Abstract
According to most historians, Christopher Columbus was born in Genoa, Italy. However, based on some key facts in the discoverer's biography, as well as in the linguistic analysis of his texts, some historians and linguists believe that Columbus could have been of Catalan origin. A Ligurian Columbus would have carried the Colombo surname, whereas he would have been called Colom if he were Catalan. In order to test whether it would be possible to discriminate between a Ligurian or a Catalan origin were Columbus' Y-chromosome haplotype to be retrieved, we genotyped 17 Y-chromosome STRs in 238 Spanish (from Catalonia, Valencia, and the Balearic Islands) and French Colom men, and 114 North Italian Colombo (from Liguria, Lombardy, and Piedmont). The Italian samples and, in particular, the Lombard Colombos were genetically as diverse as the general population, and we found little evidence of clusters of haplotypes that could indicate descent from a single founder. Colombo is actually the most frequent surname in Lombardy, where foundlings and orphans used to be given the surname Colombo. By contrast, Y-chromosome diversity was reduced in the Iberian Colom, where most of the men had Y chromosomes belonging to a few lineages. This implies that a positive identification would be more likely if Columbus were of Catalan descent. In this study, we have shown the diverse dynamics of two surnames linked by their etymology, in what is, to the best of our knowledge, the first genetic analysis of a surname in Southern Europe.
Similar content being viewed by others
Introduction
The origin of Chistopher Columbus is the object of an ongoing debate among historians. The most widely accepted hypothesis is that Christoper Columbus was Cristoforo Colombo, from Genoa, and over 200 texts document the life of a Cristoforo Colombo; the discoverer referred to himself as a Genoese in his deed of primogeniture of 1498, although many scholars contest the authenticity of this document. However, several lines of evidence, mostly linguistic, question that Columbus could be Genoese and point instead to a Catalan origin; he would be Cristòfol Colom rather than Cristoforo Colombo. Throughout Columbus's life, he was never referred as Colombo; before 1492, his contemporaries called him Colomo and Colom, and, after the first voyage, he was almost exclusively referred to with the Spanish form Cristóbal Colón. He wrote in Spanish with lexical mistakes and phonetical misspellings that are typical of a Catalan native speaker.1, 2 According to Merrill,2 the Genoese Cristoforo Colombo was a modest wool carder and cheese merchant with no maritime training, whose age does not match that of Columbus, and it is unlikely that a tradesman would marry a Portuguese noblewoman, as Columbus did.
A genetic analysis could settle this dispute. Columbus' Y-chromosome haplotype could be compared with that of extant North Italian Colombo and Catalan Colom men; a match, with the pertinent statistical assessment, could indicate which is the most likely origin of the discoverer.
Both the Italian Colombo and the Catalan Colom derive from the Latin colmbus, ‘dove’. In Italy, 21 068 telephone landlines are registered to individuals named Colombo (online search on November 2010, http://www.paginebianche.it). Most of those (16 169, 76.7%) were found in the Lombard provinces of Lecco, Monza and Brianza, Varese, Milan, and Como, which make up 9.6% of the total Italian population. In fact, Colombo is the most frequent surname among telephone customers in Lombardy. Significant numbers of Colombo were found in the Piedmontese provinces that border Lombardy, and 486 telephone directory entries for Colombo were found in Liguria (2.3%), the region around Genoa; Colombo ranks in 27th position among Ligurian telephone customers. The abundance of people named Colombo in Milan and the surrounding region can be explained by the fact (according to Mario Colombo – Gruppo Ricerche Storiche Borsano, cited by http://www.cognomiitaliani.org/cognomi/cognomi0003col.htm) that, until 1825, the orphans and foundlings hosted by the orphanage at the Ospedale Maggiore in Milan were given the surname Colombo, because a dove figured prominently in the crest of the Ospedale Maggiore.
In Spain, 4056 males and females carry Colom as a paternal surname, according to official registry figures reported by the Spanish National Statistical Institute (http://www.ine.es/fapel/FAPEL.INICIO). Of those, 1207 (29.8%) were born in the Balearic Islands, 1976 (48.7%) in Catalonia, and 564 (13.9%) in the Valencia region. Catalan is spoken in these three regions; the Balearics and Valencia were conquered by the Crown of Aragon (constituted by Catalonia and Aragon proper) from the local Muslim rulers in the 13th century and repopulated mostly with Catalans.
In 1659, the Treaty of the Pyrenees awarded the fraction of Catalonia lying north of the Pyrenees (known as the Roussillon) to France; 91 landlines are registered to Coloms in the Roussillon.
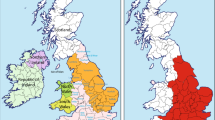
The analysis of Y-chromosome haplotypes in samples of men carrying the same surname can provide invaluable information about the genealogy linking these men, up to the point that Y chromosome analyses are being routinely used by the general public to retrace family histories.3 In England, it has been shown that it is possible to define clusters of phylogenetically related Y-chromosome haplotypes that are likely to represent the descendants of a single founder, and, from the diversity they accumulate, a time depth can be estimated that is compatible with the historical time in which paternal surname inheritance is systematized.4 In general, in England, the more frequent surnames have more diverse Y-chromosome haplotypes, as if surname frequency was driven by polyphyletism (ie, the repeated assumption of a surname by different, unrelated people).4 On the contrary, no such effect was found in Ireland,5 and the success of a surname was driven by social factors rather than by polyphyletism. It should be noted that, to the best of our knowledge, European analyses of Y-chromosome diversity in specific surnames are restricted to the British Isles (see also ref. 6).
The goal of the present research is to study the feasibility of identifying the geographical origin of Christopher Columbus in the event that his Y chromosome could be retrieved and compared with the extant Colom and Colombo men. To that effect, we have genotyped 17 Y-chromosome STRs in samples of Colombo collected in Northern Italy (Lombardy, Liguria, and Piedmont), and of Colom collected in Catalonia, the Balearic Islands, Valencia and the Roussillon. To the best of our knowledge, this is the first study of the genetics of a surname in continental Europe.
Methods
Samples
Men with the surname ‘Colom’ were sampled in Catalonia (n=126, including one sample from Andorra), the Balearic Islands (n=50), Valencia (n=45), and the Pyrénées Orientales département of Southeast France (n=17). Lists of men bearing the Colom surname were obtained from public telephone directories. Participants were shown the lists and asked to identify any relatives, in order to avoid sampling closely related men. A total of 18 men bearing the Colomb, Colom, Coulom, Coulomb, Coulon, Collon, Colon, Collomb, and Coullon surnames were sampled in Southwest France, in Bordeaux, and the rest of the Gironde département. The Colombo surname was sampled in three Italian regions: Lombardy (n=52), Liguria (n=48), and Piedmont (n=14) (Figure 1). Additionally, reference samples of 59 random Catalan males (with all eight great-grandparents born in Catalonia) and 50 North Italian men (with three generations of paternal line ancestry in Liguria and Lombardy) were gathered. In all cases, biological samples were obtained as buccal swabs, except for control Northern Italians, in which blood samples were collected at the Blood Transfusion Centre of the ‘Umberto I’ Hospital in Rome. Written informed consent was obtained from all participants.
DNA extraction and genotyping
DNA was extracted from buccal swabs using a standard organic method (proteinase K and DTT digestion, followed by phenol–chloroform extraction and Microcon 100 purification and concentration). Amplification of samples was performed with about 1 ng of target DNA. For blood samples, DNA was extracted following the salting-out procedure of Miller et al7. A total of 17 Y-chromosome STR loci (DYS19, DYS385a,b, DYS389I, DYS389II, DYS390, DYS391, DYS392, DYS393, DYS437, DYS438, DYS439, DYS448, DYS456, DYS458, DYS635, and GATAH4) contained in the AmpFlSTRYFiler PCR Amplification kit (Applied Biosystems, Foster City, CA, USA)8 were genotyped according to the manufacturer's instructions. Alleles were separated and detected using an Applied Biosystems ABI 310 genetic analyzer. Fragment sizes were analyzed using the GeneScan Analysis and Genotyper ver. 2.0 Software (Applied Biosystems). The sample run data were analyzed together with an allelic ladder and positive and negative controls. The alleles were named according to the number of repeated units based on the sequenced allelic ladder (ISFG recommendations).9
Data analysis
Basic descriptive statistics were estimated with Arlequin 3.1 (http://cmpg.unibe.ch/software/arlequin3/).10 Each individual was allocated to a haplogroup using a Bayesian approach11 as implemented in Haplogroup Predictor (http://www.hprg.com/hapest5/), with the ‘Area Selection’ field set to ‘Equal priors’; haplotypes with a posterior probability <95% were left unclassified. J2a1 and its subgroups (J2a1b, and J2a1h) were pooled, as Haplogroup Predictor often failed to discriminate among them with the current 17-STR haplotypes. We validated Haplogroup Predictor by running it through a previously published sample of 307 Y chromosomes from Catalonia, Valencia, and the Balearic Islands.12 SNPs and a widely overlapping set of Y-chromosome STRs had been typed in those samples; 14-STR-haplotypes were input in Haplogroup Predictor, with 11 (DYS19, DYS385, DYS389I, DYS389II, DYS390, DYS391, DYS392, DYS393, DYS437, DYS438, and DYS439) matching those in the present study, plus DYS460, DYS461, and DYS462.
Within each haplogroup, median joining networks13 were drawn with Network 4.5.1.6 (http://www.fluxus-engineering.com), by giving each STR a weight inversely proportional to its variance, and equating the average variance to a weight of 10. FST distances based on the number of different alleles among Y-chromosome haplotypes were computed for pairs of populations with Arlequin 3.5 (http://cmpg.unibe.ch/software/arlequin35/),14 and, after adding a small value to the entire matrix in order to remove negative values while preserving ordinality, it was plotted with multidimensional scaling as implemented in Statistica 7.0 (Statsoft, Inc., Tulsa, OK, USA).
The time depth of inherited surnames is expected to be 500–600 years in Spain and Italy;3 two extant bearers of the same surname would have at most double that time to accumulate mutations at their Y-chromosome STRs. The overall genealogical mutation rate for the 17 STRs we genotyped is 4.167 × 10−2 per generation (as compiled from multiple studies by Sascha Willuweit and Lutz Roewer and reported in http://www.yhrd.org, 22 October 2010), or, with a generation time of 25 years, 1.667 × 10−3 per year, or one mutation per 600 years. Thus, at most, bearers of the same surname are expected to differ by two mutations in their Y-chromosome STRs. Pairwise comparison of all reference haplotypes in Catalans and Italians showed that 19 out of 5886 pairs (0.32%) showed ≤2 differences, in contrast with 2.99% of the Colom/Colombo chromosome pairs. Therefore, we used these observations to construct an ad hoc heuristic to identify groups of Colom/Colombo men with shared recent ancestry. Within each haplogroup median network, haplotypes that could be linked to another haplotype by an edge at most two mutations long were grouped and were considered as descendants of a common founder. However, this algorithm has the potential to create long strings of connected haplotypes that might be false positives. We postprocessed the lineages obtained in the first pass by eliminating those with times to the most recent common ancestor >1000 year and replacing those with the lineages (if any) that could be obtained with the more stringent definition of descent lineage adopted by King and Jobling.4 Overall, we also compared the lineages obtained with our method with those produced with King and Jobling's. Time to the most common recent ancestor within lineages was computed with the ρ method15, 16 with Network 4.5.16.
Results
Haplotype, haplogroup and lineage assignment for each individual can be found in Supplementary Information. Basic descriptive statistics for the Colom/Colombo and reference samples are shown in Table 1. The Catalan and Northern Italian reference samples were quite diverse, and all haplotypes were different from each other (although one Catalan and one Northern Italian individual shared the same haplotype). Italian Colombo samples showed a slight reduction in haplotype diversity, which was more pronounced in the Iberian Colom, particularly in those from Valencia and Majorca. Pairwise differences in repeat size were reduced only in Majorcan Coloms, probably because in the remaining Colom/Colombo samples, different haplogroups were still well represented. The homogeneity of Majorcan Colom haplotypes cannot be attributed to the insular nature of the general population: the Majorcan samples in Adams et al12 showed, for a widely overlapping set of SNPs, no reduction of haplotype diversity with respect to the mainland (0.9921 vs 0.9984 in Catalans; Majorcan Coloms had H=0.5665 for the overlapping STRs).
General similarities between the samples were measured with FST and represented by means of multidimensional scaling (Figure 2a). Stress was 1.42%, below the 1st percentile for random data sets.17 The Balearic Colom appeared as extreme outliers, to the point that they might obscure the relations among other samples. When the Balearic Colom were removed from this analysis (Figure 2b), the Lombard and Ligurian Colombo, and the SW French Colomb appeared closer to the general Northern Italian and Catalan populations than other samples were.
(a) Multidimensional scaling plot based on FST distances among haplotypes. Stress was 1.4%. (b) MDS after removing the outlying Balearic Colom sample. Stress was 7.7%. Abbreviations: B, Balearic; C, Catalan; V, Valencian; FSE, SE French; FSW, SW French; LO, Lombard; LI, Ligurian; PI, Piedmontese; CTR, Catalan control sample; ITR, Northern Italian control sample.
Haplogroups were inferred for each individual based on their STR haplotypes. A previous validation study (see Methods) showed that 302 out of 307 (98.4%) Catalan, Valencian, and Balearic Y-chromosome STR haplotypes could be allocated to a haplogroup, and only three haplogroup assignments (1%) were erroneous: R1b3*, J2, and K(xP) chromosomes were called as R1a, J1, and R1b, respectively.
Haplogroup frequencies are shown in Table 1. Overall, 4 out of 479 chromosomes (0.84%) could not be classified. Most Colom/Colombo samples were similar in haplogroup frequencies to their respective reference populations. Estimated haplogroup frequencies were similar across samples, and the most salient specific features were the high frequency of J1 in the Coloms from València (31.1%, compared with 0–4% in the other samples) and the low frequency of R1b in the small Piedmontese Colombo sample (35.7% as opposed to 56–88% elsewhere).
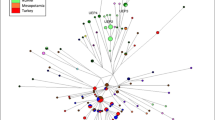
Median Joining networks for Colom/Colombo chromosomes in the major haplogroups are shown in Figures 3 and 4. These were used to detect possible founder lineages (groups of chromosomes that may descend from a single founder of the surname) as decribed in the Methods section. When we applied this approach to the reference Catalan and Northern Italian samples (which were collected regardless of surname), the 109 chromosomes represented 95 different lineages, with a maximum frequency of 5 chromosomes in Catalonia.
Median joining network for the estimated R1b chromosomes. Lineages comprising more than one haplotype are indicated with colored lines. Population color codes as in Figure 3.
In the total Colom/Colombo sample, 153 lineages were detected, most (91/135=67.4%) being represented by a single chromosome. By contrast, the eight most frequent lineages (Table 2) comprised 40.5% of the total sample, but this fraction varied from 12.5% in the Ligurian to 82% in the Balearic. Estimated ages and places of origin are shown also in Table 2, as well as the corresponding core haplotypes (descent clusters as defined more narrowly by King and Jobling4). All major lineages are clearly geographically clustered in their distribution.
Within each Colom/Colombo sample, the number and diversity of lineages varied greatly. Number of lineages and lineage diversity (computed as if it were haplogroup diversity) can be found in Table 3. Although in the Italian Colombo, the number and diversity of lineages is close to that of the general population, in the Iberian Colom (particularly in the Valencian and Balearic), a few lineages made up a sizeable portion of the chromosomes.
In the Catalan Colom sample, the four most frequent lineages comprised 40.5% of the sample; two of those have clear geographical clusterings. Lineage 77 comprised 82% of all Balearic Coloms, which is rare elsewhere and is dominated by a single haplotype covering 58.5% of the Balearic Colom in this lineage. In Valencia, four distinct lineages cover 77.8% of the chromosomes. Of note is lineage 48, which was predicted to be in haplogroup J1, which is rare in the Iberian Peninsula (0–3%12) and with 12 chromosomes carrying the distinct DYS458*16.2 allele, which has a frequency in Europe of 10/9366 (1.068 × 10−3). The Italian samples were more diverse than the Iberian ones, without frequent lineages that could point to a few discrete origins for the surname. No lineage contained more than four chromosomes in Italy.
Discussion
We have found that Colombo men in North Italy, particularly in Lombardy, carry in their Y chromosomes an array of haplotypes as diverse as that of the general population, whereas the Catalan-speaking Coloms show clear signs of founding effects, especially in València and Majorca.
In North Italy, haplotype and lineage diversity was extreme in Lombardy and less pronounced in Liguria. This observation matches the frequency of Colombo in each region: it is the most frequent surname in Lombardy, but only the 27th in Liguria. We could ask whether the high frequency by itself is sufficient to explain why a sample of Lombard Colombos is as diverse as the general population, or whether the orphanage at the Ospedale Maggiore in Milan had a role in increasing genetic diversity in the Colombos by giving that name to foundlings. Two considerations point indeed for a contribution of the foundlings: ‘Colombo’, that is, ‘dove’, does not seem a type of surname that would have the most independent origins, unlike trade names (‘Smith’) or patronyms (‘Jones’). In Campania, a South Italian region, the most frequent surname is Esposito, which was given solely to foundlings.
Genetic diversity in the Ligurian Colombos is high to the point that in a sample of 48 individuals, we found 36 different haplotypes and estimated 34 different founding events. Still, this slight reduction is sufficient to set the Ligurian Colombos apart both from the Northern Italian general population and from the Lombard Colombos, and, as seen in Figure 1, they are further apart from the Catalan Coloms; they appear closer to the Valencian Coloms, but, as discussed below, this is due just to the outlier position of the latter.
The Catalan Coloms show clear founder effects that also correspond with geographical origins; four major clusters explained 40.5% of the individuals. Colom is much less frequent than Colombo: it ranks 467th among surnames in Catalonia, and is carried by 0.027% of the population. It falls below the limit (6000 bearers) under which King and Jobling4 suggest that it is feasible to predict a surname from a Y-chromosome haplotype using the same set of STR markers we employed. The situation is more extreme in Valencia and the Balearics, and is reflected also in their position in the multidimensional scaling plot. In Valencia, four lineages covered 77.8% of the sample, again with clear geographical clustering. Given the history of the resettlement of Valencia, one could expect that those lineages would also be represented in Catalonia, but we could not identify those origins. Then, the origins of Colom in Valencia could be local: the estimated TMRCAs (Table 2) hardly overlap with the resettlement age, 750 ya. Alternatively, the Catalan descendants of the founders were not in our sample, either because we could not find them (although our sample contains ∼12% of the Catalan Colom men before excluding known relatives), or because their paternal lines became extinct. A similar situation is found in the Balearics, where the Coloms are dominated by a single lineage that comprises 82% of the sample. As in the case of the Valencian Colom, and for similar reasons, a clear founder could not be identified in the mainland.
We analyzed three other samples: the Southeast French Colom, which were linked to the Colom in Northeast Catalonia. Both regions have strong geographical, linguistic, and historic bonds. The Y chromosomes of the Southwest French Colomb reflected their mixed origins, given the variety of spellings and geographic origins gathered in that sample. Finally, the Alessandria Piedmontese Colombo seemed connected to the Ligurian Colombo, although, given its small sample size (n=5), no clear conclusions can be derived.
We have also shown that Colombo and Colom are two distinct surnames with no clear genealogical connection, and local origins in Italy and Spain. The dove, probably as a nickname, has originated surnames in many languages: Palomo (Spanish), Pigeon (French), Dove (English), Taube (German), and Golub (various Slavic languages) among others.
The main reason for this research was trying to establish whether Christopher Columbus' Y-chromosome haplotype, if retrieved, could be allocated to Liguria or to Catalonia. The most convincing evidence for either origin would be a match with a geographically specific descent cluster. If we set a simple, arbitrary threshold at a frequency of four chromosomes in each sample, then the cumulative frequencies of such lineages are 71, 87, and 82% of the Catalan, Valencian, and the Balearic Coloms, while they are only 18 and 0% in the Ligurian and Lombard Colombos, in which only lineage 109, found in four Ligurians, would provide the possibility for a specific match. On the contrary, and as also discussed above, the Colom lineages are much more geographically specific. Then, if we use as a criterion for identification a match with such lineages, a positive identification would be much more likely for a Catalan than for a Ligurian Columbus.
A match with a singleton Catalan or Ligurian haplotype should be treated with great caution, and, while indicative, would be by no means conclusive. Assessing the relative likelihood of each origin would be complex (if possible at all), and, given the sample sizes, values in favor of a particular origin would be modest. Additionally, the close similarity of the Catalan and Ligurian general populations should be taken into consideration. For instance, two different Italian reference haplotypes from our sample match two Catalan Coloms, and another Italian control matches a Catalan control.
We have shown that, although linked by their linguistic origin, Colombo and Colom are two surnames with very different histories that are reflected in the genetic diversity of their bearers, which offers a glimmer of hope for settling the dispute about Columbus' origins.
References
De Yzaguirre L : Aproximació lexicomètrica a les interferències de base fonològica en els escrits autògrafs de Cristòfor Colom: La lengua materna de Cristóbal Colón. Eivissa: Consell interinsular d'Eivissa i Formentera, 2006.
Merrill CJ : Colom of Catalonia: Origins of Christopher Columbus Revealed. Spokane, WA: Demers Books, 2008.
King TE, Jobling MA : What's in a name? Y chromosomes, surnames and the genetic genealogy revolution. Trends Genet 2009; 25: 351–360.
King TE, Jobling MA : Founders, drift, and infidelity: the relationship between Y chromosome diversity and patrilineal surnames. Mol Biol Evol 2009; 26: 1093–1102.
McEvoy B, Bradley DG : Y-chromosomes and the extent of patrilineal ancestry in Irish surnames. Hum Genet 2006; 119: 212–219.
Sykes B, Irven C : Surnames and the Y chromosome. Am J Hum Genet 2000; 66: 1417–1419.
Miller SA, Dykes DD, Polesky HF : A simple salting out procedure for extracting DNA from human nucleated cells. Nuclear Acids Research 1988; 16: 1215.
Mulero JJ, Chang CW, Calandro LM et al: Development and validation of the AmpFlSTR Yfiler PCR amplification kit: a male specific, single amplification 17 Y-STR multiplex system. J Forensic Sci 2006; 51: 64–75.
Gusmao L, Butler JM, Carracedo A et al: DNA Commission of the International Society of Forensic Genetics (ISFG): an update of the recommendations on the use of Y-STRs in forensic analysis. Forensic Sci Int 2006; 157: 187–197.
Excoffier L, Laval G, Schneider S : Arlequin (version 3.0): an integrated software package for population genetics data analysis. Evol Bioinform Online 2005; 1: 47–50.
Athey TW : Haplogroup prediction from Y-STR values using a Bayesian allele frequency approach. J Genet Geneal 2006; 2: 34–39.
Adams SM, Bosch E, Balaresque PL et al: The genetic legacy of religious diversity and intolerance: paternal lineages of Christians, Jews, and Muslims in the Iberian Peninsula. Am J Hum Genet 2008; 83: 725–736.
Bandelt HJ, Forster P, Rohl A : Median-joining networks for inferring intraspecific phylogenies. Mol Biol Evol 1999; 16: 37–48.
Excoffier L, Lischer HEL : Arlequin suite ver 3.5: a new series of programs to perform population genetics analyses under Linux and Windows. Mol Ecol Resources 2010; 10: 564–567.
Morral N, Bertranpetit J, Estivill X et al: Tracing the origin of the major cystic fibrosis mutation (ΔF508) in European populations. Nat Genet 1994; 7: 169–175.
Saillard J, Forster P, Lynnerup N, Bandelt HJ, Nørby S : mtDNA variation among Greenland Eskimos: the edge of the Beringian expansion. Am J Hum Genet 2000; 67: 718–726.
Sturrock K, Rocha J : A multidimensional scaling stress evalutation table. Field Methods 2000; 12: 49–60.
Acknowledgements
We wish to express our gratitude to all donors, and to Professor Gabriella Girelli (University of Rome ‘La Sapienza’) and Ms Irene Contini (University of Rome ‘Tor Vergata’) for their cooperation in collecting the control north Italy samples. Funding for this research was provided by Malvarrosa Media – Valencia, Fundación M. Botín and University of Granada. Sampling of the Colom men was sponsored by ‘Fausto Producciones’.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on European Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Martínez-González, L., Martínez-Espín, E., Álvarez, J. et al. Surname and Y chromosome in Southern Europe: a case study with Colom/Colombo. Eur J Hum Genet 20, 211–216 (2012). https://doi.org/10.1038/ejhg.2011.162
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2011.162
Keywords
This article is cited by
-
The Y chromosome as the most popular marker in genetic genealogy benefits interdisciplinary research
Human Genetics (2017)
-
The relationship between surname frequency and Y chromosome variation in Spain
European Journal of Human Genetics (2016)
-
Y-chromosome diversity in Catalan surname samples: insights into surname origin and frequency
European Journal of Human Genetics (2015)