as nuclear complexes on the IL6 promoter in starved cancer cells. We wanted to know whether the phosphorylation of STAT3 is necessary for both the formation of STAT3–NFκB complexes and the nuclear translocation of STAT3–NFκB complexes in starved cancer cells. STAT3 Y705F, as well as wild-type STAT3, could form complexes with NFκB in starved cancer cells. Furthermore, STAT3 Y705F could be translocated into the nucleus and had a critical role in NFκB activity during HBSS treatment (Figure 5). In a previous study, we found that phosphorylation of tyrosine 705 in STAT3 is upregulated in starved cancer cells.14 Therefore, STAT3 may have two distinct roles in starved cancer cells: being part of the primary response through the action of phosphorylated STAT3 and being a secondary part of the complete response through the action of increased amounts of unphosphorylated STAT3.13
STAT3 and NFκB control both distinct and overlapping groups of genes during tumorigenesis. These genes may contain one or both STAT3- and NFκB-binding sites, resulting in some being regulated by only one and others by both STAT3 and NFκB in a cooperative manner, respectively. Indeed, the overexpression of a constitutively active form of STAT3 in HME cells induces 427 genes, whereas tumor necrosis factor alpha (a major NFκB activator) induces the expression of 1225 genes, with only 123 genes dependent on both STAT3 and NFκB.13 The high number of overlapping target genes regulated by crosstalk between STAT3 and NFκB suggests the need for therapies to target both of these transcription factors. Through their functional interaction, STAT3 and NFκB collaboratively promote tumor development via induction of pro-tumorigenic genes, including chemokines, immunosuppressive cytokines and genes in angiogenesis and hypoxia.25, 26, 27 Thus, the inhibitors of STAT3–NFκB complexes may reduce some of the essential homeostatic functions executed by STAT3 and NFκB on their own, while inhibiting their malicious cooperation in cancer cells. The results of this study demonstrate the important roles of Rac1 in the function of STAT3–NFκB complexes in starved cancer cells. Our study highlights the importance of Rac1 and supports the idea that Rac1 may be a therapeutic target in cancer.
References
Sahai E, Marshall CJ . RHO-GTPases and cancer. Nat Rev Cancer 2002; 2: 133–142.
Wennerberg K, Der CJ . Rho-family GTPases: it's not only Rac and Rho (and I like it). J Cell Sci 2004; 117: 1301–1312.
Simon AR, Vikis HG, Stewart S, Fanburg BL, Cochran BH, Guan KL . Regulation of STAT3 by direct binding to the Rac1 GTPase. Science 2000; 290: 144–147.
Simeone-Penney MC, Severgnini M, Rozo L, Takahashi S, Cochran BH, Simon AR . PDGF-induced human airway smooth muscle cell proliferation requires STAT3 and the small GTPase Rac1. Am J Physiol Lung Cell Mol Physiol 2008; 294: L698–L704.
Kawashima T, Bao YC, Nomura Y, Moon Y, Tonozuka Y, Minoshima Y . Rac1 and a GTPase-activating protein, MgcRacGAP, are required for nuclear translocation of STAT transcription factors. J Cell Biol 2006; 175: 937–946.
Michaelson D, Abidi W, Guardavaccaro D, Zhou M, Ahearn I, Pagano M et al. Rac1 accumulates in the nucleus during the G2 phase of the cell cycle and promotes cell division. J Cell Biol 2008; 181: 485–496.
Frank DA . StAT signaling in cancer: insights into pathogenesis and treatment strategies. Cancer Treat Res 2003; 115: 267–291.
Groner B, Lucks P, Borghouts C . The function of Stat3 in tumor cells and their microenvironment. Semin Cell Dev Biol 2008; 19: 341–350.
Beg AA, Ruben SM, Scheinman RI, Haskill S, Rosen CA, Baldwin AS Jr . I kappa B interacts with the nuclear localization sequences of the subunits of NF-kappa B: a mechanism for cytoplasmic retention. Genes Dev 1992; 6: 1899–1913.
Boyer L, Travaglione S, Falzano L, Gauthier NC, Popoff MR, Lemichez E et al. Rac GTPase instructs nuclear factor-kappaB activation by conveying the SCF complex and IkBalpha to the ruffling membranes. Mol Biol Cell 2004; 15: 1124–1133.
Kesanakurti D, Chetty C, Rajasekhar Maddirela D, Gujrati M, Rao JS . Essential role of cooperative NF-kappaB and Stat3 recruitment to ICAM-1 intronic consensus elements in the regulation of radiation-induced invasion and migration in glioma. Oncogene 2013; 32: 5144–5155.
Yoon S, Woo SU, Kang JH, Kim K, Shin HJ, Gwak HS et al. NF-kappaB and STAT3 cooperatively induce IL6 in starved cancer cells. Oncogene 2012; 31: 3467–3481.
Yang J, Liao X, Agarwal MK, Barnes L, Auron PE, Stark GR . Unphosphorylated STAT3 accumulates in response to IL-6 and activates transcription by binding to NFkappaB. Genes Dev 2007; 21: 1396–1408.
Yoon S, Woo SU, Kang JH, Kim K, Kwon MH, Park S et al. STAT3 transcriptional factor activated by reactive oxygen species induces IL6 in starvation-induced autophagy of cancer cells. Autophagy 2010; 6: 1125–1138.
Miyano K, Sumimoto H . Role of the small GTPase Rac in p22phox-dependent NADPH oxidases. Biochimie 2007; 89: 1133–1144.
Mattagajasingh SN, Yang XP, Irani K, Mattagajasingh I, Becker LC . Activation of Stat3 in endothelial cells following hypoxia-reoxygenation is mediated by Rac1 and protein Kinase C. Biochim Biophys Acta 2012; 1823: 997–1006.
Yang J, Chatterjee-Kishore M, Staugaitis SM, Nguyen H, Schlessinger K, Levy DE et al. Novel roles of unphosphorylated STAT3 in oncogenesis and transcriptional regulation. Cancer Res 2005; 65: 939–947.
Bishop AL, Hall A . Rho GTPases and their effector proteins. Biochem J 2000; 348: 241–255.
Gu Y, Filippi MD, Cancelas JA, Siefring JE, Williams EP, Jasti AC et al. Hematopoietic cell regulation by Rac1 and Rac2 guanosine triphosphatases. Science 2003; 302: 445–449.
Ridley AJ . Rho-related proteins: actin cytoskeleton and cell cycle. Curr Opin Genet Dev 1995; 5: 24–30.
Diekmann D, Abo A, Johnston C, Segal AW, Hall A . Interaction of Rac with p67phox and regulation of phagocytic NADPH oxidase activity. Science 1994; 265: 531–533.
Ueyama T, Geiszt M, Leto TL . Involvement of Rac1 in activation of multicomponent Nox1- and Nox3-based NADPH oxidases. Mol Cell Biol 2006; 26: 2160–2174.
Kang J, Chong SJ, Ooi VZ, Vali S, Kumar A, Kapoor S et al. Overexpression of Bcl-2 induces STAT-3 activation via an increase in mitochondrial superoxide. Oncotarget 2015; 6: 34191–34205.
Hagihara K, Nishikawa T, Sugamata Y, Song J, Isobe T, Taga T et al. Essential role of STAT3 in cytokine-driven NF-kappaB-mediated serum amyloid A gene expression. Genes Cells 2005; 10: 1051–1063.
Bollrath J, Greten FR . IKK/NF-kappaB and STAT3 pathways: central signalling hubs in inflammation-mediated tumour promotion and metastasis. EMBO Rep 2009; 10: 1314–1319.
Atkinson GP, Nozell SE, Benveniste ET . NF-kappaB and STAT3 signaling in glioma: targets for future therapies. Expert Rev Neurother 2010; 10: 575–586.
He G, Karin M . NF-kappaB and STAT3 - key players in liver inflammation and cancer. Cell Res 2011; 21: 159–168.
Acknowledgements
This research was supported by the Basic Science Research Program through the National Research Foundation of Korea (NRF) funded by the Ministry of Education (NRF-2012R1A1A2041687 and NRF-2015R1D1A4A01020022).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on Experimental & Molecular Medicine website
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-ShareAlike 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/4.0/
About this article
Cite this article
Kim, S., Yoon, S. Activated Rac1 regulates the degradation of IκBα and the nuclear translocation of STAT3–NFκB complexes in starved cancer cells. Exp Mol Med 48, e231 (2016). https://doi.org/10.1038/emm.2016.17
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/emm.2016.17
This article is cited by
-
NF-κB and pSTAT3 synergistically drive G6PD overexpression and facilitate sensitivity to G6PD inhibition in ccRCC
Cancer Cell International (2020)