Abstract
Whole-exome sequencing (WES) has allowed the discovery of genes and variants causing rare human disease. This is often achieved by comparing nonsynonymous variants between unrelated patients, and particularly for sporadic or recessive disease, often identifies a single or few candidate genes for further consideration. However, despite the potential for this approach to elucidate the genetic cause of rare human disease, a majority of patients fail to realize a genetic diagnosis using standard exome analysis methods. Although genetic heterogeneity contributes to the difficulty of exome sequence analysis between patients, it remains plausible that rare human disease is not caused by de novo or recessive variants. Multiple human disorders have been described for which the variant was inherited from a phenotypically normal mosaic parent. Here we highlight the potential for exome sequencing to identify a reasonable number of candidate genes when dominant disease variants are inherited from a mosaic parent. We show the power of WES to identify a limited number of candidate genes using this disease model and how sequence coverage affects identification of mosaic variants by WES. We propose this analysis as an alternative to discover genetic causes of rare human disorders for which typical WES approaches fail to identify likely pathogenic variants.
Similar content being viewed by others
Introduction
Human disease-gene discovery is largely driven by advances in genomic technologies. The number of genes associated with human disease saw a significant expansion with the use of microarray technology for genomewide association studies (GWAS).1 The catalog of published GWAS maintains thousands of gene associations exceeding genomewide significance.2 However, gene associations do not imply causality, and specific variants within many associated loci are unknown.
The recent introduction of next-generation sequencing has again advanced our ability not only to identify genes causing human disease but also to directly identify specific gene variants, primarily nonsynonymous variants, in individual patients.3 Whole-exome sequencing (WES) remains a cost-effective alternative to sequencing entire genomes for studies involving modest numbers of patients with rare disease, and this method promises to identify many genes causing rare human disease in the future.4 Although WES greatly improves our ability to identify disease-causing variants, significant challenges remain in our ability both to comprehensively analyze sequence data as well as interpret the results (candidate genes) as they relate to the disease under study.
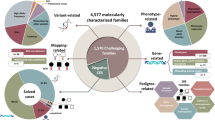
In support of this, multiple groups report varying success rates for providing a genetic diagnosis using WES.3, 5, 6, 7, 8 Success here is defined as the application of WES to identify disease-causing variants in patients. Often for rare disease, lack of a confident genetic diagnosis results from the inability to interpret potentially pathogenic variants in new disease genes, because the rationale for a gene or biological process is unclear or because there is insufficient numbers of patients to select one gene from a list of multiple equally plausible candidate genes, or because the underlying assumption of the disease model (dominant, recessive, de novo) is incorrect.
For rare disease, WES analysis often makes assumptions regarding disease inheritance (de novo vs recessive), variant frequency and genetic heterogeneity. These assumptions are interdependent and often varied between studies. For example, recessive disease analysis may be less conservative compared with sporadic analysis that includes only novel variants. Furthermore, studies allowing for allelic heterogeneity will perform differently than that restricting to shared de novo variants.
For patients without plausible genes fitting de novo or recessive models after WES analysis, focus may shift to ‘nontraditional’ analyses, such as including synonymous and intronic variants or variants in regulatory regions (5′- and 3′-UTR). Skepticism clouds the study of these variants, not undeservedly, because of our limited ability to interpret these variant classes or the potential to confound meaningful analysis with significantly higher numbers of candidate genes. However, recent studies have shown an alternative mechanism of rare/sporadic disease; rare disease may be caused by dominant variants inherited from phenotypically normal mosaic parents. Somatic mosaicism has long been implicated with diseases such as cancer; however, the extent of mosaicism in normal human tissues, particularly large chromosomal changes, are now being uncovered.9 Here we present results using WES to identify dominant sequence variants inherited from a mosaic parent. Unlike standard WES analyses for dominant or de novo variants, these variants are inherited from a phenotypically normal parent. Furthermore, this approach successfully identified the disease-causing variant in a quartet with two affected siblings in which the variant was inherited from a mosaic mother. Our results suggest that such a disease model is tractable for study using WES and may be warranted in patients with rare disease for which traditional analyses failed to identify a plausible candidate gene.
Materials and methods
Whole-exome sequencing
All research participants included in this study provided written informed consent approved by the Institutional Review Board of UT Southwestern Medical Center (UTSW). WES was performed by the Next-Generation Sequencing Core facility in the McDermott Center for Human Growth and Development at UTSW. DNA extracted from whole blood was prepared using the Illumina TruSeq kit (Illumina, San Diego, CA, USA). Exome capture was performed using the TruSeq Exome Enrichment Kit, which targets ∼62 Mb of protein-coding and regulatory sequence. Libraries were barcoded and sequenced using an Illumina HiSeq 2000, generating paired-end 100 bp reads.
Sequence analysis
Sequence reads were aligned to the human reference sequence (b37/hg19) using BWA.10 Low quality and poorly mapped reads were removed using Samtools11 and duplicate reads were removed using Picard (http://picard.sourceforge.net/). Final alignments were generated after local realignment and base quality score recalibration using the Genome Analysis Toolkit (GATK).12 Variant calling and coverage analyses were performed using GATK and Samtools. Variant quality score recalibration was performed using GATK. Variants were annotated using SeattleSeq (version 134; http://snp.gs.washington.edu/SeattleSeqAnnotation134/). Subsequent analyses were performed using custom Perl scripts (available upon request). The TUBB4A variant has been submitted to the dbSNP database (ss995812396). Two-sided Wilcoxon test was used for statistical analysis of skewed coverage distributions.
Results
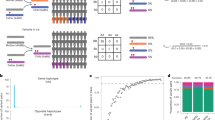
WES generated ∼50 × average gene coverage of 13 parent–child trios from seven families (Supplementary Figure 1). For high-quality heterozygous protein-coding single nucleotide variants (SNVs) with evidence of parental transmission, the distribution of allele fraction (AF, the proportion of reads with the variant allele) centered near 0.5, as expected (median=0.48; Figure 1). For all 155 470 SNPs from 13 trios, 95.30% had AF between the expected 0.3 and 0.7, a range commonly used for heterozygous variants. Mosaic variants are expected to deviate from this expectation and are often identified as those with low (<0.3) AF. We sought to identify the number of ‘candidate’ genes with rare nonsynonymous variants that fulfill a model of dominant inheritance with parental mosaicism. To determine the reproducibility in the number of candidate genes resulting from this approach, we analyzed all parent–child trios similarly. For this, we considered only rare (<1% allele frequency in the Exome Variant Server database) SNVs inherited from a single parent (the other parent with AF=0) and a heterozygous genotype in the child. No additional quality filtering was performed. Parental mosaic variants were defined as those SNVs with AF <0.3, whereas heterozygous variants in the child were required to have AF >0.3. On average, this resulted in only 17.62 genes per trio (range: 10–25; Table 1). Most candidate variants were annotated in dbSNP, suggesting they were false positives resulting from random variation in parental AF. After removing dbSNP variants, an average 6.23 novel candidate variants remained per trio (range: 4–10; Table 1). Often, quartets with two affected siblings are suspected of having rare recessive variants; however, it is possible that each sibling inherited a dominant variant from a mosaic parent. Comparing siblings, only an average 1.2 candidate genes remained (range: 0–2; Table 1).
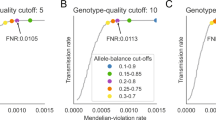
Because a majority of potential candidate genes were annotated in dbSNP and are likely false positives due to low parental AF, we sought to determine whether sequence coverage significantly affected the calculated AF. As AF is calculated, in part, from the depth of sequence coverage at each SNV position, we reasoned that low or high coverage depth would affect AF. The distribution of coverage depth was investigated separately for SNVs with low (<0.3), normal (0.3–0.7) and high (>0.7) AF. As expected, coverage depth was significantly different for SNVs with high and low AF (Figure 2). Skewed distributions were observed for the high and low AF groups, with 75th percentiles of sequence coverage of 93 and 31, respectively. Thus, sequence coverage should be investigated for all potential candidate variants.
Distribution of sequence coverage by AF category. High-quality inherited heterozygous protein-coding SNVs from 13 trios were categorized as low (<0.3), normal (0.3–0.7) or high (>0.7) AF and the distributions of sequence coverage are shown. The distributions of sequence coverage for SNVs with high and low AF were significantly different from SNVs with normal AF, with significant skewing of coverage shown by the 75th percentile of sequence coverage for each group.
To test the potential for this method to identify disease-causing variants inherited from mosaic parents, we performed WES in a quartet with two siblings diagnosed with hypomyelination with atrophy of the basal ganglia and cerebellum (HABC). The causal variant was previously reported after identifying a single de novo missense variant c.745G>A, p.(Asp249Asn) (NM_006087.2) in the TUBB4A gene in six families, as well as three additional patients for whom parental genotypes were unavailable.13 The quartet reported here is suspected to be the same as in the previous report; where the TUBB4A variant was inherited in both affected siblings from their mother who was mosaic for the variant. We sought to determine the likelihood of our approach to identify the TUBB4A variant using only WES of this quartet. The analysis identified 8 and 10 candidate genes with novel variants in each sibling. When the siblings were compared as a quartet, only TUBB4A was identified as a candidate gene (Table 1). Therefore, we recommend similar analyses be conducted in families with rare/sporadic disease for which WES failed to identify a causal variant using traditional de novo or recessive approaches.
Discussion
WES provides a powerful tool to discover new genes causing Mendelian4 and sporadic14, 15, 16, 17 disease; however, WES may be appropriate to investigate other mechanisms of disease inheritance, such as those caused by dominant variants inherited from mosaic parents. Using WES of 13 parent–child trios, we show the potential of WES to identify a modest number of candidate genes under this inheritance model, which has been shown previously to be associated with HABC,13 Kleefstra syndrome18 and X-linked severe combined immunodeficiency19 and may be relevant for seemingly recessive or sporadic disorders for which WES has failed to identify disease-causing genes. Although it is possible that variants may be inherited from mosaic fathers, in all studies mentioned above, disease-causing variants were inherited from mosaic mothers; which further reduces the number of candidate genes when applied to our families (Table 1).
Therefore, as WES continues to predominate the search for nonsynonymous sequence variants causing rare human disease, alternative analyses may help to improve genetic diagnosis in patients for which traditional analyses fail to identify a plausible candidate gene. Clinical applications for WES are evolving and face multiple challenges going forward. Not only is the use of WES expanding the variant spectrum within genes, new genes are being identified in patients with rare disease. These results suggest the growing complexity with which WES analysis will be used to provide a genetic diagnosis to allow the inclusion of new disease-causing genes or to allow reanalysis as additional suggestive evidence accumulates. Although these deal more with interpretation in the event of too many candidate genes, our analysis suggests a need for alternative methods of candidate gene discovery when traditional approaches fail to identify likely candidates. The application of next-generation sequencing in patients with rare disease will continue to expand our understanding of human disease, including alternate means of disease inheritance (that is, from mosaic parents) or genetic mechanisms involving noncoding variants.
References
McCarthy MI, Abecasis GR, Cardon LR et al: Genome-wide association studies for complex traits: consensus, uncertainty and challenges. Nat Rev Genet 2008; 9: 356–369.
Hindorff LA, Sethupathy P, Junkins HA et al: Potential etiologic and functional implications of genome-wide association loci for human diseases and traits. Proc Natl Acad Sci USA 2009; 106: 9362–9367.
Boycott KM, Vanstone MR, Bulman DE, MacKenzie AE : Rare-disease genetics in the era of next-generation sequencing: discovery to translation. Nat Rev Genet 2013; 14: 681–691.
Bamshad MJ, Ng SB, Bigham AW et al: Exome sequencing as a tool for Mendelian disease gene discovery. Nat Rev Genet 2011; 12: 745–755.
Dixon-Salazar TJ, Silhavy JL, Udpa N et al: Exome sequencing can improve diagnosis and alter patient management. Sci Transl Med 2012; 4: 138ra178.
Jacob HJ, Abrams K, Bick DP et al: Genomics in clinical practice: lessons from the front lines. Sci Transl Med 2013; 5: 194cm195.
Yang Y, Muzny DM, Reid JG et al: Clinical whole-exome sequencing for the diagnosis of mendelian disorders. N Engl J Med 2013; 369: 1502–1511.
Gahl WA, Markello TC, Toro C et al: The National Institutes of Health Undiagnosed Diseases Program: insights into rare diseases. Genet Med 2012; 14: 51–59.
Lupski JR : Genetics. Genome mosaicism–one human, multiple genomes. Science 2013; 341: 358–359.
Li H, Durbin R : Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 2009; 25: 1754–1760.
Li H, Handsaker B, Wysoker A et al: The Sequence Alignment/Map format and SAMtools. Bioinformatics 2009; 25: 2078–2079.
McKenna A, Hanna M, Banks E et al: The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res 2010; 20: 1297–1303.
Simons C, Wolf NI, McNeil N et al: A de novo mutation in the beta-tubulin gene TUBB4A results in the leukoencephalopathy hypomyelination with atrophy of the basal ganglia and cerebellum. Am J Hum Genet 2013; 92: 767–773.
Vissers LE, de Ligt J, Gilissen C et al: A de novo paradigm for mental retardation. Nat Genet 2010; 42: 1109–1112.
O'Roak BJ, Deriziotis P, Lee C et al: Exome sequencing in sporadic autism spectrum disorders identifies severe de novo mutations. Nat Genet 2011; 43: 585–589.
Neale BM, Kou Y, Liu L et al: Patterns and rates of exonic de novo mutations in autism spectrum disorders. Nature 2012; 485: 242–245.
Girard SL, Gauthier J, Noreau A et al: Increased exonic de novo mutation rate in individuals with schizophrenia. Nat Genet 2011; 43: 860–863.
Rump A, Hildebrand L, Tzschach A, Ullmann R, Schrock E, Mitter D : A mosaic maternal splice donor mutation in the EHMT1 gene leads to aberrant transcripts and to Kleefstra syndrome in the offspring. Eur J Hum Genet 2013; 21: 887–890.
Alsina L, Gonzalez-Roca E, Giner MT et al: Massively parallel sequencing reveals maternal somatic IL2RG mosaicism in an X-linked severe combined immunodeficiency family. J Allergy Clin Immunol 2013; 132: 741–743. e2.
Acknowledgements
This work was funded by Texas Scottish Rite Hospital for Children.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on European Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Rios, J., Delgado, M. Using whole-exome sequencing to identify variants inherited from mosaic parents. Eur J Hum Genet 23, 547–550 (2015). https://doi.org/10.1038/ejhg.2014.125
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2014.125