Abstract
The most frequent causes of autosomal dominant (AD) hereditary spastic paraplegias (HSP) (ADHSP) are mutations in the SPAST gene (SPG4 locus). However, roughly 60% of patients are negative for SPAST mutations, despite their family history being compatible with AD inheritance. A mutation in the gene for an acetyl-CoA transporter (SLC33A1) has recently been reported in one Chinese family to cause ADHSP-type SPG42. In this study, we screened 220 independent SPAST mutation-negative ADHSP samples for mutations in the SLC33A1 gene by high-resolution melting curve analysis. Conspicuous samples were validated by direct sequencing. Moreover, copy number variations affecting SLC33A1 were screened by multiplex ligation-dependent probe amplification assay. We could not identify potentially disease-causing mutations in our patients either by mutation scanning or by gene dosage analysis, as for the latter specific positive controls are not available to date. As our sample represents ADHSP patients for whom SPAST mutations and almost in all cases ATL1 and REEP1 mutations had been excluded, we consider SLC33A1 gene mutations as being very rare in a European ADHSP cohort, if present at all. To date, as SPG42 has still not been identified in a second, unrelated family, systematic genetic testing for SLC33A1 mutations is not recommended.
Similar content being viewed by others
INTRODUCTION
The hereditary spastic paraplegias (HSPs) are a group of neurodegenerative diseases clinically characterized by progressive lower limb spasticity, pyramidal weakness and extensor plantar responses. The main neuropathological feature of these conditions is declining degeneration of the corticospinal tracts. Pure (uncomplicated) and complicated forms of HSP have been described according to the presence of additional neurological features such as cerebellar ataxia, peripheral neuropathy and/or cognitive impairment, in addition to spastic paraparesis. In addition to clinical manifestation, age at onset and genetic background are also highly heterogeneous.1, 2 To date, at least 46 different loci have been mapped, associated with autosomal dominant (AD), autosomal recessive and X-linked mode of inheritance.1, 2, 3 A total of 17 responsible genes have been identified according to the HUGO and OMIM databases. Consistent with this high variability, HSP gene products seem to be involved in a wide range of cellular functions and pathogenic mechanisms.4 AD inheritance is the most common trait in pure HSP. At present, 18 loci have been linked to ADHSP, and 9 genes have been identified. Mutations in genes SPASTIN (MIM# 604277; SPG4, MIM# 182601), ATL1 (MIM# 606439; SPG3, MIM# 182600), KIF5A (MIM# 602821; SPG10, MIM# 604187) and REEP1 (MIM# 609139; SPG31, MIM# 610250) account for ∼40, ∼10, ∼3 and ∼8% of all ADHSP patients, respectively.2, 5, 6, 7 The remaining HSP genes seem to be relatively rare.8, 9 Spastic paraplegia type 42 (SPG42, MIM# 612539) was recently mapped to the 3q24–26 chromosomal region in a single Chinese family presenting with pure ADHSP. On the basis of the detection of a serine-to-arginine substitution at codon113 in this family, the authors reported SLC33A1 (MIM# 603690) to be the responsible gene.10 The protein product of SLC33A1 is an acetyl-CoA transporter, which serves as a substrate of acetyltransferases that modify the sialyl residues of gangliosides and glycoproteins.11 It is hypothesized that the modification of gangliosides and glycoproteins by acetylation probably has a critical role in the outgrowth and maintenance of axons of motor neurons.10
The frequency of SPG42 and its associated phenotypes is not known, as no further families with SLC33A1 mutations have been described so far.10 The purpose of this study was to determine the frequency of SPG42 by screening 220 European (German, French, Norwegian) ADHSP patients for conventional and gene dosage mutations.
PATIENTS AND METHODS
Patients
A total of 220 DNA samples of unrelated HSP patients were recruited through the European Spastic Paraplegia network (EUROSPA) and the Norwegian HSP outpatient clinic. All patients were assessed by experienced neurologists. For all patients, a family history consistent with dominant inheritance was reported. All index patients were of European descent (Germany (n=45), France (n=147) and Norwegian (n=28)). The criterion for inclusion in this study was a mutation-negative state for SPAST. Mutations in the ATL1 and REEP1 gene had previously been excluded by direct sequencing and multiplex ligation-dependent probe amplification (MLPA) at the following frequencies: SPG3A sequencing in 55% of cases, SPG3A MLPA in 85% and SPG31 (sequencing and MLPA) in 84% of cases.7, 9, 12, 13, 14 The HSP phenotype was pure in 125 (57%) and complicated in 95 (43%) patients. Informed consent was obtained in all cases.
Methods
Analysis of the SLC33A1 gene
DNA was extracted from peripheral blood samples following standard protocols. The six coding exons of the SLC33A1 gene (ENSG00000169359, ENST00000359479) were screened by high-resolution melting curve analysis (HRM) in all patients. The primers were designed to flank the coding regions and to amplify fragments of an average size of 250 bp (with the largest amplicon spanning 298 bp). PCR and HRM were performed in a single run on a LightCycler 480 instrument (Roche Diagnostics, Mannheim, Germany). A normal control DNA, which had been completely sequenced, served as the HRM baseline sample in each fragment. Samples with an aberrant melting profile were assigned for validation sequencing. These samples were reamplified from genomic DNA and directly sequenced to identify or exclude sequence variants. Forward and reverse sequence reactions were performed with the Big Dye Terminator Cycle Sequencing Ready Reaction Kit (Applied Biosystems Inc., Foster City, CA, USA) using the same primers. The sequence products were analyzed on an ABI3100 Genetic Analyzer (Applied Biosystems Inc.). See supplement for detailed experimental procedures and primer sequences (Supplementary Table 1).
To screen for copy number aberrations affecting SLC33A1, we designed an MLPA assay. It targets the coding sequence of each exon, as well as the promoter sequence of the SLC33A1 gene, with one probe each. Three probes localizing to different chromosomes are included as references (probe sequences are available on request). Pertinent synthetic oligonucleotides (MWG Biotech, Ebersberg, Germany) and reagents from the EK1 kit provided by MRC-Holland (Amsterdam, The Netherlands) were used. MLPA reactions were performed according to the instructions of the manufacturers. Analysis of MLPA data was carried out as described previously.15
RESULTS AND DISCUSSION
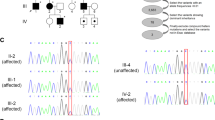
HRM analysis detected an aberrant melting profile caused by an A>G substitution at position 512 of the coding sequence that causes a p. D171G substitution in 16 index patients (Figure 1). This sequence abnormality represents the known single-nucleotide polymorphism (SNP) rs3804769 according to the SNP database.16 No other sequence alterations were detected. As HRM analysis is a screening method with ∼99% sensitivity,17, 18 we handled the results with care. Along with this high sensitivity, HRM is a simple, rapid and low-cost method to screen for unknown sequence variants in a high-throughput modality. Compared with other methods, such as denaturing high-performance liquid chromatography, HRM offers at least two additional benefits: (1) the temperature gradient covers all potential melting domains of amplicons, which provides superior detection sensitivity; and (2) the closed-tube method reduces post-PCR manipulations and the risk of pipetting errors and contaminations.17, 18 Moreover, our identification of the rs3804769 SNP represents a positive control and argues for the general validity of our approach.
Known SNP rs3804769 in SLC33A1. In total, 16 patients were found to be heterozygous for rs3804769 in SLC33A1 exon 1 (c.512A>G, p.D171G). A total of 16 variant alleles (7.0%) were present in 440 sample chromosomes, as assessed by high-resolution melting curve analysis (left part). Difference plot analysis revealed the heterozygous group (red), in addition to the homozygous samples (blue). Heterozygosity has been validated by conventional sequencing (right part).
Similar to all PCR-based techniques, large insertions or deletions are usually not detected during gene scanning by HRM.17 This mutational class is frequent (∼20%) in SPG4 and is also relevant for SPG31.7, 14, 19 Both forms are associated with pure ADHSP. To assess quantitative changes in SLC33A1 copy number, MLPA was used but no rearrangements were identified. As no duplications or deletions of SLC33A1 have been reported so far, no positive control was available. However, the technique can be regarded as rather robust, and homemade probe sets developed by other groups (ie, Ganesamoorthy et al20), as well as by our groups (unpublished), have been successfully used in unambiguously detecting copy number alterations. Moreover, high overall data quality (data not shown) argues for the applicability of this assay; therefore, we believe that our findings are not likely to be false negative.
Screening for gene dosage seemed especially important in SPG42, as haploinsufficiency is supposed to be the pathogenic disease mechanism.10 In addition, SLC33A1 has an unusually high content of Alu sequences (38.1% compared with the ∼10% genomic average21). In other genes, such high values are associated with an increased susceptibility to genomic deletions (ie, MSH2).22 Nevertheless, MLPA failed to identify this potential pathological mutation class in our study.
The 220 index patients screened in this study represent a cohort of more than 500 consecutive ADHSP index patients, as mutations in common HSP genes (SPASTIN, ATL1 and REEP1), which account for >50% of our ADHSP diagnoses, had already been excluded. On the basis of data published previously by Lin et al,10 the results of this study can be interpreted in two ways. First, the lack of SLC33A1 mutations in our sample may indicate that SPG42 is an extremely rare form of ADHSP, at least in our European cohort. An alternative explanation for the negative finding of our study may consider the SLC33A1 gene to not be the true disease gene for the SPG42 HSP subtype. As only a single SPG42 family has been described so far, it is difficult to draw any genotype/phenotype correlations. In this Chinese family, 20 affected subjects presented with a ‘pure’ form of HSP starting between 4 and 42 years of age; only some had wasting of lower limbs mentioned, apart from pyramidal symptoms.
Identification of additional SPG42 families would help to decide whether SPG42 represents a pure spastic paraplegia or whether a complicated course might be seen in other patients with different mutations. Moreover, a next-generation sequencing approach encompassing the whole locus in the only SPG42 family known so far should help to search for sequence alterations in the nearly 125 genes that have not been screened yet.
References
Fink JK : Hereditary spastic paraplegia. Curr Neurol Neurosci Rep 2006; 6: 65–76.
Depienne C, Stevanin G, Brice A, Durr A : Hereditary spastic paraplegias: an update. Curr Opin Neurol 2007; 20: 674–680.
Harding AE : Classification of the hereditary ataxias and paraplegias. Lancet 1983; 1: 1151–1155.
Salinas S, Proukakis C, Crosby A, Warner TT : Hereditary spastic paraplegia: clinical features and pathogenetic mechanisms. Lancet Neurol 2008; 7: 1127–1138.
Sauter SM, Engel W, Neumann LM, Kunze J, Neesen J : Novel mutations in the Atlastin gene (SPG3A) in families with autosomal dominant hereditary spastic paraplegia and evidence for late onset forms of HSP linked to the SPG3A locus. Hum Mutat 2004; 23: 98.
Schule R, Kremer BP, Kassubek J et al: SPG10 is a rare cause of spastic paraplegia in European families. J Neurol Neurosurg Psychiatry 2008; 79: 584–587.
Beetz C, Schule R, Deconinck T et al: REEP1 mutation spectrum and genotype/phenotype correlation in hereditary spastic paraplegia type 31. Brain 2008; 131 (Part 4): 1078–1086.
Hansen J, Svenstrup K, Ang D et al: A novel mutation in the HSPD1 gene in a patient with hereditary spastic paraplegia. J Neurol 2007; 254: 897–900.
Beetz C, Schule R, Klebe S et al: Screening of hereditary spastic paraplegia patients for alterations at NIPA1 mutational hotspots. J Neurol Sci 2008; 268: 131–135.
Lin P, Li J, Liu Q et al: A missense mutation in SLC33A1, which encodes the acetyl-CoA transporter, causes autosomal-dominant spastic paraplegia (SPG42). Am J Hum Genet 2008; 83: 752–759.
Hirabayashi Y, Kanamori A, Nomura KH, Nomura K : The acetyl-CoA transporter family SLC33. Pflugers Arch 2004; 447: 760–762.
Namekawa M, Ribai P, Nelson I et al: SPG3A is the most frequent cause of hereditary spastic paraplegia with onset before age 10 years. Neurology 2006; 66: 112–114.
Goizet C, Boukhris A, Mundwiller E et al: Complicated forms of autosomal dominant hereditary spastic paraplegia are frequent in SPG10. Hum Mutat 2009; 30: E376–E385.
Depienne C, Fedirko E, Forlani S et al: Exon deletions of SPG4 are a frequent cause of hereditary spastic paraplegia. J Med Genet 2007; 44: 281–284.
Schule R, Brandt E, Karle KN et al: Analysis of CYP7B1 in non-consanguineous cases of hereditary spastic paraplegia. Neurogenetics 2008; 10: 97–104.
Sherry ST, Ward MH, Kholodov M et al: dbSNP: the NCBI database of genetic variation. Nucleic Acids Res 2001; 29: 308–311.
Wittwer CT : High-resolution DNA melting analysis: advancements and limitations. Hum Mutat 2009; 30: 857–859.
Vossen RH, Aten E, Roos A, den Dunnen JT : High-resolution melting analysis (HRMA) – more than just sequence variant screening. Hum Mutat 2009; 30: 860–866.
Beetz C, Nygren AO, Schickel J et al: High frequency of partial SPAST deletions in autosomal dominant hereditary spastic paraplegia. Neurology 2006; 67: 1926–1930.
Ganesamoorthy D, Bruno DL, Schoumans J et al: Development of a multiplex ligation-dependent probe amplification assay for diagnosis and estimation of the frequency of spinocerebellar ataxia type 15. Clin Chem 2009; 55: 1415–1418.
Lander ES, Linton LM, Birren B et al: Initial sequencing and analysis of the human genome. Nature 2001; 409: 860–921.
Charbonnier F, Baert-Desurmont S, Liang P et al: The 5′ region of the MSH2 gene involved in hereditary non-polyposis colorectal cancer contains a high density of recombinogenic sequences. Hum Mutat 2005; 26: 255–261.
Acknowledgements
We thank the patients for participating and the DNA and Cell bank of the Centre de Recherche de l’Institut du Cerveau et de la Moelle épinière (Paris, France). This study was supported by the German Ministry of Education and research (BMBF) funding the German Network for Movement Disorders (GeNeMove) Grant 01GM0603, the Programme Hospitalier de Recherche Clinique (to AD) and by the E-RARE program of the EU funding the European Network of Spastic Paraplegia (EUROSPA) Grant 01GM0807. CB was supported by the Foundation for Pathobiochemistry and Molecular Diagnostics.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Web resources
The URLs for data presented herein are as follows:
HUGO Gene Nomenclature Committee. http://www.genenames.org
Online Medelian Inheritance in Man (OMIM), http://www.ncbi.nlm.nih.gov/omim/
UCSC Genome Browser, http://genome.ucsc.edu
Supplementary Information accompanies the paper on European Journal of Human Genetics website
Supplementary information
Rights and permissions
About this article
Cite this article
Schlipf, N., Beetz, C., Schüle, R. et al. A total of 220 patients with autosomal dominant spastic paraplegia do not display mutations in the SLC33A1 gene (SPG42). Eur J Hum Genet 18, 1065–1067 (2010). https://doi.org/10.1038/ejhg.2010.68
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2010.68
Keywords
This article is cited by
-
Application of a custom NGS gene panel revealed a high diagnostic utility for molecular testing of hereditary ataxias
Journal of Applied Genetics (2022)
-
Challenges and Controversies in the Genetic Diagnosis of Hereditary Spastic Paraplegia
Current Neurology and Neuroscience Reports (2021)
-
Huppke-Brendel syndrome in a seven months old boy with a novel 2-bp deletion in SLC33A1
Metabolic Brain Disease (2016)
-
Hereditary spastic paraplegia: clinico-pathologic features and emerging molecular mechanisms
Acta Neuropathologica (2013)
-
A high-throughput resequencing microarray for autosomal dominant spastic paraplegia genes
neurogenetics (2012)